Phylogenetic Analysis of Trichoderma Species Associated with Green Mold Disease on Mushrooms and Two New Pathogens on Ganoderma sichuanense
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Isolation
2.2. Growth Characterization
2.3. Morphological Study
2.4. DNA Extraction, PCR, and Sequencing
2.5. Phylogenetic Analyses
3. Results
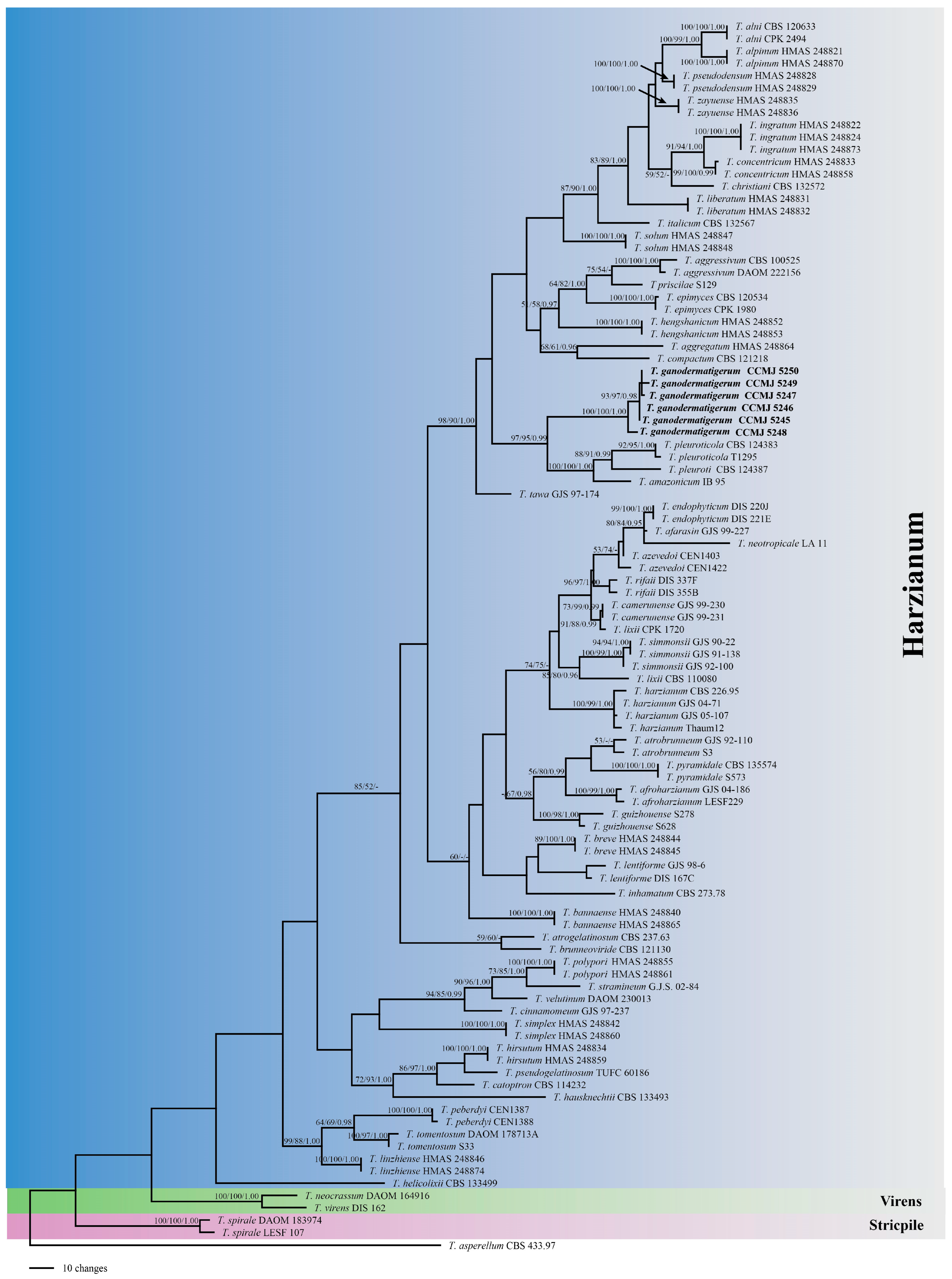
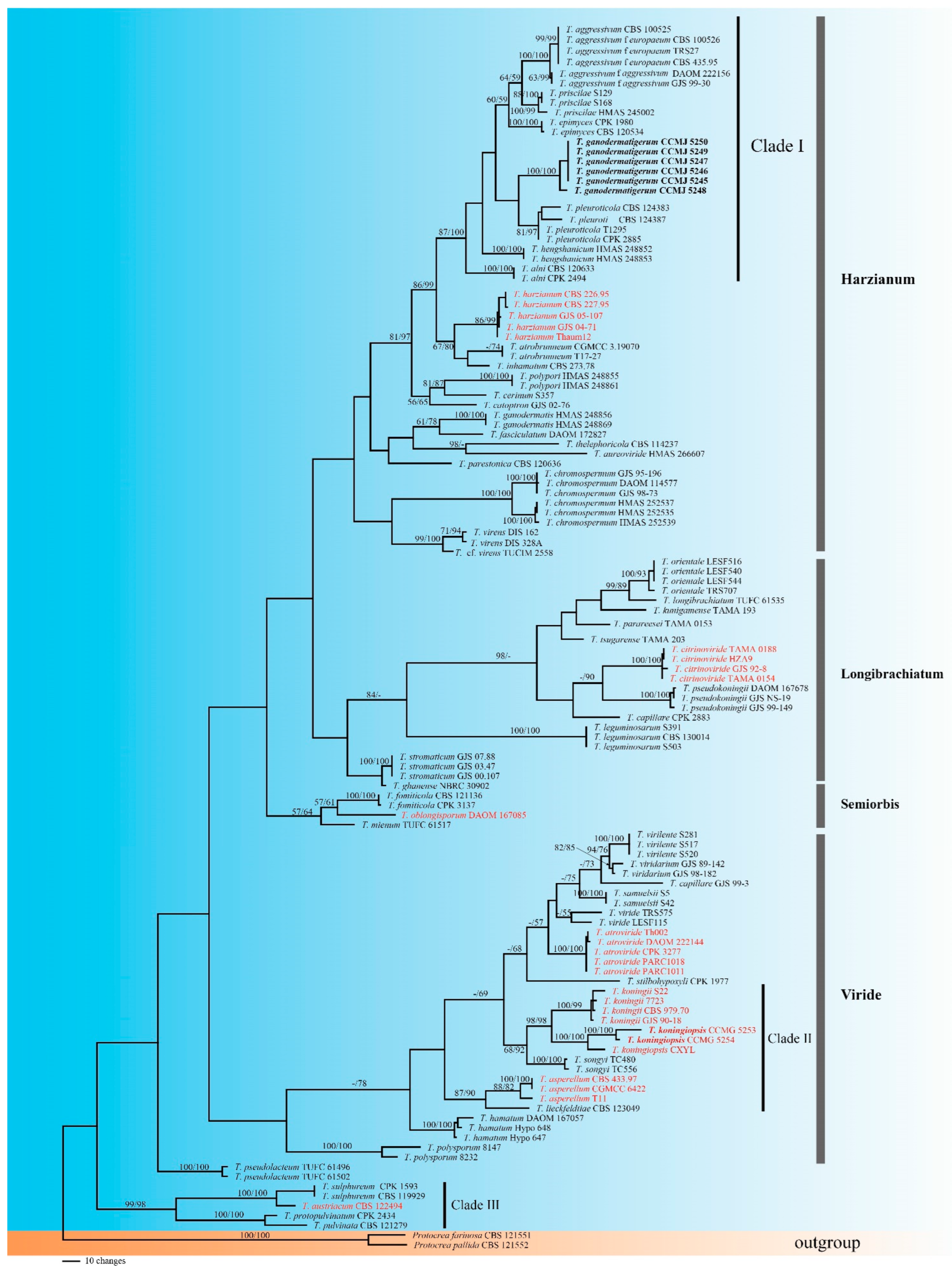
3.1. Molecular Phylogeny
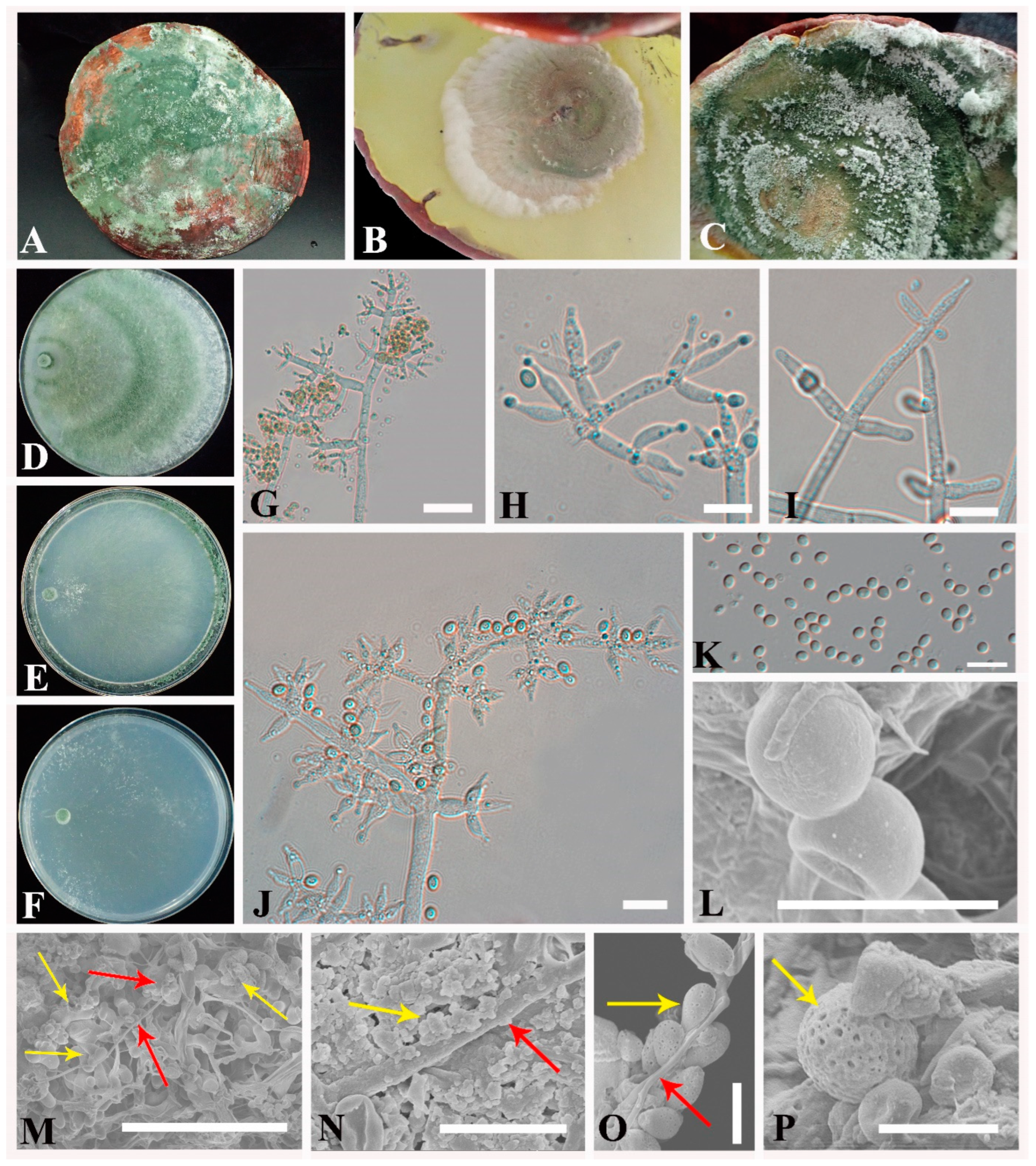
3.2. Taxonomy
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Dong, C.H.; Guo, S.P.; Wang, W.F.; Liu, X.Z. Cordyceps industry in China. Mycology 2015, 6, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.X.; Chen, Q.; Huang, C.Y.; Gao, W.; Qu, J.B. History, current situation and trend of edible mushroom industry development. Mycosystema 2015, 34, 524–540. [Google Scholar] [CrossRef]
- Andri Wihastuti, T.; Sargowo, D.; Heriansyah, T.; Eka Aziza, Y.; Puspitarini, D.; Nur Iwana, A.; Astrida Evitasari, L. The reduction of aorta histopathological images through inhibition of reactive oxygen species formation in hypercholesterolemia rattus norvegicus treated with polysaccharide peptide of Ganoderma lucidum. Iran. J. Basic Med. Sci. 2015, 18, 514–519. [Google Scholar] [PubMed]
- Nguyen, V.T.; Tung, N.T.; Cuong, T.D.; Hung, T.M.; Kim, J.A.; Woo, M.H.; Choi, J.S.; Lee, J.H.; Min, B.S. Cytotoxic and anti-angiogenic effects of lanostane triterpenoids from Ganoderma lucidum. Phytochem. Lett. 2015, 12, 69–74. [Google Scholar] [CrossRef]
- Cao, Y.; Wu, S.; Dai, Y. Species clarification of the prize medicinal Ganoderma mushroom “Lingzhi”. Fungal Divers. 2012, 56, 49–62. [Google Scholar] [CrossRef]
- Kim, S.; Song, J.; Choi, H.T. Genetic transformation and mutant isolation in Ganoderma lucidum by restriction enzyme-mediated integration. FEMS Microbiol. Lett. 2004, 233, 201–204. [Google Scholar] [CrossRef][Green Version]
- Vitak, T.Y.; Wasser, S.P.; Nevo, E.; Sybirna, N.O. The effect of the medicinal mushrooms Agaricus brasiliensis and Ganoderma lucidum (higher Basidiomycetes) on the erythron system in normal and streptozotocin-induced diabetic rats. Int. J. Med. Mushrooms 2015, 17, 277–286. [Google Scholar] [CrossRef]
- Zhao, L.Y.; Dong, Y.H.; Chen, G.T.; Hu, Q.H. Extraction, purification, characterization and antitumor activity of polysaccharides from Ganoderma lucidum. Carbohydr. Polym. 2010, 80, 783–789. [Google Scholar] [CrossRef]
- Lu, B.; Zuo, B.; Liu, X.; Feng, J.; Wang, Z.; Gao, J. Trichoderma harzianum causing green mold disease on cultivated Ganoderma lucidum in Jilin province, China. Plant Dis. 2016, 100, 2524. [Google Scholar] [CrossRef]
- Yan, Y.H.; Zhang, C.L.; Moodley, O.; Zhang, L.; Xu, J.Z. Green mold caused by Trichoderma atroviride on the Lingzhi medicinal mushroom, Ganoderma lingzhi (Agaricomycetes). Int. J. Med. Mushrooms 2019, 21, 515–521. [Google Scholar] [CrossRef]
- Zuo, B.; Lu, B.; Liu, X.; Wang, Y.; Ma, G.; Gao, J. First report of Cladobotryum mycophilum causing cobweb on Ganoderma lucidum cultivated in Jilin province, China. Plant Dis. 2016, 100, 1239. [Google Scholar] [CrossRef]
- Persoon, C.H. Neuer versuch einer systematischen eintheilung der schwämme. Römer’s Neues Mag. Bot. 1794, 1, 63–128. [Google Scholar] [CrossRef]
- Barrera, V.A.; Iannone, L.; Romero, A.I.; Chaverri, P. Expanding the Trichoderma harzianum species complex: Three new species from Argentine natural and cultivated ecosystems. Mycologia 2021, 113, 1136–1155. [Google Scholar] [CrossRef] [PubMed]
- Bissett, J.; Gams, W.; Jaklitsch, W.; Samuels, G.J. Accepted Trichoderma names in the year 2015. IMA Fungus 2015, 6, 263–295. [Google Scholar] [CrossRef] [PubMed]
- Cai, F.; Druzhinina, I.S. In honor of John Bissett: Authoritative guidelines on molecular identification of Trichoderma. Fungal Divers. 2021, 107, 1–69. [Google Scholar] [CrossRef]
- Chaverri, P.; Castlebury, L.A.; Samuels, G.J.; Geiser, D.M. Multilocus phylogenetic structure within the Trichoderma harzianum/Hypocrea lixii complex. Mol. Phylogenet. Evol. 2003, 27, 302–313. [Google Scholar] [CrossRef]
- Druzhinina, I.S.; Komoń-Zelazowska, M.; Ismaiel, A.; Jaklitsch, W.; Mullaw, T.; Samuels, G.J.; Kubicek, C.P. Molecular phylogeny and species delimitation in the section Longibrachiatum of Trichoderma. Fungal Genet. Biol. 2012, 49, 358–368. [Google Scholar] [CrossRef]
- Jaklitsch, W.M. European species of Hypocrea Part I. The green-spored species. Stud. Mycol. 2009, 63, 1–91. [Google Scholar] [CrossRef]
- Jaklitsch, W.M. European species of Hypocrea part II: Species with hyaline ascospores. Fungal Divers. 2011, 48, 1–250. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Voglmayr, H. Biodiversity of Trichoderma (Hypocreaceae) in Southern Europe and Macaronesia. Stud. Mycol. 2015, 80, 1–87. [Google Scholar] [CrossRef]
- Kullnig-Gradinger, C.M.; Szakacs, G.; Kubicek, C.P. Phylogeny and evolution of the genus Trichoderma: A multigene approach. Mycol. Res. 2002, 106, 757–767. [Google Scholar] [CrossRef]
- Pani, S.; Kumar, A.; Sharma, A. Trichoderma harzianum: An overview. Bull. Environ. Pharmacol. Life Sci. 2021, 10, 32–39. [Google Scholar]
- Samuels, G.J.; Dodd, S.L.; Lu, B.S.; Petrini, O.; Schroers, H.J.; Druzhinina, I.S. The Trichoderma koningii aggregate species. Stud. Mycol. 2006, 56, 67–133. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.Z.; Liu, X.Z.; McKenzie, E.H.C.; Jeewon, R.; Liu, J.K.; Zhang, X.L.; Zhao, Q.; Hyde, K.D. Fungicolous fungi: Terminology, diversity, distribution, evolution, and species checklist. Fungal Divers. 2019, 95, 337–430. [Google Scholar] [CrossRef]
- Zhu, Z.X.; Zhuang, W.Y. Trichoderma (Hypocrea) species with green ascospores from China. Persoonia 2015, 34, 113–129. [Google Scholar] [CrossRef]
- Zhu, Z.X.; Zhuang, W.Y.; Li, Y. A new species of the Longibrachiatum Clade of Trichoderma (Hypocreaceae) from Northeast China. Nova Hedwigia 2018, 106, 441–453. [Google Scholar] [CrossRef]
- Atanasova, L.; Druzhinina, I.S.; Jaklitsch, W.M. Two hundred Trichoderma species recognized on the basis of molecular phylogeny. In Trichoderma: Biology and Applications; Mukherjee, P.K., Horwitz, B.A., Singh, U.S., Mukherjee, M., Schmoll, M., Eds.; CABI Publishing: Croydon, UK, 2013; pp. 10–42. [Google Scholar]
- Bissett, J. A revision of the genus Trichoderma. II. Infrageneric classification. Can. J. Bot. 1991, 69, 2357–2372. [Google Scholar] [CrossRef]
- Bissett, J. A revision of the genus Trichoderma. IV. Additional notes on section Longibrachiatum. Can. J. Bot. 1991, 69, 2418–2420. [Google Scholar] [CrossRef]
- Aamir, M.; Kashyap, S.P.; Zehra, A.; Dubey, M.K.; Singh, V.K.; Ansari, W.A.; Upadhyay, R.S.; Singh, S. Trichoderma erinaceum bio-priming modulates the WRKYs defense programming in tomato against the Fusarium oxysporum f. sp. lycopersici (Fol) challenged condition. Front. Plant Sci. 2019, 10, 911. [Google Scholar] [CrossRef]
- Erazo, J.G.; Palacios, S.A.; Pastor, N.; Giordano, F.D.; Rovera, M.; Reynoso, M.M.; Venisse, J.S.; Torres, A.M. Biocontrol mechanisms of Trichoderma harzianum ITEM 3636 against peanut brown root rot caused by Fusarium solani RC 386. Biol. Control 2021, 164, 104774. [Google Scholar] [CrossRef]
- John, R.P.; Tyagi, R.D.; Prévost, D.; Brar, S.K.; Pouleur, S.; Surampalli, R.Y. Mycoparasitic Trichoderma viride as a biocontrol agent against Fusarium oxysporum f. sp. adzuki and Pythium arrhenomanes and as a growth promoter of soybean. Crop Prot. 2010, 29, 1452–1459. [Google Scholar] [CrossRef]
- Wonglom, P.; Ito, S.; Sunpapao, A. Volatile organic compounds emitted from endophytic fungus Trichoderma asperellum T1 mediate antifungal activity, defense response and promote plant growth in lettuce (Lactuca sativa). Fungal Ecol. 2020, 43, 100867. [Google Scholar] [CrossRef]
- Chomnunti, P.; Hongsanan, S.; Aguirre-Hudson, B.; Tian, Q.; Peršoh, D.; Dhami, M.K.; Alias, A.S.; Xu, J.C.; Liu, X.Z.; Stadler, M.; et al. The sooty moulds. Fungal Divers. 2014, 66, 1–36. [Google Scholar] [CrossRef]
- Nirenberg, H.I. Neuerscheinung. Untersuchungen über die morphologische und biologische Differenzierung in der Fusarium-Sektion Liseola, von Dr. Helgard Nirenberg (Inst. f. Mykologie). Z. Pflanzenernahr. Bodenkd. 1977, 140, 243. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Samuels, G.J.; Dodd, S.L.; Lu, B.S.; Druzhinina, I.S. Hypocrea rufa/Trichoderma viride: A reassessment, and description of five closely related species with and without warted conidia. Stud. Mycol. 2006, 56, 135–177. [Google Scholar] [CrossRef]
- Rifai, M.A. A revision of the genus Trichoderma. Mycol. Pap. 1969, 116, 1–56. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods And Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Komon, M.; Kubicek, C.P.; Druzhinina, I.S. Hypocrea voglmayrii sp. nov. from the Austrian Alps represents a new phylogenetic clade in Hypocrea/Trichoderma. Mycologia 2005, 97, 1365–1378. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef] [PubMed]
- Samuels, G.J.; Dodd, S.L.; Gams, W.; Castlebury, L.A.; Petrini, O. Trichoderma species associated with the green mold epidemic of commercially grown Agaricus bisporus. Mycologia 2002, 94, 146–170. [Google Scholar] [CrossRef] [PubMed]
- Montoya, Q.V.; Meirelles, L.A.; Chaverri, P.; Rodrigues, A. Unraveling Trichoderma species in the attine ant environment: Description of three new taxa. Antonie Van Leeuwenhoek 2016, 109, 633–651. [Google Scholar] [CrossRef]
- Chen, K.; Zhuang, W.Y. Discovery from a large-scaled survey of Trichoderma in soil of China. Sci. Rep. 2017, 7, 9090. [Google Scholar] [CrossRef]
- Chaverri, P.; Castlebury, L.A.; Overton, B.E.; Samuels, G.J. Hypocrea/Trichoderma: Species with conidiophore elongations and green conidia. Mycologia 2003, 95, 1100–1140. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Kubicek, C.P.; Druzhinina, I.S. Three European species of Hypocrea with reddish brown stromata and green ascospores. Mycologia 2008, 100, 796–815. [Google Scholar] [CrossRef]
- Chaverri, P.; Gazis, R.O.; Samuels, G.J. Trichoderma amazonicum, a new endophytic species on Hevea brasiliensis and H. guianensis from the Amazon basin. Mycologia 2011, 103, 139–151. [Google Scholar] [CrossRef]
- Hanada, R.E.; de Jorge Souza, T.; Pomella, A.W.V.; Hebbar, K.P.; Pereira, J.O.; Ismaiel, A.; Samuels, G.J. Trichoderma martiale sp. nov., a new endophyte from sapwood of Theobroma cacao with a potential for biological control. Mycol. Res. 2008, 112, 1335–1343. [Google Scholar] [CrossRef]
- Inglis, P.W.; Mello, S.C.M.; Martins, I.; Silva, J.B.T.; Macêdo, K.; Sifuentes, D.N.; Valadares-Inglis, M.C. Trichoderma from Brazilian garlic and onion crop soils and description of two new species: Trichoderma azevedoi and Trichoderma peberdyi. PLoS ONE 2020, 15, e0228485. [Google Scholar] [CrossRef]
- Chaverri, P.; Samuels, G.J. Hypocrea/Trichoderma (Ascomycota, Hypocreales, Hypocreaceae): Species with green ascospores. Stud. Mycol. 2003, 48, 1–116. [Google Scholar]
- Jaklitsch, W.M.; Lechat, C.; Voglmayr, H. The rise and fall of Sarawakus (Hypocreaceae, Ascomycota). Mycologia 2014, 106, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Chaverri, P.; Branco-Rocha, F.; Jaklitsch, W.; Gazis, R.; Degenkolb, T.; Samuels, G.J. Systematics of the Trichoderma harzianum species complex and the re-identification of commercial biocontrol strains. Mycologia 2015, 107, 558–590. [Google Scholar] [CrossRef] [PubMed]
- Gazis, R.; Rehner, S.; Chaverri, P. Species delimitation in fungal endophyte diversity studies and its implications in ecological and biogeographic inferences. Mol. Ecol. 2011, 20, 3001–3013. [Google Scholar] [CrossRef] [PubMed]
- Hoyos-Carvajal, L.; Orduz, S.; Bissett, J. Genetic and metabolic biodiversity of Trichoderma from Colombia and adjacent neotropic regions. Fungal Genet. Biol. 2009, 46, 615–631. [Google Scholar] [CrossRef]
- Kim, C.S.; Yu, S.H.; Nakagiri, A.; Shirouzu, T.; Sotome, K.; Kim, S.C.; Maekawa, N. Re-evaluation of Hypocrea pseudogelatinosa and H. pseudostraminea isolated from shiitake mushroom (Lentinula edodes) cultivation in Korea and Japan. Plant Pathol. J. 2012, 28, 341–356. [Google Scholar] [CrossRef]
- Samuels, G. Trichoderma: Systematics, the sexual state, and ecology. Phytopathology 2006, 96, 195–206. [Google Scholar] [CrossRef]
- Ospina-Giraldo, M.D.; Royse, D.J.; Chen, X.; Romaine, C.P. Molecular phylogenetic analyses of biological control strains of Trichoderma harzianum and other biotypes of Trichoderma spp. associated with mushroom green mold. Phytopathology 1999, 89, 308–313. [Google Scholar] [CrossRef]
- Qiu, Z.H.; Wu, X.L.; Zhang, J.X.; Huang, C.Y. High temperature enhances the ability of Trichoderma asperellum to infect Pleurotus ostreatus mycelia. PLoS ONE 2017, 12, e0187055. [Google Scholar] [CrossRef]
- Liu, X.M.; Wu, X.L.; Chen, Q.; Qiu, Z.H.; Zhang, J.X.; Huang, C.Y. Effects of heat stress on Pleurotus eryngii mycelial growth and its resistance to Trichoderma asperellum. Mycosystema 2017, 36, 1566–1574. [Google Scholar] [CrossRef]
- Jiang, H.; Zhang, L.; Zhang, J.Z.; Ojaghian, M.R.; Hyde, K.D. Antagonistic interaction between Trichoderma asperellum and Phytophthora capsici in vitro. J. Zhejiang Univ. Sci. B 2016, 17, 271–281. [Google Scholar] [CrossRef]
- Sun, J.Z.; Liu, X.Z.; Jeewon, R.; Li, Y.L.; Lin, C.G.; Tian, Q.; Zhao, Q.; Xiao, X.P.; Hyde, K.D.; Nilthong, S. Fifteen fungicolous Ascomycetes on edible and medicinal mushrooms in China and Thailand. Asian J. Mycol. 2019, 2, 129–169. [Google Scholar] [CrossRef]
- Rees, H.J.; Bashir, N.; Drakulic, J.; Cromey, M.G.; Bailey, A.M.; Foster, G.D. Identification of native endophytic Trichoderma spp. for investigation of in vitro antagonism towards Armillaria mellea using synthetic- and plant-based substrates. J. Appl. Microbiol. 2021, 131, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Kredics, L.; Kocsubé, S.; Nagy, L.; Komoń-Zelazowska, M.; Manczinger, L.; Sajben, E.; Nagy, A.; Vágvölgyi, C.; Kubicek, C.P.; Druzhinina, I.S.; et al. Molecular identification of Trichoderma species associated with Pleurotus ostreatus and natural substrates of the oyster mushroom. FEMS Microbiol. Lett. 2009, 300, 58–67. [Google Scholar] [CrossRef] [PubMed]
- Kosanović, D.; Potocnik, I.; Duduk, B.; Vukojevic, J.; Stajic, M.; Rekanovic, E.; Milijašević-Marčić, S. Trichoderma species on Agaricus bisporus farms in Serbia and their biocontrol. Ann. Appl. Biol. 2013, 163, 218–230. [Google Scholar] [CrossRef]
- Ma, X.L.; Fan, X.L.; Wang, G.Z.; Xu, R.P.; Yan, L.L.; Zhou, Y.; Gong, Y.H.; Xiao, Y.; Bian, Y.B. Enhanced expression of thaumatin-like protein gene (LeTLP1) endows resistance to Trichoderma atroviride in Lentinula edodes. Life 2021, 11, 863. [Google Scholar] [CrossRef]
- Lee, S.H.; Jung, H.J.; Hong, S.B.; Choi, J.I.; Ryu, J.S. Molecular markers for detecting a wide range of Trichoderma spp. that might potentially cause green mold in Pleurotus eryngii. Mycobiology 2020, 48, 313–320. [Google Scholar] [CrossRef]
- Úrbez-Torres, J.R.; Tomaselli, E.; Pollaed-Flamand, J.; Boule, J.; Gerin, D.; Pollastro, S. Characterization of Trichoderma isolates from southern Italy, and their potential biocontrol activity against grapevine trunk disease fungi. Phytopathol. Mediterr. 2020, 59, 425–439. [Google Scholar] [CrossRef]
- Dodd, S.L.; Lieckfeldt, E.; Samuels, G.J. Hypocrea atroviridis sp. nov., the teleomorph of Trichoderma atroviride. Mycologia 2003, 95, 27–40. [Google Scholar] [CrossRef]
- Smith, A.; Beltrán, C.A.; Kusunoki, M.; Cotes, A.M.; Motohashi, K.; Kondo, T.; Deguchi, M. Diversity of soil-dwelling Trichoderma in Colombia and their potential as biocontrol agents against the phytopathogenic fungus Sclerotinia sclerotiorum (Lib.) de Bary. J. Gen. Plant Pathol. 2013, 79, 74–85. [Google Scholar] [CrossRef]
- Choi, I.Y.; Choi, J.N.; Sharma, P.K.; Lee, W.H. Isolation and identification of mushroom pathogens from Agrocybe aegerita. Mycobiology 2010, 38, 310–315. [Google Scholar] [CrossRef] [PubMed]
- Samuels, G.J.; Ismaiel, A.; Mulaw, T.B.; Szakacs, G.; Druzhinina, I.S.; Kubicek, C.P.; Jaklitsch, W.M. The Longibrachiatum Clade of Trichoderma: A revision with new species. Fungal Divers. 2012, 55, 77–108. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.H. Research on Identification of Trichoderma of Mushrooms and Control of Trichoderma, Mycogone cervine. Master Thesis, Fujian Agriculture and Forestry University, Fuzhou, China, 2011. [Google Scholar]
- Yabuki, T.; Miyazaki, K.; Okuda, T. Japanese species of the Longibrachiatum Clade of Trichoderma. Mycoscience 2014, 55, 196–212. [Google Scholar] [CrossRef]
- Kim, J.Y.; Kwon, H.W.; Lee, D.H.; Ko, H.K.; Kim, S.H. Isolation and characterization of airborne mushroom damaging Trichoderma spp. from indoor air of cultivation houses used for oak wood mushroom production using sawdust media. Plant Pathol. J. 2019, 35, 674–683. [Google Scholar] [CrossRef] [PubMed]
- Tomah, A.A.; Abd Alamer, I.S.; Li, B.; Zhang, J.Z. A new species of Trichoderma and gliotoxin role: A new observation in enhancing biocontrol potential of T. virens against Phytophthora capsici on chili pepper. Biol. Control 2020, 145, 104261. [Google Scholar] [CrossRef]
- Samuels, G.J.; Suarez, C.; Solis, K.; Holmes, K.A.; Thomas, S.E.; Ismaiel, A.; Evans, H.C. Trichoderma theobromicola and T. paucisporum: Two new species isolated from cacao in South America. Mycol. Res. 2006, 110, 381–392. [Google Scholar] [CrossRef]
- Bissett, J. A revision of the genus Trichoderma. III. Section Pachybasium. Can. J. Bot. 1991, 69, 2373–2417. [Google Scholar] [CrossRef]
- Górski, R.; Sobieralski, K.; Siwulski, M.; Frąszczak, B.; Sas-Golak, I. The effect of Trichoderma isolates, from family mushroom growing farms, on the yield of four Agaricus bisporus (Lange) Imbach strains. J. Plant Prot. Res. 2014, 54, 102–105. [Google Scholar] [CrossRef][Green Version]
- Xu, Y.B.; Wen, C.J. Studies of contaminative fungi in the substituted cultivation of Pleurotus ostreatus and Lentinus edodes. Edible Fungi China 2004, 23, 45–47. [Google Scholar]
- Wang, G.Z.; Cao, X.T.; Ma, X.L.; Guo, M.P.; Liu, C.H.; Yan, L.L.; Bian, Y.B. Diversity and effect of Trichoderma spp. associated with green mold disease on Lentinula edodes in China. MicrobiologyOpen 2016, 5, 709–718. [Google Scholar] [CrossRef]
- Reper, C.; Penninckx, M.J. Inhibition of Pleurotus ostreatus growth and fructification by a diffusible toxin from Trichoderma hamatum. J. Food Sci. Technol. 1987, 20, 291–292. [Google Scholar]
- He, Z.D.; Sun, H.Q.; Gao, Y.F. Identification of Trichoderma species on mushrooms. J. Hebei Norm. Univ. Sci. Technol. 2008, 22, 41–45. [Google Scholar]
- Zhou, Y.; Wang, J.; Laying, Y.; Guo, L.J.; He, S.T.; Zhou, W.; Huang, J.S. Development of species/genus specific primers for identification of three Trichoderma species and for detection of Trichoderma genus. Research Square, 2021; preprint. [Google Scholar] [CrossRef]
- Cai, M.Z.; Idrees, M.; Zhou, Y.; Zhang, C.L.; Xu, J.Z. First report of green mold disease caused by Trichoderma hengshanicum on Ganoderma lingzhi. Mycobiology 2020, 48, 427–430. [Google Scholar] [CrossRef] [PubMed]
- Song, X.X.; Wang, Q.; Jun, J.X.; Zhang, J.J.; Chen, H.; Chen, M.J.; Huang, J.C.; Xie, B.Q. Study on accurate identification of four Trichoderma diseases in Agaricus bisporus in factory cultivation. Edible Fungi 2019, 41, 67–72. [Google Scholar]
- Zhu, Z.X.; Zhuang, W.Y. Three new species of Trichoderma with hyaline ascospores from China. Mycologia 2015, 107, 328–345. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Samuels, G.J.; Ismaiel, A.; Voglmayr, H. Disentangling the Trichoderma viridescens complex. Persoonia 2013, 31, 112–146. [Google Scholar] [CrossRef]
- Chen, X.Y.L.; Zhou, X.H.; Zhao, J.; Tang, X.L.; Pasquali, M.; Migheli, Q.; Berg, G.; Cernava, T. Occurrence of green mold disease on Dictyophora rubrovolvata caused by Trichoderma koningiopsis. J. Plant Pathol. 2021, 103, 981–984. [Google Scholar] [CrossRef]
- Samuels, G.J.; Ismaiel, A. Trichoderma evansii and T. lieckfeldtiae: Two new T. hamatum-like species. Mycologia 2009, 101, 142–156. [Google Scholar] [CrossRef]
- Choi, I.Y.; Joung, G.T.; Ryu, J.; Choi, J.S.; Choi, Y.G. Physiological characteristics of green mold (Trichoderma spp.) isolated from oyster mushroom (Pleurotus spp.). Mycobiology 2003, 31, 139–144. [Google Scholar] [CrossRef]
- Kim, C.S.; Shirouzu, T.; Nakagiri, A.; Sotome, K.; Nagasawa, E.; Maekawa, N. Trichoderma mienum sp. nov., isolated from mushroom farms in Japan. Antonie Van Leeuwenhoek 2012, 102, 629–641. [Google Scholar] [CrossRef]
- Błaszczyk, L.; Siwulski, M.; Sobieralski, K.; Frużyńska-Jóźwiak, D. Diversity of Trichoderma spp. causing Pleurotus green mould diseases in Central Europe. Folia Microbiol. 2013, 58, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.M.; Bak, W.C.; Lee, B.H.; Park, H.; Ka, K.H. Breeding and screening of Lentinula edodes strains resistant to Trichoderma spp. Mycobiology 2008, 36, 270–273. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Al-Rubaiey, W.; Al-Juboory, H.H. Molecular identification of Trichoderma longibrachiatum causing green mold in Pleurotus eryngii culture media. Plant Archives 2020, 20, 181–184. [Google Scholar] [CrossRef]
- Choi, I.Y.; Lee, W.; Choi, J.S. Forest green mold disease caused by Trichoderma pseudokoningii in winter mushroom, Flammulina velutipes. Korean J. Mycol. 1998, 26, 531–537. [Google Scholar]
- Druzhinina, I.S.; Kopchinskiy, A.G.; Komoń, M.; Bissett, J.; Szakacs, G.; Kubicek, C.P. An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet. Biol. 2005, 42, 813–828. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.S.; Shirouzu, T.; Nakagiri, A.; Sotome, K.; Maekawa, N. Trichoderma eijii and T. pseudolacteum, two new species from Japan. Mycol. Prog. 2012, 12, 739–753. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Stadler, M.; Voglmayr, H. Blue pigment in Hypocrea caerulescens sp. nov. and two additional new species in sect. Trichoderma. Mycologia 2012, 104, 925–941. [Google Scholar] [CrossRef]
- Park, M.S.; Oh, S.Y.; Cho, H.J.; Fong, J.J.; Cheon, W.J.; Lim, Y.W. Trichoderma songyi sp. nov., a new species associated with the pine mushroom (Tricholoma matsutake). Antonie Van Leeuwenhoek 2014, 106, 593–603. [Google Scholar] [CrossRef]
- Samuels, G.J.; Ismaiel, A.; de Souza, J.; Chaverri, P. Trichoderma stromaticum and its overseas relatives. Mycol. Prog. 2012, 11, 215–254. [Google Scholar] [CrossRef]
- Choi, I.Y.; Hong, S.B.; Yadav, M.C. Molecular and morphological characterization of green mold, Trichoderma spp. isolated from oyster mushrooms. Mycobiology 2003, 31, 74–80. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Põldmaa, K.; Samuels, G.J. Reconsideration of Protocrea (Hypocreales, Hypocreaceae). Mycologia 2008, 100, 962–984. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP*: Phylogenetic Analysis Using Parsimony (*and other methods); Version 4.0b10; Sinauer Associates: Sunderland, UK, 2002. [Google Scholar]
- Edler, D.; Klein, J.; Antonelli, A.; Silvestro, D. raxmlGUI 2.0: A graphical interface and toolkit for phylogenetic analyses using RAxML. Methods Ecol. Evol. 2021, 12, 373–377. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Nylander, J.A.A. MrModeltest v2. Program Distributed by the Author. Ph.D. Thesis, Uppsala University, Uppsala, Sweden, 2004. [Google Scholar]
- Harman, G.E.; Kubicek, C.P. Trichoderma and Gliocladium, Volume 2: Enzymes, Biological Control and Commercial Applications, 1st ed.; CRC Press: London, UK, 1998. [Google Scholar]
- Kubicek, C.P.; Mikus, M.; Schuster, A.; Schmoll, M.; Seiboth, B. Metabolic engineering strategies for the improvement of cellulase production by Hypocrea jecorina. Biotechnol. Biofuels 2009, 2, 19. [Google Scholar] [CrossRef] [PubMed]
- Chaverri, P.; Samuels, G.J. Evolution of habitat preference and nutrition mode in a cosmopolitan fungal genus with evidence of interkingdom host jumps and major shifts in ecology. Evolution 2013, 67, 2823–2837. [Google Scholar] [CrossRef] [PubMed]
- Samuels, G.J. Trichoderma: A review of biology and systematics of the genus. Mycol. Res. 1996, 100, 923–935. [Google Scholar] [CrossRef]
- Li, Q.R.; Tan, P.; Jiang, Y.L.; Hyde, K.D.; McKenzie, E.; Bahkali, A.; Kang, J.C.; Wang, Y. A novel Trichoderma species isolated from soil in Guizhou, T. guizhouense. Mycol. Prog. 2012, 12, 167–172. [Google Scholar] [CrossRef]
- Evans, H.C.; Holmes, K.A.; Thomas, S.E. Endophytes and mycoparasites associated with an indigenous forest tree, Theobroma gileri, in Ecuador and a preliminary assessment of their potential as biocontrol agents of cocoa diseases. Mycol. Prog. 2003, 2, 149–160. [Google Scholar] [CrossRef]
- Gazis, R.; Chaverri, P. Diversity of fungal endophytes in leaves and stems of wild rubber trees (Hevea brasiliensis) in Peru. Fungal Ecol. 2010, 3, 240–254. [Google Scholar] [CrossRef]
- Rossman, A.Y.; Samuels, G.J.; Rogerson, C.T.; Lowen, R. Genera of bionectriaceae, Hypocreaceae and Nectriaceae (Hypocreales, Ascomycetes). Stud. Mycol. 1999, 42, 1–248. [Google Scholar]
- Hatvani, L. Mushroom Pathogenic Trichoderma Species, Occurrence, Biodiversity, Diagnosis and Extracellular Enzyme Production. Ph.D. Thesis, University of Szeged, Szeged, Hungary, 2008. [Google Scholar]
- Castle, A.; Speranzini, D.; Rghei, N.; Alm, G.; Rinker, D.; Bissett, J. Morphological and molecular identification of Trichoderma isolates on North American mushroom farms. Appl. Environ. Microbiol. 1998, 64, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Park, M.S.; Bae, K.S.; Yu, S.H. Two new species of Trichoderma associated with green mold of oyster mushroom cultivation in Korea. Mycobiology 2006, 34, 111–113. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.T.; Gaze, R.H. Mushroom Pest and Disease Control: A Color Handbook, 1st ed.; CRC Press: London, UK, 2007. [Google Scholar]
- Mamoun, M.L.; Lapicco, R.; Savoie, J.M.; Olivier, J.M. Green mould disease in France: Trichoderma harzianum Th2 and other species causing damages on mushroom farms. In Proceedings of the 15th International Congress on the Science and Cultivation of Edible Fungi, Maastricht, The Netherlands, 15–19 May 2000. [Google Scholar]
- Kim, C.S.; Park, M.S.; Kim, S.C.; Maekawa, N.; Yu, S.H. Identification of Trichoderma, a competitor of shiitake mushroom (Lentinula edodes), and competition between Lentinula edodes and Trichoderma species in Korea. Plant Pathol. J. 2012, 28, 137–148. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Voglmayr, H. New combinations in Trichoderma (Hypocreaceae, Hypocreales). Mycotaxon 2013, 126, 143–156. [Google Scholar] [CrossRef]
- Sinden, J.W.; Hauser, E. Nature and control of three mildew diseases of mushrooms in America. Mushroom Sei. 1953, 2, 177–180. [Google Scholar]
- Elad, Y. Biocontrol of foliar pathogens: Mechanisms and application. Commun Agric. Appl. Biol. Sci. 2003, 68, 17–24. [Google Scholar]
- Mumpuni, A.; Sharma, H.S.S.; Brown, A.E. Effect of metabolites produced by Trichoderma harzianum biotypes and Agaricus bisporus on their respective growth radii in culture. Appl. Environ. Microbiol. 1998, 64, 5053–5056. [Google Scholar] [CrossRef]
- Neethling, D.; Nevalainen, H. Mycoparasitic species of Trichoderma produce lectins. Can. J. Microbiol. 1996, 42, 141–146. [Google Scholar] [CrossRef]
- Williams, J.; Clarkson, J.M.; Mills, P.R.; Cooper, R.M. Saprotrophic and mycoparasitic components of aggressiveness of Trichoderma harzianum groups toward the commercial mushroom Agaricus bisporus. Appl. Environ. Microbiol. 2003, 69, 4192–4199. [Google Scholar] [CrossRef]
- Abubaker, K.S.; Sjaarda, C.; Castle, A.J. Regulation of three genes encoding cell-wall-degrading enzymes of Trichoderma aggressivum during interaction with Agaricus bisporus. Can. J. Microbiol. 2013, 59, 417–424. [Google Scholar] [CrossRef]
- Weindling, R. Studies on a lethal principle effective in the parasitic action of Trichoderma lignorum on Rhizoctonia solani and other soil fungi. Phytopathology 1934, 24, 1153–1179. [Google Scholar]
- Castrillo, M.L.; Bich, G.A.; Zapata, P.D.; Villalba, L.L. Biocontrol of Leucoagaricus gongylophorus of leaf-cutting ants with the mycoparasitic agent Trichoderma koningiopsis. Mycosphere 2016, 7, 810–819. [Google Scholar] [CrossRef]
- Kubicek, C.P.; Herrera-Estrella, A.; Seidl-Seiboth, V.; Martínez, D.; Druzhinina, I.S.; Thon, M.R.; Zeilinger, S.; Casas-Flores, S.; Horwitz, B.A.; Mukherjee, P.K.; et al. Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. Genome Biol. 2011, 12, R40. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Luo, X. Trichoderma koningiopsis controls Fusarium oxysporum causing damping-off in Pinus massoniana seedlings by regulating active oxygen metabolism, osmotic potential, and the rhizosphere microbiome. Biol. Control 2020, 150, 104352. [Google Scholar] [CrossRef]
| Species | Strains | GenBank Accession Number | References | |
|---|---|---|---|---|
| TEF1 | RPB2 | |||
| T. afarasin | GJS 99-227 | AF348093 | — | [45] |
| T. afroharzianum | LESF229 | KT279013 | KT278945 | [46] |
| T. afroharzianum | GJS04-186 (T) | FJ463301 | FJ442691 | In GenBank |
| T. aggregatum | HMAS248864 | KY688063 | KY688002 | [47] |
| T. aggressivum | CBS100525 | AF534614 | AF545541 | [48] |
| T. aggressivum | DAOM222156 | AF348098 | FJ442752 | [45] |
| T. alni | CPK2494 | EU498313 | EU498350 | [49] |
| T. alni | CBS120633 = CPK1982 (T) | EU498312 | EU498349 | [49] |
| T. alpinum | HMAS248870 | KY688017 | KY687963 | [47] |
| T. alpinum | HMAS248821 (T) | KY688012 | KY687958 | [47] |
| T. amazonicum | IB95 | HM142377 | HM142368 | [50] |
| T. asperellum | CBS433.97 = TR3 (T) | AF456907 | EU248617 | [51] |
| T. atrobrunneum | S3 | KJ665376 | KJ665241 | [20] |
| T. atrobrunneum | GJS92-110 (T) | AF443942 | — | [16] |
| T. atrogelatinosum | CBS237.63 (T) | — | KJ842201 | In GenBank |
| T. azevedoi | CEN1403 | MK696638 | MK696800 | [52] |
| T. azevedoi | CEN1422 | MK696660 | MK696821 | [52] |
| T. bannaense | HMAS248865 | KY688038 | KY688003 | [47] |
| T. bannaense | HMAS248840 (T) | KY688037 | KY687979 | [47] |
| T. breve | HMAS248845 | KY688046 | KY687984 | [47] |
| T. breve | HMAS248844 (T) | KY688045 | KY687983 | [47] |
| T. brunneoviride | CBS121130 = CPK2014 | EU498316 | EU498357 | [49] |
| T. camerunense | GJS99-231 | AF348108 | — | [45] |
| T. camerunense | GJS99-230 (T) | AF348107 | — | [45] |
| T. catoptron | GJS02-76 = CBS114232 (T) | AY391963 | AY391900 | [53] |
| T. christiani | CBS132572 = S442 (T) | KJ665439 | KJ665244 | [20] |
| T. cinnamomeum | GJS97-237 (T) | AY391979 | AY391920 | [53] |
| T. compactum | CBS121218 | KF134798 | KF134789 | [54] |
| T. concentricum | HMAS248858 | KY688028 | KY687997 | [47] |
| T. concentricum | HMAS248833 (T) | KY688027 | KY687971 | [47] |
| T. endophyticum | DIS220J | FJ463330 | FJ442690 | [55] |
| T. endophyticum | DIS221E | FJ463316 | FJ442775 | In GenBank |
| T. epimyces | CPK1980 | EU498319 | EU498359 | [49] |
| T. epimyces | CBS120534 = CPK1981 (T) | EU498320 | EU498360 | [49] |
| T. ganodermatigerum | CCMJ5245 (T) | ON567195 | ON567189 | This study |
| T. ganodermatigerum | CCMJ5246 | ON567196 | ON567190 | This study |
| T. ganodermatigerum | CCMJ5247 | ON567197 | ON567191 | This study |
| T. ganodermatigerum | CCMJ5248 | ON567198 | ON567192 | This study |
| T. ganodermatigerum | CCMJ5249 | ON567199 | ON567193 | This study |
| T. ganodermatigerum | CCMJ5250 | ON567200 | ON567194 | This study |
| T. guizhouense | S278 | KF134799 | KF134791 | [54] |
| T. guizhouense | S628 | KJ665511 | KJ665273 | [20] |
| T. harzianum | GJS05-107 | FJ463329 | FJ442708 | In GenBank |
| T. harzianum | GJS04-71 | FJ463396 | FJ442779 | In GenBank |
| T. harzianum | Thaum12 | MT081433 | MT118248 | In GenBank |
| T. harzianum | CBS226.95 (T) | AF534621 | AF545549 | [48] |
| T. hausknechtii | Hypo649 = CBS133493 (T) | KJ665515 | KJ665276 | [20] |
| T. helicolixii | S640 = CBS133499 (T) | KJ665517 | KJ665278 | [20] |
| T. hengshanicum | HMAS248853 | KY688055 | KY687992 | [47] |
| T. hengshanicum | HMAS248852 (T) | KY688054 | KY687991 | [47] |
| T. hirsutum | HMAS248859 | KY688030 | KY687998 | [47] |
| T. hirsutum | HMAS248834 (T) | KY688029 | KY687972 | [47] |
| T. ingratum | HMAS248824 | KY688019 | KY687964 | [47] |
| T. ingratum | HMAS248873 | KY688022 | KY688010 | [47] |
| T. ingratum | HMAS248822 (T) | KY688018 | KY687973 | [47] |
| T. inhamatum | CBS273.78 (T) | AF348099 | FJ442725 | [45] |
| T. italicum | S131 = CBS132567 (T) | KJ665525 | KJ665282 | [20] |
| T. lentiforme | DIS167C | FJ463309 | FJ442689 | In GenBank |
| T. lentiforme | GJS98-6 (T) | AF469195 | — | [16] |
| T. liberatum | HMAS248832 | KY688026 | KY687970 | [47] |
| T. liberatum | HMAS248831 (T) | KY688025 | KY687969 | [47] |
| T. linzhiense | HMAS248874 | KY688048 | KY688011 | [47] |
| T. linzhiense | HMAS248846 (T) | KY688047 | KY687985 | [47] |
| T. lixii | CBS110080 = GJS97-96 | FJ716622 | KJ665290 | [20] |
| T. neocrassum | DAOM164916 = CBS336.93 (T) | AF534615 | AF545542 | [48] |
| T. neotropicale | LA11 | HQ022771 | — | [56] |
| T. peberdyi | CEN1387 | MK696619 | MK696781 | [52] |
| T. peberdyi | CEN1388 | MK696620 | MK696782 | [52] |
| T. pleuroticola | T1295 | EU279973 | — | [57] |
| T. pleuroticola | CBS124383 (T) | HM142381 | HM142371 | [50] |
| T. pleuroti | CBS124387 (T) | HM142382 | HM142372 | [50] |
| T. polypori | HMAS248855 | KY688058 | KY687994 | [47] |
| T. polypori | HMAS248861 | KY688059 | KY688000 | [47] |
| T. priscilae | S129 | KJ665689 | KJ665332 | [20] |
| T. pseudodensum | HMAS248829 | KY688024 | KY687968 | [47] |
| T. pseudodensum | HMAS248828 (T) | KY688023 | KY687967 | [47] |
| T. pseudogelatinosum | TUFC60186 (T) | JQ797397 | JQ797405 | [58] |
| T. pyramidale | S573 | KJ665698 | — | [20] |
| T. pyramidale | S73 = CBS135574 (T) | KJ665699 | KJ665334 | [20] |
| T. rifaii | DIS337F | FJ463321 | FJ442720 | In GenBank |
| T. rifaii | DIS355B (T) | FJ463324 | — | In GenBank |
| T. simmonsii | GJS90-22 | AY391984 | AY391925 | [53] |
| T. simmonsii | GJS92-100 | AF443937 | FJ442710 | [16] |
| T. simmonsii | GJS91-138 | AF443935 | FJ442757 | [16] |
| T. simplex | HMAS248860 | KY688042 | KY687999 | [47] |
| T. simplex | HMAS248842 (T) | KY688041 | KY687981 | [47] |
| T. solum | HMAS248848 | KY688050 | KY687987 | [47] |
| T. solum | HMAS248847 (T) | KY688049 | KY687986 | [47] |
| T. spirale | DAOM183974 | EU280049 | — | [57] |
| T. spirale | LESF107 | KT279022 | KT278956 | [46] |
| T. stramineum | GJS02-84 = CBS114248 (T) | AY391999 | AY391945 | [53] |
| T. tawa | GJS97-174 = CBS114233 (T) | AY392004 | AY391956 | [53] |
| T. tomentosum | S33 | KF134801 | KF134793 | [54] |
| T. tomentosum | DAOM178713A (T) | AF534630 | AF545557 | [48] |
| T. velutinum | DAOM230013 = CPK298 | AY937415 | KF134794 | [59] |
| T. virens | DIS162 | FJ463367 | FJ442696 | In GenBank |
| T. zayuense | HMAS248836 | KY688032 | KY687975 | [47] |
| T. zayuense | HMAS248835 (T) | KY688031 | KY687974 | [47] |
| Species | Host Range | Isolates | GenBank Accession Number | References | |
|---|---|---|---|---|---|
| TEF1 | RPB2 | ||||
| T. aggressivum | Agaricus bisporus | CBS100525 | AF534614 | AF545541 | [48] |
| T. aggressivum f. aggressivum | Agaricus bisporus | GJS99-30 | AF348109 | — | [60] |
| DAOM222156 | AF348098 | FJ442752 | [45] | ||
| T. aggressivum f. europaeum | Agaricus bisporus | CBS100526 (T) | KP008993 | KP009166 | [45] |
| — | TRS27 | KP008994 | KP009163 | In GenBank | |
| — | CBS435.95 | KP008998 | KP009169 | In GenBank | |
| T. alni | Macrotyphula cf. contorta | CBS120633 | EU498312 | EU498349 | [49] |
| CPK2494 | EU498313 | EU498350 | |||
| T. asperellum | Pleurotus ostreatus | T11 (ACCC32725) | MF049065 | — | [61] |
| Pleurotus eryngii | — | — | — | [62] | |
| — | CGMCC6422 | KF425756 | KF425755 | [63] | |
| — | CBS433.97 = TR3 (T) | AF456907 | EU248617 | In GenBank | |
| T. atrobrunneum | Ganoderma sichuanense | CGMCC3.19070 | MH464779 | — | [64] |
| — | T17-27 | MW232537 | MW232508 | [65] | |
| T. atroviride | Pleurotus ostreatus | CPK3277 | EU918154 | — | [66] |
| Ganoderma sichuanense | 2015005 | — | — | [10] | |
| Agaricus bisporus | T33 | — | — | [67] | |
| Lentinula edodes | T25 | — | — | [68] | |
| Pleurotus eryngii | — | — | — | [69] | |
| — | PARC1011 | MT454114 | MT454130 | [70] | |
| — | PARC1018 | MT454121 | MT454137 | ||
| — | DAOM222144 | AF456889 | FJ442754 | [71] | |
| — | Th002 | AB558906 | AB558915 | [72] | |
| T. aureoviride | Pleurotus ostreatus | HMAS266607 | KF923280 | KF923306 | [73] |
| T. austriacum | Peziza sp. | CBS122494 (T) | FJ860619 | FJ860525 | [19] |
| T. capillare | Agaricus bisporus | CPK2883 | JN182283 | JN182312 | [74] |
| GJS99-3 | JN175584 | JN175529 | |||
| T. catoptron | Aphyllophorales s. l. | GJS02-76 (T) | AY391963 | AY391900 | [53] |
| T. cerinum | Lentinula edodes | S357 | KF134797 | KF134788 | [75] |
| T. chromospermum | black mycelium and black pyrenomycete | GJS95-196 | AY391975 | AY391914 | [53] |
| GJS98-73 | AY391976 | AY391915 | |||
| GJS94-68 = CBS114577 | — | AY391913 | |||
| — | HMAS252537 | KF729986 | KF730004 | [25] | |
| — | HMAS252539 | KF923287 | KF923314 | ||
| — | HMAS252535 | KF923292 | KF923315 | ||
| T. citrinoviride | Lentinula edodes | TAMA0154 | AB807641 | AB807653 | [76] |
| Pleurotus ostreatus | GJS92-8 | JN175595 | JN175544 | [77] | |
| Pleurotus eryngii | GJS01-364 | AY225860 | AF545565 | [69] | |
| Polypore mushroom | TAMA0188 | AB807644 | AB807656 | [76] | |
| — | HZA9 | MK850831 | MK962804 | [78] | |
| T. epimyces | Polyporus umbellatus | CPK1980 | EU498319 | EU498359 | [49] |
| CBS120534 (T) | EU498320 | EU498360 | |||
| T. erinaceum | — | DIS7 | DQ109547 | EU248604 | [79] |
| T. fasciculatum | Hypocrea ascospores | CBS118.72 | — | — | [80] |
| — | DAOM172827 | AF534628 | AF545555 | [48] | |
| T. fomiticola | Fomes fomentarius | CBS121136 | FJ860639 | FJ860538 | [18] |
| CPK3137 | FJ860640 | FJ860539 | |||
| T. ghanense | Agaricus bisporus | NBRC30902 | AB807638 | AB807650 | [76] |
| T. ganodermatis | Ganoderma sichuanense | HMAS248856 | KY688060 | KY687995 | [47] |
| HMAS248869 | KY688061 | KY688007 | [47] | ||
| T. ganodermatigerum | Ganoderma sichuanense | CCMJ5245(T) | ON567195 | ON567189 | This study |
| CCMJ5246 | ON567196 | ON567190 | |||
| CCMJ5247 | ON567197 | ON567191 | |||
| CCMJ5248 | ON567198 | ON567192 | |||
| CCMJ5249 | ON567199 | ON567193 | |||
| CCMJ5250 | ON567200 | ON567194 | |||
| T. ghanense | Agaricus bisporus | NBRC30902 | AB807638 | — | [76] |
| T. hamatum | Agaricus bisporus | Tham20-3 | — | — | [81] |
| Lentinula edodes | — | — | — | [82] | |
| — | DAOM167057 (T) | EU279965 | AF545548 | [57] | |
| — | Hypo647 = WU31629 | KJ665513 | KJ665274 | [20] | |
| — | Hypo648 = CBS132565 | KJ665514 | KJ665275 | [20] | |
| T. harzianum | Pleurotus ostreatus | KACC40558 | — | — | [66] |
| Cyclocybe aegerita | JB1 | — | — | [73] | |
| Lentinula edodes | T50 | — | — | [83] | |
| Pleurotus eryngii | KACC40784 | — | — | [69] | |
| Pleurotus ostreatus | |||||
| Agaricus bisporus | — | — | — | [45] | |
| Pleurotus ostreatus | — | — | — | [84] | |
| Polypores/Corticiaceous | — | — | — | [18] | |
| Pleurotus tuoliensis | — | — | — | [85] | |
| Tremella fuciformis | |||||
| Flammulina filiformis | |||||
| — | CBS226.95 | AF348101 | AF545549 | [48] | |
| — | Thaum12 | MT081433 | MT118248 | [86] | |
| — | CBS227.95 | AF348100 | — | [45] | |
| — | GJS05-107 | FJ463329 | FJ442708 | In GenBank | |
| — | GJS04-71 | FJ463396 | FJ442779 | In GenBank | |
| T. hengshanicum | Ganoderma sichuanense | 1009 | — | — | [87] |
| — | HMAS248852 (T) | KY688054 | KY687991 | [47] | |
| — | HMAS248853 | KY688055 | KY687992 | ||
| T. inhamatum | Agaricus bisporus | CBS273.78 (T) | AF348099 | FJ442725 | [81] |
| Pleurotus tuoliensis | — | — | — | [85] | |
| T. koningii | Pleurotus eryngii | — | — | — | [69] |
| Agaricus bisporus | — | — | — | [88] | |
| Lentinula edodes | — | — | — | [85] | |
| Pleurotus ostreatus | |||||
| Pleurotus tuoliensis | |||||
| Flammulina filiformis | |||||
| Volvariella volvacea | |||||
| Hypsizygus marmoreus | |||||
| Ganoderma sichuanense | TFl040917 | — | — | [75] | |
| Tremella fuciformis | TGy040604 | — | — | ||
| — | 7723 | KJ634753 | KJ634720 | [89] | |
| — | GJS90-18 | DQ289007 | EU248600 | [23] | |
| — | CBS979.70 | AY665703 | EU248601 | In GenBank | |
| — | S22 | KC285595 | KC285749 | [90] | |
| T. koningiopsis | Phaiius rubrovolvata | CXYL | MN135988 | MT038997 | [91] |
| Ganoderma sichuanense | CCMJ5253 | ON567187 | ON567201 | This study | |
| CCMJ5254 | ON567188 | ON567202 | |||
| T. kunigamense | Lentinula edodes | TAMA193 | AB807645 | AB807657 | [76] |
| T. leguminosarum | dark corticiaceous fungus | S391 | KJ665548 | KJ665287 | [20] |
| CBS130014 | KJ665551 | KJ665288 | |||
| S503 | KJ665552 | KJ665289 | |||
| T. lieckfeldtiae | Moniliophthora roreri | GJS00-14 = CBS123049 (T) | EU856326 | EU883562 | [92] |
| T. longibrachiatum | Pleurotus ostreatus | TUFC61535 = CBS816.68(T) | EU401591 | DQ087242 | [40] |
| Agrocybe aegerita | JB4 | — | — | [73] | |
| Lentinula edodes | T57 | — | — | [83] | |
| Ganoderma sichuanense | TFl040921 | — | — | [75] | |
| Pleurotus eryngii | — | — | — | [93] | |
| Agaricus bisporus | — | — | — | [81] | |
| Pleurotus tuoliensis | — | — | — | [85] | |
| Hypsizygus marmoreus | |||||
| Volvariella volvacea | |||||
| T. mienum | Lentinula edodes | TUFC61517 | JQ621975 | JQ621965 | [94] |
| T. orientale | Ganoderma applanatum | LESF516 | KT279041 | KT278976 | [46] |
| Ganoderma applanatum | LESF540 | KT279042 | KT278977 | ||
| Ganoderma applanatum | LESF544 | KT279043 | KT278978 | ||
| Ganoderma applanatum | TRS707 | KP008888 | KP009202 | ||
| T. oblongisporum | Lentinula edodes | T37 | — | — | [83] |
| — | DAOM167085 | AF534623 | AF545551 | [48] | |
| T. parareesei | Pleurotus eryngii | TAMA0153 | AB807640 | AB807652 | [76] |
| T. parestonica | Hymenochaete tabacina | CBS120636 (T) | FJ860667 | FJ860565 | [18] |
| T. pleuroticola | Pleurotus ostreatus | CBS124383 (T) | HM142381 | HM142371 | [66] |
| CPK2885 | EU918161 | EU918141 | |||
| Pleurotus eryngii | CAF-TP3 | — | — | [69] | |
| Lentinula edodes | T22 | — | — | [83] | |
| Cyclocybe aegerita | JB7 | — | — | [73] | |
| — | T1295 | EU279973 | — | [57] | |
| T. pleuroti | Pleurotus ostreatus | KACC44537 | — | — | [69] |
| Pleurotus eryngii var. ferulae | — | — | — | [95] | |
| — | CBS124387 (T) | HM142382 | HM142372 | [50] | |
| T. polypori | Lentinula edodes | HMAS248861 | KY688059 | KY688000 | [47] |
| Polyporus sp. | HMAS248855 (T) | KY688058 | KY687994 | ||
| T. polysporum | Lentinula edodes | — | — | — | [96] |
| — | 8232 | KJ634779 | KJ634746 | [89] | |
| — | 8147 | KJ634771 | KJ634738 | ||
| T. priscilae | Crepidotus sp. | S168 = CBS131487 (T) | KJ665691 | KJ665333 | [20] |
| Stereum sp. | S129 | KJ665689 | KJ665332 | ||
| — | HMAS245002 | KT343760 | KT343764 | In GenBank | |
| T. protopulvinatum | Fomitopsis pinicola | CPK2434 | FJ860677 | FJ860574 | [18] |
| T. pulvinatum | Fomitopsis pinicola | CBS121279 | FJ860683 | FJ860577 | [18] |
| T. pseudokoningii | Lentinula edodes | DUCC4021 | KX431217 | — | [77] |
| Cyclocybe aegerita | TGc050619 | — | — | [75] | |
| Ganoderma sichuanense | TFl040926 | — | — | ||
| Pleurotus eryngii | — | — | — | [97] | |
| Flammulina filiformis | — | — | — | [98] | |
| Pleurotus tuoliensis | — | — | — | [85] | |
| Volvariella volvacea | |||||
| Hypsizygus marmoreus | |||||
| — | DAOM167678 | AY865641 | KJ842214 | [99] | |
| — | GJS99-149 | JN175589 | JN175536 | [17] | |
| — | GJSNS19 | JN175588 | JN175535 | ||
| T. pseudolacteum | Lentinula edodes | TUFC61496 | JX238494 | JX238479 | [100] |
| TUFC61502 | JX238480 | JX238471 | |||
| T. samuelsii | Hymenochaete sp. | S5 = CBS130537 | JN715651 | JN715599 | [101] |
| S42 | JN715652 | JN715598 | |||
| T. songyi | Tricholoma matsutake | TC556 | KX266244 | KX266250 | [102] |
| TC480 | KX266243 | KX266249 | |||
| T. stilbohypoxyli | Stilbohypoxylon moelleri | Hypo256 = CPK1977 | FJ860702 | FJ860592 | [23] |
| T. stromaticum | Agaricus bisporus | GJS97-181 | AY937447 | HQ342227 | [59] |
| — | GJS07-88 | HQ342195 | HQ342258 | [103] | |
| — | GJS03-47 | HQ342201 | HQ342264 | ||
| — | GJS00-107 | HQ342202 | HQ342265 | ||
| T. sulphureum | Laetiporus sulphureus | CBS119929 | FJ860710 | FJ179620 | [18] |
| CPK1593 | FJ860709 | FJ860599 | |||
| Thelephora sp. | GJS95-135 = CBS114237 | AY392006 | AY391958 | [53] | |
| T. tsugarense | Lentinula edodes | TAMA203 (T) | AB807647 | AB807659 | [76] |
| T. viride | Lentinula edodes | T13 | — | — | [83] |
| Pleurotus ostreatus | — | — | — | [82] | |
| Tremella fuciformis | TGc040905 | — | — | [75] | |
| Ganoderma sichuanense | TFl080706 | — | — | [75] | |
| Flammulina filiformis | TFj10010 | — | — | [75] | |
| Cyclocybe aegerita | TGc040905 | — | — | [75] | |
| Phallus indusiatus | TFl080706 | — | — | [75] | |
| Tremella fuciformis | TGc040905 | — | — | [75] | |
| Agaricus bisporus | — | — | — | [88] | |
| Pleurotus eryngii | — | — | [69] | ||
| — | TRS575 | KP008931 | KP009081 | In GenBank | |
| — | LESF115 | KT278989 | KT278921 | [46] | |
| T. virens | Agaricus bisporus | — | — | — | [88] |
| Pleurotus eryngii | — | — | — | ||
| — | DIS162 | FJ463367 | FJ442696 | In GenBank | |
| — | DIS328A | FJ463363 | FJ442738 | In GenBank | |
| T. cf. virens | Pleurotus eryngii | KACC40783 | — | — | [69] |
| Pleurotus ostreatus | TUCIM2558 | KX655776 | — | [104] | |
| T. viridarium | Steccherinum ochraceum | GJS89-142 | AY376049 | EU241495 | [51] |
| Nemania sp. | GJS98-182 | DQ307511 | EU252011 | [23] | |
| Protocrea farinosa | — | CBS121551 | EU703889 | EU703935 | [105] |
| Protocrea pallida | — | CBS121552 | EU703897 | EU703944 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
An, X.-Y.; Cheng, G.-H.; Gao, H.-X.; Li, X.-F.; Yang, Y.; Li, D.; Li, Y. Phylogenetic Analysis of Trichoderma Species Associated with Green Mold Disease on Mushrooms and Two New Pathogens on Ganoderma sichuanense. J. Fungi 2022, 8, 704. https://doi.org/10.3390/jof8070704
An X-Y, Cheng G-H, Gao H-X, Li X-F, Yang Y, Li D, Li Y. Phylogenetic Analysis of Trichoderma Species Associated with Green Mold Disease on Mushrooms and Two New Pathogens on Ganoderma sichuanense. Journal of Fungi. 2022; 8(7):704. https://doi.org/10.3390/jof8070704
Chicago/Turabian StyleAn, Xiao-Ya, Guo-Hui Cheng, Han-Xing Gao, Xue-Fei Li, Yang Yang, Dan Li, and Yu Li. 2022. "Phylogenetic Analysis of Trichoderma Species Associated with Green Mold Disease on Mushrooms and Two New Pathogens on Ganoderma sichuanense" Journal of Fungi 8, no. 7: 704. https://doi.org/10.3390/jof8070704
APA StyleAn, X.-Y., Cheng, G.-H., Gao, H.-X., Li, X.-F., Yang, Y., Li, D., & Li, Y. (2022). Phylogenetic Analysis of Trichoderma Species Associated with Green Mold Disease on Mushrooms and Two New Pathogens on Ganoderma sichuanense. Journal of Fungi, 8(7), 704. https://doi.org/10.3390/jof8070704







