Associations between Cryptococcus Genotypes, Phenotypes, and Clinical Parameters of Human Disease: A Review
Abstract
1. Introduction
2. Cryptococcus Genotypes and Virulence Phenotypes
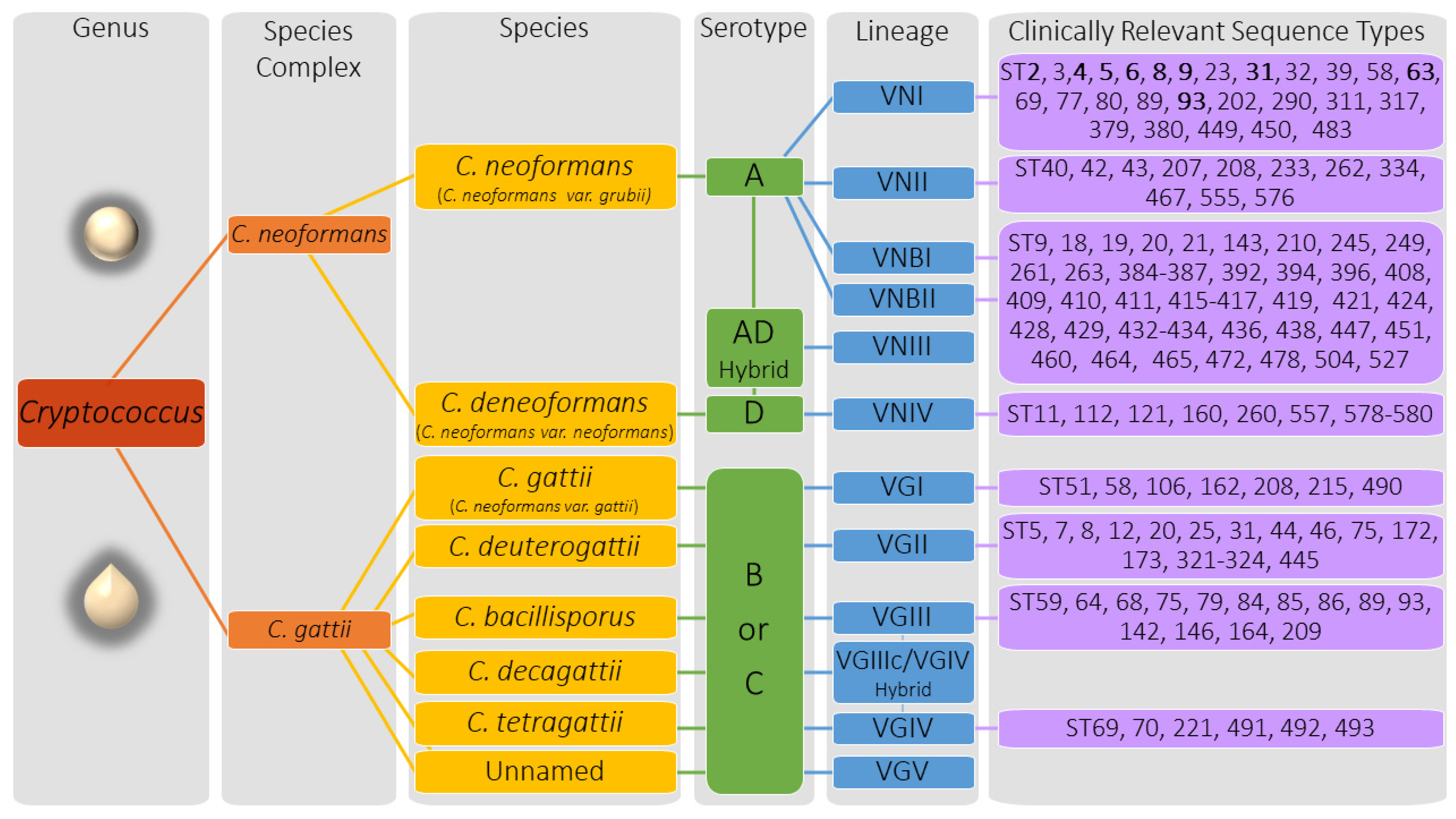
2.1. Cryptococcus Genetics and Genotypic Descriptors
2.2. Cryptococcus Virulence Phenotypes
2.2.1. Capsule
2.2.2. Cell Wall and Melanin
2.2.3. Secretion of Degradative Enzymes
2.2.4. Novel Virulence Phenotype: Yeast Cell Size
3. Cryptococcosis Clinical Diagnostics and Measures of Disease Severity in Humans
3.1. Clinical Diagnostics to Detect Cryptococcus in Humans
3.1.1. Cell Culture, India Ink Staining, and Quantitative Yeast Counts
3.1.2. Cryptococcus Antigen
3.2. Human Clinical Phenotypes: Clinical Measures of Disease Severity
Symptoms, Radiology, and Clinical Diagnostics
4. Associations between Cryptococcal Genotypes and Clinical Presentation
4.1. Mating Type, Species, and Serotype
4.2. Molecular Type
4.3. Sequence Type
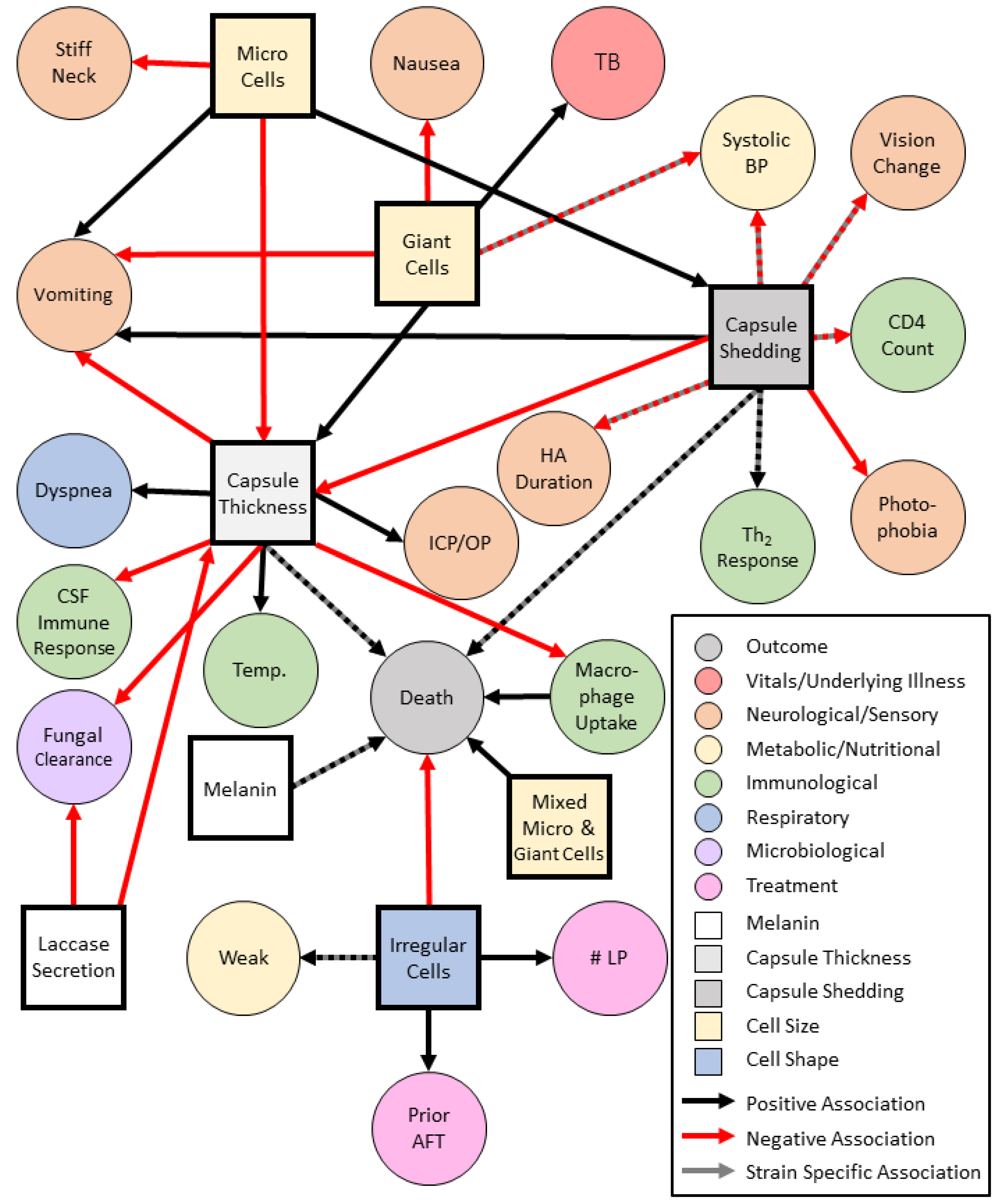
5. Associations between Cryptococcal Phenotypes and Clinical Presentation
Cryptococcal Virulence Phenotypes and Clinical Presentation
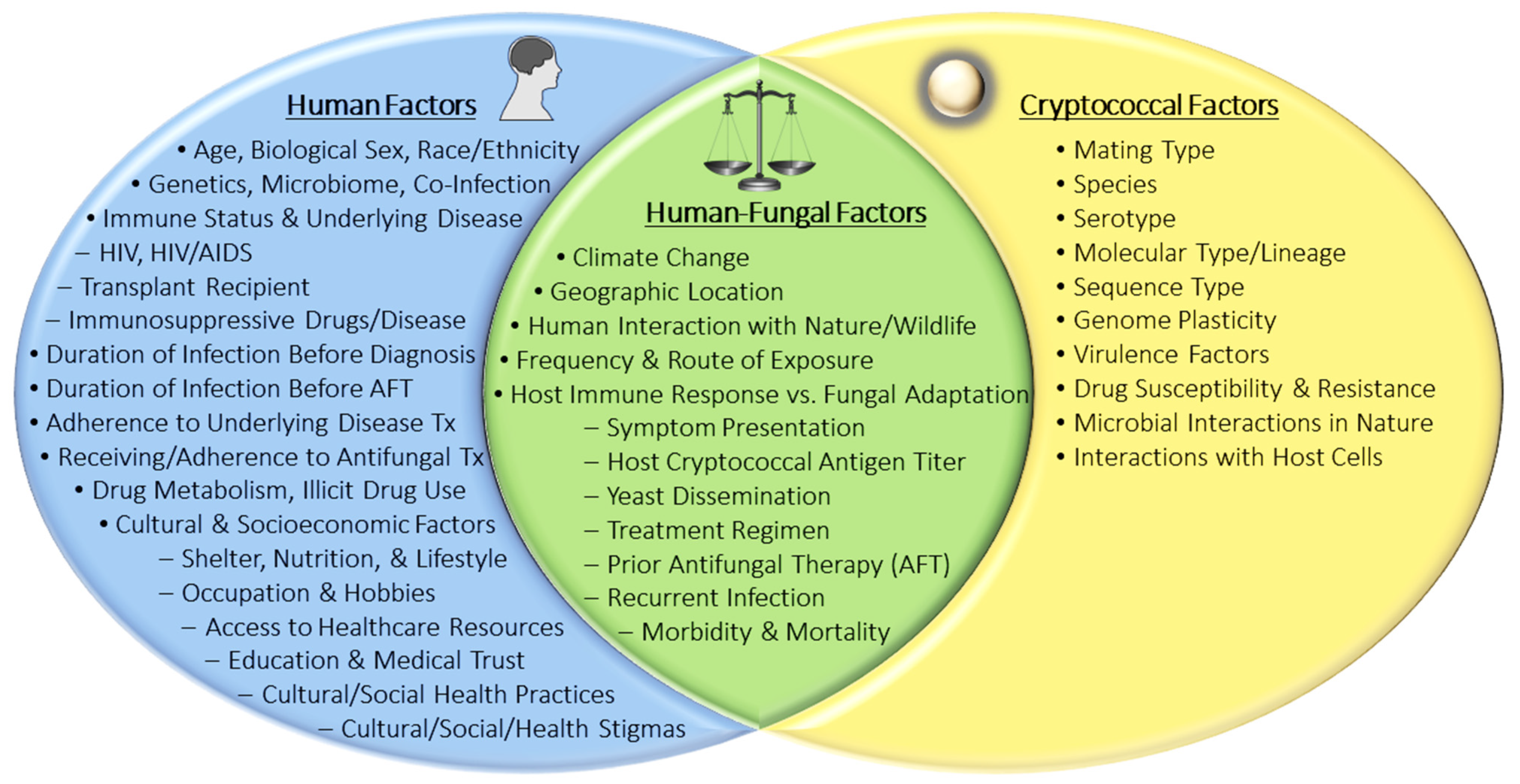
6. Associations between Human Attributes and Cryptococcosis
6.1. Associations between Human Genetics and Cryptococcosis
6.1.1. Mannose-Binding Lectin
6.1.2. Immunoglobulins
6.1.3. Dectin-2, Pentraxin-3, Interleukin
6.1.4. Anti-Granulocyte-Macrophage Colony-Stimulating Factor Autoantibodies
6.1.5. Biological Sex and Hormones
6.2. Associations between Human Phenotypes and Cryptococcosis
6.2.1. Pediatric Cryptococcosis
6.2.2. Illicit Drug Use
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
References
- May, R.C.; Stone, N.R.; Wiesner, D.L.; Bicanic, T.; Nielsen, K. Cryptococcus: From environmental saprophyte to global pathogen. Nat. Rev. Microbiol. 2016, 14, 106–117. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Fraser, J.A.; Doering, T.L.; Wang, Z.; Janbon, G.; Idnurm, A.; Bahn, Y.S. Cryptococcus neoformans and Cryptococcus gattii, the etiologic agents of cryptococcosis. Cold Spring Harb. Perspect. Med. 2014, 4, a019760. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Heitman, J. The biology of the Cryptococcus neoformans species complex. Annu. Rev. Microbiol. 2006, 60, 69–105. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, T.G.; Castañeda, E.; Nielsen, K.; Wanke, B.; Lazéra, M.S. Environmental Niches for Cryptococcus neoformans and Cryptococcus gattii. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Zonios, D.I.; Falloon, J.; Huang, C.Y.; Chaitt, D.; Bennett, J.E. Cryptococcosis and idiopathic CD4 lymphocytopenia. Medicine 2007, 86, 78–92. [Google Scholar] [CrossRef] [PubMed]
- Saijo, T.; Chen, J.; Chen, S.C.; Rosen, L.B.; Yi, J.; Sorrell, T.C.; Bennett, J.E.; Holland, S.M.; Browne, S.K.; Kwon-Chung, K.J. Anti-granulocyte-macrophage colony-stimulating factor autoantibodies are a risk factor for central nervous system infection by Cryptococcus gattii in otherwise immunocompetent patients. mBio 2014, 5, e00912–e00914. [Google Scholar] [CrossRef]
- Rajasingham, R.; Smith, R.M.; Park, B.J.; Jarvis, J.N.; Govender, N.P.; Chiller, T.M.; Denning, D.W.; Loyse, A.; Boulware, D.R. Global burden of disease of HIV-associated cryptococcal meningitis: An updated analysis. Lancet. Infect. Dis. 2017, 17, 873–881. [Google Scholar] [CrossRef]
- Dromer, F.; Casadevall, A.; Perfect, J.; Sorrell, T. Cryptococcus neoformans: Latency and Disease. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Setianingrum, F.; Rautemaa-Richardson, R.; Denning, D.W. Pulmonary cryptococcosis: A review of pathobiology and clinical aspects. Med. Mycol. 2019, 57, 133–150. [Google Scholar] [CrossRef]
- Walsh, N.M.; Botts, M.R.; McDermott, A.J.; Ortiz, S.C.; Wüthrich, M.; Klein, B.; Hull, C.M. Infectious particle identity determines dissemination and disease outcome for the inhaled human fungal pathogen Cryptococcus. PLoS Pathog. 2019, 15, e1007777. [Google Scholar] [CrossRef] [PubMed]
- Goldman, D.L.; Khine, H.; Abadi, J.; Lindenberg, D.J.; Pirofski, L.; Niang, R.; Casadevall, A. Serologic evidence for Cryptococcus neoformans infection in early childhood. Pediatrics 2001, 107, E66. [Google Scholar] [CrossRef] [PubMed]
- Speed, B.R.; Kaldor, J. Rarity of cryptococcal infection in children. Pediatric Infect. Dis. J. 1997, 16, 536–537. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Hermoso, D.; Janbon, G.; Dromer, F. Epidemiological evidence for dormant Cryptococcus neoformans infection. J. Clin. Microbiol. 1999, 37, 3204–3209. [Google Scholar] [CrossRef] [PubMed]
- Saha, D.C.; Goldman, D.L.; Shao, X.; Casadevall, A.; Husain, S.; Limaye, A.P.; Lyon, M.; Somani, J.; Pursell, K.; Pruett, T.L.; et al. Serologic evidence for reactivation of cryptococcosis in solid-organ transplant recipients. Clin. Vaccine Immunol. 2007, 14, 1550–1554. [Google Scholar] [CrossRef] [PubMed]
- Crump, J.R.; Elner, S.G.; Elner, V.M.; Kauffman, C.A. Cryptococcal endophthalmitis: Case report and review. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 1992, 14, 1069–1073. [Google Scholar] [CrossRef]
- Sharma, M.; Khatib, R.; Jones, B.A.; Fakih, M.G. Cryptococcus neoformans myositis without dissemination. Scand. J. Infect. Dis. 2002, 34, 858–859. [Google Scholar] [CrossRef]
- Noguchi, H.; Matsumoto, T.; Kimura, U.; Hiruma, M.; Kusuhara, M.; Ihn, H. Cutaneous Cryptococcosis. Med. Mycol. J. 2019, 60, 101–107. [Google Scholar] [CrossRef]
- Zhou, H.X.; Lu, L.; Chu, T.; Wang, T.; Cao, D.; Li, F.; Ning, G.; Feng, S. Skeletal cryptococcosis from 1977 to 2013. Front. Microbiol. 2014, 5, 740. [Google Scholar] [CrossRef]
- Cuomo, C.A.; Rhodes, J.; Desjardins, C.A. Advances in Cryptococcus genomics: Insights into the evolution of pathogenesis. Mem. Do Inst. Oswaldo Cruz. 2018, 113, e170473. [Google Scholar] [CrossRef] [PubMed]
- Esher, S.K.; Zaragoza, O.; Alspaugh, J.A. Cryptococcal pathogenic mechanisms: A dangerous trip from the environment to the brain. Mem. Do Inst. Oswaldo Cruz 2018, 113, e180057. [Google Scholar] [CrossRef] [PubMed]
- Litvintseva, A.P.; Mitchell, T.G. Most environmental isolates of Cryptococcus neoformans var. grubii (serotype A) are not lethal for mice. Infect. Immun. 2009, 77, 3188–3195. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, K.E.; Brockway, A.; Haverkamp, M.; Cuomo, C.A.; van Ogtrop, F.; Perfect, J.R.; Carter, D.A. Phenotypic Variability Correlates with Clinical Outcome in Cryptococcus Isolates Obtained from Botswanan HIV/AIDS Patients. mBio 2018, 9. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Boekhout, T.; Wickes, B.L.; Fell, J.W. Systematics of the Genus Cryptococcus and Its Type Species, C. neoformans. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Bennett, J.E.; Wickes, B.L.; Meyer, W.; Cuomo, C.A.; Wollenburg, K.R.; Bicanic, T.A.; Castañeda, E.; Chang, Y.C.; Chen, J.; et al. The Case for Adopting the “Species Complex” Nomenclature for the Etiologic Agents of Cryptococcosis. mSphere 2017, 2, e00357–e00316. [Google Scholar] [CrossRef] [PubMed]
- Farrer, R.A.; Chang, M.; Davis, M.J.; van Dorp, L.; Yang, D.H.; Shea, T.; Sewell, T.R.; Meyer, W.; Balloux, F.; Edwards, H.M.; et al. A New Lineage of Cryptococcus gattii (VGV) Discovered in the Central Zambezian Miombo Woodlands. mBio 2019, 10. [Google Scholar] [CrossRef]
- Meyer, W.; Aanensen, D.M.; Boekhout, T.; Cogliati, M.; Diaz, M.R.; Esposto, M.C.; Fisher, M.; Gilgado, F.; Hagen, F.; Kaocharoen, S.; et al. Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med. Mycol. 2009, 47, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Perfect, J.R.; Lang, S.D.; Durack, D.T. Chronic cryptococcal meningitis: A new experimental model in rabbits. Am. J. Pathol. 1980, 101, 177–194. [Google Scholar]
- Desjardins, C.A.; Giamberardino, C.; Sykes, S.M.; Yu, C.H.; Tenor, J.L.; Chen, Y.; Yang, T.; Jones, A.M.; Sun, S.; Haverkamp, M.R.; et al. Population genomics and the evolution of virulence in the fungal pathogen Cryptococcus neoformans. Genome Res. 2017, 27, 1207–1219. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Edman, J.C.; Wickes, B.L. Genetic association of mating types and virulence in Cryptococcus neoformans. Infect. Immun. 1992, 60, 602–605. [Google Scholar] [CrossRef]
- Farrer, R.A.; Desjardins, C.A.; Sakthikumar, S.; Gujja, S.; Saif, S.; Zeng, Q.; Chen, Y.; Voelz, K.; Heitman, J.; May, R.C.; et al. Genome Evolution and Innovation across the Four Major Lineages of Cryptococcus gattii. mBio 2015, 6, e00868-15. [Google Scholar] [CrossRef]
- Hagen, F.; Khayhan, K.; Theelen, B.; Kolecka, A.; Polacheck, I.; Sionov, E.; Falk, R.; Parnmen, S.; Lumbsch, H.T.; Boekhout, T. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal. Genet. Biol. 2015, 78, 16–48. [Google Scholar] [CrossRef] [PubMed]
- Velegraki, A.; Kiosses, V.G.; Pitsouni, H.; Toukas, D.; Daniilidis, V.D.; Legakis, N.J. First report of Cryptococcus neoformans var. gattii serotype B from Greece. Med. Mycol. 2001, 39, 419–422. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, C.A.; Kronstad, J.W.; Taylor, G.; Warren, R.; Yuen, M.; Hu, G.; Jung, W.H.; Sham, A.; Kidd, S.E.; Tangen, K.; et al. Genome Variation in Cryptococcus gattii, an Emerging Pathogen of Immunocompetent Hosts. mBio 2011, 2, e00342-10. [Google Scholar] [CrossRef]
- Kidd, S.E.; Hagen, F.; Tscharke, R.L.; Huynh, M.; Bartlett, K.H.; Fyfe, M.; Macdougall, L.; Boekhout, T.; Kwon-Chung, K.J.; Meyer, W. A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc. Natl. Acad. Sci. USA 2004, 101, 17258–17263. [Google Scholar] [CrossRef] [PubMed]
- Varma, A.; Kwon-Chung, K.J. Heteroresistance of Cryptococcus gattii to fluconazole. Antimicrob Agents Chemother 2010, 54, 2303–2311. [Google Scholar] [CrossRef] [PubMed]
- Probert, M.; Zhou, X.; Goodall, M.; Johnston, S.A.; Bielska, E.; Ballou, E.R.; May, R.C. A Glucuronoxylomannan Epitope Exhibits Serotype-Specific Accessibility and Redistributes towards the Capsule Surface during Titanization of the Fungal Pathogen Cryptococcus neoformans. Infect. Immun. 2019, 87, e00731-18. [Google Scholar] [CrossRef]
- Meyer, W.; Castañeda, A.; Jackson, S.; Huynh, M.; Castañeda, E. Molecular typing of IberoAmerican Cryptococcus neoformans isolates. Emerg. Infect. Dis. 2003, 9, 189–195. [Google Scholar] [CrossRef]
- Janbon, G.; Doering, T.L. Biosynthesis and Genetics of the Cryptococcus Capsule. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Rodrigues, M.L.; Casadevall, A.; Zaragoza, O. The Architecture and Antigenic Composition of the Polysaccharide Capsule. Cryptococcus 2011. [Google Scholar] [CrossRef]
- O’Meara, T.R.; Alspaugh, J.A. The Cryptococcus neoformans capsule: A sword and a shield. Clin. Microbiol. Rev. 2012, 25, 387–408. [Google Scholar] [CrossRef]
- Wang, Z.A.; Li, L.X.; Doering, T.L. Unraveling synthesis of the cryptococcal cell wall and capsule. Glycobiology 2018, 28, 719–730. [Google Scholar] [CrossRef] [PubMed]
- Clancy, C.J.; Nguyen, M.H.; Alandoerffer, R.; Cheng, S.; Iczkowski, K.; Richardson, M.; Graybill, J.R. Cryptococcus neoformans var. grubii isolates recovered from persons with AIDS demonstrate a wide range of virulence during murine meningoencephalitis that correlates with the expression of certain virulence factors. Microbiology 2006, 152, 2247–2255. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liaw, S.J.; Wu, H.C.; Hsueh, P.R. Microbiological characteristics of clinical isolates of Cryptococcus neoformans in Taiwan: Serotypes, mating types, molecular types, virulence factors, and antifungal susceptibility. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2010, 16, 696–703. [Google Scholar] [CrossRef] [PubMed]
- Robertson, E.J.; Najjuka, G.; Rolfes, M.A.; Akampurira, A.; Jain, N.; Anantharanjit, J.; von Hohenberg, M.; Tassieri, M.; Carlsson, A.; Meya, D.B.; et al. Cryptococcus neoformans ex vivo capsule size is associated with intracranial pressure and host immune response in HIV-associated cryptococcal meningitis. J. Infect. Dis. 2014, 209, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Sabiiti, W.; Robertson, E.; Beale, M.A.; Johnston, S.A.; Brouwer, A.E.; Loyse, A.; Jarvis, J.N.; Gilbert, A.S.; Fisher, M.C.; Harrison, T.S.; et al. Efficient phagocytosis and laccase activity affect the outcome of HIV-associated cryptococcosis. J. Clin. Investig. 2014, 124, 2000–2008. [Google Scholar] [CrossRef]
- Denham, S.T.; Verma, S.; Reynolds, R.C.; Worne, C.L.; Daugherty, J.M.; Lane, T.E.; Brown, J.C.S. Regulated Release of Cryptococcal Polysaccharide Drives Virulence and Suppresses Immune Cell Infiltration into the Central Nervous System. Infect. Immun. 2018, 86, e00662-17. [Google Scholar] [CrossRef]
- Chang, Y.C.; Kwon-Chung, K.J. Complementation of a capsule-deficient mutation of Cryptococcus neoformans restores its virulence. Mol. Cell. Biol. 1994, 14, 4912–4919. [Google Scholar] [CrossRef]
- Chang, Y.C.; Penoyer, L.A.; Kwon-Chung, K.J. The second capsule gene of Cryptococcus neoformans, CAP64, is essential for virulence. Infect. Immun. 1996, 64, 1977–1983. [Google Scholar] [CrossRef]
- Chang, Y.C.; Kwon-Chung, K.J. Isolation of the third capsule-associated gene, CAP60, required for virulence in Cryptococcus neoformans. Infect. Immun. 1998, 66, 2230–2236. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.C.; Kwon-Chung, K.J. Isolation, characterization, and localization of a capsule-associated gene, CAP10, of Cryptococcus neoformans. J. Bacteriol. 1999, 181, 5636–5643. [Google Scholar] [CrossRef] [PubMed]
- Zaragoza, O.; Chrisman, C.J.; Castelli, M.V.; Frases, S.; Cuenca-Estrella, M.; Rodríguez-Tudela, J.L.; Casadevall, A. Capsule enlargement in Cryptococcus neoformans confers resistance to oxidative stress suggesting a mechanism for intracellular survival. Cell. Microbiol. 2008, 10, 2043–2057. [Google Scholar] [CrossRef] [PubMed]
- Johnston, S.A.; May, R.C. Cryptococcus interactions with macrophages: Evasion and manipulation of the phagosome by a fungal pathogen. Cell. Microbiol. 2013, 15, 403–411. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, N.M.; Lodge, J.K.; Specht, C.A. The Cell Wall of Cryptococcus. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Trofa, D.; Casadevall, A.; Nosanchuk, J.D. Melanin: Structure, Function, and Biosynthesis in Cryptococcus. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Frazão, S.O.; Sousa, H.R.; Silva, L.G.D.; Folha, J.D.S.; Gorgonha, K.C.M.; Oliveira, G.P., Jr.; Felipe, M.S.S.; Silva-Pereira, I.; Casadevall, A.; Nicola, A.M.; et al. Laccase Affects the Rate of Cryptococcus neoformans Nonlytic Exocytosis from Macrophages. mBio 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A.; Coelho, C.; Alanio, A. Mechanisms of Cryptococcus neoformans-Mediated Host Damage. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Ghannoum, M.A. Potential Role of Phospholipases in Virulence and Fungal Pathogenesis. Clin. Microbiol. Rev. 2000, 13, 122–143. [Google Scholar] [CrossRef]
- Cox, G.M.; Mukherjee, J.; Cole, G.T.; Casadevall, A.; Perfect, J.R. Urease as a virulence factor in experimental cryptococcosis. Infect. Immun. 2000, 68, 443–448. [Google Scholar] [CrossRef]
- Ganendren, R.; Carter, E.; Sorrell, T.; Widmer, F.; Wright, L. Phospholipase B activity enhances adhesion of Cryptococcus neoformans to a human lung epithelial cell line. Microbes Infect. 2006, 8, 1006–1015. [Google Scholar] [CrossRef]
- Djordjevic, J.T. Role of phospholipases in fungal fitness, pathogenicity, and drug development-lessons from Cryptococcus neoformans. Front. Microbiol. 2010, 1, 125. [Google Scholar] [CrossRef]
- Cox, G.M.; McDade, H.C.; Chen, S.C.; Tucker, S.C.; Gottfredsson, M.; Wright, L.C.; Sorrell, T.C.; Leidich, S.D.; Casadevall, A.; Ghannoum, M.A.; et al. Extracellular phospholipase activity is a virulence factor for Cryptococcus neoformans. Mol. Microbiol. 2001, 39, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, R.; Zoellner, H.; Sorrell, T.; Wilson, C.; Donald, C.; Djordjevic, J.; Shounan, Y.; Wright, L. Role of extracellular phospholipases and mononuclear phagocytes in dissemination of cryptococcosis in a murine model. Infect. Immun. 2004, 72, 2229–2239. [Google Scholar] [CrossRef] [PubMed]
- Olszewski, M.A.; Noverr, M.C.; Chen, G.H.; Toews, G.B.; Cox, G.M.; Perfect, J.R.; Huffnagle, G.B. Urease expression by Cryptococcus neoformans promotes microvascular sequestration, thereby enhancing central nervous system invasion. Am. J. Pathol. 2004, 164, 1761–1771. [Google Scholar] [CrossRef]
- Love, G.L.; Boyd, G.D.; Greer, D.L. Large Cryptococcus neoformans isolated from brain abscess. J. Clin. Microbiol. 1985, 22, 1068–1070. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.M.; Zhou, Q.; Cai, H.R.; Zhuang, Y.; Zhang, Y.F.; Xin, X.Y.; Meng, F.Q.; Wang, Y.P. Clinicopathological features of pulmonary cryptococcosis with cryptococcal titan cells: A comparative analysis of 27 cases. Int. J. Clin. Exp. Pathol. 2014, 7, 4837–4846. [Google Scholar]
- Gerstein, A.C.; Fu, M.S.; Mukaremera, L.; Li, Z.; Ormerod, K.L.; Fraser, J.A.; Berman, J.; Nielsen, K. Polyploid titan cells produce haploid and aneuploid progeny to promote stress adaptation. mBio 2015, 6, e01340-15. [Google Scholar] [CrossRef] [PubMed]
- Mukaremera, L.; Lee, K.K.; Wagener, J.; Wiesner, D.L.; Gow, N.A.R.; Nielsen, K. Titan cell production in Cryptococcus neoformans reshapes the cell wall and capsule composition during infection. Cell Surf. (Amst. Neth.) 2018, 1, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Okagaki, L.H.; Nielsen, K. Titan cells confer protection from phagocytosis in Cryptococcus neoformans infections. Eukaryot. Cell 2012, 11, 820–826. [Google Scholar] [CrossRef]
- Crabtree, J.N.; Okagaki, L.H.; Wiesner, D.L.; Strain, A.K.; Nielsen, J.N.; Nielsen, K. Titan cell production enhances the virulence of Cryptococcus neoformans. Infect. Immun. 2012, 80, 3776–3785. [Google Scholar] [CrossRef]
- Feldmesser, M.; Kress, Y.; Casadevall, A. Dynamic changes in the morphology of Cryptococcus neoformans during murine pulmonary infection. Microbiology 2001, 147, 2355–2365. [Google Scholar] [CrossRef]
- Xie, S.; Sao, R.; Braun, A.; Bottone, E.J. Difference in Cryptococcus neoformans cellular and capsule size in sequential pulmonary and meningeal infection: A postmortem study. Diagn. Microbiol. Infect. Dis. 2012, 73, 49–52. [Google Scholar] [CrossRef]
- Skipper, C.; Abassi, M.; Boulware, D.R. Diagnosis and Management of Central Nervous System Cryptococcal Infections in HIV-Infected Adults. J. Fungi (Baselswitzerland) 2019, 5, 65. [Google Scholar] [CrossRef]
- Diaz, M.R.; Nguyen, M.H. Diagnostic Approach Based on Capsular Antigen, Capsule Detection, β-Glucan, and DNA Analysis. Cryptococcus 2011. [Google Scholar] [CrossRef]
- Nalintya, E.; Kiggundu, R.; Meya, D. Evolution of Cryptococcal Antigen Testing: What is new? Curr. Fungal Infect. Rep. 2016, 10, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Rajasingham, R.; Wake, R.M.; Beyene, T.; Katende, A.; Letang, E.; Boulware, D.R. Cryptococcal Meningitis Diagnostics and Screening in the Era of Point-of-Care Laboratory Testing. J. Clin. Microbiol. 2019, 57, e01238-01218. [Google Scholar] [CrossRef] [PubMed]
- Saha, D.C.; Xess, I.; Jain, N. Evaluation of conventional & serological methods for rapid diagnosis of cryptococcosis. Indian J. Med. Res. 2008, 127, 483–488. [Google Scholar]
- Boulware, D.R.; Rolfes, M.A.; Rajasingham, R.; von Hohenberg, M.; Qin, Z.; Taseera, K.; Schutz, C.; Kwizera, R.; Butler, E.K.; Meintjes, G.; et al. Multisite validation of cryptococcal antigen lateral flow assay and quantification by laser thermal contrast. Emerg. Infect. Dis. 2014, 20, 45–53. [Google Scholar] [CrossRef]
- Snow, R.M.; Dismukes, W.E. Cryptococcal Meningitis: Diagnostic Value of Cryptococcal Antigen in Cerebrospinal Fluid. Arch. Intern. Med. 1975, 135, 1155–1157. [Google Scholar] [CrossRef]
- Cohen, J. Comparison of the sensitivity of three methods for the rapid identification of Cryptococcus neoformans. J. Clin. Pathol. 1984, 37, 332–334. [Google Scholar] [CrossRef]
- Marcos, J.Y.; Pincus, D.H. Fungal diagnostics: Review of commercially available methods. Methods Mol. Biol. (Cliftonn.J.) 2013, 968, 25–54. [Google Scholar] [CrossRef]
- Jarvis, J.N.; Bicanic, T.; Loyse, A.; Namarika, D.; Jackson, A.; Nussbaum, J.C.; Longley, N.; Muzoora, C.; Phulusa, J.; Taseera, K.; et al. Determinants of mortality in a combined cohort of 501 patients with HIV-associated Cryptococcal meningitis: Implications for improving outcomes. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2014, 58, 736–745. [Google Scholar] [CrossRef] [PubMed]
- Boulware, D.R.; Meya, D.B.; Muzoora, C.; Rolfes, M.A.; Huppler Hullsiek, K.; Musubire, A.; Taseera, K.; Nabeta, H.W.; Schutz, C.; Williams, D.A.; et al. Timing of antiretroviral therapy after diagnosis of cryptococcal meningitis. N. Engl. J. Med. 2014, 370, 2487–2498. [Google Scholar] [CrossRef]
- Bicanic, T.; Meintjes, G.; Wood, R.; Hayes, M.; Rebe, K.; Bekker, L.G.; Harrison, T. Fungal burden, early fungicidal activity, and outcome in cryptococcal meningitis in antiretroviral-naive or antiretroviral-experienced patients treated with amphotericin B or fluconazole. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2007, 45, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Montezuma-Rusca, J.M.; Powers, J.H.; Follmann, D.; Wang, J.; Sullivan, B.; Williamson, P.R. Early Fungicidal Activity as a Candidate Surrogate Endpoint for All-Cause Mortality in Cryptococcal Meningitis: A Systematic Review of the Evidence. PLoS ONE 2016, 11, e0159727. [Google Scholar] [CrossRef] [PubMed]
- Temfack, E.; Rim, J.J.B.; Spijker, R.; Loyse, A.; Chiller, T.; Pappas, P.G.; Perfect, J.; Sorell, T.C.; Harrison, T.S.; Cohen, J.F.; et al. Cryptococcal antigen in serum and cerebrospinal fluid for detecting cryptococcal meningitis in adults living with HIV: Systematic review and meta-analysis of diagnostic test accuracy studies. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2020. [Google Scholar] [CrossRef]
- Rajasingham, R.; Meya, D.B.; Boulware, D.R. Integrating cryptococcal antigen screening and pre-emptive treatment into routine HIV care. J. Acquir. Immune Defic. Syndr. 2012, 59, e85–e91. [Google Scholar] [CrossRef]
- Meya, D.B.; Kiragga, A.N.; Nalintya, E.; Morawski, B.M.; Rajasingham, R.; Park, B.J.; Mubiru, A.; Kaplan, J.E.; Manabe, Y.C.; Boulware, D.R. Reflexive Laboratory-Based Cryptococcal Antigen Screening and Preemptive Fluconazole Therapy for Cryptococcal Antigenemia in HIV-Infected Individuals With CD4 < 100 Cells/µL: A Stepped-Wedge, Cluster-Randomized Trial. J. Acquir. Immune Defic. Syndr. 2019, 80, 182–189. [Google Scholar] [CrossRef]
- Wake, R.M.; Britz, E.; Sriruttan, C.; Rukasha, I.; Omar, T.; Spencer, D.C.; Nel, J.S.; Mashamaite, S.; Adelekan, A.; Chiller, T.M.; et al. High Cryptococcal Antigen Titers in Blood Are Predictive of Subclinical Cryptococcal Meningitis Among Human Immunodeficiency Virus-Infected Patients. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2018, 66, 686–692. [Google Scholar] [CrossRef]
- Beyene, T.; Zewde, A.G.; Balcha, A.; Hirpo, B.; Yitbarik, T.; Gebissa, T.; Rajasingham, R.; Boulware, D.R. Inadequacy of High-Dose Fluconazole Monotherapy Among Cerebrospinal Fluid Cryptococcal Antigen (CrAg)-Positive Human Immunodeficiency Virus-Infected Persons in an Ethiopian CrAg Screening Program. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2017, 65, 2126–2129. [Google Scholar] [CrossRef] [PubMed]
- Nalintya, E.; Meya, D.B.; Lofgren, S.; Huppler Hullsiek, K.; Boulware, D.R.; Rajasingham, R. A Prospective Evaluation of a Multisite Cryptococcal Screening and Treatment Program in HIV Clinics in Uganda. J. Acquir. Immune Defic. Syndr. 2018, 78, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, S.W.; Magambo, K.A.; Kalluvya, S.E.; Fitzgerald, D.W.; Peck, R.N.; Downs, J.A. Six-month outcomes of HIV-infected patients given short-course fluconazole therapy for asymptomatic cryptococcal antigenemia. Aids (Lond. Engl.) 2015, 29, 2473–2478. [Google Scholar] [CrossRef][Green Version]
- Longley, N.; Jarvis, J.N.; Meintjes, G.; Boulle, A.; Cross, A.; Kelly, N.; Govender, N.P.; Bekker, L.G.; Wood, R.; Harrison, T.S. Cryptococcal Antigen Screening in Patients Initiating ART in South Africa: A Prospective Cohort Study. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2016, 62, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Denning, D.W.; Armstrong, R.W.; Lewis, B.H.; Stevens, D.A. Elevated cerebrospinal fluid pressures in patients with cryptococcal meningitis and acquired immunodeficiency syndrome. Am. J. Med. 1991, 91, 267–272. [Google Scholar] [CrossRef]
- Bicanic, T.; Brouwer, A.E.; Meintjes, G.; Rebe, K.; Limmathurotsakul, D.; Chierakul, W.; Teparrakkul, P.; Loyse, A.; White, N.J.; Wood, R.; et al. Relationship of cerebrospinal fluid pressure, fungal burden and outcome in patients with cryptococcal meningitis undergoing serial lumbar punctures. Aids (Lond. Engl.) 2009, 23, 701–706. [Google Scholar] [CrossRef]
- Rosen, L.B.; Freeman, A.F.; Yang, L.M.; Jutivorakool, K.; Olivier, K.N.; Angkasekwinai, N.; Suputtamongkol, Y.; Bennett, J.E.; Pyrgos, V.; Williamson, P.R.; et al. Anti-GM-CSF autoantibodies in patients with cryptococcal meningitis. J. Immunol. (Baltim. Md.) 2013, 190, 3959–3966. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.; Jian, C.; Zhang, H.; Xu, S. Comparison of Clinical Features and Prognostic Factors of Cryptococcal Meningitis Caused by Cryptococcus neoformans in Patients With and Without Pulmonary Nodules. Mycopathologia 2019, 184, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Li, M.; Luo, Y.; Wang, D.; Yang, J.; Chen, Q.; Lao, J.; Chen, N.; Yang, M.; Wang, Q. A retrospective study of contributing factors for prognosis and survival length of cryptococcal meningoencephalitis in Southern part of China (1998–2013). BMC Infect. Dis. 2015, 15, 77. [Google Scholar] [CrossRef]
- Chen, S.C.A.; Meyer, W.; Sorrell, T.C. Cryptococcus gattii infections. Clin. Microbiol. Rev. 2014, 27, 980–1024. [Google Scholar] [CrossRef]
- Chen, S.C.; Slavin, M.A.; Heath, C.H.; Playford, E.G.; Byth, K.; Marriott, D.; Kidd, S.E.; Bak, N.; Currie, B.; Hajkowicz, K.; et al. Clinical manifestations of Cryptococcus gattii infection: Determinants of neurological sequelae and death. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2012, 55, 789–798. [Google Scholar] [CrossRef]
- Altamirano, S.; Jackson, K.M.; Nielsen, K. The interplay of phenotype and genotype in Cryptococcus neoformans disease. Biosci. Rep. 2020, 40. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.; Cox, G.M.; Litvintseva, A.P.; Mylonakis, E.; Malliaris, S.D.; Benjamin, D.K., Jr.; Giles, S.S.; Mitchell, T.G.; Casadevall, A.; Perfect, J.R.; et al. Cryptococcus neoformans α strains preferentially disseminate to the central nervous system during coinfection. Infect. Immun. 2005, 73, 4922–4933. [Google Scholar] [CrossRef]
- Chang, Y.C.; Wickes, B.L.; Miller, G.F.; Penoyer, L.A.; Kwon-Chung, K.J. Cryptococcus neoformans STE12alpha regulates virulence but is not essential for mating. J. Exp. Med. 2000, 191, 871–882. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Bennett, J.E. Distribution of alpha and alpha mating types of Cryptococcus neoformans among natural and clinical isolates. Am. J. Epidemiol. 1978, 108, 337–340. [Google Scholar] [CrossRef]
- Wickes, B.L. The role of mating type and morphology in Cryptococcus neoformans pathogenesis. Int. J. Med Microbiol. Ijmm 2002, 292, 313–329. [Google Scholar] [CrossRef]
- Prakash, A.; Sundar, G.; Sharma, B.; Hagen, F.; Meis, J.F.; Chowdhary, A. Genotypic diversity in clinical and environmental isolates of Cryptococcus neoformans from India using multilocus microsatellite and multilocus sequence typing. Mycoses 2020, 63, 284–293. [Google Scholar] [CrossRef]
- Wongsuk, T.; Homkaew, A.; Faksri, K.; Thongnak, C. Multi-locus Sequence Typing and Whole Genome Sequence Analysis of Cryptococcus neoformans Isolated from Clinical Specimens in Vajira Hospital, Bangkok, Thailand. Mycopathologia 2020, 185, 503–514. [Google Scholar] [CrossRef]
- Nielsen, K.; Marra, R.E.; Hagen, F.; Boekhout, T.; Mitchell, T.G.; Cox, G.M.; Heitman, J. Interaction Between Genetic Background and the Mating-Type Locus in Cryptococcus neoformans Virulence Potential. Genetics 2005, 171, 975–983. [Google Scholar] [CrossRef]
- Lin, X.; Nielsen, K.; Patel, S.; Heitman, J. Impact of Mating Type, Serotype, and Ploidy on the Virulence of Cryptococcus neoformans. Infect. Immun. 2008, 76, 2923–2938. [Google Scholar] [CrossRef] [PubMed]
- Desnos-Ollivier, M.; Patel, S.; Raoux-Barbot, D.; Heitman, J.; Dromer, F. Cryptococcosis Serotypes Impact Outcome and Provide Evidence of Cryptococcus neoformans Speciation. mBio 2015, 6, e00311–e00315. [Google Scholar] [CrossRef] [PubMed]
- Wiesner, D.L.; Moskalenko, O.; Corcoran, J.M.; McDonald, T.; Rolfes, M.A.; Meya, D.B.; Kajumbula, H.; Kambugu, A.; Bohjanen, P.R.; Knight, J.F.; et al. Cryptococcal genotype influences immunologic response and human clinical outcome after meningitis. mBio 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Beale, M.A.; Sabiiti, W.; Robertson, E.J.; Fuentes-Cabrejo, K.M.; O’Hanlon, S.J.; Jarvis, J.N.; Loyse, A.; Meintjes, G.; Harrison, T.S.; May, R.C.; et al. Genotypic Diversity Is Associated with Clinical Outcome and Phenotype in Cryptococcal Meningitis across Southern Africa. PLoS Negl. Trop. Dis. 2015, 9, e0003847. [Google Scholar] [CrossRef]
- Xess, I.; Pandey, M.; Dabas, Y.; Agarwal, R.; Das, S.; Srivastava, P.M.V.; Thakur, R.; Sharma, S.; Mani, P.; Biswas, A.; et al. Multilocus Sequence Typing of Clinical Isolates of Cryptococcus from India. Mycopathologia 2021. [Google Scholar] [CrossRef]
- Ponzio, V.; Chen, Y.; Rodrigues, A.M.; Tenor, J.L.; Toffaletti, D.L.; Medina-Pestana, J.O.; Colombo, A.L.; Perfect, J.R. Genotypic diversity and clinical outcome of cryptococcosis in renal transplant recipients in Brazil. Emerg. Microbes Infect. 2019, 8, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Mukaremera, L.; McDonald, T.R.; Nielsen, J.N.; Molenaar, C.J.; Akampurira, A.; Schutz, C.; Taseera, K.; Muzoora, C.; Meintjes, G.; Meya, D.B.; et al. The Mouse Inhalation Model of Cryptococcus neoformans Infection Recapitulates Strain Virulence in Humans and Shows that Closely Related Strains Can Possess Differential Virulence. Infect. Immun. 2019, 87. [Google Scholar] [CrossRef]
- Brito-Santos, F.; Trilles, L.; Firacative, C.; Wanke, B.; Carvalho-Costa, F.A.; Nishikawa, M.M.; Campos, J.P.; Junqueira, A.C.V.; Souza, A.C.; Lazéra, M.D.S.; et al. Indoor Dust as a Source of Virulent Strains of the Agents of Cryptococcosis in the Rio Negro Micro-Region of the Brazilian Amazon. Microorganisms 2020, 8, 682. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wang, Y.; Li, Y.; Hong, N.; Zhu, X.; Pan, W.; Liao, W.; Xu, J.; Du, J.; Chen, J. Genotypic diversity and antifungal susceptibility of environmental isolates of Cryptococcus neoformans from the Yangtze River Delta region of East China. Med. Mycol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.H.; Yu, F.; Bian, Z.Y.; Hong, J.M.; Zhang, N.; Zhong, Q.S.; Hang, Y.P.; Xu, J.; Hu, L.H. Multilocus Sequence Typing Reveals both Shared and Unique Genotypes of Cryptococcus neoformans in Jiangxi Province, China. Sci. Rep. 2018, 8, 1495. [Google Scholar] [CrossRef]
- Corrêa Pinheiro, M.; Dos Reis, D.S.T.; de Brito, M.; Simões Quaresma, J.A. Cryptococcosis in the Amazon: A current overview and future perspectives. Acta Trop. 2019, 197, 105023. [Google Scholar] [CrossRef]
- Day, J.N.; Qihui, S.; Thanh, L.T.; Trieu, P.H.; Van, A.D.; Thu, N.H.; Chau, T.T.H.; Lan, N.P.H.; Chau, N.V.V.; Ashton, P.M.; et al. Comparative genomics of Cryptococcus neoformans var. grubii associated with meningitis in HIV infected and uninfected patients in Vietnam. PLoS Negl. Trop. Dis. 2017, 11, e0005628. [Google Scholar] [CrossRef]
- Dou, H.T.; Xu, Y.C.; Wang, H.Z.; Li, T.S. Molecular epidemiology of Cryptococcus neoformans and Cryptococcus gattii in China between 2007 and 2013 using multilocus sequence typing and the DiversiLab system. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2015, 34, 753–762. [Google Scholar] [CrossRef]
- Fang, L.F.; Zhang, P.P.; Wang, J.; Yang, Q.; Qu, T.T. Clinical and microbiological characteristics of cryptococcosis at an university hospital in China from 2013 to Braz. J. Infect. Dis. Off. Publ. Braz. Soc. Infect. Dis. 2020, 24, 7–12. [Google Scholar] [CrossRef]
- Hatthakaroon, C.; Pharkjaksu, S.; Chongtrakool, P.; Suwannakarn, K.; Kiratisin, P.; Ngamskulrungroj, P. Molecular epidemiology of cryptococcal genotype VNIc/ST5 in Siriraj Hospital, Thailand. PLoS ONE 2017, 12, e0173744. [Google Scholar] [CrossRef]
- Hong, N.; Chen, M.; Xu, N.; Al-Hatmi, A.M.S.; Zhang, C.; Pan, W.H.; Hagen, F.; Boekhout, T.; Xu, J.; Zou, X.B.; et al. Genotypic diversity and antifungal susceptibility of Cryptococcus neoformans isolates from paediatric patients in China. Mycoses 2019, 62, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zou, M.; Yin, J.; Liu, Z.; Lu, B. Microbiological, Epidemiological, and Clinical Characteristics of Patients With Cryptococcal Meningitis at a Tertiary Hospital in China: A 6-Year Retrospective Analysis. Front. Microbiol. 2020, 11, 1837. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Choi, S.C.; Lee, K.W.; Kim, M.N.; Hwang, S.M. Genotypes of Clinical and Environmental Isolates of Cryptococcus neoformans and Cryptococcus gattii in Korea. Mycobiology 2015, 43, 360–365. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Kim, M.; Joo, S.I.; Hwang, S.M. Molecular Epidemiology of Clinical Cryptococcus neoformans Isolates in Seoul, Korea. Mycobiology 2014, 42, 73–78. [Google Scholar] [CrossRef]
- Pharkjaksu, S.; Kwon-Chung, K.J.; Bennett, J.E.; Ngamskulrungroj, P. Population diversity and virulence characteristics of Cryptococcus neoformans/C. gattii species complexes isolated during the pre-HIV-pandemic era. PLoS Negl. Trop. Dis. 2020, 14, e0008651. [Google Scholar] [CrossRef]
- Pinheiro, S.B.; Sousa, E.S.; Cortez, A.C.A.; da Silva Rocha, D.F.; Menescal, L.S.F.; Chagas, V.S.; Gómez, A.S.P.; Cruz, K.S.; Santos, L.O.; Alves, M.J.; et al. Cryptococcal meningitis in non-HIV patients in the State of Amazonas, Northern Brazil. Braz. J. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Rocha, D.F.S.; Cruz, K.S.; Santos, C.; Menescal, L.S.F.; Neto, J.; Pinheiro, S.B.; Silva, L.M.; Trilles, L.; Braga de Souza, J.V. MLST reveals a clonal population structure for Cryptococcus neoformans molecular type VNI isolates from clinical sources in Amazonas, Northern-Brazil. PLoS ONE 2018, 13, e0197841. [Google Scholar] [CrossRef]
- Thanh, L.T.; Phan, T.H.; Rattanavong, S.; Nguyen, T.M.; Duong, A.V.; Dacon, C.; Hoang, T.N.; Nguyen, L.P.H.; Tran, C.T.H.; Davong, V.; et al. Multilocus sequence typing of Cryptococcus neoformans var. grubii from Laos in a regional and global context. Med. Mycol. 2018, 57, 557–565. [Google Scholar] [CrossRef] [PubMed]
- Thanh, L.T.; Toffaletti, D.L.; Tenor, J.L.; Giamberardino, C.; Sempowski, G.D.; Asfaw, Y.; Phan, H.T.; Van Duong, A.; Trinh, N.M.; Thwaites, G.E.; et al. Assessing the virulence of Cryptococcus neoformans causing meningitis in HIV infected and uninfected patients in Vietnam. Med. Mycol. 2020, 58, 1149–1161. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hong, J.J.; Zhang, P.P.; Yang, M.F.; Yang, Q.; Qu, T.T. Cryptococcal pleuritis with pleural effusion as the only clinical presentation in a patient with hepatic cirrhosis: A case report and literature review. Medicine 2019, 98, e16354. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.Y.; Kang, M.; Liu, Y.; Chen, Z.X.; Xiao, Y.L.; He, C.; Ma, Y. Molecular epidemiology and antifungal susceptibilities of Cryptococcus species isolates from HIV and non-HIV patients in Southwest China. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2021, 40, 287–295. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.Y.; Lei, Y.; Kang, M.; Xiao, Y.L.; Chen, Z.X. Molecular characterisation of clinical Cryptococcus neoformans and Cryptococcus gattii isolates from Sichuan province, China. Mycoses 2015, 58, 280–287. [Google Scholar] [CrossRef]
- Ferreira-Paim, K.; Andrade-Silva, L.; Fonseca, F.M.; Ferreira, T.B.; Mora, D.J.; Andrade-Silva, J.; Khan, A.; Dao, A.; Reis, E.C.; Almeida, M.T.; et al. MLST-Based Population Genetic Analysis in a Global Context Reveals Clonality amongst Cryptococcus neoformans var. grubii VNI Isolates from HIV Patients in Southeastern Brazil. PLoS Negl. Trop. Dis. 2017, 11, e0005223. [Google Scholar] [CrossRef]
- Khayhan, K.; Hagen, F.; Pan, W.; Simwami, S.; Fisher, M.C.; Wahyuningsih, R.; Chakrabarti, A.; Chowdhary, A.; Ikeda, R.; Taj-Aldeen, S.J.; et al. Geographically structured populations of Cryptococcus neoformans Variety grubii in Asia correlate with HIV status and show a clonal population structure. PLoS ONE 2013, 8, e72222. [Google Scholar] [CrossRef]
- Gerstein, A.C.; Jackson, K.M.; McDonald, T.R.; Wang, Y.; Lueck, B.D.; Bohjanen, S.; Smith, K.D.; Akampurira, A.; Meya, D.B.; Xue, C.; et al. Identification of Pathogen Genomic Differences That Impact Human Immune Response and Disease during Cryptococcus neoformans Infection. mBio 2019, 10. [Google Scholar] [CrossRef]
- Alanio, A.; Desnos-Ollivier, M.; Dromer, F. Dynamics of Cryptococcus neoformans-macrophage interactions reveal that fungal background influences outcome during cryptococcal meningoencephalitis in humans. mBio 2011, 2. [Google Scholar] [CrossRef]
- Rivera, J.; Feldmesser, M.; Cammer, M.; Casadevall, A. Organ-Dependent Variation of Capsule Thickness inCryptococcus neoformans during Experimental Murine Infection. Infect. Immun. 1998, 66, 5027–5030. [Google Scholar] [CrossRef]
- Chen, Y.; Toffaletti, D.L.; Tenor, J.L.; Litvintseva, A.P.; Fang, C.; Mitchell, T.G.; McDonald, T.R.; Nielsen, K.; Boulware, D.R.; Bicanic, T.; et al. The Cryptococcus neoformans transcriptome at the site of human meningitis. mBio 2014, 5, e01087-13. [Google Scholar] [CrossRef]
- Hansakon, A.; Mutthakalin, P.; Ngamskulrungroj, P.; Chayakulkeeree, M.; Angkasekwinai, P. Cryptococcus neoformans and Cryptococcus gattii clinical isolates from Thailand display diverse phenotypic interactions with macrophages. Virulence 2019, 10, 26–36. [Google Scholar] [CrossRef]
- Ma, H.; Hagen, F.; Stekel, D.J.; Johnston, S.A.; Sionov, E.; Falk, R.; Polacheck, I.; Boekhout, T.; May, R.C. The fatal fungal outbreak on Vancouver Island is characterized by enhanced intracellular parasitism driven by mitochondrial regulation. Proc. Natl. Acad. Sci. USA 2009, 106, 12980–12985. [Google Scholar] [CrossRef]
- Eisen, D.P.; Minchinton, R.M. Impact of mannose-binding lectin on susceptibility to infectious diseases. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2003, 37, 1496–1505. [Google Scholar] [CrossRef]
- Shen, L.; Zheng, J.; Wang, Y.; Zhu, M.; Zhu, H.; Cheng, Q.; Li, Q. Increased activity of the complement system in cerebrospinal fluid of the patients with Non-HIV Cryptococcal meningitis. BMC Infect. Dis. 2017, 17, 7. [Google Scholar] [CrossRef]
- Ou, X.T.; Wu, J.Q.; Zhu, L.P.; Guan, M.; Xu, B.; Hu, X.P.; Wang, X.; Weng, X.H. Genotypes coding for mannose-binding lectin deficiency correlated with cryptococcal meningitis in HIV-uninfected Chinese patients. J. Infect. Dis. 2011, 203, 1686–1691. [Google Scholar] [CrossRef] [PubMed]
- Eisen, D.P.; Dean, M.M.; O’Sullivan, M.V.; Heatley, S.; Minchinton, R.M. Mannose-binding lectin deficiency does not appear to predispose to cryptococcosis in non-immunocompromised patients. Med. Mycol. 2008, 46, 371–375. [Google Scholar] [CrossRef]
- Yanagisawa, K.; Wichukchinda, N.; Tsuchiya, N.; Yasunami, M.; Rojanawiwat, A.; Tanaka, H.; Saji, H.; Ogawa, Y.; Handa, H.; Pathipvanich, P.; et al. Deficiency of mannose-binding lectin is a risk of Pneumocystis jirovecii pneumonia in a natural history cohort of people living with HIV/AIDS in Northern Thailand. PLoS ONE 2020, 15, e0242438. [Google Scholar] [CrossRef] [PubMed]
- Iseki, M.; Anzo, M.; Yamashita, N.; Matsuo, N. Hyper-IgM immunodeficiency with disseminated cryptococcosis. Acta Paediatr. 1994, 83, 780–782. [Google Scholar] [CrossRef] [PubMed]
- Tabone, M.D.; Leverger, G.; Landman, J.; Aznar, C.; Boccon-Gibod, L.; Lasfargues, G. Disseminated lymphonodular cryptococcosis in a child with X-linked hyper-IgM immunodeficiency. Pediatric Infect. Dis. J. 1994, 13, 77–79. [Google Scholar] [CrossRef]
- Pacharn, P.; Phongsamart, W.; Boonyawat, B.; Jirapongsananuruk, O.; Visitsunthorn, N.; Chokephaibulkit, K. Disseminated cryptococcosis in two boys with novel mutation of CD40 Ligand-Associated X-linked hyper-IgM syndrome. Asian Pac. J. Allergy Immunol. 2018. [Google Scholar] [CrossRef]
- Suzuki, S.M.L.; Morelli, F.; Negri, M.; Bonfim-Mendonça, P.; Kioshima, É., S.; Salci, T.; Voidaleski, M.F.; Vicente, V.A.; Svidzinski, T. FATAL cryptococcal meningitis in a child with hyper-immunoglobulin M syndrome, with an emphasis on the agent. J. Mycol. Med. 2019, 29, 273–277. [Google Scholar] [CrossRef]
- Acker, K.P.; Fetch, A.; Schnell, S.A.; Hammond, J.; Herrera, C.; Niedt, G.; Ratner, A.J.; Lauren, C.T. Scalp Lesions in a Pediatric Patient with Hyper IgM Syndrome: Clinical and Histologic Mimicry of Cryptococcus neoformans Infection. J. Pediatrics 2018, 192, 256–258. [Google Scholar] [CrossRef]
- Escárcega-Barbosa, D.; Ortiz-Jiménez, M.P.; Juárez-García, J.; Miranda-Feria, A.J. [Hyper-IgM syndrome: Mucocutaneous lesions and neutropenia]. Rev. Alerg. Mex. 2002, 49, 57–59. [Google Scholar]
- Jo, E.K.; Kim, H.S.; Lee, M.Y.; Iseki, M.; Lee, J.H.; Song, C.H.; Park, J.K.; Hwang, T.J.; Kook, H. X-linked hyper-IgM syndrome associated with Cryptosporidium parvum and Cryptococcus neoformans infections: The first case with molecular diagnosis in Korea. J. Korean Med. Sci. 2002, 17, 116–120. [Google Scholar] [CrossRef]
- França, T.T.; Leite, L.F.B.; Maximo, T.A.; Lambert, C.G.; Zurro, N.B.; Forte, W.C.N.; Condino-Neto, A. A Novel de Novo Mutation in the CD40 Ligand Gene in a Patient With a Mild X-Linked Hyper-IgM Phenotype Initially Diagnosed as CVID: New Aspects of Old Diseases. Front. Pediatrics 2018, 6, 130. [Google Scholar] [CrossRef]
- Trevijano-Contador, N.; Pianalto, K.M.; Nichols, C.B.; Zaragoza, O.; Alspaugh, J.A.; Pirofski, L.-A. Human IgM Inhibits the Formation of Titan-Like Cells in Cryptococcus neoformans. Infect. Immun. 2020, 88, e00046-20. [Google Scholar] [CrossRef] [PubMed]
- Rohatgi, S.; Gohil, S.; Kuniholm, M.H.; Schultz, H.; Dufaud, C.; Armour, K.L.; Badri, S.; Mailliard, R.B.; Pirofski, L.-A. Fc Gamma Receptor 3A Polymorphism and Risk for HIV-Associated Cryptococcal Disease. mBio 2013, 4, e00573-13. [Google Scholar] [CrossRef] [PubMed]
- Meletiadis, J.; Walsh, T.J.; Hwa Choi, E.; Pappas, P.G.; Ennis, D.; Douglas, J.; Pankey, G.A.; Larsen, R.A.; Hamill, R.J.; Chanock, S. Study of common functional genetic polymorphisms of FCGR2A, 3A and 3B genes and the risk for cryptococcosis in HIV-uninfected patients. Med. Mycol. 2007, 45, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.P.; Wu, J.Q.; Zhu, L.P.; Wang, X.; Xu, B.; Wang, R.Y.; Ou, X.T.; Weng, X.H. Association of Fcγ receptor IIB polymorphism with cryptococcal meningitis in HIV-uninfected Chinese patients. PLoS ONE 2012, 7, e42439. [Google Scholar] [CrossRef]
- Hu, X.P.; Wang, R.Y.; Wang, X.; Cao, Y.H.; Chen, Y.Q.; Zhao, H.Z.; Wu, J.Q.; Weng, X.H.; Gao, X.H.; Sun, R.H.; et al. Dectin-2 polymorphism associated with pulmonary cryptococcosis in HIV-uninfected Chinese patients. Med. Mycol. 2015, 53, 810–816. [Google Scholar] [CrossRef]
- Kanazawa, N.; Tashiro, K.; Inaba, K.; Lutz, M.B.; Miyachi, Y. Molecular cloning of human dectin-2. J. Investig. Dermatol. 2004, 122, 1522–1524. [Google Scholar] [CrossRef]
- Willment, J.A.; Brown, G.D. C-type lectin receptors in antifungal immunity. Trends Microbiol. 2008, 16, 27–32. [Google Scholar] [CrossRef]
- Bottazzi, B.; Garlanda, C.; Salvatori, G.; Jeannin, P.; Manfredi, A.; Mantovani, A. Pentraxins as a key component of innate immunity. Curr. Opin. Immunol. 2006, 18, 10–15. [Google Scholar] [CrossRef]
- Zhang, W.; Liao, Q.; Liu, Y.; Wu, S.; Deng, J.; Xiao, Y.; Ma, Y.; Xie, Y.; Kang, M. PTX3 gene polymorphism associated with cryptococcosis in HIV-uninfected Chinese patients. Mycoses 2020. [Google Scholar] [CrossRef]
- Tang, T.; Dai, Y.; Zeng, Q.; Bu, S.; Huang, B.; Xiao, Y.; Wei, Z.; Lin, X.; Huang, L.; Jiang, S. Pentraxin-3 polymorphisms and pulmonary fungal disease in non-neutropenic patients. Ann. Transl. Med. 2020, 8, 1142. [Google Scholar] [CrossRef] [PubMed]
- Jirapongsananuruk, O.; Luangwedchakarn, V.; Niemela, J.E.; Pacharn, P.; Visitsunthorn, N.; Thepthai, C.; Vichyanond, P.; Piboonpocanun, S.; Fleisher, T.A. Cryptococcal osteomyelitis in a child with a novel compound mutation of the IL12RB1 gene. Asian Pac. J. Allergy Immunol. 2012, 30, 79–82. [Google Scholar] [PubMed]
- Levitz, S.M. Activation of human peripheral blood mononuclear cells by interleukin-2 and granulocyte-macrophage colony-stimulating factor to inhibit Cryptococcus neoformans. Infect. Immun. 1991, 59, 3393–3397. [Google Scholar] [CrossRef] [PubMed]
- Collins, H.L.; Bancroft, G.J. Cytokine enhancement of complement-dependent phagocytosis by macrophages: Synergy of tumor necrosis factor-alpha and granulocyte-macrophage colony-stimulating factor for phagocytosis of Cryptococcus neoformans. Eur. J. Immunol. 1992, 22, 1447–1454. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.H.; Curtis, J.L.; Mody, C.H.; Christensen, P.J.; Armstrong, L.R.; Toews, G.B. Effect of granulocyte-macrophage colony-stimulating factor on rat alveolar macrophage anticryptococcal activity in vitro. J. Immunol. (Baltim. Md.) 1994, 152, 724–734. [Google Scholar]
- Viola, G.M.; Malek, A.E.; Rosen, L.B.; DiNardo, A.R.; Nishiguchi, T.; Okhuysen, P.C.; Holland, S.M.; Kontoyiannis, D.P. Disseminated cryptococcosis and anti-granulocyte-macrophage colony-stimulating factor autoantibodies: An underappreciated association. Mycoses 2021. [Google Scholar] [CrossRef]
- Huynh, J.; Saddi, V.; Cooper, P.; Cheng, A.T.; Meyer, W.; Chen, S.; Isaacs, D. Unusual Presentation of Severe Endobronchial Obstruction Caused by Cryptococcus gattii in a Child. J. Pediatric Infect. Dis. Soc. 2020, 9, 67–70. [Google Scholar] [CrossRef]
- Applen Clancey, S.; Ciccone, E.J.; Coelho, M.A.; Davis, J.; Ding, L.; Betancourt, R.; Glaubiger, S.; Lee, Y.; Holland, S.M.; Gilligan, P.; et al. Cryptococcus deuterogattii VGIIa Infection Associated with Travel to the Pacific Northwest Outbreak Region in an Anti-Granulocyte-Macrophage Colony-Stimulating Factor Autoantibody-Positive Patient in the United States. mBio 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, B.; Bundell, C.; Mulrennan, S.; McLean-Tooke, A.; Murray, R.; Brusch, A. The significance of anti-granulocyte-macrophage colony-stimulating factor antibodies in cryptococcal infection: Case series and review of antibody testing. Intern. Med. J. 2019, 49, 1446–1450. [Google Scholar] [CrossRef]
- Kuo, C.Y.; Wang, S.Y.; Shih, H.P.; Tu, K.H.; Huang, W.C.; Ding, J.Y.; Lin, C.H.; Yeh, C.F.; Ho, M.W.; Chang, S.C.; et al. Disseminated Cryptococcosis Due to Anti-Granulocyte-Macrophage Colony-Stimulating Factor Autoantibodies in the Absence of Pulmonary Alveolar Proteinosis. J. Clin. Immunol. 2017, 37, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Guess, T.E.; Rosen, J.A.; McClelland, E.E. An Overview of Sex Bias in C. neoformans Infections. J. Fungi 2018, 4, 49. [Google Scholar] [CrossRef] [PubMed]
- McClelland, E.E.; Hobbs, L.M.; Rivera, J.; Casadevall, A.; Potts, W.K.; Smith, J.M.; Ory, J.J. The role of host gender in the pathogenesis of Cryptococcus neoformans infections. PLoS ONE 2013, 8, e63632. [Google Scholar] [CrossRef]
- Guess, T.E.; Rosen, J.; Castro-Lopez, N.; Wormley, F.L., Jr.; McClelland, E.E. An inherent T cell deficit in healthy males to C. neoformans infection may begin to explain the sex susceptibility in incidence of cryptococcosis. Biol. Sex Differ. 2019, 10, 44. [Google Scholar] [CrossRef]
- Tucker, J.S.; Guess, T.E.; McClelland, E.E. The Role of Testosterone and Gibberellic Acid in the Melanization of Cryptococcus neoformans. Front. Microbiol. 2020, 11, 1921. [Google Scholar] [CrossRef]
- Miglia, K.J.; Govender, N.P.; Rossouw, J.; Meiring, S.; Mitchell, T.G. Analyses of Pediatric Isolates of Cryptococcus neoformans from South Africa. J. Clin. Microbiol. 2011, 49, 307–314. [Google Scholar] [CrossRef]
- Liu, L.; Guo, L.; Liu, Y.; Chen, T.; Li, S.; Yang, Y.; Liu, G. Clinical characteristics and prognosis of pediatric cryptococcosis in Beijing Children’s Hospital, 2002–2014. Eur. J. Pediatrics 2017, 176, 1235–1244. [Google Scholar] [CrossRef]
- Lizarazo, J.; Escandón, P.; Agudelo, C.I.; Castañeda, E. Cryptococcosis in Colombian children and literature review. Mem. Do Inst. Oswaldo Cruz 2014, 109, 797–804. [Google Scholar] [CrossRef]
- Gao, L.W.; Jiao, A.X.; Wu, X.R.; Zhao, S.Y.; Ma, Y.; Liu, G.; Yin, J.; Xu, B.P.; Shen, K.L. Clinical characteristics of disseminated cryptococcosis in previously healthy children in China. BMC Infect. Dis. 2017, 17, 359. [Google Scholar] [CrossRef] [PubMed]
- Luo, F.L.; Tao, Y.H.; Wang, Y.M.; Li, H. Clinical study of 23 pediatric patients with cryptococcosis. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 3801–3810. [Google Scholar] [PubMed]
- Marwaha, R.K.; Trehan, A.; Jayashree, K.; Vasishta, R.K. Hypereosinophilia in disseminated cryptococcal disease. Pediatric Infect. Dis. J. 1995, 14, 1102–1103. [Google Scholar]
- Hsu, A.P.; Davis, J.; Puck, J.M.; Holland, S.M.; Freeman, A.F. STAT3 Hyper IgE Syndrome. In GeneReviews(®); Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Goldman, D.L.; Li, X.; Tsirilakis, K.; Andrade, C.; Casadevall, A.; Vicencio, A.G. Increased chitinase expression and fungal-specific antibodies in the bronchoalveolar lavage fluid of asthmatic children. Clin. Exp. Allergy J. Br. Soc. Allergy Clin. Immunol. 2012, 42, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Aslanyan, L.; Lee, H.H.; Ekhar, V.V.; Ramos, R.L.; Martinez, L.R. Methamphetamine Impairs IgG1-Mediated Phagocytosis and Killing of Cryptococcus neoformans by J774.16 Macrophage- and NR-9640 Microglia-Like Cells. Infect. Immun. 2019, 87. [Google Scholar] [CrossRef]
- Tallóczy, Z.; Martinez, J.; Joset, D.; Ray, Y.; Gácser, A.; Toussi, S.; Mizushima, N.; Nosanchuk, J.D.; Goldstein, H.; Loike, J.; et al. Methamphetamine inhibits antigen processing, presentation, and phagocytosis. PLoS Pathog. 2008, 4, e28. [Google Scholar] [CrossRef]
- Patel, D.; Desai, G.M.; Frases, S.; Cordero, R.J.B.; DeLeon-Rodriguez, C.M.; Eugenin, E.A.; Nosanchuk, J.D.; Martinez, L.R. Methamphetamine Enhances Cryptococcus neoformans Pulmonary Infection and Dissemination to the Brain. mBio 2013, 4, e00400-13. [Google Scholar] [CrossRef]
- Eugenin, E.A.; Greco, J.M.; Frases, S.; Nosanchuk, J.D.; Martinez, L.R. Methamphetamine alters blood brain barrier protein expression in mice, facilitating central nervous system infection by neurotropic Cryptococcus neoformans. J. Infect. Dis. 2013, 208, 699–704. [Google Scholar] [CrossRef]
- Dietrich, J.B. Alteration of blood-brain barrier function by methamphetamine and cocaine. Cell Tissue Res. 2009, 336, 385–392. [Google Scholar] [CrossRef]
- Polk, C.; Meredith, J.; Kuprenas, A.; Leonard, M. Cryptococcus meningitis mimicking cerebral septic emboli, a case report series demonstrating injection drug use as a risk factor for development of disseminated disease. BMC Infect. Dis. 2020, 20, 381. [Google Scholar] [CrossRef]
| Strain | MAT | Species | Lineage | Serotype | Genome Size (Mb) | Source | Origin | Ref. |
|---|---|---|---|---|---|---|---|---|
| H99 | α | C. neoformans | VNI | A | 18.9 | Clinical | USA | [27] |
| Bt63 | a | C. neoformans | VNBI | A | 18.4 | Clinical | Africa | [28] |
| Bt1 | α | C. neoformans | VNBII | A | 18.3 | Clinical | Africa | [28] |
| JEC21 | α | C. deneoformans | VNIV | D | 19.1 | Lab Derived | Multiple 1 | [29] |
| B-3501A | α | C. deneoformans | VNIV | D | 18.5 | Lab Derived | Multiple 1 | [29] |
| WM276 | α | C. gattii | VGI | B | 15.8 | Environment | Australia | [30] |
| E566 | a | C. gattii | VGI | B | 17.7 | Environment | Australia | [30] |
| RU294 | α | C. gattii | VGI | B | 17.7 | Environment | Africa | [30] |
| CBS10089 | α | C. deuterogattii | VGII | B | 17.0 | Clinical | Greece | [31] |
| CBS1930 | a | C. deuterogattii | VGII | B | 17.4 | Clinical | Greece | [30,32] |
| R265 | α | C. deuterogattii | VGIIa | B | 17.2 | Clinical | Canada | [33,34] |
| RAM5 | α | C. deuterogattii | VGIIb | B | 17.3 | Clinical | Australia | [30] |
| B8571 | α | C. deuterogattii | VGIIc | B | 17.1 | Clinical | USA | [30] |
| NIH312 | α | C. bacillisporus | VGIII | C | 15.9 | Clinical | USA | [35] |
| CA1873 | a | C. bacillisporus | VGIII | B | 17.4 | Clinical | USA | [30,36] |
| CBS10101 | α | C. tetragattii | VGIV | C | 15.8 | Veterinary | Africa | [30,31,37] |
| MF34 | α | Unnamed | VGV | B | 17.9 | Environment | Africa | [25] |
| Crypto. Genotype | Associations with Human Clinical Presentation | Assoc. with Crypto. Phenotype |
|---|---|---|
| Mating Type | Clinical Isolate Details: MATα > MATa | |
| Species Complex C. neoformans (Serotype A, D, AD) | Population: Immunocompromised> Immunocompetent, HIV-infected > HIV-uninfected PC Symptoms: Cough, Dyspnea, ↑Respiration Rate, Chest Pain, Fever CM Symptoms: Headache, Fever, Neck Pain/Stiffness, Nausea/Vomiting, AMS, Confusion, BC, Photophobia, VC, Seizure, Night Sweats, Limb Weakness, | |
| C. neoformans | Population: Immunocompromised Disease: CM > PC, Skin Lesions < (versus C. deneoformans) and > (versus C. gattii) Yeast Localization: CNS > Blood, Lungs, Skin, Other organs | |
| Serotype A | Population: Immunocompromised > Immunocompetent Risk Factors for HIV-negative Individuals: Malignancy, Other Immunocompromising Conditions Diagnostic Tests: ↑ICP/OP Antifungal Therapy: AmB + 5FC Induction therapy > (versus Serotype D) | |
| VNI (Serotype A) | Symptoms: ↑Vomiting, ↑ICP/OP Clinical Tests: ↓CD4 Count | Cell Size: Micro Cells, Normal Capsule: ↑Capsule Shedding |
| VNII (Serotype A) | Melanin: ↑ Laccase Production (vs. VNI or VNB), ↑Survival in ex-vivo CSF | |
| VNB (Serotype A) | Disease: CM Symptoms: Skin Lesions Patient Outcome: ↑Death | |
| VNBI (Serotype A) | Symptoms: ↑Fever (versus VNI), ↑ICP/OP, ↓Neck Stiffness, ↓Diastolic BP, Clinical Tests: ↓CD4 Count | Capsule: ↑Capsule Shedding |
| VNBII (Serotype A) | Clinical Tests: ↓CD4 Count | |
| Crypto. Genotype | Associations with Human Clinical Presentation | Assoc. Crypto. Phenotype |
| VNIII (Serotype AD) | Population: Immunocompromised, HIV-Infected > HIV-Uninfected Clinical Isolate Details: αADa > aADα > αADα Yeast Localization: ↓Dissemination to CNS Symptoms: ↓Lung involvement, Diagnostic Tests: ↓CrAg Titers, ↑CSF Fungal Clearance, ↑Th2 Cytokine Response Antifungal Therapy: AmB + 5FC Induction therapy > (versus Serotype D) Patient Outcome: ↑Death | ↑Melanin Production |
| C. deneoformans VNIV (Serotype D) | Population: Immunocompromised, HIV-Infected > HIV-Uninfected (vs. AD hybrid) Risk factors for HIV-Uninfected Individuals: Malignancy, No Underlying Conditions Disease: CM > PC, Skin Lesions > (versus C. neoformans or C. gattii) Diagnostic Tests: ↓Blood CrAg Titers, ↓Brain Abnormality (Radiology) Antifungal Therapy: AmB + 5FC Induction therapy < (versus Serotype A or AD) | |
| Species Complex C. gattii (Serotype B, C) | Population: Immunocompetent > Immunocompromised Disease: PC > CM, Skin Lesions < (versus C. deneoformans and C. gattii) PC Symptoms: Cough (Productive or Unproductive), Dyspnea, Chest Pain, Fever, Hemoptysis, Pulmonary Nodules/Mass Lesions (Patient can be asymptomatic) CM Symptoms: Headache, Fever, AMS, Confusion, BC, Neck Pain/Stiffness, Nausea/Vomiting, Photophobia, VC, ONS, Hearing Loss, Cerebellar Abnormalities, Limb Weakness, Seizure, Cranial Nerve Abnormalities, Papilledema | |
| C. gattii–VGI | Population: Immunocompetent, HIV-Uninfected | |
| C. deuterogattii–VGII | Population: Immunocompetent, HIV-Uninfected Patient Outcome: ↑Death (Renal Transplant Patients) | |
| C. bacillisporus–VGIII | Population: Immunocompromised, HIV-Infected | |
| C. tetragattii–VGIV | Population: Immunocompromised, HIV-Infected Disease: CM > PC Diagnostic Tests: ↑CD4 Count, ↓ICP/OP Symptoms: ↓Nausea, ↓Vomiting, ↑BP, Night Sweats, VC, Headache, AMS, Confusion, BC, Photophobia, Fever | Cell Size: Giant/Titan cells, Normal Size, No Micro Cells |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Montoya, M.C.; Magwene, P.M.; Perfect, J.R. Associations between Cryptococcus Genotypes, Phenotypes, and Clinical Parameters of Human Disease: A Review. J. Fungi 2021, 7, 260. https://doi.org/10.3390/jof7040260
Montoya MC, Magwene PM, Perfect JR. Associations between Cryptococcus Genotypes, Phenotypes, and Clinical Parameters of Human Disease: A Review. Journal of Fungi. 2021; 7(4):260. https://doi.org/10.3390/jof7040260
Chicago/Turabian StyleMontoya, Marhiah C., Paul M. Magwene, and John R. Perfect. 2021. "Associations between Cryptococcus Genotypes, Phenotypes, and Clinical Parameters of Human Disease: A Review" Journal of Fungi 7, no. 4: 260. https://doi.org/10.3390/jof7040260
APA StyleMontoya, M. C., Magwene, P. M., & Perfect, J. R. (2021). Associations between Cryptococcus Genotypes, Phenotypes, and Clinical Parameters of Human Disease: A Review. Journal of Fungi, 7(4), 260. https://doi.org/10.3390/jof7040260







