Genome-Wide Expression Profiling of Small RNAs in Indian Strain of Rhizoctonia solani AG1-1A Reveals Differential Regulation of milRNAs during Pathogenesis and Crosstalk of Gene Regulation
Abstract
:1. Introduction
2. Material and Methods
2.1. Fungal and Plant Material
2.2. RNA Isolation and Sequencing
2.3. milRNAs Identification in R. solani
2.4. The Target Gene Prediction in R. solani and Rice Genome
2.5. Expression Analysis of Fungal milRNAs and Their Target Genes
3. Results
3.1. Sequence Statistics
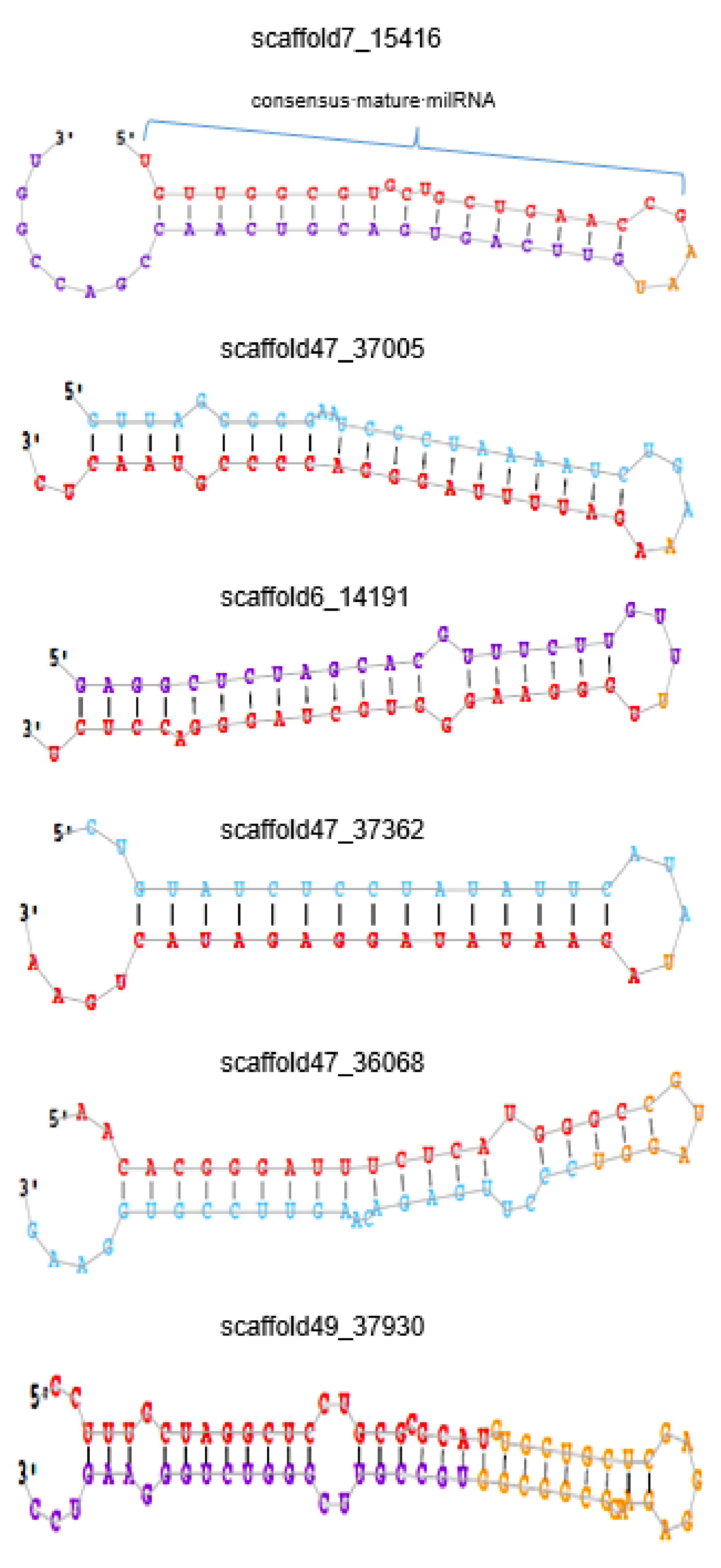
3.2. Identification of Known and Novel milRNAs
3.3. Identification of Target Genes in R. solani Genome
3.4. Identification of R. solani-milRNAs Target Genes in Oryza Sativa Genome
3.5. qRT-PCR Expression Analysis of milRNAs
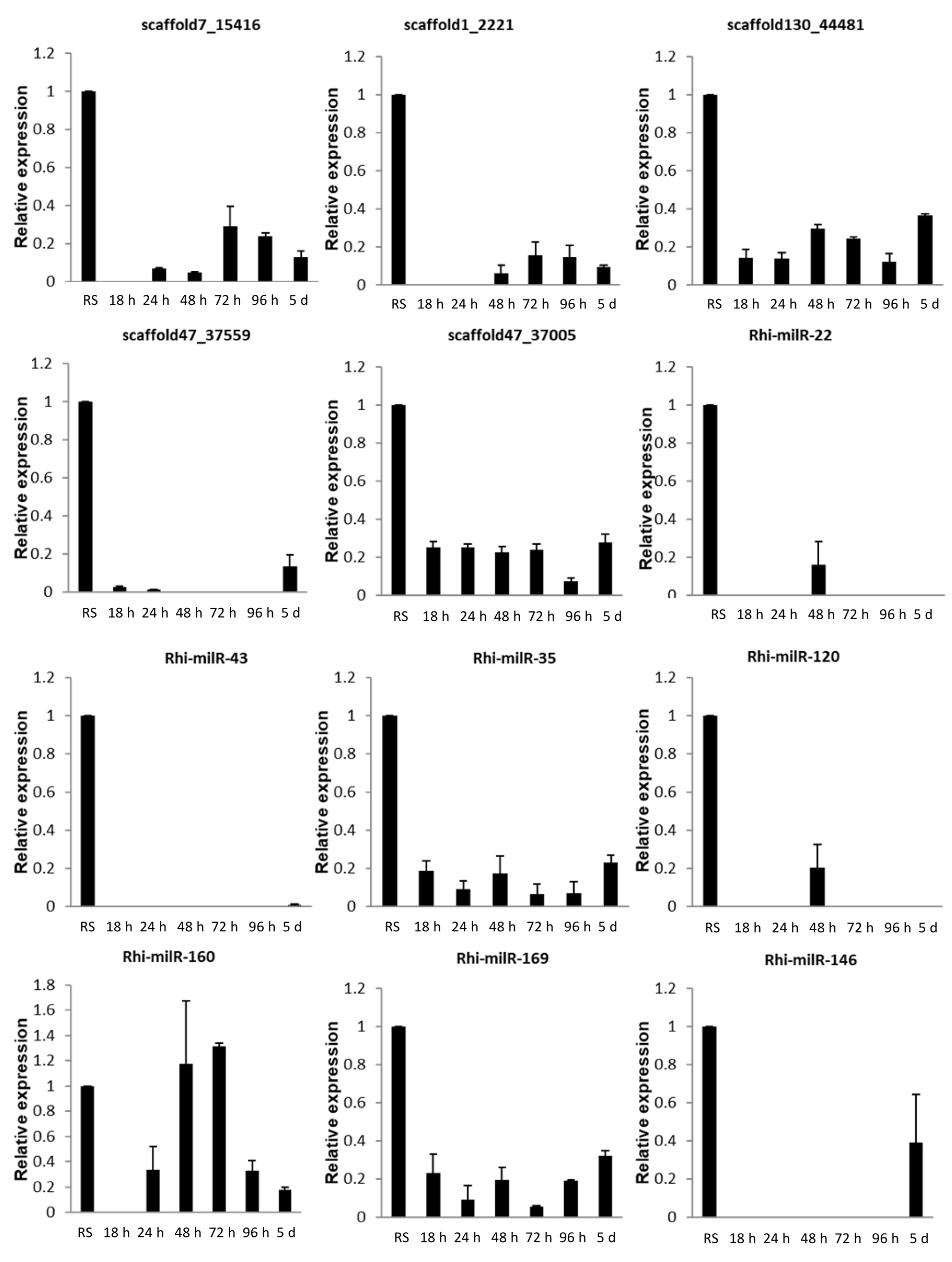
3.5.1. At Different Time Points of Inoculation
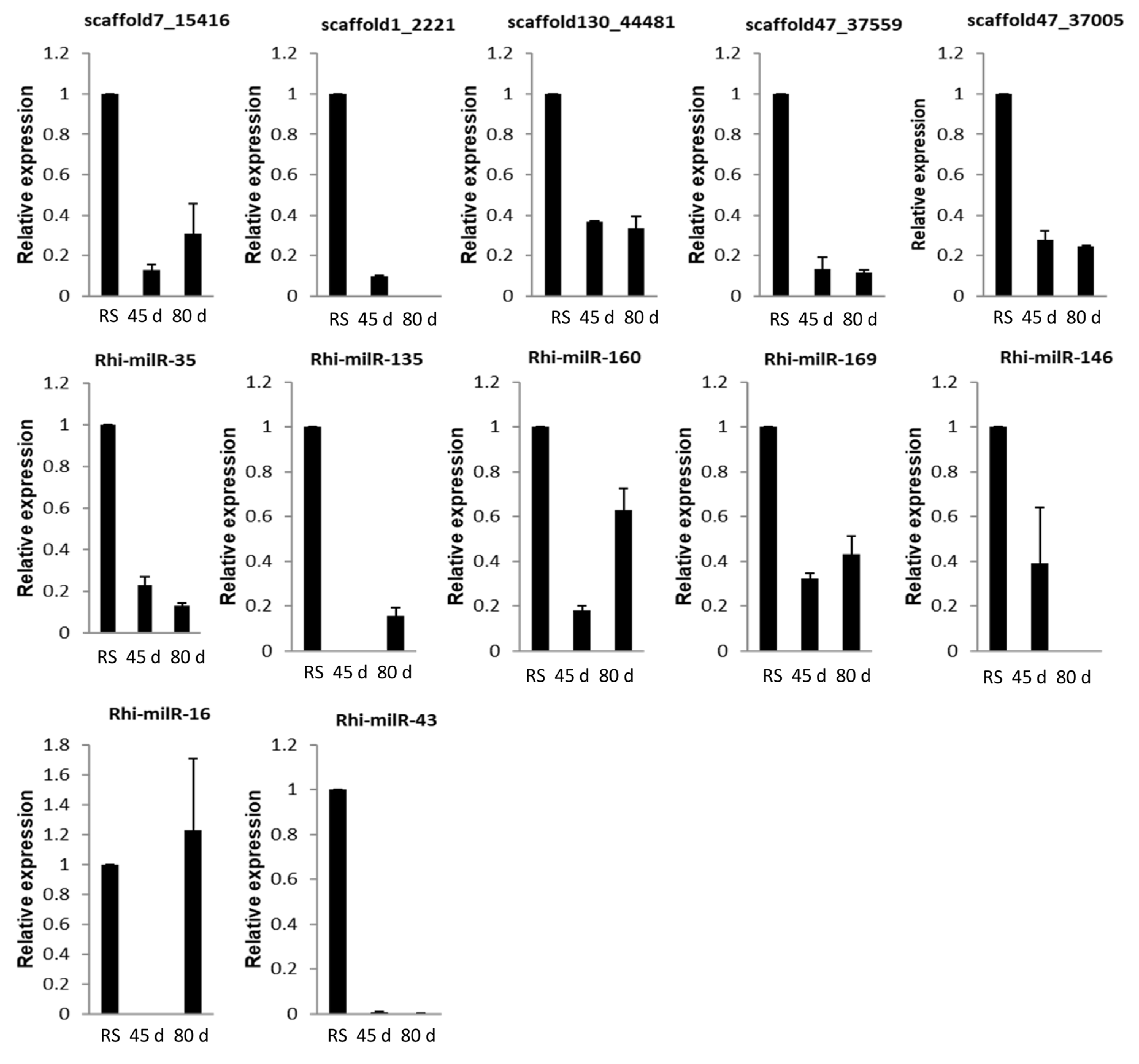
3.5.2. In Different Growth Stages of Host Plant
3.5.3. In Different Rice Genotypes
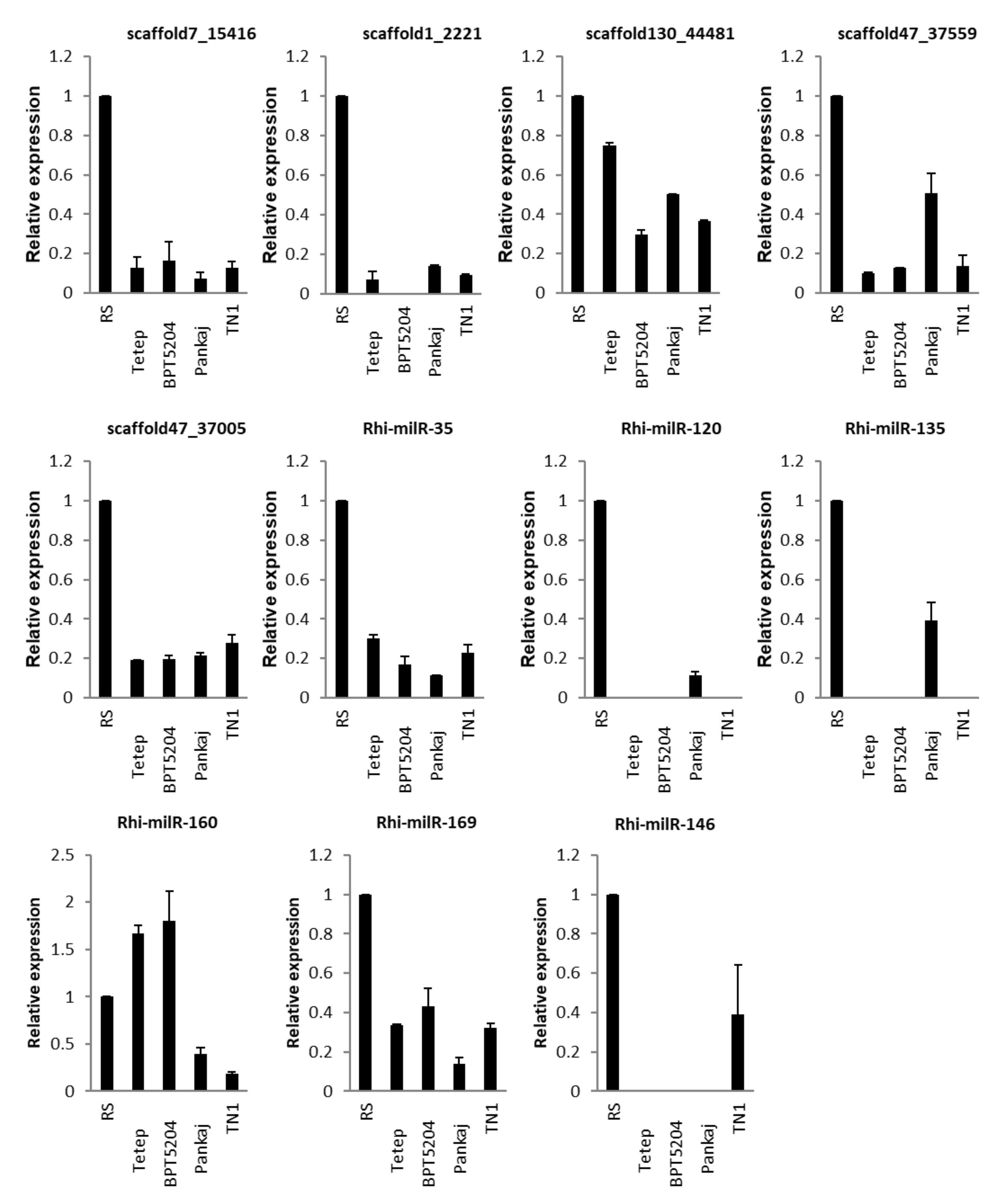
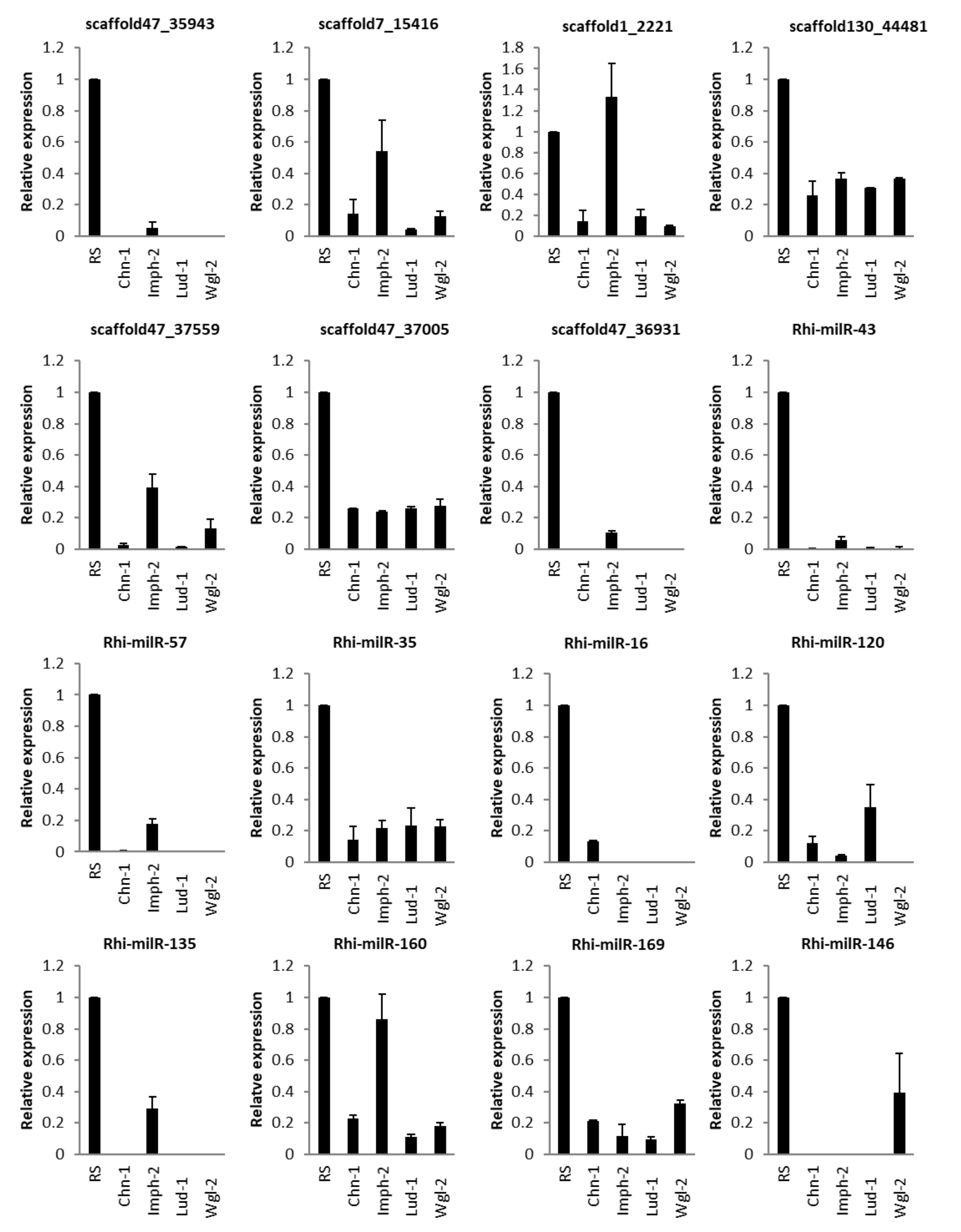
3.5.4. In Different Strains of Fungus
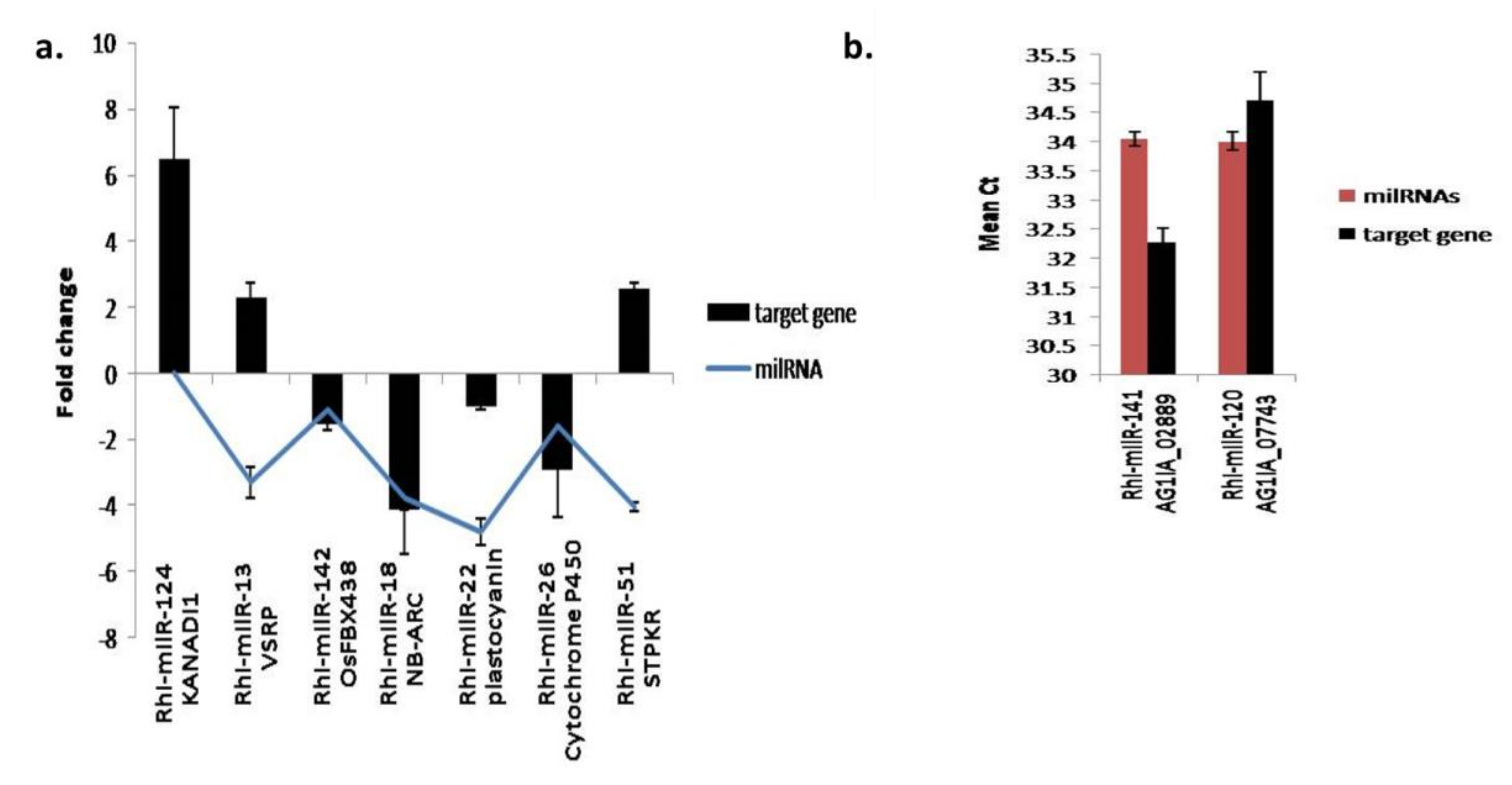
3.5.5. Expression Analysis of Fungal and Plant Target Genes
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bernardes-De-Assis, J.; Storari, M.; Zala, M.; Wang, W.; Jiang, D.; Shidong, L.; Jin, M.; McDonald, B.; Ceresini, P. Genetic Structure of Populations of the Rice-Infecting Pathogen Rhizoctonia solani AG-1 IA from China. Phytopathology 2009, 99, 1090–1099. [Google Scholar] [CrossRef] [Green Version]
- Boukaew, S.; Prasertsan, P. Suppression of rice sheath blight disease using a heat stable culture filtrate from Streptomyces philanthi RM-1-138. Crop Prot. 2014, 61, 1–10. [Google Scholar] [CrossRef]
- Meyers, B.C.; Axtell, M.J.; Bartel, B.; Bartel, D.P.; Baulcombe, D.; Bowman, J.L.; Cao, X.; Carrington, J.C.; Chen, X.; Green, P.J.; et al. Criteria for Annotation of Plant MicroRNAs. Plant Cell 2008, 20, 3186–3190. [Google Scholar] [CrossRef]
- Rao, T.B.; Chopperla, R.; Methre, R.; Punniakotti, E.; Venkatesh, V.; Sailaja, B.; Raghurami Reddy, M.; Yugander, A.; Laha, G.S.; SheshuMadhav, M.; et al. Pectin induced transcriptome of a Rhizoctonia solani strain causing sheath blight disease in rice reveals insights on key genes and RNAi machinery for development of pathogen derived resistance. Plant Mol. Biol. 2019, 100, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Zheng, A.; Lin, R.; Zhang, D.; Qin, P.; Xu, L.; Ai, P.; Ding, L.; Wang, Y.; Chen, Y.; Liu, Y.; et al. The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat. Commun. 2013, 4, 1424. [Google Scholar] [CrossRef]
- Lee, B.-M.; Park, Y.-J.; Park, D.-S.; Kang, H.-W.; Kim, J.-H.; Song, E.-S.; Park, I.-C.; Yoon, U.-H.; Hahn, J.-H.; Koo, B.-S.; et al. The genome sequence of Xanthomonas oryzae pathovar oryzae KACC10331, the bacterial blight pathogen of rice. Nucleic Acids Res. 2005, 33, 577–586. [Google Scholar] [CrossRef] [Green Version]
- Mücke, S.; Reschke, M.; Erkes, A.; Schwietzer, C.-A.; Becker, S.; Streubel, J.; Morgan, R.D.; Wilson, G.G.; Grau, J.; Boch, J. Transcriptional Reprogramming of Rice Cells by Xanthomonas oryzae TALEs. Front. Plant Sci. 2019, 10, 162. [Google Scholar] [CrossRef]
- Oliva, R.; Ji, C.; Atienza-Grande, G.; Huguet-Tapia, J.C.; Perez-Quintero, A.; Li, T.; Eom, J.-S.; Li, C.; Nguyen, H.; Liu, B.; et al. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat. Biotechnol. 2019, 37, 1344–1350. [Google Scholar] [CrossRef] [Green Version]
- Sundaram, R.M.; Vishnupriya, M.R.; Biradar, S.K.; Laha, G.S.; Reddy, G.A.; Rani, N.S.; Sarma, N.P.; Sonti, R.V. Marker assisted introgression of bacterial blight resistance in Samba Mahsuri, an elite indica rice variety. Euphytica 2008, 160, 411–422. [Google Scholar] [CrossRef]
- Rampersad, S.N. Pathogenomics and Management of Fusarium Diseases in Plants. Pathogens 2020, 9, 340. [Google Scholar] [CrossRef]
- Dong, O.X.; Ronald, P.C. Genetic Engineering for Disease Resistance in Plants: Recent Progress and Future Perspectives. Plant Physiol. 2019, 180, 26–38. [Google Scholar] [CrossRef] [Green Version]
- Cools, H.J.; Hammond-Kosack, K. Exploitation of genomics in fungicide research: Current status and future perspectives. Mol. Plant Pathol. 2012, 14, 197–210. [Google Scholar] [CrossRef]
- Bhogireddy, S.; Mangrauthia, S.K.; Kumar, R.; Pandey, A.K.; Singh, S.; Jain, A.; Budak, H.; Varshney, R.K.; Kudapa, H. Regulatory non-coding RNAs: A new frontier in regulation of plant biology. Funct. Integr. Genom. 2021, 1–18. [Google Scholar] [CrossRef]
- Wani, S.H.; Kumar, V.; Khare, T.; Tripathi, P.; Shah, T.; Ramakrishna, C.; Aglawe, S.; Mangrauthia, S.K. miRNA applications for engineering abiotic stress tolerance in plants. Biologia 2020, 75, 1063–1081. [Google Scholar] [CrossRef]
- Vazquez, F.; Vaucheret, H.; Rajagopalan, R.; Lepers, C.; Gasciolli, V.; Mallory, A.C.; Hilbert, J.-L.; Bartel, D.P.; Crété, P. Endogenous trans-Acting siRNAs Regulate the Accumulation of Arabidopsis mRNAs. Mol. Cell 2004, 16, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Bartel, D.P. Small RNAs Correspond to Centromere Heterochromatic Repeats. Science 2002, 297, 1831. [Google Scholar] [CrossRef] [Green Version]
- Zilberman, D.; Cao, X.; Jacobsen, S.E. ARGONAUTE4 Control of Locus-Specific siRNA Accumulation and DNA and Histone Methylation. Science 2003, 299, 716–719. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taverna, S.D.; Coyne, R.S.; Allis, C.D. Methylation of Histone H3 at Lysine 9 Targets Programmed DNA Elimination in Tetrahymena. Cell 2002, 110, 701–711. [Google Scholar] [CrossRef] [Green Version]
- Chen, R.; Jiang, N.; Jiang, Q.; Sun, X.; Wang, Y.; Zhang, H.; Hu, Z. Exploring MicroRNA-Like Small RNAs in the Filamentous Fungus Fusarium oxysporum. PLoS ONE 2014, 9, e104956. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Heikkinen, L.; Wang, C.; Yang, Y.; Sun, H.; Wong, G. Trends in the development of miRNA bioinformatics tools. Brief. Bioinform. 2019, 20, 1836–1852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, R.; He, L.; He, J.; Qin, P.; Wang, Y.; Deng, Q.; Yang, X.; Li, S.; Wang, S.; Wang, W.; et al. Comprehensive analysis of microRNA-Seq and target mRNAs of rice sheath blight pathogen provides new insights into pathogenic regulatory mechanisms. DNA Res. 2016, 23, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Lan, F.; Yang, W.; Zhang, F.; Yang, K.; Li, Z.; Gao, P.; Wang, S. sRNA profiling in Aspergillus flavus reveals differentially expressed miRNA-like RNAs response to water activity and temperature. Fungal Genet. Biol. 2015, 81, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Yang, F. Genome-wide analysis of small RNAs in the wheat pathogenic fungus Zymoseptoria tritici. Fungal Biol. 2015, 119, 631–640. [Google Scholar] [CrossRef]
- Lee, H.-C.; Li, L.; Gu, W.; Xue, Z.; Crosthwaite, S.K.; Pertsemlidis, A.; Lewis, Z.; Freitag, M.; Selker, E.U.; Mello, C.C.; et al. Diverse Pathways Generate MicroRNA-like RNAs and Dicer-Independent Small Interfering RNAs in Fungi. Mol. Cell 2010, 38, 803–814. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, H.; Wang, S.; Yang, W.; Ding, X.; Li, N.; Chu, Z.; Li, X. Identification of virulence associated milRNAs and their bidirectional targets in Rhizoctonia solani and maize during infection. BMC Plant Biol. 2021, 21, 155. [Google Scholar] [CrossRef]
- Yugander, A.; Ladhalakshmi, D.; Prakasham, V.; Mangrauthia, S.K.; Prasad, M.S.; Krishnaveni, D.; Madhav, M.S.; Sundaram, R.M.; Laha, G.S. Pathogenic and Genetic Variation among the Isolates of Rhizoctonia solani (AG 1-IA), the Rice Sheath Blight Pathogen. J. Phytopathol. 2015, 163, 465–474. [Google Scholar] [CrossRef]
- Chen, L.; Ai, P.; Zhang, J.; Deng, Q.; Wang, S.; Li, S.; Zhu, J.; Li, P.; Zheng, A. RSIADB, a collective resource for genome and transcriptome analyses in Rhizoctonia solani AG1 IA. Database 2016, 2016, baw031. [Google Scholar] [CrossRef] [Green Version]
- Axtell, M.J.; Meyers, B.C. Revisiting Criteria for Plant MicroRNA Annotation in the Era of Big Data. Plant Cell 2018, 30, 272–284. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef] [Green Version]
- Mangrauthia, S.K.; Bhogireddy, S.; Agarwal, S.; Prasanth, V.V.; Voleti, S.R.; Neelamraju, S.; Subrahmanyam, D. Genome-wide changes in microRNA expression during short and prolonged heat stress and recovery in contrasting rice cultivars. J. Exp. Bot. 2017, 68, 2399–2412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rao, T.B.; Chopperla, R.; Prathi, N.B.; Balakrishnan, M.; Prakasam, V.; Laha, G.S.; Balachandran, S.M.; Mangrauthia, S.K. A Comprehensive Gene Expression Profile of Pectin Degradation Enzymes Reveals the Molecular Events during Cell Wall Degradation and Pathogenesis of Rice Sheath Blight Pathogen Rhizoctonia solani AG1-IA. J. Fungi 2020, 6, 71. [Google Scholar] [CrossRef]
- Chang, S.-S.; Zhang, Z.; Liu, Y. RNA Interference Pathways in Fungi: Mechanisms and Functions. Annu. Rev. Microbiol. 2012, 66, 305–323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Molla, K.A.; Karmakar, S.; Molla, J.; Bajaj, P.; Varshney, R.K.; Datta, S.K.; Datta, K. Understanding sheath blight resistance in rice: The road behind and the road ahead. Plant Biotechnol. J. 2020, 18, 895–915. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Shu, X.; Niu, X.; Zhao, W.; Ai, P.; Li, P.; Zheng, A. Comparison of gene co-networks analysis provide a systems view of rice (Oryza sativa L.) response to Tilletia horrida infection. PLoS ONE 2018, 13, e0202309. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhao, W.; Fu, R.; Fu, C.; Wang, L.; Liu, H.; Li, S.; Deng, Q.; Wang, S.; Zhu, J.; et al. Comparison of gene co-networks reveals the molecular mechanisms of the rice (Oryza sativa L.) response to Rhizoctonia solani AG1 IA infection. Funct. Integr. Genom. 2018, 18, 545–557. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xu, X.; Yang, J.; Chen, L.; Liu, B.; Liu, T.; Jin, Q. Integrated microRNA and mRNA analysis in the pathogenic filamentous fungus Trichophyton rubrum. BMC Genom. 2018, 19, 933. [Google Scholar] [CrossRef] [PubMed]
- Xia, Z.; Wang, Z.; Kav, N.N.; Ding, C.; Liang, Y. Characterization of microRNA-like RNAs associated with sclerotial development in Sclerotinia sclerotiorum. Fungal Genet. Biol. 2020, 144, 103471. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Zhao, J.-H.; Zhao, P.; Zhang, T.; Wang, S.; Guo, H.-S. A fungal milRNA mediates epigenetic repression of a virulence gene in Verticillium dahliae. Philos. Trans. R. Soc. B 2019, 374, 20180309. [Google Scholar] [CrossRef] [Green Version]
- Shao, Y.; Tang, J.; Chen, S.; Wu, Y.; Wang, K.; Ma, B.; Zhou, Q.; Chen, A.; Wang, Y. milR4 and milR16 Mediated Fruiting Body Development in the Medicinal Fungus Cordyceps militaris. Front. Microbiol. 2019, 10, 83. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, M.; Guo, Y.; Tian, R.; Gao, C.; Guo, F.; Voegele, R.T.; Bao, J.; Li, C.; Jia, C.; Feng, H.; et al. Adaptive regulation of virulence genes by microRNA-like RNAs in Valsamali. New Phytol. 2020, 227, 899–913. [Google Scholar] [CrossRef]
- Mathur, M.; Nair, A.; Kadoo, N. Plant-pathogen interactions: MicroRNA-mediated trans-kingdom gene regulation in fungi and their host plants. Genomics 2020, 112, 3021–3035. [Google Scholar] [CrossRef]
- Wang, M.; Dean, R.A. Movement of small RNAs in and between plants and fungi. Mol. Plant Pathol. 2020, 21, 589–601. [Google Scholar] [CrossRef]
- Cui, C.; Wang, Y.; Liu, J.; Zhao, J.; Sun, P.; Wang, S. A fungal pathogen deploys a small silencing RNA that attenuates mosquito immunity and facilitates infection. Nat. Commun. 2019, 10, 4298. [Google Scholar] [CrossRef]
- Derbyshire, M.; Mbengue, M.; Barascud, M.; Navaud, O.; Raffaele, S. Small RNAs from the plant pathogenic fungus Sclerotinia sclerotiorum highlight host candidate genes associated with quantitative disease resistance. Mol. Plant Pathol. 2019, 20, 1279–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, T.; Harrar, Y.; Lin, C.; Reinhart, B.; Newell, N.R.; Talavera-Rauh, F.; Hokin, S.A.; Barton, M.K.; Kerstetter, R.A. Arabidopsis KANADI1 Acts as a Transcriptional Repressor by Interacting with a Specific cis-Element and Regulates Auxin Biosynthesis, Transport, and Signaling in Opposition to HD-ZIPIII Factors. Plant Cell 2014, 26, 246–262. [Google Scholar] [CrossRef] [Green Version]
- Ludwig-Müller, J. Bacteria and fungi controlling plant growth by manipulating auxin: Balance between development and defense. J. Plant Physiol. 2015, 172, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Wiedmer, J.; Palma-Guerrero, J. Small RNA Bidirectional Crosstalk During the Interaction Between Wheat and Zymoseptoria tritici. Front. Plant Sci. 2020, 10, 1669. [Google Scholar] [CrossRef]
- Ruano, G.; Scheuring, D. Plant Cells under Attack: Unconventional Endomembrane Trafficking during Plant Defense. Plants 2020, 9, 389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Zhao, S.-L.; Li, J.-L.; Hu, X.-H.; Wang, H.; Cao, X.-L.; Xu, Y.-J.; Zhao, Z.; Xiao, Z.-Y.; Yang, N.; et al. Osa-miR169 Negatively Regulates Rice Immunity against the Blast Fungus Magnaporthe oryzae. Front. Plant Sci. 2017, 8, 2. [Google Scholar] [CrossRef] [Green Version]
- Cheng, Q.; Li, N.; Dong, L.; Zhang, D.; Fan, S.; Jiang, L.; Wang, X.; Xu, P.; Zhang, S. Overexpression of Soybean Isoflavone Reductase (GmIFR) Enhances Resistance to Phytophthora sojae in Soybean. Front. Plant Sci. 2015, 6, 1024. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, H.; Zhang, Q.; Wang, B.; Fu, Y.; Huang, L.; Wang, X.; Kang, Z. Exploration of microRNAs and their targets engaging in the resistance interaction between wheat and stripe rust. Front. Plant Sci. 2015, 6, 469. [Google Scholar] [CrossRef] [Green Version]
- Goff, K.E.; Ramonell, K.M. The role and regulation of receptor-like kinases in plant defense. Gene Regul. Syst. Biol. 2007, 1, 167–175. [Google Scholar] [CrossRef] [Green Version]
- van den Burg, H.A.; Tsitsigiannis, D.I.; Rowland, O.; Lo, J.; Rallapalli, G.; MacLean, D.; Takken, F.L.W.; Jones, J.D.G. The F-Box Protein ACRE189/ACIF1 Regulates Cell Death and Defense Responses Activated during Pathogen Recognition in Tobacco and Tomato. Plant Cell 2008, 20, 697–719. [Google Scholar] [CrossRef] [Green Version]
- Wen, Z.; Yao, L.; Wan, R.; Li, Z.; Liu, C.; Wang, X. Ectopic Expression in Arabidopsis thaliana of an NB-ARC Encoding Putative Disease Resistance Gene from Wild Chinese Vitis pseudoreticulata Enhances Resistance to Phytopathogenic Fungi and Bacteria. Front. Plant Sci. 2015, 6, 1087. [Google Scholar] [CrossRef] [Green Version]
- Weiberg, A.; Wang, M.; Lin, F.-M.; Zhao, H.; Zhang, Z.; Kaloshian, I.; Huang, H.-D.; Jin, H. Fungal Small RNAs Suppress Plant Immunity by Hijacking Host RNA Interference Pathways. Science 2013, 342, 118–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dubey, H.; Kiran, K.; Jaswal, R.; Jain, P.; Kayastha, A.M.; Bhardwaj, S.C.; Mondal, T.K.; Sharma, T.R. Discovery and profiling of small RNAs from Puccinia triticina by deep sequencing and identification of their potential targets in wheat. Funct. Integr. Genom. 2019, 19, 391–407. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Sun, Y.; Song, N.; Zhao, M.; Liu, R.; Feng, H.; Wang, X.; Kang, Z. Puccinia striiformis f. sp. tritici mi croRNA -like RNA 1 (Pst -milR1), an important pathogenicity factor of Pst, impairs wheat resistance to Pst by suppressing the wheat pathogenesis-related 2 gene. New Phytol. 2017, 215, 338–350. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, X.; Qiao, F.; Long, Y.; Cong, H.; Sun, H. MicroRNA-like RNAs in plant pathogenic fungus Fusarium oxysporum f. sp. niveum are involved in toxin gene expression fine tuning. 3 Biotech 2017, 7, 354. [Google Scholar] [CrossRef]
- Zhou, J.; Fu, Y.; Xie, J.; Li, B.; Jiang, D.; Li, G.; Cheng, J. Identification of microRNA-like RNAs in a plant pathogenic fungus Sclerotinia sclerotiorum by high-throughput sequencing. Mol. Genet. Genom. 2012, 287, 275–282. [Google Scholar] [CrossRef]
| S. No. | Novel milRNA | miRDeep2 Score | Estimated Probability That the miRNA Candidate Is a True Positive | Total Read Count | Consensus Mature Sequence | Precursor Coordinate |
|---|---|---|---|---|---|---|
| 1 | scaffold47_37407 | 193.1 | 70 +/− 24% | 379 | aaucccuaggaucuccacuug | scaffold47:62019..62062:− |
| 2 | scaffold47_37523 | 193 | 70 +/− 24% | 379 | aaucccuaggaucuccacuug | scaffold47:127383..127426:− |
| 3 | scaffold47_37341 | 192.9 | 70 +/− 24% | 379 | aaucccuaggaucuccacuug | scaffold47:26292..26335:− |
| 4 | scaffold47_36931 | 30.3 | 70 +/− 24% | 80 | Uggacaauguucugugggguu | scaffold47:118388..118435:+ |
| 5 | scaffold145_44745 | 17.3 | 70 +/− 24% | 37 | Gacuaaucagcguugggcgcauuu | scaffold145:16023..16102:+ |
| 6 | scaffold6_14191 | 6 | 66 +/− 23% | 11 | Ugggaagggugcuagggaccucu | scaffold6:504695..504742:− |
| 7 | scaffold10_19054 | 1.3 | 35 +/− 26% | 3 | Ugccgacugugcucccgcccaucg | scaffold10:751369..751434:− |
| 8 | scaffold1_2221 | 0.9 | 16 +/− 15% | 5 | Cagcacacuggaccgagagcucu | scaffold1:3047379..3047427:+ |
| 9 | scaffold47_37005 | 0.8 | 16 +/− 15% | 249 | Agauuuuagggaccccguaacuc | scaffold47:124971..125020:+ |
| 10 | scaffold12_20737 | 0.8 | 16 +/− 15% | 27 | gaguuuuggacuggcaccc | scaffold12:518031..518072:+ |
| 11 | scaffold7_15416 | 0.6 | 16 +/− 15% | 2 | uguuggcgugcugcugaaccg | scaffold7:443987..444035:− |
| 12 | scaffold47_35943 | 0.4 | 16 +/− 15% | 29 | Aaggaaauucucguagggcuucu | scaffold47:16996..17045:+ |
| 13 | scaffold47_36418 | 0.4 | 16 +/− 15% | 94 | Uuucuauggaccaccacgaguacu | scaffold47:65345..65392:+ |
| 14 | scaffold49_37930 | 0.4 | 16 +/− 15% | 2 | Ccuuugcuaggcuccugcgcgcau | scaffold49:58540..58612:+ |
| 15 | scaffold48_37687 | 0.3 | 16 +/− 15% | 14 | Ucucgaagcgcgacucuguccuu | scaffold48:142724..142771:+ |
| 16 | scaffold30_30807 | 0.3 | 16 +/− 15% | 30 | Agcgcaacucgaccucugaucacg | scaffold30:89871..89933:− |
| 17 | scaffold34_32450 | 0.2 | 16 +/− 15% | 52 | Aucgcugacugcgguguccucu | scaffold34:218833..218913:− |
| 18 | scaffold13_22000 | 0 | 16 +/− 15% | 6028 | Aaggugccggaauauacgcucau | scaffold13:463575..463652:− |
| 19 | scaffold130_44481 | 0 | 16 +/− 15% | 2 | ccgugcaacggacgaucgac | scaffold130:10887..10931:− |
| 20 | scaffold47_37559 | 0 | 16 +/− 15% | 148 | Cguggacgggccgcauccc | scaffold47:139094..139138:− |
| 21 | scaffold22_27474 | 0 | 16 +/− 15% | 2 | Ucgggcgagacgagugcuuucc | scaffold22:336752..336810:+ |
| 22 | scaffold1_2114 | 0 | 16 +/− 15% | 3 | Ccugucgcugcucgugaagccucu | scaffold1:2908975..2909036:+ |
| milRNA Id | Target Gene | Gene Description |
|---|---|---|
| Carbohydrate Active Enzymes | ||
| Rhi-milR-56 | AG1IA_01218 | Beta-glucosidase (EC 3.2.1.21) |
| Rhi-milR-120 | AG1IA_01405 | 1,3-beta-glucan synthase component GLS2 |
| Rhi-milR-122 | AG1IA_01406 | Sterol 3-beta-glucosyltransferase (EC 2.4.1.173) (Autophagy-related protein 26) |
| Rhi-milR-91 | AG1IA_02027 | Endoplasmic reticulum protein |
| Rhi-milR-169 | AG1IA_02441 | Beta-xylosidase |
| Rhi-milR-146 | AG1IA_02513 | Chitin deacetylase |
| Rhi-milR-120 | AG1IA_02835 | Putative 1,4-alpha-glucan branching enzyme from glycoside hydrolase family GH13 |
| Rhi-milR-141 | AG1IA_02889 | Pectinesterase (EC 3.1.1.11) |
| Rhi-milR-150 | AG1IA_03463 | Glycosyltransferase family 2 protein |
| Rhi-milR-36 | AG1IA_03939 | Beta-mannosidase |
| Rhi-milR-91 | AG1IA_04214 | Trehalose 6-phosphate phosphatase, glycosyltransferase family 20 protein |
| Rhi-milR-81 | AG1IA_04527 | Uridine Di Phosphate-N-acetylglucosaminyltransferase |
| Rhi-milR-68 | AG1IA_04727 | Farnesyltransferase subunit beta |
| Rhi-milR-111 | AG1IA_04740 | Polysaccharide lyase family 1 protein |
| Rhi-milR-27 | AG1IA_04862 | 1,3-beta-glucan synthase component GLS2 |
| Rhi-milR-45 | AG1IA_05653 | Glycoside hydrolase family 51 protein |
| Rhi-milR-52 | AG1IA_05719 | Chitin synthase D |
| Rhi-milR-119 | AG1IA_05754 | Glycogen phosphorylase |
| Rhi-milR-42 | AG1IA_05803 | Alpha-galactosidase (EC 3.2.1.22) (Melibiase) |
| Rhi-milR-139 | AG1IA_05807 | Adenylosuccinate synthetase (AMPSase) (AdSS) (EC 6.3.4.4) (IMP-aspartate ligase) |
| Rhi-milR-122 | AG1IA_06014 | Glycoside hydrolase family 51 protein |
| Rhi-milR-41 | AG1IA_06294 | Glycoside hydrolase family 3 protein |
| Rhi-milR-58 | AG1IA_06593 | Alpha glucosidase II, alpha subunit, putative |
| Rhi-milR-167 | AG1IA_07255 | Killer toxin alpha/beta |
| Rhi-milR-51 | AG1IA_07341 | Galactan 1,3-beta-galactosidase |
| Rhi-milR-120 | AG1IA_07743 | Pectinesterase (EC 3.1.1.11) |
| Rhi-milR-162 | AG1IA_07787 | Glycoside hydrolase family 31 protein |
| Rhi-milR-168 | AG1IA_07905 | Glycoside hydrolase family 95 protein |
| Rhi-milR-97 | AG1IA_08771 | Exo-beta-1,3-glucanase |
| Secretory Proteins | ||
| Rhi-milR-150 | AG1IA_00157 | Polysaccharide deacetylase domain-containing protein |
| Rhi-milR-98 | AG1IA_01858 | Uncharacterized protein |
| Rhi-milR-124 | AG1IA_01958 | Rad1 domain-containing protein |
| Rhi-milR-95 | AG1IA_02532 | Lipase domain-containing protein |
| Rhi-milR-144 | AG1IA_03100 | Glycosyl hydrolase family 61 domain-containing protein |
| Rhi-milR-122 | AG1IA_03118 | Uncharacterized protein |
| Rhi-milR-141 | AG1IA_03171 | Copper/zinc superoxide dismutase domain-containing protein |
| Rhi-milR-165 | AG1IA_05741 | Uncharacterized protein |
| Rhi-milR-141 | AG1IA_06494 | Uncharacterized protein |
| Rhi-milR-122 | AG1IA_07117 | Uncharacterized protein |
| Rhi-milR-52 | AG1IA_07216 | Uncharacterized protein |
| Rhi-milR-19 | AG1IA_07698 | Transcription initiation factor TFIID complex 60 kDa subunit |
| Rhi-milR-54 | AG1IA_08056 | Tyrosinase domain-containing protein |
| Rhi-milR-144 | AG1IA_08227 | Uncharacterized protein |
| Rhi-milR-90 | AG1IA_08293 | Cytochrome P450 domain-containing protein |
| Rhi-milR-146 | AG1IA_08653 | Uncharacterized protein |
| Rhi-milR-122 | AG1IA_08711 | Uncharacterized protein |
| Rhi-milR-144 | AG1IA_09802 | Protein tyrosine kinase domain-containing protein |
| Rhi-milR-130 | AG1IA_10060 | Cytochrome P450 domain-containing protein |
| S. No. | Novel milRNA | Target Gene | Gene Description |
|---|---|---|---|
| 1 | scaffold47_37407 | AG1IA_04067 | Ubiquitin conjugating enzyme family protein |
| 2 | scaffold47_37407 | AG1IA_02707 | MFS transporter, putative |
| 3 | scaffold47_37523 | AG1IA_04067 | Ubiquitin conjugating enzyme family protein |
| 4 | scaffold47_37523 | AG1IA_02707 | MFS transporter, putative |
| 5 | scaffold47_37341 | AG1IA_04067 | Ubiquitin conjugating enzyme family protein |
| 6 | scaffold47_37341 | AG1IA_02707 | MFS transporter, putative |
| 7 | scaffold47_36931 | AG1IA_08169 | Phosphatidylinositol 3-kinase tor2 |
| 8 | scaffold47_36931 | AG1IA_05744 | DNA-directed RNA polymerase subunit (EC 2.7.7.6) |
| 9 | scaffold47_36931 | AG1IA_06882 | EOS1 domain-containing protein |
| 10 | scaffold47_36931 | AG1IA_06165 | ABC transporter |
| 11 | scaffold47_36931 | AG1IA_07629 | Uncharacterized protein |
| 12 | scaffold47_36931 | AG1IA_07663 | ATP-dependent rRNA helicase RRP3 |
| 13 | scaffold47_36931 | AG1IA_04769 | Fungal zn(2)-Cys(6) binuclear cluster domain-containing protein |
| 14 | scaffold47_36931 | AG1IA_08613 | BMR1 protein |
| 15 | scaffold47_36931 | AG1IA_00053 | Uncharacterized protein |
| 16 | scaffold47_36931 | AG1IA_06822 | Uncharacterized protein |
| 17 | scaffold47_36931 | AG1IA_07843 | Uncharacterized protein |
| 18 | scaffold47_36931 | AG1IA_08221 | Molybdenum cofactor biosynthesis protein |
| 19 | scaffold145_44745 | AG1IA_06624 | Uncharacterized protein |
| 20 | scaffold145_44745 | AG1IA_10187 | RNase H domain-containing protein |
| 21 | scaffold145_44745 | AG1IA_10310 | TFIIA domain-containing protein |
| 22 | scaffold145_44745 | AG1IA_02688 | GPI transamidase component PIG-S |
| 23 | scaffold145_44745 | AG1IA_05493 | Uncharacterized protein |
| 24 | scaffold145_44745 | AG1IA_04447 | HLH domain-containing protein |
| milRNAs | Target Gene Accession | Expect | Gene Description | Inhibition |
|---|---|---|---|---|
| Rhi-milR-1 | LOC_Os01g47740.2 | 3 | cDNA|zinc finger, C3HC4 type domain containing protein, expressed | Translation |
| Rhi-milR-111 | LOC_Os07g48720.3 | 2.5 | cDNA|MAG2, putative, expressed | Cleavage |
| Rhi-milR-124 | LOC_Os09g23200.1 | 2.5 | cDNA|KANADI1, putative, expressed | Cleavage |
| Rhi-milR-13 | LOC_Os10g20630.1 | 1 | cDNA|vacuolar-sorting receptor precursor, putative, expressed | Cleavage |
| LOC_Os11g02464.1 | 2.5 | cDNA|vacuolar-sorting receptor precursor, putative, expressed | Cleavage | |
| Rhi-milR-130 | LOC_Os06g16140.1 | 2.5 | cDNA|expressed protein | Cleavage |
| Rhi-milR-131 | LOC_Os12g42400.4 | 2.5 | cDNA|nuclear transcription factor Y subunit, putative, expressed | Cleavage |
| LOC_Os12g42400.1 | 2.5 | cDNA|nuclear transcription factor Y subunit, putative, expressed | Cleavage | |
| LOC_Os02g58790.5 | 3 | cDNA|cell division inhibitor, putative, expressed | Cleavage | |
| LOC_Os02g58790.2 | 3 | cDNA|cell division inhibitor, putative, expressed | Cleavage | |
| LOC_Os02g58790.1 | 3 | cDNA|cell division inhibitor, putative, expressed | Cleavage | |
| LOC_Os02g58790.4 | 3 | cDNA|cell division inhibitor, putative, expressed | Cleavage | |
| LOC_Os02g58790.3 | 3 | cDNA|cell division inhibitor, putative, expressed | Cleavage | |
| LOC_Os01g55200.1 | 3 | cDNA|potassium channel KAT1, putative, expressed | Cleavage | |
| Rhi-milR-135 | LOC_Os12g16290.1 | 2.5 | cDNA|isoflavone reductase, putative, expressed | Cleavage |
| LOC_Os05g45180.1 | 3 | cDNA|anthocyanidin 5,3-O-glucosyltransferase, putative, expressed | Cleavage | |
| Rhi-milR-142 | LOC_Os12g03740.1 | 2.5 | cDNA|OsFBX438-F-box domain containing protein, expressed | Cleavage |
| Rhi-milR-159 | LOC_Os12g43720.1 | 3 | cDNA|early-responsive to dehydration protein-related, putative, expressed | Cleavage |
| Rhi-milR-160 | LOC_Os03g24410.1 | 1.5 | cDNA|expressed protein | Cleavage |
| Rhi-milR-18 | LOC_Os08g30660.1 | 2.5 | cDNA|NB-ARC domain containing protein, expressed | Cleavage |
| LOC_Os12g02570.2 | 2.5 | cDNA|expressed protein | Cleavage | |
| LOC_Os12g02570.1 | 2.5 | cDNA|expressed protein | Cleavage | |
| Rhi-milR-20 | LOC_Os01g25430.1 | 2 | cDNA|expressed protein | Cleavage |
| LOC_Os07g48200.2 | 2.5 | cDNA|B3 DNA binding domain containing protein, putative, expressed | Cleavage | |
| LOC_Os07g48200.1 | 2.5 | cDNA|B3 DNA binding domain containing protein, putative, expressed | Cleavage | |
| Rhi-milR-22 | LOC_Os02g49350.1 | 2.5 | cDNA|plastocyanin-like domain containing protein, putative, expressed | Cleavage |
| Rhi-milR-26 | LOC_Os03g55250.1 | 2.5 | cDNA|cytochrome P450 81E1, putative, expressed | Cleavage |
| Rhi-milR-43 | LOC_Os04g28160.1 | 3 | cDNA|response regulator receiver domain containing protein, expressed | Cleavage |
| Rhi-milR-51 | LOC_Os04g34390.1 | 3 | cDNA|serine/threonine-protein kinase receptor precursor, putative, expressed | Cleavage |
| Rhi-milR-61 | LOC_Os10g26660.2 | 2.5 | cDNA|expressed protein | Translation |
| LOC_Os08g40620.1 | 2.5 | cDNA|rabGAP/TBC domain-containing protein, putative, expressed | Cleavage | |
| LOC_Os08g40620.2 | 2.5 | cDNA|rabGAP/TBC domain-containing protein, putative, expressed | Cleavage | |
| Rhi-milR-66 | LOC_Os05g11414.1 | 2.5 | cDNA|OsMADS58-MADS-box family gene with MIKCc type-box, expressed | Cleavage |
| Rhi-milR-81 | LOC_Os05g50910.2 | 3 | cDNA|extra-large G-protein-related, putative, expressed | Cleavage |
| LOC_Os05g50910.1 | 3 | cDNA|extra-large G-protein-related, putative, expressed | Cleavage | |
| Rhi-milR-89 | LOC_Os06g33020.1 | 2.5 | cDNA|retrotransposon protein, putative, Ty3-gypsy subclass, expressed | Cleavage |
| LOC_Os06g32890.1 | 2.5 | cDNA|retrotransposon protein, putative, Ty3-gypsy subclass, expressed | Cleavage | |
| Rhi-milR-9 | LOC_Os02g35820.2 | 2.5 | cDNA|retrotransposon protein, putative, unclassified, expressed | Cleavage |
| LOC_Os02g35820.1 | 2.5 | cDNA|retrotransposon protein, putative, unclassified, expressed | Cleavage | |
| LOC_Os07g04230.1 | 2.5 | cDNA|retrotransposon protein, putative, unclassified, expressed | Cleavage | |
| Rhi-milR-97 | LOC_Os07g47950.1 | 2.5 | cDNA|expressed protein | Cleavage |
| scaffold130_44481 | LOC_Os09g29390.1 | 2.5 | cDNA|plastocyanin-like domain containing protein, putative, expressed | Cleavage |
| scaffold47_37559 | LOC_Os04g52164.1 | 2 | cDNA|transferase family protein, putative, expressed | Cleavage |
| LOC_Os04g52164.2 | 2 | cDNA|transferase family protein, putative, expressed | Cleavage |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prathi, N.B.; Durga Rani, C.V.; Balachandran, S.M.; Prakasam, V.; Chandra Mohan, Y.; Nagalakshmi, S.; Srivastava, S.K.; Sundaram, R.M.; Mangrauthia, S.K. Genome-Wide Expression Profiling of Small RNAs in Indian Strain of Rhizoctonia solani AG1-1A Reveals Differential Regulation of milRNAs during Pathogenesis and Crosstalk of Gene Regulation. J. Fungi 2021, 7, 561. https://doi.org/10.3390/jof7070561
Prathi NB, Durga Rani CV, Balachandran SM, Prakasam V, Chandra Mohan Y, Nagalakshmi S, Srivastava SK, Sundaram RM, Mangrauthia SK. Genome-Wide Expression Profiling of Small RNAs in Indian Strain of Rhizoctonia solani AG1-1A Reveals Differential Regulation of milRNAs during Pathogenesis and Crosstalk of Gene Regulation. Journal of Fungi. 2021; 7(7):561. https://doi.org/10.3390/jof7070561
Chicago/Turabian StylePrathi, Naresh Babu, Chagamreddy Venkata Durga Rani, Sena Munuswamy Balachandran, Vellaisamy Prakasam, Yeshala Chandra Mohan, Sanivarapu Nagalakshmi, Sunil K. Srivastava, Raman Meenakshi Sundaram, and Satendra K. Mangrauthia. 2021. "Genome-Wide Expression Profiling of Small RNAs in Indian Strain of Rhizoctonia solani AG1-1A Reveals Differential Regulation of milRNAs during Pathogenesis and Crosstalk of Gene Regulation" Journal of Fungi 7, no. 7: 561. https://doi.org/10.3390/jof7070561






