Proteomic Analysis of the Responses of Candida albicans during Infection of Galleria mellonella Larvae
Abstract
1. Introduction
2. Materials and Methods
2.1. Larval Culture and Inoculation
2.2. Yeast Strain
2.3. Effect of Candida albicans Infection on G. mellonella Larvae
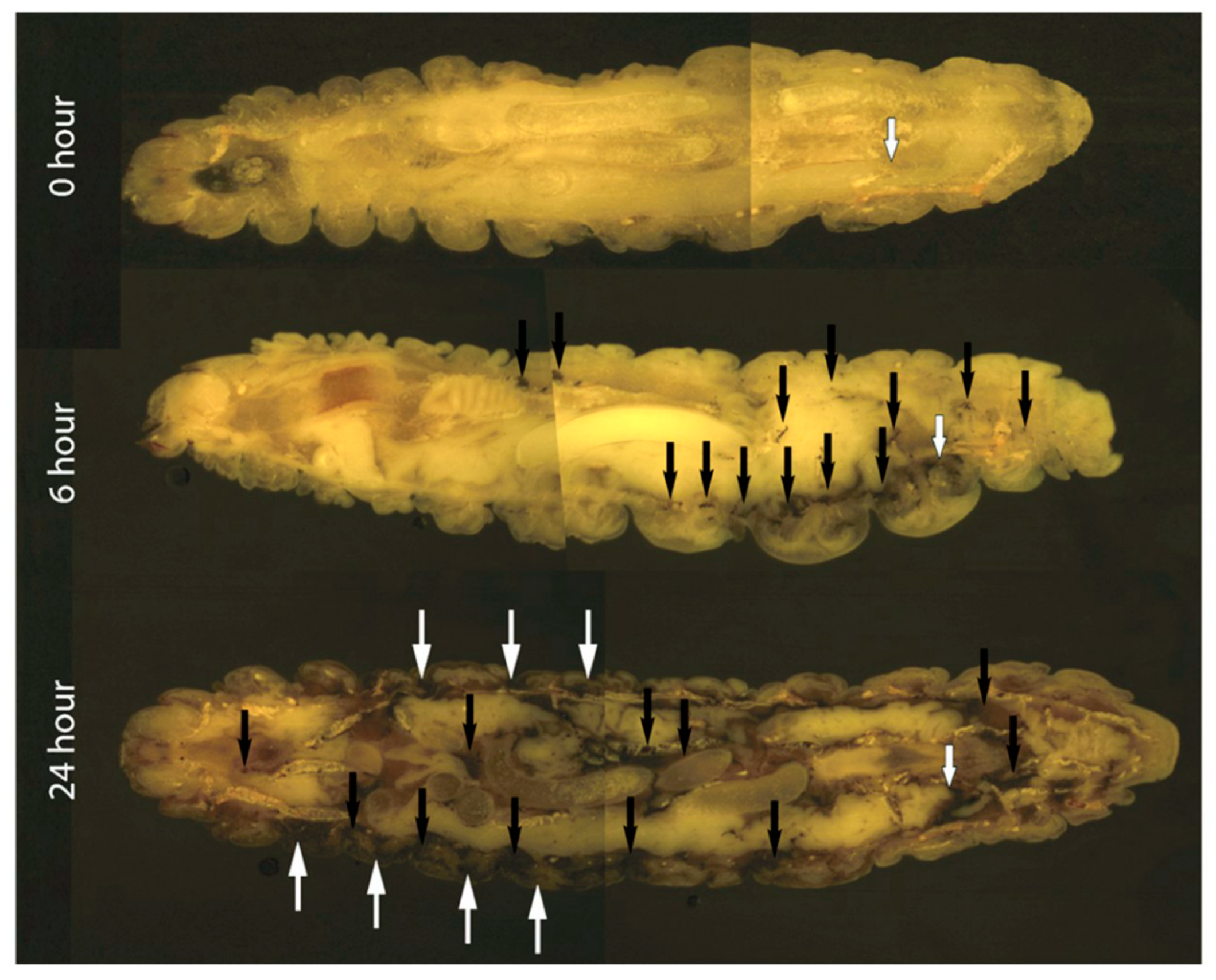
2.4. Cryo-Imaging of C. albicans Infection in G. mellonella
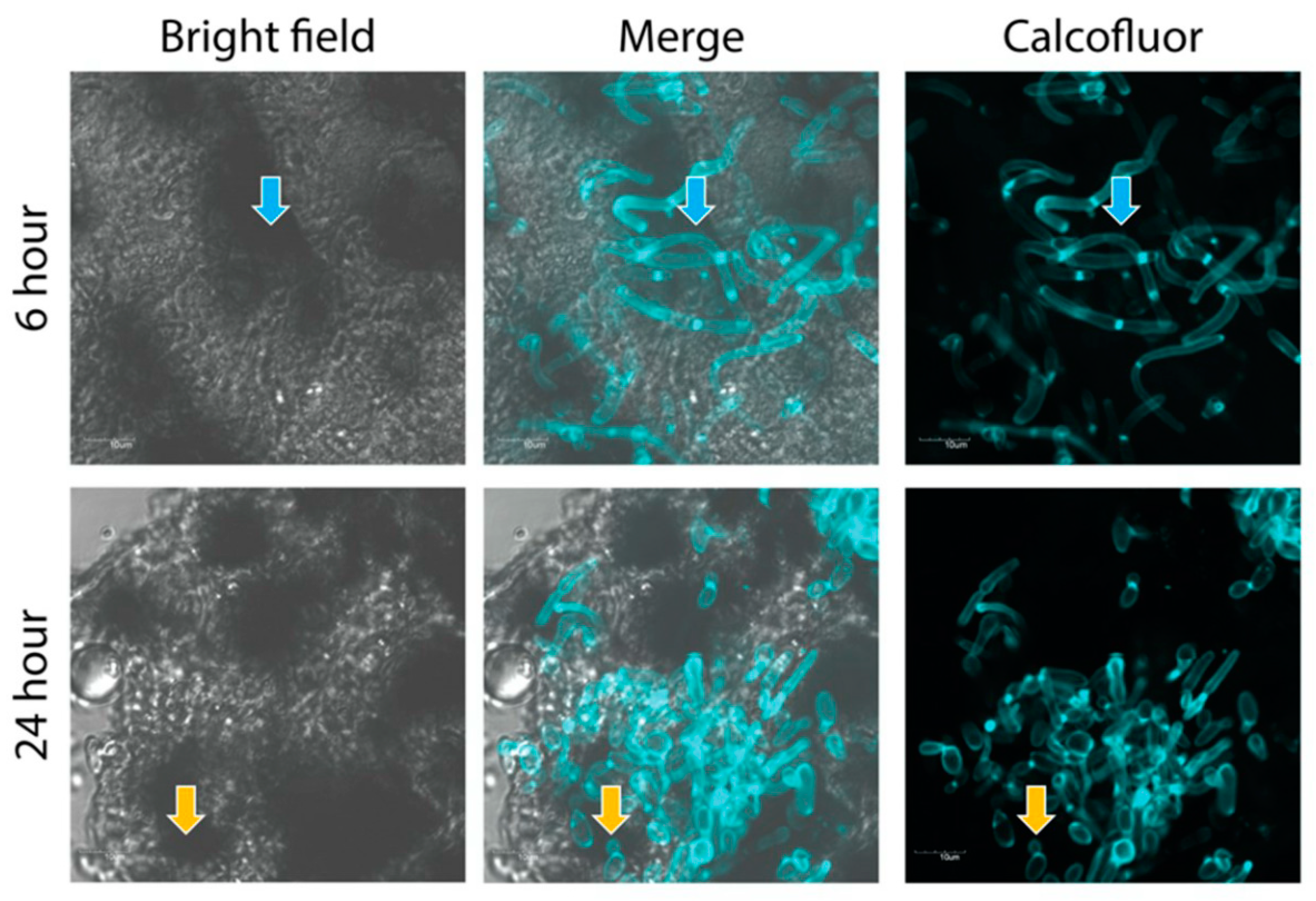
2.5. Confocal Imaging of Fungal Nodules
2.6. Identification of C. albicans Proteins Secreted during Infection of G. mellonella Larvae
2.7. Ex Vivo Hemolymph Fungicidal Activity Assay
2.8. Protein Isolation and Purification from C. albicans Exposed to Larval Hemolymph
2.9. Label Free Quantitative Proteomics Workflow
2.10. Statistical Analysis
2.11. Data Availability
3. Results
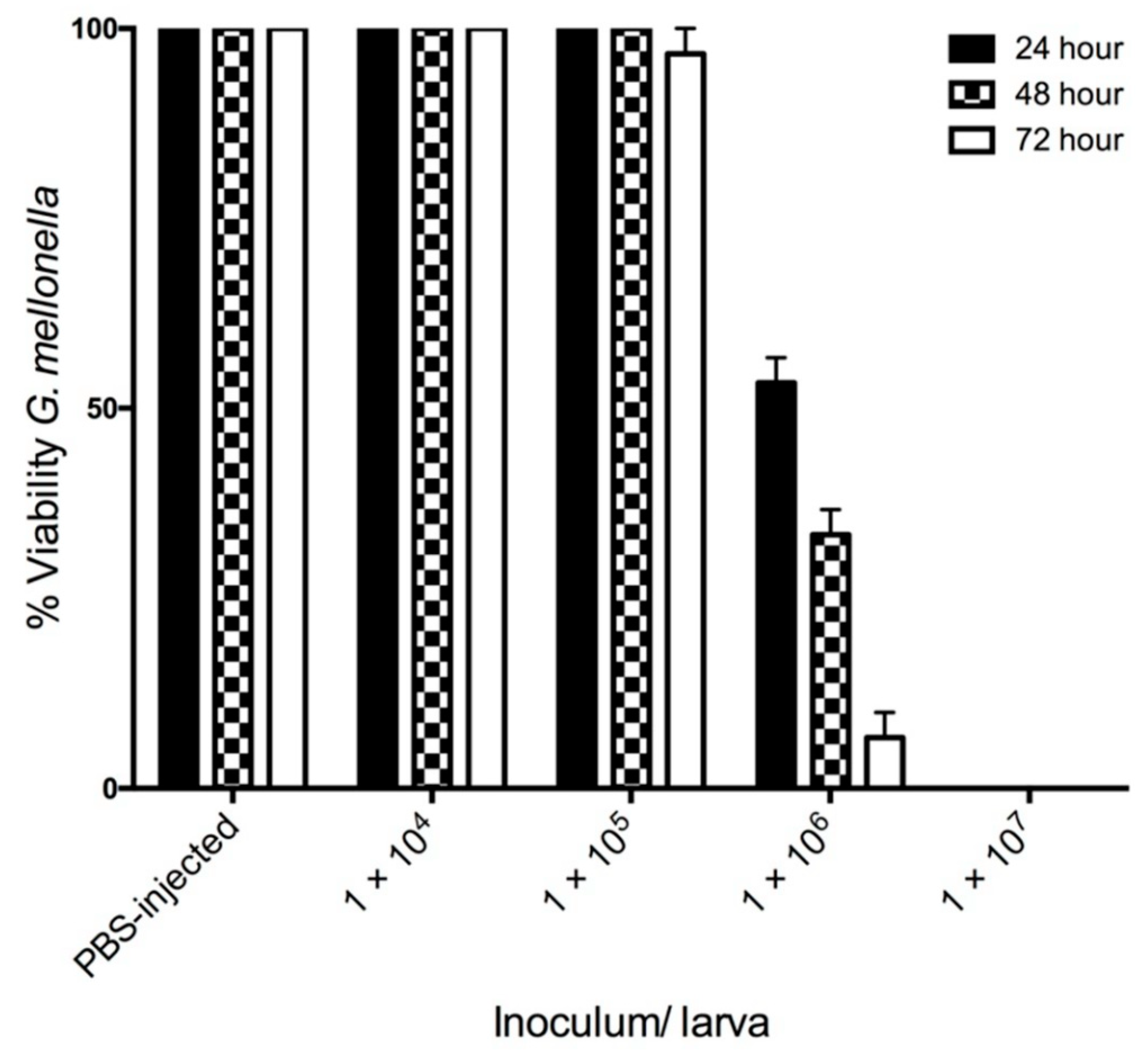
3.1. The Effect of C. albicans Infection on G. mellonella Larvae
3.2. Characterization of C. albicans Proteins Secreted during Infection of G. mellonella Larvae
3.3. Assessment of Fungicidal Activity of Larval Hemolymph on Candida albicans (Ex Vivo)
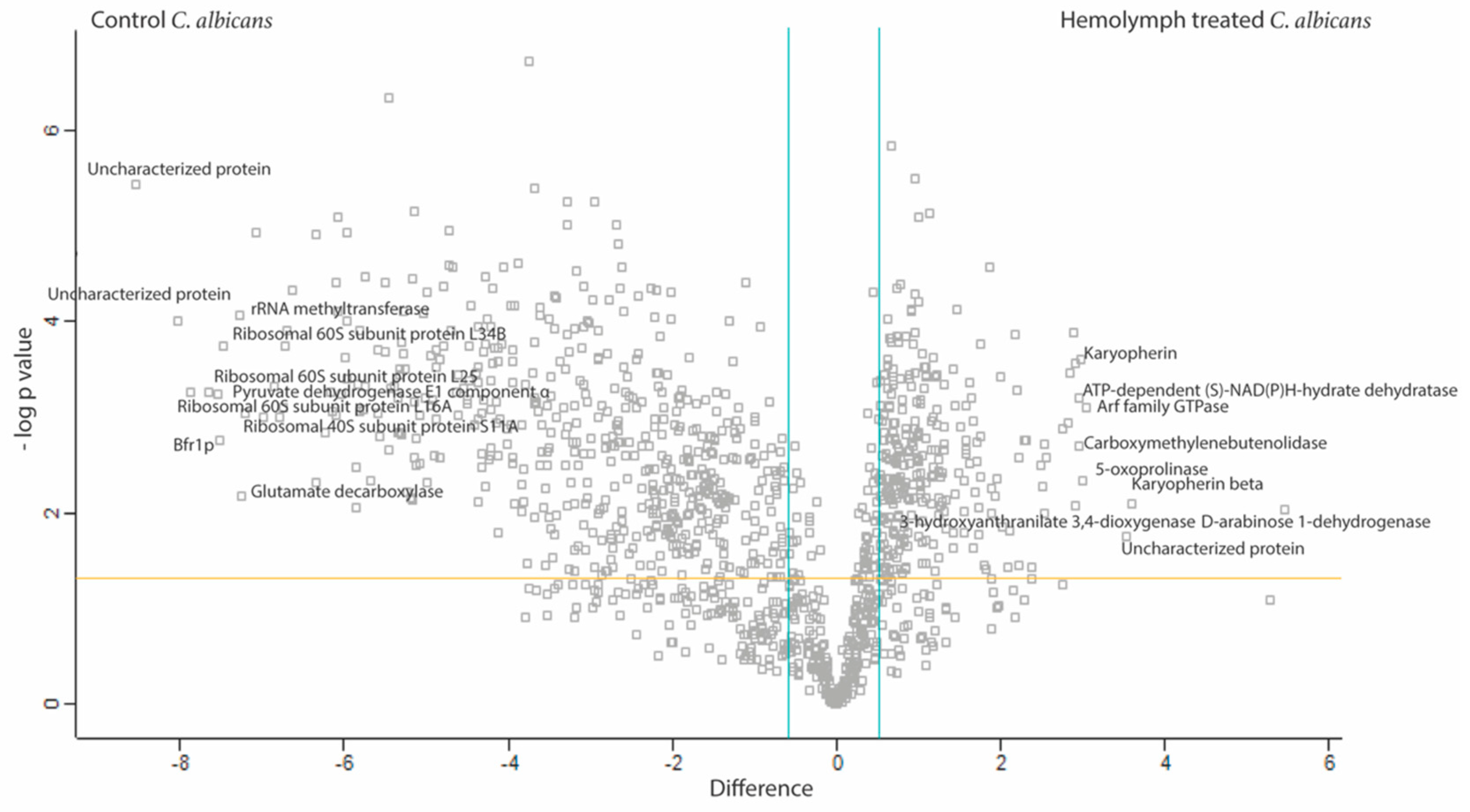
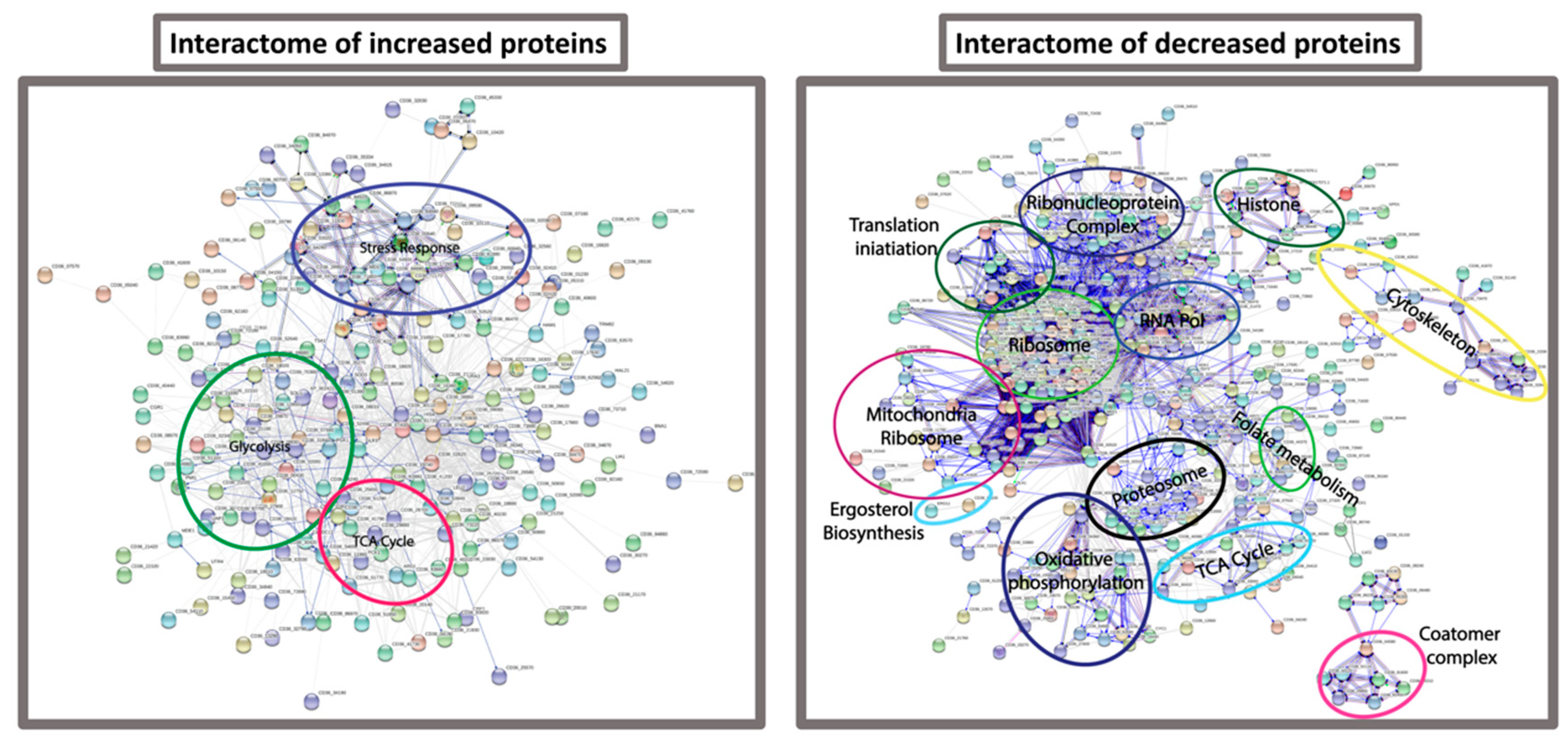
3.4. Proteomic Response of C. albicans to G. mellonella Hemolymph (Ex Vivo)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cotter, G.; Doyle, S.; Kavanagh, K. Development of an insect model for the in vivo pathogenicity testing of yeasts. FEMS Immunol. Med. Microbiol. 2000, 27, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Brennan, M.; Thomas, D.Y.; Whiteway, M.; Kavanagh, K. Correlation between virulence of Candida albicans mutants in mice and Galleria mellonella larvae. FEMS Immunol. Med. Microbiol. 2002, 34, 153–157. [Google Scholar] [CrossRef] [PubMed]
- Slater, J.L.; Gregson, L.; Denning, D.W.; Warn, P.A. Pathogenicity of Aspergillus fumigatus mutants assessed in Galleria mellonella matches that in mice. Med. Mycol. 2011, 49, S107–S113. [Google Scholar] [CrossRef] [PubMed]
- Reeves, E.P.; Messina, C.G.M.; Doyle, S.; Kavanagh, K. Correlation between gliotoxin production and virulence of Aspergillus fumigatus in Galleria mellonella. Mycopathologia 2004, 158, 73–79. [Google Scholar] [CrossRef] [PubMed]
- O’Hanlon, K.A.; Cairns, T.; Stack, D.; Schrettl, M.; Bignell, E.M.; Kavanagh, K.; Miggin, S.M.; O’Keeffe, G.; Larsen, T.O.; Doyle, S. Targeted disruption of nonribosomal peptide synthetase Pes3 augments the virulence of Aspergillus fumigatus. Infect. Immun. 2011, 79, 3978–3992. [Google Scholar] [CrossRef] [PubMed]
- Jander, G.; Rahme, L.G.; Ausubel, F.M. Positive correlation between virulence of Pseudomonas aeruginosa mutants in mice and insects. J. Bacteriol. 2000, 182, 3843–3845. [Google Scholar] [CrossRef] [PubMed]
- Senior, N.J.; Bagnall, M.C.; Champion, O.L.; Reynolds, S.E.; la Ragione, R.M.; Woodward, M.J.; Salguero, F.J.; Titball, R.W. Galleria mellonella as an infection model for campylobacter jejuni virulence. J. Med. Microbiol. 2011, 60, 661–669. [Google Scholar] [CrossRef]
- Kavanagh, K.; Reeves, E.P. Exploiting the potential of insects for in vivo pathogenicity testing of microbial pathogens. FEMS Microbiol. Rev. 2004, 28, 101–112. [Google Scholar] [CrossRef]
- Kavanagh, K.; Sheehan, G. The Use of Galleria mellonella Larvae to Identify Novel Antimicrobial Agents against Fungal Species of Medical Interest. J. Fungi 2018, 4, 113. [Google Scholar] [CrossRef]
- Sheehan, G.; Clarke, G.; Kavanagh, K. Characterisation of the cellular and proteomic response of Galleria mellonella larvae to the development of invasive aspergillosis. BMC Microbiol. 2018, 18. [Google Scholar] [CrossRef]
- Fuchs, B.B.; O’Brien, E.; El Khoury, J.B.; Mylonakis, E. Methods for using Galleria mellonella as a model host to study fungal pathogenesis. Virulence 2010, 1, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, G.; Garvey, A.; Croke, M.; Kavanagh, K. Innate humoral immune defences in mammals and insects: The same, with differences? Virulence 2018, 9, 1625–1639. [Google Scholar] [CrossRef] [PubMed]
- Schuhmann, B.; Seitz, V.; Vilcinskas, A.; Podsiadlowski, L. Cloning and expression of gallerimycin, an antifungal peptide expressed in immune response of greater wax moth larvae, Galleria mellonella. Arch. Insect Biochem. Physiol. 2003, 53, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Yun, E.K.; Jang, W.S.; Kim, I.; Lee, J.H.; Park, S.Y.; Ryu, K.S.; Seo, S.J.; Kim, C.H.; Lee, I.H. Purification, cDNA cloning and expression of an insect defensin from the great wax moth, Galleria mellonella. Insect Mol. Biol. 2004, 13, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Seitz, V.; Clermont, A.; Wedde, M.; Hummel, M.; Vilcinskas, A.; Schlatterer, K.; Podsiadlowski, L. Identification of immunorelevant genes from greater wax moth (Galleria mellonella) by a subtractive hybridization approach. Dev. Comp. Immunol. 2003, 27, 207–215. [Google Scholar] [CrossRef]
- Bolouri Moghaddam, M.R.; Tonk, M.; Schreiber, C.; Salzig, D.; Czermak, P.; Vilcinskas, A.; Rahnamaeian, M. The potential of the Galleria mellonella innate immune system is maximized by the co-presentation of diverse antimicrobial peptides. Biol. Chem. 2016, 397, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Brown, S.E.; Howard, A.; Kasprzak, A.B.; Gordon, K.H.; East, P.D. The discovery and analysis of a diverged family of novel antifungal moricin-like peptides in the wax moth Galleria mellonella. Insect Biochem. Mol. Biol. 2008, 38, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, G.; Kavanagh, K. Analysis of the early cellular and humoral responses of Galleria mellonella larvae to infection by Candida albicans. Virulence 2018, 9, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Yun, J.E.; Lee, D.G. Cecropin A-induced apoptosis is regulated by ion balance and glutathione antioxidant system in Candida albicans. IUBMB Life 2016, 652–662. [Google Scholar] [CrossRef]
- Lee, E.; Shin, A.; Kim, Y. Anti-inflammatory activities of cecropin A and its mechanism of action. Arch. Insect Biochem. Physiol. 2015, 88, 31–44. [Google Scholar] [CrossRef]
- Moazeni, M.; Asgari, S.; Nabili, M. Nosocomial fungal infections: Epidemiology, diagnosis, treatment and prevention. J. Maz. Univ. Med. Sci. 2018, 28, 182–212. [Google Scholar]
- Brown, G.D.; Denning, D.W.; Gow, N.A.R.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden Killers: Human Fungal Infections. Sci. Transl. Med. 2012, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Wilson, D.; Hube, B. Candida albicans pathogenicity mechanisms. Virulence 2013, 4, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Priebe, S.; Linde, J.; Albrecht, D.; Guthke, R.; Brakhage, A.A. FungiFun: A web-based application for functional categorization of fungal genes and proteins. Fungal Genet. Biol. 2011, 48, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Jensen, L.J.; Kuhn, M.; Stark, M.; Chaffron, S.; Creevey, C.; Muller, J.; Doerks, T.; Julien, P.; Roth, A.; Simonovic, M.; et al. STRING 8—A global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 2009, 37. [Google Scholar] [CrossRef] [PubMed]
- Côté, R.G.; Griss, J.; Dianes, J.A.; Wang, R.; Wright, J.C.; van den Toorn, H.W.P.; van Breukelen, B.; Heck, A.J.R.; Hulstaert, N.; Martens, L.; et al. The PRoteomics IDEntification (PRIDE) Converter 2 Framework: An Improved Suite of Tools to Facilitate Data Submission to the PRIDE Database and the ProteomeXchange Consortium. Mol. Cell. Proteom. 2012, 11, 1682–1689. [Google Scholar] [CrossRef] [PubMed]
- Calderone, R.A.; Fonzi, W.A. Virulence factors of Candida albicans. Trends Microbiol. 2001, 9, 327–335. [Google Scholar] [CrossRef]
- Ruiz-Herrera, J.; Victoria Elorza, M.; Valentín, E.; Sentandreu, R. Molecular organization of the cell wall of Candida albicans and its relation to pathogenicity. FEMS Yeast Res. 2006, 6, 14–29. [Google Scholar] [CrossRef]
- Misme-Aucouturier, B.; Albassier, M.; Alvarez-Rueda, N.; Pape, P. Le Specific human and Candida cellular interactions lead to controlled or persistent infection outcomes during granuloma-like formation. Infect. Immun. 2017, 85, e00807-16. [Google Scholar] [CrossRef]
- Weissman, Z.; Kornitzer, D. A family of Candida cell surface haem-binding proteins involved in haemin and haemoglobin-iron utilization. Mol. Microbiol. 2004, 53, 1209–1220. [Google Scholar] [CrossRef]
- Sigel, H.C.; Thewes, S.; Niewerth, M.; Korting, H.C.; Schäder-Korting, M.; Hube, B. Oxygen accessibility and iron levels are critical factors for the antifungal action of ciclopirox against Candida albicans. J. Antimicrob. Chemother. 2005, 55, 663–673. [Google Scholar] [CrossRef] [PubMed]
- Pérez, A.; Pedrós, B.; Murgui, A.; Casanova, M.; López-Ribot, J.L.; Martínez, J.P. Biofilm formation by Candida albicans mutants for genes coding fungal proteins exhibiting the eight-cysteine-containing CFEM domain. FEMS Yeast Res. 2006, 6, 1074–1084. [Google Scholar] [CrossRef]
- Urban, C.; Xiong, X.; Sohn, K.; Schröppel, K.; Brunner, H.; Rupp, S. The moonlighting protein Tsa1p is implicated in oxidative stress response and in cell wall biogenesis in Candida albicans. Mol. Microbiol. 2005, 57, 1318–1341. [Google Scholar] [CrossRef]
- Jong, A.Y.; Chen, S.H.M.; Stins, M.F.; Kim, K.S.; Tuan, T.L.; Huang, S.H. Binding of Candida albicans enolase to plasmin(ogen) results in enhanced invasion of human brain microvascular endothelial cells. J. Med. Microbiol. 2003, 52, 615–622. [Google Scholar] [CrossRef] [PubMed]
- Li, X.S.; Reddy, M.S.; Baev, D.; Edgerton, M. Candida albicans Ssa1/2p is the cell envelope binding protein for human salivary histatin 5. J. Biol. Chem. 2003, 278, 28553–28561. [Google Scholar] [CrossRef] [PubMed]
- Pietrella, D.; Bistoni, G.; Corbucci, C.; Perito, S.; Vecchiarelli, A. Candida albicans mannoprotein influences the biological function of dendritic cells. Cell. Microbiol. 2006, 8, 602–612. [Google Scholar] [CrossRef] [PubMed]
- Sandini, S.; Stringaro, A.; Arancia, S.; Colone, M.; Mondello, F.; Murtas, S.; Girolamo, A.; Mastrangelo, N.; De Bernardis, F. The MP65 gene is required for cell wall integrity, adherence to epithelial cells and biofilm formation in Candida albicans. BMC Microbiol. 2011, 11. [Google Scholar] [CrossRef] [PubMed]
- Braun, B.R.; Head, W.S.; Wang, M.X.; Johnson, A.D. Identification and characterization of TUP1-regulated genes in Candida albicans. Genetics 2000, 156, 31–44. [Google Scholar] [CrossRef] [PubMed]
- Fradin, C.; Kretschmar, M.; Nichterlein, T.; Gaillardin, C.; D’Enfert, C.; Hube, B. Stage-specific gene expression of Candida albicans in human blood. Mol. Microbiol. 2003, 47, 1523–1543. [Google Scholar] [CrossRef]
- Gyurko, C.; Lendenmann, U.; Troxler, R.F.; Oppenheim, F.G. Candida albicans mutants deficient in respiration are resistant to the small cationic salivary antimicrobial peptide histatin 5. Antimicrob. Agents Chemother. 2000. [Google Scholar] [CrossRef]
- Geraghty, P.; Kavanagh, K. Disruption of mitochondrial function in Candida albicans leads to reduced cellular ergosterol levels and elevated growth in the presence of amphotericin B. Arch. Microbiol. 2003. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.N.; Solis, N.V.; Phan, Q.T.; Bajwa, J.S.; Kashleva, H.; Thompson, A.; Liu, Y.; Dongari-Bagtzoglou, A.; Edgerton, M.; Filler, S.G. Host cell invasion and virulence mediated by Candida albicans Ssa1. PLoS Pathog. 2010, 6. [Google Scholar] [CrossRef] [PubMed]
- Cho, T.; Toyoda, M.; Sudoh, M.; Nakashima, Y.; Calderone, R.A.; Kaminishi, H. Isolation and sequencing of the Candida albicans MS13, a putative novel member of the HSP70 family. Yeast 2003, 20, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Nagao, J.I.; Cho, T.; Uno, J.; Ueno, K.; Imayoshi, R.; Nakayama, H.; Chibana, H.; Kaminishi, H. Candida albicans Msi3p, a homolog of the Saccharomyces cerevisiae Sse1p of the Hsp70 family, is involved in cell growth and fluconazole tolerance. FEMS Yeast Res. 2012, 12, 728–737. [Google Scholar] [CrossRef] [PubMed]
- Hornbach, A.; Heyken, A.; Schild, L.; Hube, B.; Löffler, J.; Kurzai, O. The glycosylphosphatidylinositol-anchored protease Sap9 modulates the interaction of Candida albicans with human neutrophils. Infect. Immun. 2009, 77, 5216–5224. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheehan, G.; Kavanagh, K. Proteomic Analysis of the Responses of Candida albicans during Infection of Galleria mellonella Larvae. J. Fungi 2019, 5, 7. https://doi.org/10.3390/jof5010007
Sheehan G, Kavanagh K. Proteomic Analysis of the Responses of Candida albicans during Infection of Galleria mellonella Larvae. Journal of Fungi. 2019; 5(1):7. https://doi.org/10.3390/jof5010007
Chicago/Turabian StyleSheehan, Gerard, and Kevin Kavanagh. 2019. "Proteomic Analysis of the Responses of Candida albicans during Infection of Galleria mellonella Larvae" Journal of Fungi 5, no. 1: 7. https://doi.org/10.3390/jof5010007
APA StyleSheehan, G., & Kavanagh, K. (2019). Proteomic Analysis of the Responses of Candida albicans during Infection of Galleria mellonella Larvae. Journal of Fungi, 5(1), 7. https://doi.org/10.3390/jof5010007





