Candida albicans Hyphae: From Growth Initiation to Invasion
Abstract
1. Introduction
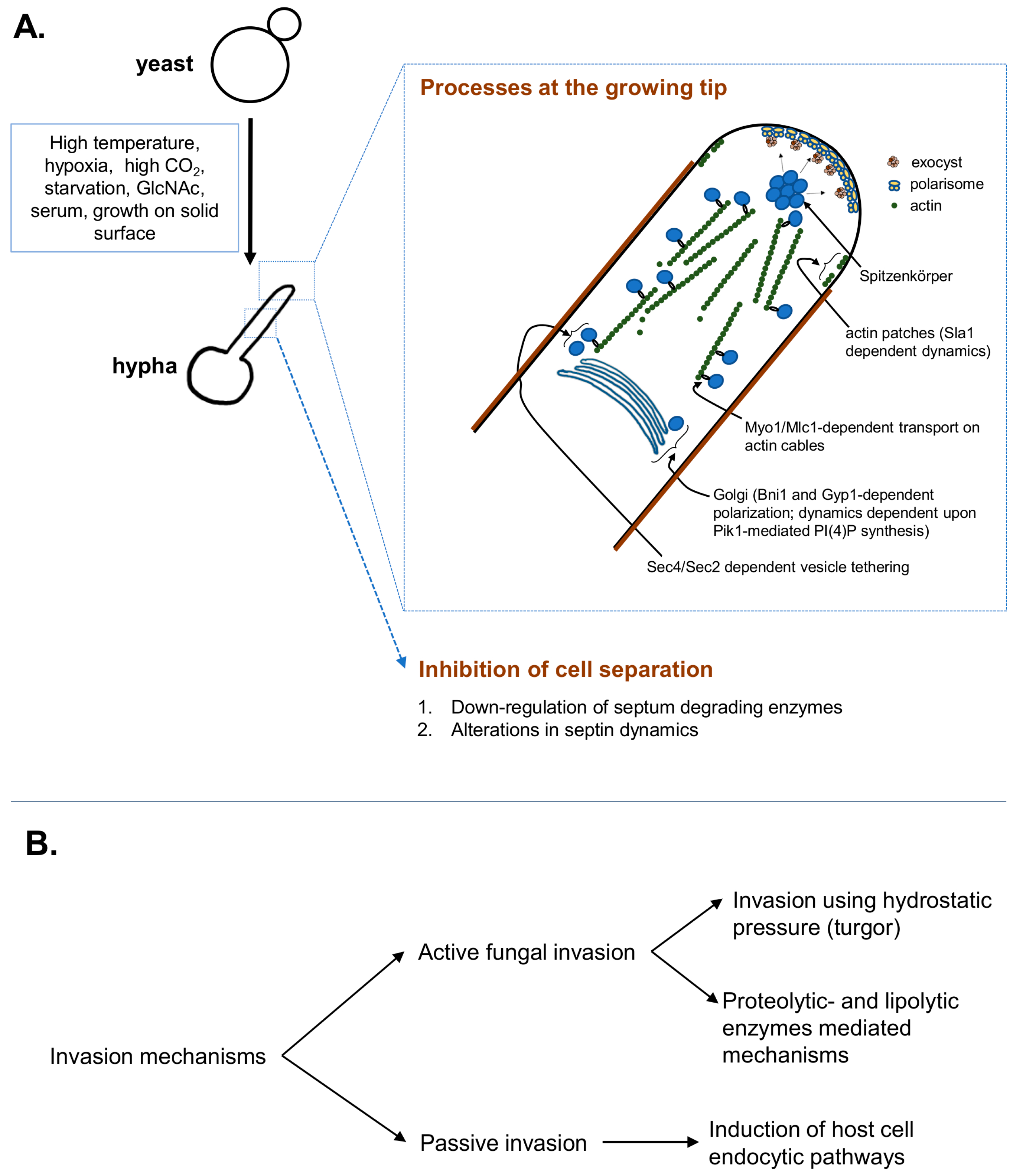
2. Hyphal Initiation, Elongation and Directionality Maintenance
3. Hyphal Invasion
Acknowledgments
Conflicts of Interest
References
- Sanati Nezhad, A.; Geitmann, A. The cellular mechanics of an invasive lifestyle. J. Exp. Bot. 2013, 64, 4709–4728. [Google Scholar] [CrossRef] [PubMed]
- LeGoff, L.; Lecuit, T. Mechanical Forces and Growth in Animal Tissues. Cold Spring Harb. Perspect. Biol. 2015, 8, a019232. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.D.; Forn, I.; Gadelha, C.; Egan, M.J.; Bass, D.; Massana, R.; Richards, T.A. Discovery of novel intermediate forms redefines the fungal tree of life. Nature 2011, 474, 200–203. [Google Scholar] [CrossRef] [PubMed]
- Harris, S.D. Hyphal morphogenesis: An evolutionary perspective. Fungal Biol. 2011, 115, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Moyes, D.L.; Richardson, J.P.; Naglik, J.R. Candida albicans-epithelial interactions and pathogenicity mechanisms: Scratching the surface. Virulence 2015, 6, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Jabra-Rizk, M.A.; Kong, E.F.; Tsui, C.; Nguyen, M.H.; Clancy, C.J.; Fidel, P.L., Jr.; Noverr, M. Candida albicans Pathogenesis: Fitting within the Host-Microbe Damage Response Framework. Infect. Immun. 2016, 84, 2724–2739. [Google Scholar] [CrossRef] [PubMed]
- Kadosh, D. Morphogenesis in C. albicans. In Candida albicans: Cellular and Molecular Biology; Prasad, R., Ed.; Springer International Publishing: Cham, Switzerland, 2017; pp. 41–62. [Google Scholar]
- Sudbery, P.E. Growth of Candida albicans hyphae. Nat. Rev. Microbiol. 2011, 9, 737–748. [Google Scholar] [CrossRef] [PubMed]
- Gale, C.; Gerami-Nejad, M.; McClellan, M.; Vandoninck, S.; Longtine, M.S.; Berman, J. Candida albicans Int1p interacts with the septin ring in yeast and hyphal cells. Mol. Biol. Cell 2001, 12, 3538–3549. [Google Scholar] [CrossRef] [PubMed]
- Yaar, L.; Mevarech, M.; Koltin, Y. A Candida albicans RAS-related gene (CaRSR1) is involved in budding, cell morphogenesis and hypha development. Microbiology 1997, 143 Pt 9, 3033–3044. [Google Scholar] [CrossRef] [PubMed]
- Hausauer, D.L.; Gerami-Nejad, M.; Kistler-Anderson, C.; Gale, C.A. Hyphal guidance and invasive growth in Candida albicans require the Ras-like GTPase Rsr1p and its GTPase-activating protein Bud2p. Eukaryot. Cell 2005, 4, 1273–1286. [Google Scholar] [CrossRef] [PubMed]
- Warenda, A.J.; Konopka, J.B. Septin function in Candida albicans morphogenesis. Mol. Biol. Cell 2002, 13, 2732–2746. [Google Scholar] [CrossRef] [PubMed]
- Sudbery, P.E. The germ tubes of Candida albicans hyphae and pseudohyphae show different patterns of septin ring localization. Mol. Microbiol. 2001, 41, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Srikantha, T.; Tsai, L.; Daniels, K.; Enger, L.; Highley, K.; Soll, D.R. The two-component hybrid kinase regulator CaNIK1 of Candida albicans. Microbiology 1998, 144 Pt 10, 2715–2729. [Google Scholar] [CrossRef] [PubMed]
- Crampin, H.; Finley, K.; Gerami-Nejad, M.; Court, H.; Gale, C.; Berman, J.; Sudbery, P. Candida albicans hyphae have a Spitzenkorper that is distinct from the polarisome found in yeast and pseudohyphae. J. Cell Sci. 2005, 118 Pt 13, 2935–2947. [Google Scholar] [CrossRef] [PubMed]
- Thomson, D.D.; Berman, J.; Brand, A.C. High frame-rate resolution of cell division during Candida albicans filamentation. Fungal Genet. Biol. 2016, 88, 54–58. [Google Scholar] [CrossRef] [PubMed]
- Azadmanesh, J.; Gowen, A.M.; Creger, P.E.; Schafer, N.D.; Blankenship, J.R. Filamentation Involves Two Overlapping, but Distinct, Programs of Filamentation in the Pathogenic Fungus Candida albicans. G3 (Bethesda) 2017, 7, 3797–3808. [Google Scholar] [CrossRef] [PubMed]
- Nantel, A.; Dignard, D.; Bachewich, C.; Harcus, D.; Marcil, A.; Bouin, A.P.; Sensen, C.W.; Hogues, H.; van het Hoog, M.; Gordon, P.; et al. Transcription profiling of Candida albicans cells undergoing the yeast-to-hyphal transition. Mol. Biol. Cell 2002, 13, 3452–3465. [Google Scholar] [CrossRef] [PubMed]
- Rida, P.C.; Nishikawa, A.; Won, G.Y.; Dean, N. Yeast-to-hyphal transition triggers formin-dependent Golgi localization to the growing tip in Candida albicans. Mol. Biol. Cell 2006, 17, 4364–4378. [Google Scholar] [CrossRef] [PubMed]
- Ghugtyal, V.; Garcia-Rodas, R.; Seminara, A.; Schaub, S.; Bassilana, M.; Arkowitz, R.A. Phosphatidylinositol-4-phosphate-dependent membrane traffic is critical for fungal filamentous growth. Proc. Natl. Acad. Sci. USA 2015, 112, 8644–8649. [Google Scholar] [CrossRef] [PubMed]
- Jones, L.A.; Sudbery, P.E. Spitzenkorper, exocyst, and polarisome components in Candida albicans hyphae show different patterns of localization and have distinct dynamic properties. Eukaryot. Cell 2010, 9, 1455–1465. [Google Scholar] [CrossRef] [PubMed]
- Bishop, A.; Lane, R.; Beniston, R.; Chapa-y-Lazo, B.; Smythe, C.; Sudbery, P. Hyphal growth in Candida albicans requires the phosphorylation of Sec2 by the Cdc28-Ccn1/Hgc1 kinase. EMBO J. 2010, 29, 2930–2942. [Google Scholar] [CrossRef] [PubMed]
- Caballero-Lima, D.; Kaneva, I.N.; Watton, S.P.; Sudbery, P.E.; Craven, C.J. The spatial distribution of the exocyst and actin cortical patches is sufficient to organize hyphal tip growth. Eukaryot. Cell 2013, 12, 998–1008. [Google Scholar] [CrossRef] [PubMed]
- Arkowitz, R.A.; Bassilana, M. Regulation of hyphal morphogenesis by Ras and Rho small GTPases. Fungal Biol. Rev. 2015, 29, 7–19. [Google Scholar] [CrossRef]
- Fortwendel, J.R. Ras-Mediated Signal Transduction and Virulence in Human Pathogenic Fungi. Fungal Genom. Biol. 2012, 2, 105. [Google Scholar] [CrossRef] [PubMed]
- Hazan, I.; Liu, H. Hyphal tip-associated localization of Cdc42 is F-actin dependent in Candida albicans. Eukaryot. Cell 2002, 1, 856–864. [Google Scholar] [CrossRef] [PubMed]
- Pulver, R.; Heisel, T.; Gonia, S.; Robins, R.; Norton, J.; Haynes, P.; Gale, C.A. Rsr1 focuses Cdc42 activity at hyphal tips and promotes maintenance of hyphal development in Candida albicans. Eukaryot. Cell 2013, 12, 482–495. [Google Scholar] [CrossRef] [PubMed]
- Park, H.O.; Kang, P.J.; Rachfal, A.W. Localization of the Rsr1/Bud1 GTPase involved in selection of a proper growth site in yeast. J. Biol. Chem. 2002, 277, 26721–26724. [Google Scholar] [CrossRef] [PubMed]
- Brand, A.C.; Morrison, E.; Milne, S.; Gonia, S.; Gale, C.A.; Gow, N.A. Cdc42 GTPase dynamics control directional growth responses. Proc. Natl. Acad. Sci. USA 2014, 111, 811–816. [Google Scholar] [CrossRef] [PubMed]
- Kozubowski, L.; Saito, K.; Johnson, J.M.; Howell, A.S.; Zyla, T.R.; Lew, D.J. Symmetry-breaking polarization driven by a Cdc42p GEF-PAK complex. Curr. Biol. 2008, 18, 1719–1726. [Google Scholar] [CrossRef] [PubMed]
- Brand, A.; Shanks, S.; Duncan, V.M.; Yang, M.; Mackenzie, K.; Gow, N.A. Hyphal orientation of Candida albicans is regulated by a calcium-dependent mechanism. Curr. Biol. 2007, 17, 347–352. [Google Scholar] [CrossRef] [PubMed]
- Guo, P.P.; Yong, J.Y.; Wang, Y.M.; Li, C.R. Sec15 links bud site selection to polarised cell growth and exocytosis in Candida albicans. Sci. Rep. 2016, 6, 26464. [Google Scholar] [CrossRef] [PubMed]
- Thomson, D.D.; Wehmeier, S.; Byfield, F.J.; Janmey, P.A.; Caballero-Lima, D.; Crossley, A.; Brand, A.C. Contact-induced apical asymmetry drives the thigmotropic responses of Candida albicans hyphae. Cell Microbiol. 2015, 17, 342–354. [Google Scholar] [CrossRef] [PubMed]
- Hope, H.; Bogliolo, S.; Arkowitz, R.A.; Bassilana, M. Activation of Rac1 by the guanine nucleotide exchange factor Dck1 is required for invasive filamentous growth in the pathogen Candida albicans. Mol. Biol. Cell 2008, 19, 3638–3651. [Google Scholar] [CrossRef] [PubMed]
- Hope, H.; Schmauch, C.; Arkowitz, R.A.; Bassilana, M. The Candida albicans ELMO homologue functions together with Rac1 and Dck1, upstream of the MAP Kinase Cek1, in invasive filamentous growth. Mol. Microbiol. 2010, 76, 1572–1590. [Google Scholar] [CrossRef] [PubMed]
- Berman, J. Morphogenesis and cell cycle progression in Candida albicans. Curr. Opin. Microbiol. 2006, 9, 595–601. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Huang, Z.X.; Au Yong, J.Y.; Zou, H.; Zeng, G.; Gao, J.; Wang, Y.; Wong, A.H.; Wang, Y. CDK phosphorylates the polarisome scaffold Spa2 to maintain its localization at the site of cell growth. Mol. Microbiol. 2016, 101, 250–264. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.D.; Lee, R.T.; Wang, Y.M.; Lin, Q.S.; Wang, Y. Phosphorylation of Rga2, a Cdc42 GAP, by CDK/Hgc1 is crucial for Candida albicans hyphal growth. EMBO J. 2007, 26, 3760–3769. [Google Scholar] [CrossRef] [PubMed]
- Zeng, G.; Wang, Y.M.; Wang, Y. Cdc28-Cln3 phosphorylation of Sla1 regulates actin patch dynamics in different modes of fungal growth. Mol. Biol. Cell 2012, 23, 3485–3497. [Google Scholar] [CrossRef] [PubMed]
- Caballero-Lima, D.; Sudbery, P.E. In Candida albicans, phosphorylation of Exo84 by Cdk1-Hgc1 is necessary for efficient hyphal extension. Mol. Biol. Cell 2014, 25, 1097–1110. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Escribano, P.; Gonzalez-Novo, A.; Suarez, M.B.; Li, C.R.; Wang, Y.; de Aldana, C.R.; Correa-Bordes, J. CDK-dependent phosphorylation of Mob2 is essential for hyphal development in Candida albicans. Mol. Biol. Cell 2011, 22, 2458–2469. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; McQuilken, M.; Gladfelter, A.S. Septins and Generation of Asymmetries in Fungal Cells. Annu. Rev. Microbiol. 2015, 69, 487–503. [Google Scholar] [CrossRef] [PubMed]
- Blankenship, J.R.; Cheng, S.; Woolford, C.A.; Xu, W.; Johnson, T.M.; Rogers, P.D.; Fanning, S.; Nguyen, M.H.; Clancy, C.J.; Mitchell, A.P. Mutational analysis of essential septins reveals a role for septin-mediated signaling in filamentation. Eukaryot. Cell 2014, 13, 1403–1410. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Novo, A.; Correa-Bordes, J.; Labrador, L.; Sanchez, M.; Vazquez de Aldana, C.R.; Jimenez, J. Sep7 is essential to modify septin ring dynamics and inhibit cell separation during Candida albicans hyphal growth. Mol. Biol. Cell 2008, 19, 1509–1518. [Google Scholar] [CrossRef] [PubMed]
- Sinha, I.; Wang, Y.-M.; Philp, R.; Li, C.-R.; Yap, W.H.; Wang, Y. Cyclin-Dependent Kinases Control Septin Phosphorylation in Candida albicans Hyphal Development. Dev. Cell 2007, 13, 421–432. [Google Scholar] [CrossRef] [PubMed]
- Brand, A.; Vacharaksa, A.; Bendel, C.; Norton, J.; Haynes, P.; Henry-Stanley, M.; Wells, C.; Ross, K.; Gow, N.A.; Gale, C.A. An internal polarity landmark is important for externally induced hyphal behaviors in Candida albicans. Eukaryot. Cell 2008, 7, 712–720. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Raniga, P.P.; Lane, S.; Lu, Y.; Liu, H. Hyphal chain formation in Candida albicans: Cdc28-Hgc1 phosphorylation of Efg1 represses cell separation genes. Mol. Cell. Biol. 2009, 29, 4406–4416. [Google Scholar] [CrossRef] [PubMed]
- Calderon-Norena, D.M.; Gonzalez-Novo, A.; Orellana-Munoz, S.; Gutierrez-Escribano, P.; Arnaiz-Pita, Y.; Duenas-Santero, E.; Suarez, M.B.; Bougnoux, M.E.; Del Rey, F.; Sherlock, G.; et al. A single nucleotide polymorphism uncovers a novel function for the transcription factor Ace2 during Candida albicans hyphal development. PLoS Genet. 2015, 11, e1005152. [Google Scholar] [CrossRef] [PubMed]
- Lassak, T.; Schneider, E.; Bussmann, M.; Kurtz, D.; Manak, J.R.; Srikantha, T.; Soll, D.R.; Ernst, J.F. Target specificity of the Candida albicans Efg1 regulator. Mol. Microbiol. 2011, 82, 602–618. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Mao, X.; Raniga, P.P.; Liu, H.; Chen, J. Efg1-mediated recruitment of NuA4 to promoters is required for hypha-specific Swi/Snf binding and activation in Candida albicans. Mol. Biol. Cell 2008, 19, 4260–4272. [Google Scholar] [CrossRef] [PubMed]
- Stoldt, V.R.; Sonneborn, A.; Leuker, C.E.; Ernst, J.F. Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J. 1997, 16, 1982–1991. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, R.S.; Uppuluri, P.; Zaas, A.K.; Collins, C.; Senn, H.; Perfect, J.R.; Heitman, J.; Cowen, L.E. Hsp90 orchestrates temperature-dependent Candida albicans morphogenesis via Ras1-PKA signaling. Curr. Biol. 2009, 19, 621–629. [Google Scholar] [CrossRef] [PubMed]
- Hall, R.A.; De Sordi, L.; Maccallum, D.M.; Topal, H.; Eaton, R.; Bloor, J.W.; Robinson, G.K.; Levin, L.R.; Buck, J.; Wang, Y.; et al. CO(2) acts as a signalling molecule in populations of the fungal pathogen Candida albicans. PLoS Pathog. 2010, 6, e1001193. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Liu, H. Candida albicans hyphal initiation and elongation. Trends Microbiol. 2014, 22, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Unoje, O.; Liu, H. Quorum sensing controls hyphal initiation in Candida albicans through Ubr1-mediated protein degradation. Proc. Natl. Acad. Sci. USA 2014, 111, 1975–1980. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Kim, J.M.; Kang, W.K.; Yang, H.; Kim, J.Y. The NDR Kinase Cbk1 Downregulates the Transcriptional Repressor Nrg1 through the mRNA-Binding Protein Ssd1 in Candida albicans. Eukaryot. Cell 2015, 14, 671–683. [Google Scholar] [CrossRef] [PubMed]
- Davis, D.A. How human pathogenic fungi sense and adapt to pH: The link to virulence. Curr. Opin. Microbiol. 2009, 12, 365–370. [Google Scholar] [CrossRef] [PubMed]
- Zucchi, P.C.; Davis, T.R.; Kumamoto, C.A. A Candida albicans cell wall-linked protein promotes invasive filamentation into semi-solid medium. Mol. Microbiol. 2010, 76, 733–748. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.X.; Wang, H.; Wang, Y.M.; Wang, Y. Novel mechanism coupling cyclic AMP-protein kinase A signaling and golgi trafficking via Gyp1 phosphorylation in polarized growth. Eukaryot. Cell 2014, 13, 1548–1556. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Liu, H. A GATA transcription factor recruits Hda1 in response to reduced Tor1 signaling to establish a hyphal chromatin state in Candida albicans. PLoS Pathog. 2012, 8, e1002663. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Wang, A.; Liu, H. Hyphal development in Candida albicans requires two temporally linked changes in promoter chromatin for initiation and maintenance. PLoS Biol. 2011, 9, e1001105. [Google Scholar] [CrossRef]
- Woolford, C.A.; Lagree, K.; Aleynikov, T.; Mitchell, A.P. Negative control of Candida albicans filamentation-associated gene expression by essential protein kinase gene KIN28. Curr. Genet. 2017, 63, 1073–1079. [Google Scholar] [CrossRef] [PubMed]
- Woolford, C.A.; Lagree, K.; Xu, W.; Aleynikov, T.; Adhikari, H.; Sanchez, H.; Cullen, P.J.; Lanni, F.; Andes, D.R.; Mitchell, A.P. Bypass of Candida albicans Filamentation/Biofilm Regulators through Diminished Expression of Protein Kinase Cak1. PLoS Genet. 2016, 12, e1006487. [Google Scholar] [CrossRef] [PubMed]
- Lew, R.R. How does a hypha grow? The biophysics of pressurized growth in fungi. Nat. Rev. Microbiol. 2011, 9, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Wilson, D.; Hube, B. Candida albicans pathogenicity mechanisms. Virulence 2013, 4, 119–128. [Google Scholar] [CrossRef] [PubMed]
- De Jong, J.C.; McCormack, B.J.; Smirnoff, N.; Talbot, N.J. Glycerol generates turgor in rice blast. Nature 1997, 389, 244. [Google Scholar] [CrossRef]
- Bonhomme, J.; Chauvel, M.; Goyard, S.; Roux, P.; Rossignol, T.; d’Enfert, C. Contribution of the glycolytic flux and hypoxia adaptation to efficient biofilm formation by Candida albicans. Mol. Microbiol. 2011, 80, 995–1013. [Google Scholar] [CrossRef] [PubMed]
- Desai, J.V.; Cheng, S.; Ying, T.; Nguyen, M.H.; Clancy, C.J.; Lanni, F.; Mitchell, A.P. Coordination of Candida albicans Invasion and Infection Functions by Phosphoglycerol Phosphatase Rhr2. Pathogens 2015, 4, 573–589. [Google Scholar] [CrossRef] [PubMed]
- Wachtler, B.; Wilson, D.; Haedicke, K.; Dalle, F.; Hube, B. From attachment to damage: Defined genes of Candida albicans mediate adhesion, invasion and damage during interaction with oral epithelial cells. PLoS ONE 2011, 6, e17046. [Google Scholar] [CrossRef] [PubMed]
- Phan, Q.T.; Myers, C.L.; Fu, Y.; Sheppard, D.C.; Yeaman, M.R.; Welch, W.H.; Ibrahim, A.S.; Edwards, J.E., Jr.; Filler, S.G. Als3 is a Candida albicans invasin that binds to cadherins and induces endocytosis by host cells. PLoS Biol. 2007, 5, e64. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Phan, Q.T.; Boontheung, P.; Solis, N.V.; Loo, J.A.; Filler, S.G. EGFR and HER2 receptor kinase signaling mediate epithelial cell invasion by Candida albicans during oropharyngeal infection. Proc. Natl. Acad. Sci. USA 2012, 109, 14194–14199. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Ruiz, E.; Galan-Diez, M.; Zhu, W.; Fernandez-Ruiz, E.; d’Enfert, C.; Filler, S.G.; Cossart, P.; Veiga, E. Candida albicans internalization by host cells is mediated by a clathrin-dependent mechanism. Cell Microbiol. 2009, 11, 1179–1189. [Google Scholar] [CrossRef] [PubMed]
- Phan, Q.T.; Eng, D.K.; Mostowy, S.; Park, H.; Cossart, P.; Filler, S.G. Role of endothelial cell septin 7 in the endocytosis of Candida albicans. mBio 2013, 4, e00542-13. [Google Scholar] [CrossRef] [PubMed]
- Solis, N.V.; Swidergall, M.; Bruno, V.M.; Gaffen, S.L.; Filler, S.G. The Aryl Hydrocarbon Receptor Governs Epithelial Cell Invasion during Oropharyngeal Candidiasis. mBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Desai, J.V.; van de Veerdonk, F.L.; Lionakis, M.S. Understanding the role of host immune responses in invasive candidiasis. Intensive Care Med. 2017. [Google Scholar] [CrossRef] [PubMed]
- Lionakis, M.S.; Iliev, I.D.; Hohl, T.M. Immunity against fungi. JCI Insight 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Erwig, L.P.; Gow, N.A. Interactions of fungal pathogens with phagocytes. Nat. Rev. Microbiol. 2016, 14, 163–176. [Google Scholar] [CrossRef] [PubMed]
- O’Meara, T.R.; Veri, A.O.; Ketela, T.; Jiang, B.; Roemer, T.; Cowen, L.E. Global analysis of fungal morphology exposes mechanisms of host cell escape. Nat. Commun. 2015, 6, 6741. [Google Scholar] [CrossRef] [PubMed]
- Vylkova, S.; Lorenz, M.C. Phagosomal Neutralization by the Fungal Pathogen Candida albicans Induces Macrophage Pyroptosis. Infect. Immun. 2017, 85. [Google Scholar] [CrossRef] [PubMed]
- Wachtler, B.; Citiulo, F.; Jablonowski, N.; Forster, S.; Dalle, F.; Schaller, M.; Wilson, D.; Hube, B. Candida albicans-epithelial interactions: Dissecting the roles of active penetration, induced endocytosis and host factors on the infection process. PLoS ONE 2012, 7, e36952. [Google Scholar] [CrossRef] [PubMed]
- Dalle, F.; Wachtler, B.; L’Ollivier, C.; Holland, G.; Bannert, N.; Wilson, D.; Labruere, C.; Bonnin, A.; Hube, B. Cellular interactions of Candida albicans with human oral epithelial cells and enterocytes. Cell Microbiol. 2010, 12, 248–271. [Google Scholar] [CrossRef] [PubMed]
- Desai, J.V.; Bruno, V.M.; Ganguly, S.; Stamper, R.J.; Mitchell, K.F.; Solis, N.; Hill, E.M.; Xu, W.; Filler, S.G.; Andes, D.R.; et al. Regulatory role of glycerol in Candida albicans biofilm formation. mBio 2013, 4, e00637-12. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Desai, J.V. Candida albicans Hyphae: From Growth Initiation to Invasion. J. Fungi 2018, 4, 10. https://doi.org/10.3390/jof4010010
Desai JV. Candida albicans Hyphae: From Growth Initiation to Invasion. Journal of Fungi. 2018; 4(1):10. https://doi.org/10.3390/jof4010010
Chicago/Turabian StyleDesai, Jigar V. 2018. "Candida albicans Hyphae: From Growth Initiation to Invasion" Journal of Fungi 4, no. 1: 10. https://doi.org/10.3390/jof4010010
APA StyleDesai, J. V. (2018). Candida albicans Hyphae: From Growth Initiation to Invasion. Journal of Fungi, 4(1), 10. https://doi.org/10.3390/jof4010010



