Coniochaeta massiliensis sp. nov. Isolated from a Clinical Sampl28
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Strains
2.2. Macroscopic Characterisation
2.3. Microscopic Characterisation
2.4. MALDI-TOF MS Identification
2.5. Antifungal Susceptibility Testing (AFST)
2.6. Physiological Analyses
2.6.1. Energy-Dispersive X-ray Spectroscopy (EDX) Analysis
2.6.2. BiologTM Phenotypic Analysis
2.7. DNA Extraction and Sequencing
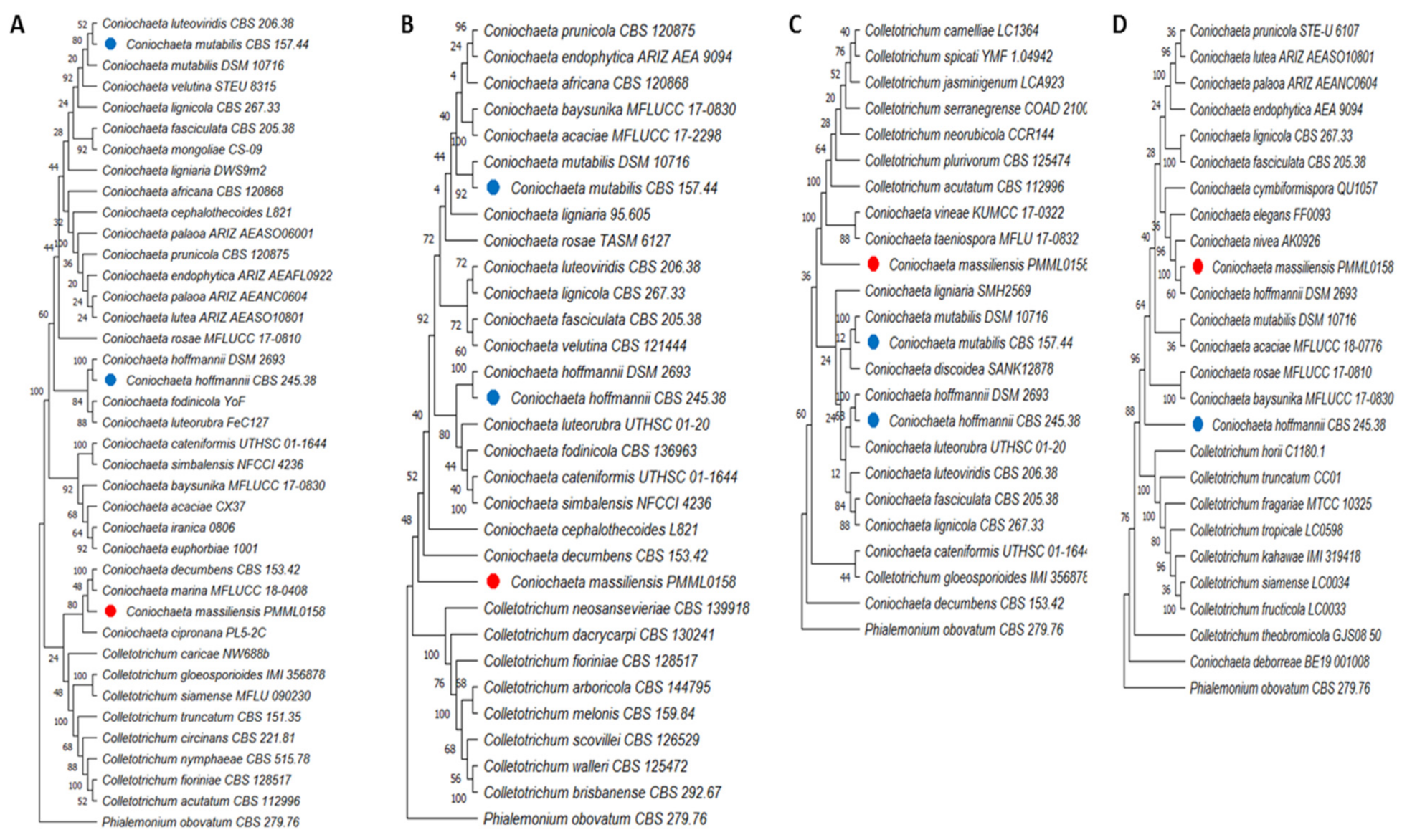
2.8. Phylogenetic Analyses
3. Results
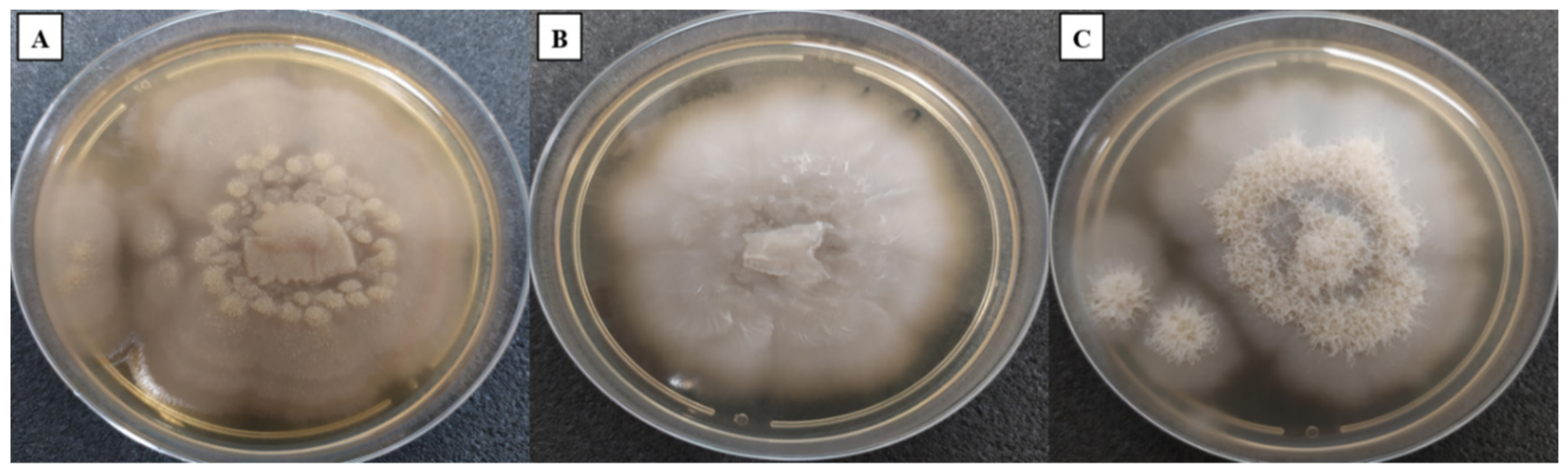
3.1. Macroscopic Characterisation
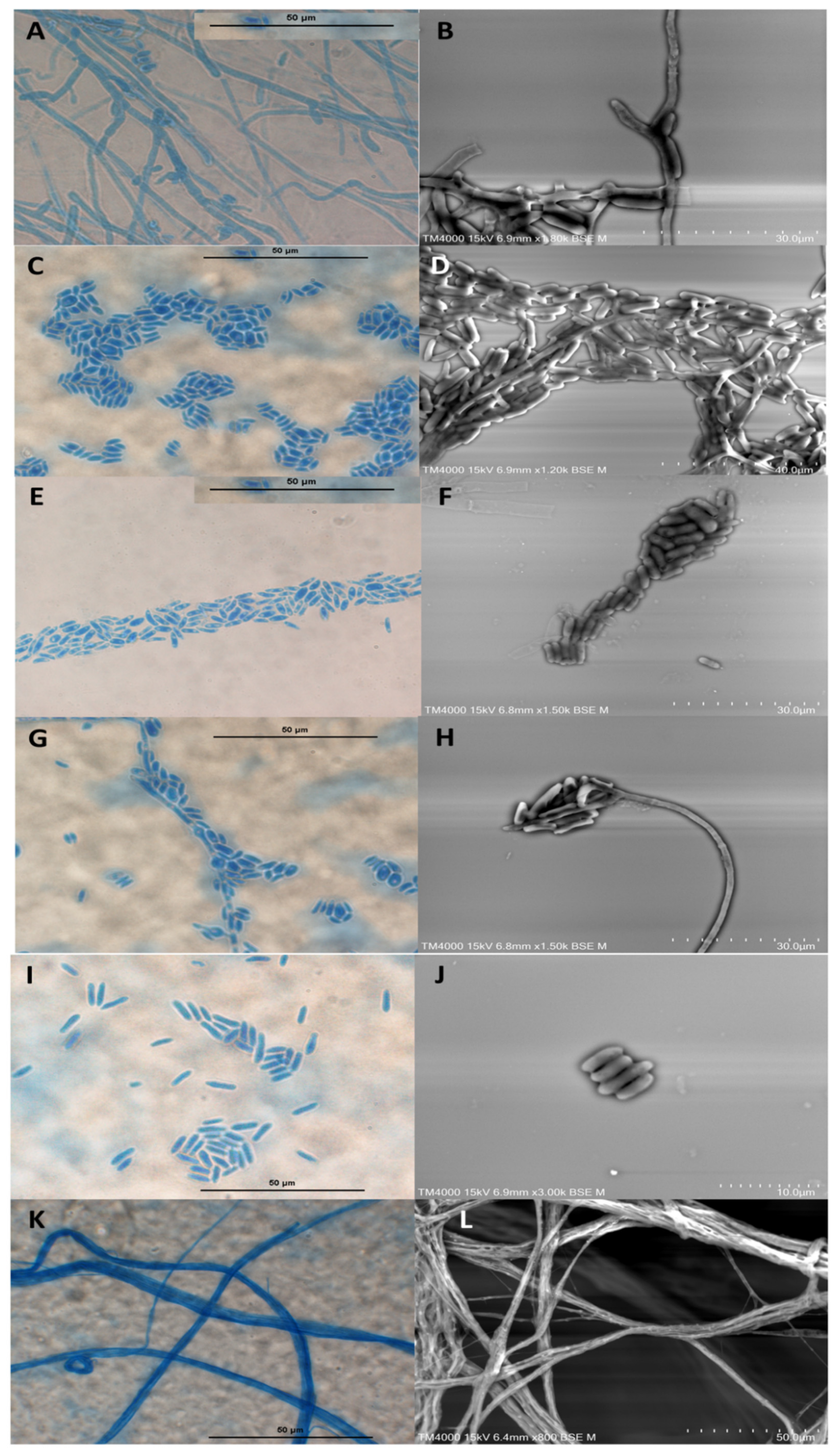
3.2. Microscopic Characterisation
3.3. Antifungal Susceptibility Testing (AFST)
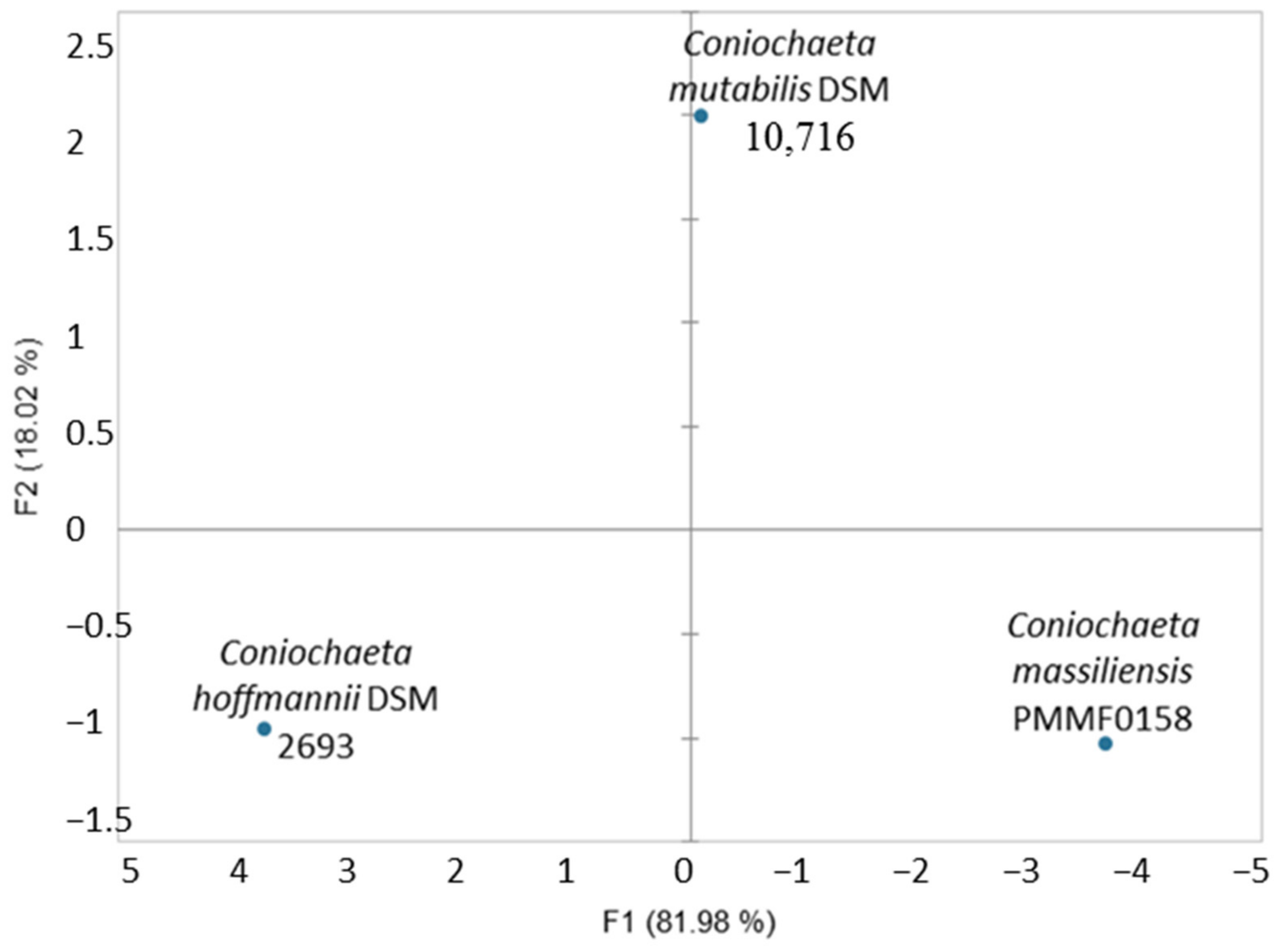
3.4. MALDI-TOF MS Identification
3.5. Physiological Analysis
3.5.1. EDX (Energy-Dispersive X-ray Spectroscopy)
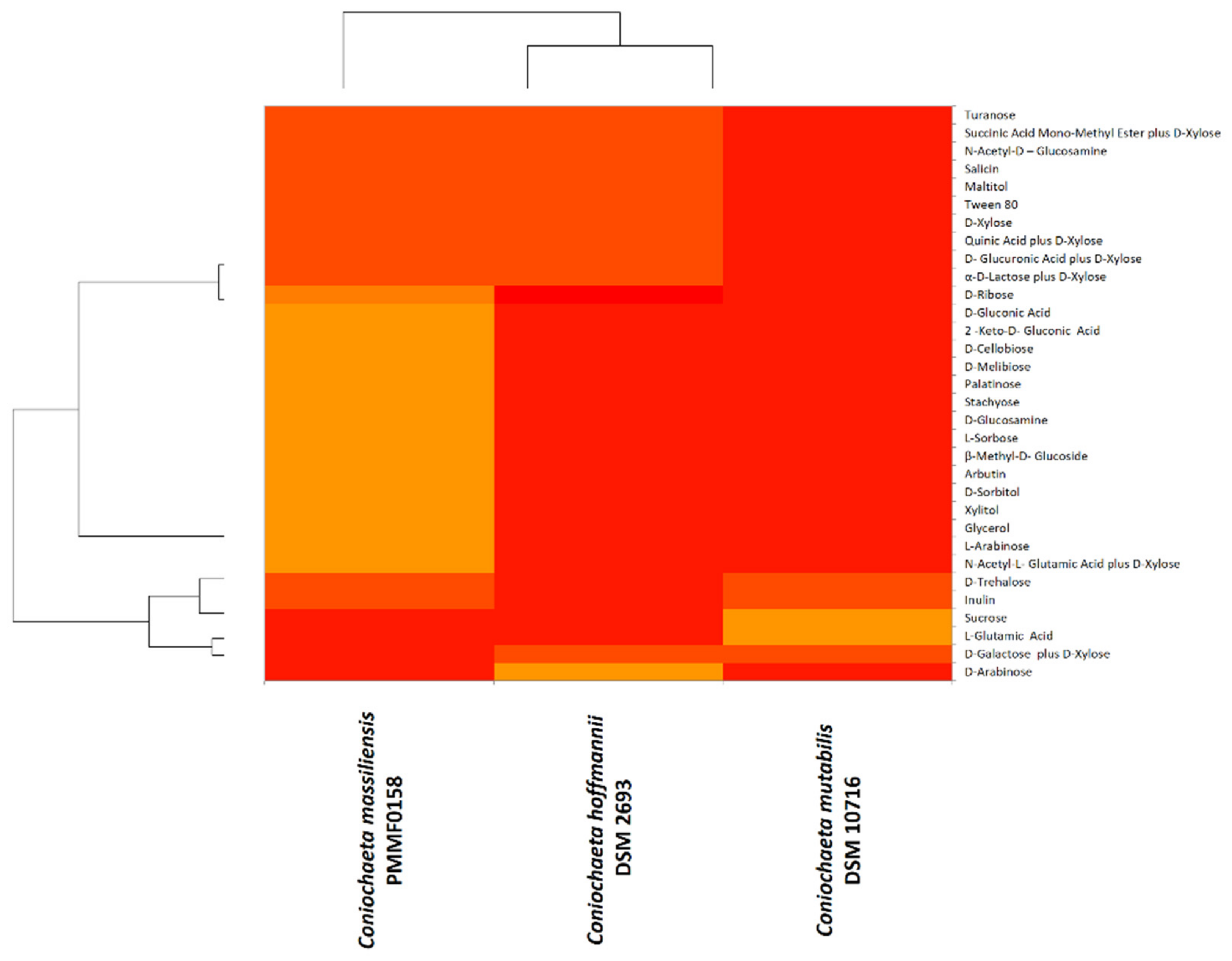
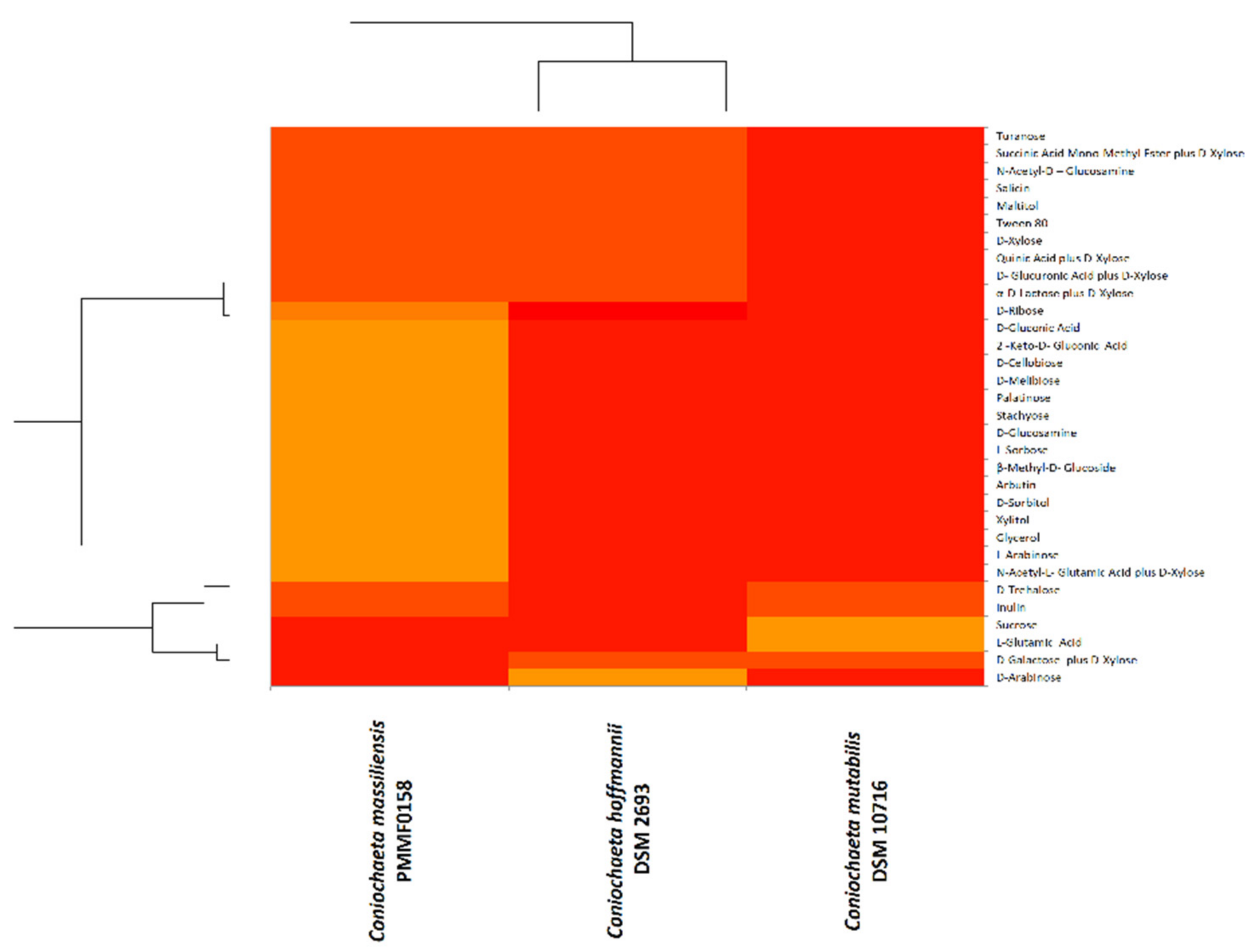
3.5.2. BiologTM System
3.6. DNA Sequencing
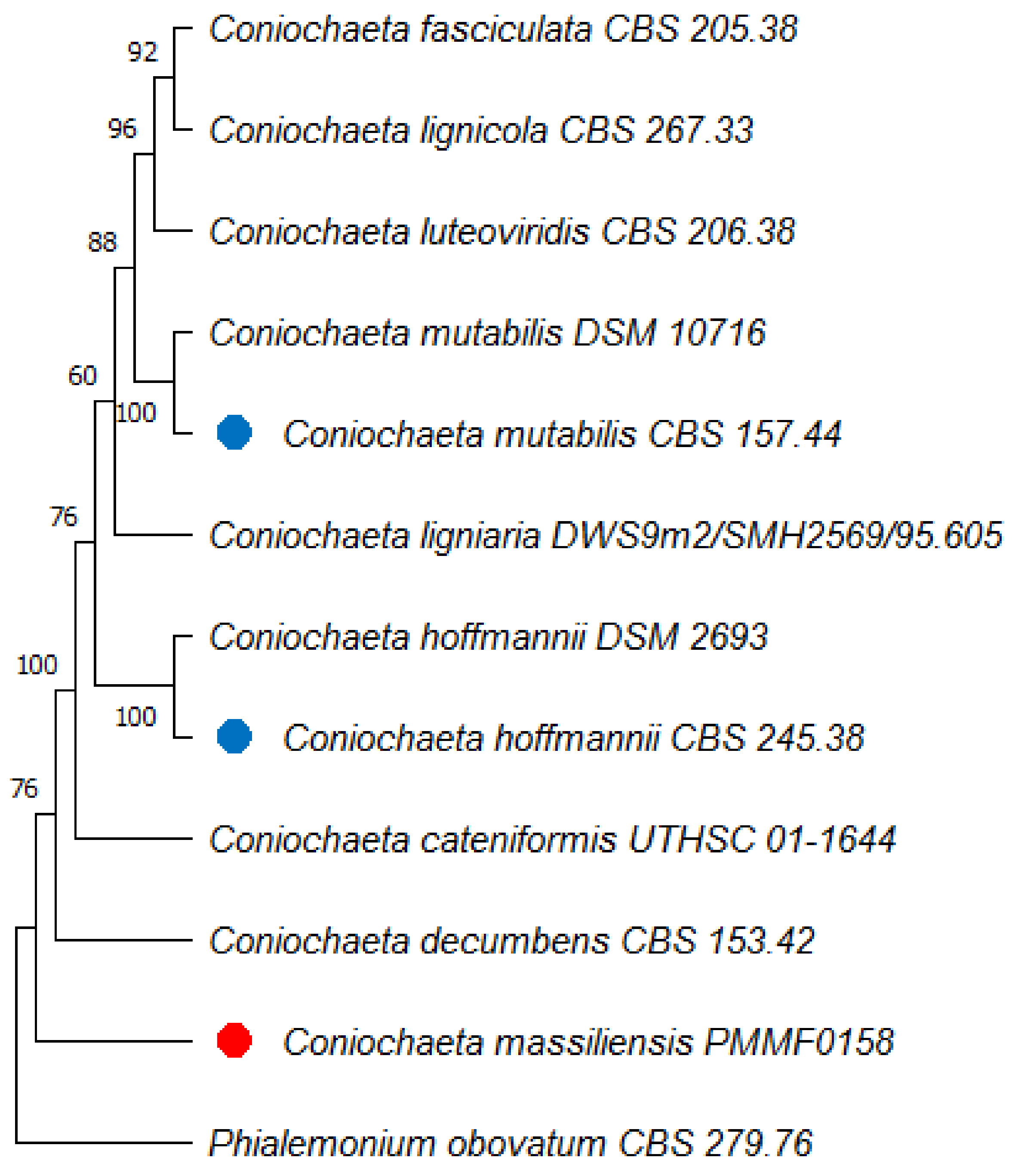
3.7. Phylogenetic Analyses
3.8. Taxonomy
Host: Human
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 5-FC | 5-fluorocytosine |
| AMB | Amphotericin B |
| AND | Anidulafungin |
| CAS | Caspofungin |
| DNA | Deoxyribonucleic acid |
| EDX | Energy-dispersive X-ray spectroscopy |
| FL | Fluconazole |
| GC | Gentamycin and chloramphenicol |
| IHU Méditerranée Infection | Institut Hospitalo-Universitaire Méditerranée Infection |
| ITC | Itraconazole |
| ITS | Internal transcribed spacers of the rRNA |
| SDA | Sabouraud Dextrose Agar |
| LSU | Large subunit of rRNA |
| MALDI-TOF MS | Matrix-assisted laser desorption/ionization–time of flight mass spectrometry |
| MICs | Minimum inhibitory concentrations |
| MIF | Micafungin |
| NA | Not available |
| POS | Posaconazole |
| rRNA | Ribosomal ribonucleic acid |
| SSU | Small subunit of rRNA |
| TEF-1α | Partial translation elongation factor 1-alpha gene |
| B-tub2 | Partial β-tubulin 2 gene |
| VOR | Voriconazole |
References
- Weber, E.G. The Lecythophora-Coniochaeta Complex: I. Morphological Studies on Lecythophora Species Isolated from Picea Abies. Nova Hedwig. 2002, 74, 85–159. [Google Scholar] [CrossRef]
- Nosanchuk, J.D.; Casadevall, A. Impact of Melanin on Microbial Virulence and Clinical Resistance to Antimicrobial Compounds. Antimicrob. Agents Chemother. 2006, 50, 28–3519. [Google Scholar] [CrossRef] [PubMed]
- Guarro, J.; Gené, J.; Ai-Bader, S.M.; Abdullah, S.K. A New Species of Coniochaetidium from Soil. Mycoscience 1997, 38, 25–123. [Google Scholar] [CrossRef]
- Kamiya, S.; Uchiyama, S.; Udagawa, S. Two New Species of Coniochaeta with a Cephaiothecoid Peridium Wall. Mycoscience 1995, 36, 83–377. [Google Scholar] [CrossRef]
- Yuan, Z.Q.; Mohamed, C. New species and new records of Ascomycetes of stems of Eucalyptus from Tasmania, Australia. Mycotaxon 1997, 63, 9–23. [Google Scholar]
- Mahoney, D.P.; Lafavre, J.S. Coniochaeta extramundana, with a synopsis of other Coniochaeta species. Mycologia 1981, 73, 931–952. [Google Scholar] [CrossRef]
- Crane, J.L.; Shearer, C.A. A new Coniochaeta from fresh water. Mycotaxon 1995, 54, 107–110. [Google Scholar]
- Perdomo, H.D.; García, D.; Gené, J.; Cano, J.; Sutton, D.A.; Summerbell, R.C.; Guarro, J. Phialemoniopsis, a New Genus of Sordariomycetes, and New Species of Phialemonium and Lecythophora. Mycologia 2013, 105, 398–421. [Google Scholar] [CrossRef]
- Khan, Z.; Gené, J.; Ahmad, S.; Cano, J.; Al-Sweih, N.; Joseph, L.; Chandy, R.; Guarro, J. Coniochaeta Polymorpha, a New Species from Endotracheal Aspirate of a Preterm Neonate, and Transfer of Lecythophora Species to Coniochaeta. Antonie Van Leeuwenhoek 2013, 104, 52–243. [Google Scholar] [CrossRef]
- Weber, E.G.; Begerow, D. The Lecythophora-Coniochaeta Complex: II. Molecular Studies Based on Sequences of the Large Subunit of Ribosomal DNA. Nova Hedwig. 2002, 74, 187–200. [Google Scholar] [CrossRef]
- De Hoog, G.S.; Guarro, J.; Gené, J.; Figueras, M.J. Atlas of Clinical Fungi, 2nd ed.; Universitat Rovira i Virgili Reus: Tarragona, Spain; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2000. [Google Scholar]
- Gams, W.; McGinnis, M.R. Phialemonium, A New Anamorph Genus Intermediate Between Phialophora and Acremonium. Mycologia 1983, 75, 977–987. [Google Scholar] [CrossRef]
- Fintelmann, R.E. Recurrent Lecythophora mutabilis Keratitis and Endophthalmitis after Deep Anterior Lamellar Keratoplasty. Arch. Ophthalmol. 2011, 129, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Marriott, D.J.; Wong, K.H.; Aznar, E.; Harkness, J.L.; Cooper, D.A.; Muir, D. Scytalidium dimidiatum and Lecythophora hoffmannii: Unusual Causes of Fungal Infections in a Patient with AIDS. J. Clin. Microbiol. 1997, 35, 2949–2952. [Google Scholar] [CrossRef] [PubMed]
- Sakaeyama, S.; Sano, A.; Murata, Y.; Kamei, K.; Nishimura, K.; Hatai, K. Lecythophora hoffmannii Isolated from a Case of Canine Osteomyelitis in Japan. Med. Mycol. 2007, 45, 72–267. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ahmad, S.; Johnson, R.J.; Hillier, S.; Shelton, W.R.; Rinaldi, M.G. Fungal Peritonitis Caused by Lecythophora mutabilis. J. Clin. Microbiol. 1985, 22, 86–182. [Google Scholar] [CrossRef]
- Slifkin, M.; Bowers, H.M. Phialophora mutabilis Endocarditis. Am. J. Clin. Pathol. 1975, 63, 30–120. [Google Scholar] [CrossRef]
- Drees, M.; Wickes, B.L.; Gupta, M.; Hadley, S. Lecythophora mutabilis Prosthetic Valve Endocarditis in a Diabetic Patient. Med. Mycol. 2007, 45, 67–463. [Google Scholar] [CrossRef][Green Version]
- Taniguchi, Y.; Taketani, T.; Moriyama, H.; Moriki, S.; Nishimura, K.; Sato, E.; Notsu, Y.; Higuchi, T.; Sugitani, Y.; Yasuda, K.; et al. Septic Shock Induced by Lecythophora mutabilis in a Patient with Mitochondrial Encephalomyopathy. J. Med. Microbiol. 2009, 58, 58–1255. [Google Scholar] [CrossRef]
- Scott, I.U.; Cruz-Villegas, V.; Flynn, H.W.; Miller, D. Delayed-Onset, Bleb-Associated Endophthalmitis Caused by Lecythophora Mutabilis. Am. J. Ophthalmol. 2004, 3, 85–583. [Google Scholar] [CrossRef]
- Marcus, D.M.; Hull, D.S.; Rubin, R.M.; Newman, C.L. Lecythophora mutabilis endophthalmitis after long-term corneal cyanoacrylate. Retina 1999, 19, 351–353. [Google Scholar] [CrossRef]
- Ho, R.H.T.; Bernard, P.J.; McClellan, K.A. Phialophora mutabilis Keratomycosis. Am. J. Ophthalmol. 1991, 112, 29–728. [Google Scholar] [CrossRef]
- Cassagne, C.; Normand, A.N.; L’Ollivier, C.; Ranque, S.; Piarroux, R. Performance of MALDI-TOF MS Platforms for Fungal Identification. Mycoses 2016, 59, 90–678. [Google Scholar] [CrossRef] [PubMed]
- Normand, A.C.; Cassagne, C.; Gautier, M.; Becker, P.; Ranque, S.; Hendrickx, M.; Piarroux, R. Decision Criteria for MALDI-TOF MS-Based Identification of Filamentous Fungi Using Commercial and in-House Reference Databases. BMC Microbiol. 2017, 17, 17–25. [Google Scholar] [CrossRef]
- Bochner, B.R. Phenotype MicroArrays for High-Throughput Phenotypic Testing and Assay of Gene Function. Genome Res. 2001, 11, 55–1246. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplificarion and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc. 1990, 18, 22–315. [Google Scholar]
- Glass, N.L.; Donaldson, G.C. Development of Primer Sets Designed for Use with the PCR to Amplify Conserved Genes from Filamentous Ascomycetes. Appl. Environ. Microbiol. 1995, 61, 30–1323. [Google Scholar] [CrossRef] [PubMed]
- Carbone, I.; Kohn, L.M. A Method for Designing Primer Sets for Speciation Studies in Filamentous Ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Perdomo, H.D.; Sutton, A.; Garcia, D.; Fothergill, A.W.; Gené, J.; Cano, J.; Summerbell, R.C.; Rinaldi, M.G.; Guarro, J. Molecular and Phenotypic Characterization of Phialemonium and Lecythophora Isolates from Clinical Samples. J. Clin. Microbiol. 2011, 49, 1209–1216. [Google Scholar] [CrossRef]

| Primers | Sequences | Targeted Regions | References |
|---|---|---|---|
| ITS1 | TCCGTAGGTGAACCTGCGG | 18S-5.8S | [26] |
| ITS2 | GCTGCGTTCTTCATCGATGC | 18S-5.8S | [26] |
| ITS3 | GCATCGATGAAGAACGCAGC | 5.8S-28S | [26] |
| ITS4 | TCCTCCGCTTATTGATATGC | 5.8S-28S | [26] |
| ITS1 | TCCGTAGGTGAACCTGCGG | 18S-5.8S, 5.8S-28S | [26] |
| ITS4 | TCCTCCGCTTATTGATATGC | 18S-5.8S, 5.8S-28S | [26] |
| Bt-2a | GGTAACCAAATCGGTGCTGCTTTC | B-tub2 | [27] |
| Bt-2b | ACCCTCAGTGTAGTGACCCTTGGC | B-tub2 | [27] |
| EF1-728F | CATCGAGAAGTTCGAGAAGG | TEF1 | [28] |
| EF1-986R | TACTTGAAGGAACCCTTACC | TEF1 | [28] |
| D1 | AACTTAAGCATATCAATAAGCGGAGGA | 28S | [11] |
| D2 | GGT CCG TGT TTC AAG ACG G | 28S | [11] |
| Species | Collection ID | GenBank Accession Numbers | |||
|---|---|---|---|---|---|
| ITS | B-tub2 | D1/D2 | TEF1 | ||
| Coniochaeta massiliensis | PMML0158 | OM366153 | ON000097 | OM366268 | OM640093 |
| Coniochaeta mutabilis | DSM 10716 | OM366154 | ON000098 | OM366269 | OM640094 |
| Coniochaeta hoffmannii | DSM 2693 | OM366155 | ON000099 | OM366270 | OM640095 |
| Coniochaeta fasciculata | CBS 205.38 | HE610336 | HE610350 | FR691988 | MK693152 |
| Coniochaeta lignicola | CBS 267.33 | HE610335 | HE610353 | FR691986 | MK693154 |
| Coniochaeta luteoviridis | CBS 206.38 | HE610333 | HE610351 | FR691987 | NA * |
| Coniochaeta hoffmannii | CBS 245.38 | HE610332 | HE610352 | FR691982 | MK693150 |
| Coniochaeta mutabilis | CBS 157.44 | HE610334 | HE610349 | AF353604 | NA |
| Coniochaeta lignaria | DWS9m2/SMH2569/95.605 | KJ188673 | AY780113 | AF353584 | NA |
| Coniochaeta cateniformis | UTHSC 01-1644 | HE610331 | HE610347 | HE610329 | NA |
| Coniochaeta decumbens | CBS 153.42 | HE610337 | HE610348 | HE610463 | NA |
| Coniochaeta velutina | CBS 121444 | NA | NA | GQ154605 | NA |
| Coniochaeta velutina | STEU 8315 | KY312638 | NA | NA | NA |
| Coniochaeta simbalensis | NFCCI 4236 | MG825743 | NA | NG_068555 | NA |
| Coniochaeta fodinicola | YoF/CBS 136963 | JQ904607 | NA | NG_064287 | NA |
| Coniochaeta prunicola | CBS 120875 | GQ154540 | NA | NG_066151 | NA |
| Coniochaeta prunicola | STE-U 6107 | NA * | NA | NA | MK693162 |
| Coniochaeta africana | CBS 120868 | GQ154539 | NA | NG_066150 | NA |
| Coniochaeta palaoa | ARIZ AEASO06001 | MZ241149 | NA | NA | NA |
| Coniochaeta palaoa | ARIZ AEANC0604 | MK458764 | NA | NA | MZ241188 |
| Coniochaeta marina | MFLUCC 18-0408 | MF422164 | NA | NA | NA |
| Coniochaeta cipronana | PL5-2C | MG828883 | NA | NA | NA |
| Coniochaeta rosae | MFLUCC/17-0810 | MG828880 | NA | NA | MG829197 |
| Coniochaeta rosae | TASM 6127 | NA | NA | NG_066204 | NA |
| Coniochaeta baysunika | MFLUCC/17-0830 | MW750761 | NA | MG828996 | MG829196 |
| Coniochaeta acaciae | CX37 | MW750756 | NA | NA | NA |
| Coniochaeta acaciae | MFLUCC 18-0776 | NA | NA | NA | MT503199 |
| Coniochaeta acaciae | MFLUCC 17-2298 | NA | NA | MG062737 | NA |
| Coniochaeta fibrosae | CX04D1 | MW077645 | NA | NA | NA |
| Coniochaeta mongoliae | CS-09 | KP941078 | NA | NA | NA |
| Coniochaeta iranica | 806 | KP941076 | NA | NA | NA |
| Coniochaeta euphorbiae | 1001 | KY064029 | NA | NA | NA |
| Coniochaeta cephalothecoides | L821 | MW447035 | NA | KY064030 | NA |
| Coniochaeta luteorubra | FeC127 | MZ241160 | NA | NA | NA |
| Coniochaeta luteorubra | UTHSC 01-20 | NA | HE610346 | HE610328 | NA |
| Coniochaeta lutea | AEASO10801 | MZ241150 | NA | NA | MZ241193 |
| Coniochaeta endophytica | ARIZ AEAFL0922 | MZ241147 | NA | NA | NA |
| Coniochaeta endophytica | ARIZ AEA 9094 | NA | NA | NG_075158 | NA |
| Coniochaeta vineae | KUMCC 17-0322 | NA | MN485898 | NA | NA |
| Coniochaeta taeniospora | MFLU 17-0832 | NA | MN509784 | NA | NA |
| Coniochaeta discoidea | SANK12878 | NA | AY780134 | NA | NA |
| Coniochaeta elegans | FF0093 | NA | NA | NA | MZ267815 |
| Coniochaeta nivea | AK0926 | NA | NA | NA | MZ267793 |
| Coniochaeta deborreae | BE19_001008 | NA | NA | NA | MW890087 |
| Coniochaeta cymbiformispora | QU1057 | NA | NA | NA | MZ267839 |
| Coniochaeta endophytica | AEA 9094 | NA | NA | NG_075158 | MK693159 |
| Colletotrichum nymphaeae | CBS 515.78 | NR_111736 | NA | NA | NA |
| Colletotrichum caricae | NW688b | NR_111736 | NA | NA | NA |
| Colletotrichum fioriniae | CBS 151.35 | NR_111458 | NA | NG_069002 | NA |
| Colletotrichum circinans | CBS 128517 | NR_111747 | NA | NA | NA |
| Colletotrichum gloeosporioides | CBS 221.81 | NR_111457 | NA | NA | NA |
| Colletotrichum gloeosporioides | IMI 356878 | NA | AJ409291 | NA | NA |
| Colletotrichum acutatum | IMI 356878 | NR_160754 | NA | NA | NA |
| Colletotrichum acutatum | CBS 112996 | NA | AJ409296 | NA | NA |
| Colletotrichum siamense | CBS 112996 | NR_144794 | NA | NA | NA |
| Colletotrichum siamense | LC0034 | NA | NA | NA | JQ071904 |
| Colletotrichum truncatum | MFLU 090230 | NR_144784 | NA | NA | NA |
| Colletotrichum truncatum | CC01 | NA | NA | NA | MW030430 |
| Colletotrichum plurivorum | CBS 125474 | NA | MG600985 | NA | NA |
| Colletotrichum jasminigenum | LCA923 | NA | HM153770 | NA | NA |
| Colletotrichum serranegrense | COAD 2100 | NA | KY407896 | NA | NA |
| Colletotrichum neorubicola | CCR144 | NA | MN186400 | NA | NA |
| Colletotrichum spicati | YMF 1.04942 | NA | OL981226 | NA | NA |
| Colletotrichum fructicola | LC0033 | NA | NA | NA | JQ071903 |
| Colletotrichum theobromicola | GJS08_50 | NA | NA | NA | GU994506 |
| Colletotrichum horii | C1180.1 | NA | NA | NA | GQ329693 |
| Colletotrichum tropicale | LC0598 | NA | NA | NA | JQ071909 |
| Colletotrichum kahawae | IMI 319418 | NA | NA | NA | JQ071908 |
| Colletotrichum fragariae | MTCC 10325 | NA | NA | NA | JQ071906 |
| Colletotrichum camelliae | LC1364 | NA | JN936976 | NA | NA |
| Colletotrichum dacrycarpi | CBS 130241 | NA | NA | NG_073638 | NA |
| Colletotrichum neosansevieriae | CBS 139918 | NA | NA | NG_070628 | NA |
| Colletotrichum arboricola | CBS 144795 | NA | NA | NG_070064 | NA |
| Colletotrichum scovillei | CBS 126529 | NA | NA | NG_070041 | NA |
| Colletotrichum walleri | CBS 125472 | NA | NA | NG_070040 | NA |
| Colletotrichum melonis | CBS 159.84 | NA | NA | NG_070037 | NA |
| Colletotrichum brisbanense | CBS 292.67 | NA | NA | NG_070034 | NA |
| Phialemonium obovatum | CBS 279.76 | HE610365 | HE599334 | FR691997 | LT634003 |
| MIC * (mg/L) Read at 48 h | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| AMB | AND | CAS | 5-FC | FL | ITC | MIF | POS | VOR ** | |
| Coniochaeta massiliensis PMML0158 | 0.25 | 0.5 | 1 | 2 | 8 | 0.25 | 1 | 0.25 | 0.12 |
| Coniochaeta hoffmannii DSM 2693 | 0.12 | 4 | 4 | 1 | 4 | 0.06 | >32 | 0.12 | 0.12 |
| Coniochaeta mutabilis DSM 10716 | 0.12 | 4 | 8 | 1 | 2 | 0.015 | >32 | 0.03 | 0.03 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kabtani, J.; Militello, M.; Ranque, S. Coniochaeta massiliensis sp. nov. Isolated from a Clinical Sampl28. J. Fungi 2022, 8, 999. https://doi.org/10.3390/jof8100999
Kabtani J, Militello M, Ranque S. Coniochaeta massiliensis sp. nov. Isolated from a Clinical Sampl28. Journal of Fungi. 2022; 8(10):999. https://doi.org/10.3390/jof8100999
Chicago/Turabian StyleKabtani, Jihane, Muriel Militello, and Stéphane Ranque. 2022. "Coniochaeta massiliensis sp. nov. Isolated from a Clinical Sampl28" Journal of Fungi 8, no. 10: 999. https://doi.org/10.3390/jof8100999
APA StyleKabtani, J., Militello, M., & Ranque, S. (2022). Coniochaeta massiliensis sp. nov. Isolated from a Clinical Sampl28. Journal of Fungi, 8(10), 999. https://doi.org/10.3390/jof8100999






