Abstract
In the standard genetic code, the CUG triplet is translated as leucine. The pathogenic yeast Candida albicans and other CTG-clade yeasts contain tRNACAG, which is recognized by both leucine- and serine-tRNA synthetases. The CUG codon in these yeasts is translated most often as serine, and only in 3–5% of cases as leucine. Therefore, CTG Candida species have unstable proteomes. The effect of serine–leucine exchange on the structure and function of proteins has only been experimentally examined for a few cases. In C. albicans, CUG codons occur even in genes deemed to be essential. This means that serine–leucine ambiguity either does not affect the structure and function of the respective proteins, or that the presence of these amino acids at specific positions is associated with meaningful alteration of the proteins’ function. This study employed AlphaFold2 to evaluate the potential effects of serine-to-leucine exchange in 12 proteins encoded by essential genes lacking orthologs in other yeasts and human genomes. The low homology with known proteins allowed us to make only low-confidence predictions. The analyzed proteins could be grouped into subsets based on the structural outcomes. Structural changes were observed only in four proteins. The remaining eight proteins showed no significant differences between serine and leucine variants.
1. Introduction
The genetic code is a universal principle uniting all living organisms. It is generally regarded as stable; nevertheless, the number of observed deviations from the standard decoding rules continues to grow. The most common coding alteration identified so far is translating one or two stop codons as an amino acid [1]. This occurs in bacteria, mitochondrial genomes of metazoa, or nuclear genomes of ciliates [1,2,3,4]. The use of all three stop codons as sense codons also occurs in nature. It has been observed in the nuclear genome of a parasitic protist, Blastocrithidia, and in eukaryotic marine microorganisms, Condylostoma magnum and Parduzcia orbis, where the stop codons can terminate translation only with the help of an adjacent poly(A) tail [5,6,7].
Sense-to-sense codon changes are rare, especially in eukaryotic nuclear genomes. Moreover, they are challenging to identify and experimentally verify [8]. Despite this, changes in the meaning of the CUG codon have been known since the end of the 1980s. In the standard genetic code, the CUG codon is translated as leucine. But in the set of yeast species of the subphylum Saccharomycotina, it is decoded as serine. This alteration was first discovered in Candida cylindracea [9] and then found in more Candida species, including the prominent human pathogens Candida albicans, Candida parapsilosis, and Candida tropicalis [10,11]. The set of yeast species possessing this feature has been denominated as the CUG (or CTG) clade. Later on, decoding of the CUG as alanine was observed in the yeast Pachysolen tannophilus, known for its ability to produce ethanol from xylose [12,13,14]. Then, Krassowski et al. reported on an additional CUG-Ser clade within the Saccharomycotina phylogenetic tree [13].
The key elements ensuring translation accuracy and stability are tRNA molecules. In C. albicans, tRNACAG has structural determinants that can be recognized by both seryl- and leucyl-tRNA synthetases. In 93–95% of cases, tRNACAG is charged with serine, while in the remaining cases, it is charged with leucine [4,15]. The dual meaning of the CUG codon in C. albicans is yet another intriguing alteration to the standard genetic code, challenging the traditional view that each codon corresponds to a single amino acid. Evolutionarily, the dual meaning of the CUG codon affected its usage. Potentially adverse effects resulting from an unstable proteome can be avoided by eliminating the altered codon from protein-coding sequences [16]. Indeed, a comparative genomics analysis of C. albicans, Saccharomyces cerevisiae, and Schizosaccharomyces pombe revealed that CUG codon ambiguity led to the disappearance of approximately 26,000–30,000 ancestral CUG codons in Candida spp. Only 2% of C. albicans CUG codons correspond to the leucine codons in S. cerevisiae and S. pombe. However, there is one more twist: over the last 272 ± 25 million years, new Ser-Leu-CUG codons have emerged, accounting for approximately 17,000 extant CUG codons in Candida. These codons align with those for serine- or conserved-serine-related amino acids in related yeast species [17,18]. The C. albicans proteome must have experienced periods of considerable instability over geological time and remains unstable to this day [15,17].
Can such instability be advantageous? One approach to addressing this question involved transforming S. cerevisiae with C. albicans Ser-tRNACAG, leading to the translation of Saccharomyces CUG codons as both leucine and serine. The resulting S. cerevisiae exhibited greater tolerance to a range of stress conditions. This suggests that the synthesis of aberrant proteins due to CUG’s dual meaning induced the expression of molecular chaperones, which then protected the cells against various stresses. Indeed, Hsp104 and Hsp70 expression was increased in S. cerevisiae with CUG codon ambiguity [19,20]. Conversely, C. albicans with increased incorporation of leucine at CUG sites displayed more variable morphology and tolerance to higher levels of azole antifungals than the parental strain [21]. C. albicans is known for its ability to withstand stress conditions, a trait considered important for its pathogenicity [22]. The polysemous CUG codon may, therefore, be one of the factors contributing to Candida’s significance as a public health threat.
C. albicans is the leading cause of yeast infections [23]. It can coexist with healthy hosts as a harmless commensal, but when the constraints imposed by the host’s immune system or competing microflora are removed, it can cause disease, termed candidiasis [24]. Individuals at risk of Candida infections include patients who have undergone organ transplantation, chemotherapy, or other medical treatments that weaken the immune system. Long-term use of broad-spectrum antibiotics, which reduce prokaryotic microflora, is another risk factor that facilitates the development of candidiasis [25]. The disease can be relatively mild, with superficial and mucosal manifestations, but in deeply immunocompromised patients, the yeast can enter the bloodstream and spread to internal organs, causing life-threatening conditions [26]. Since C. albicans can be part of the human microbiome, candidiasis can have an endogenous origin. However, it can also be acquired externally, with nosocomial transmission being a particular concern.
Several classes of drugs have been developed to treat candidiasis. They include polyenes, azoles, echinocandins, allylamines, and pyrimidine analogs, with azoles being the most frequently used [26]. While these treatments are often effective for superficial Candida infections, deep-seated candidiasis is often difficult to cure. Another challenge is that the widespread use of antifungal agents exerts selective pressure, enabling the rapid evolution of C. albicans strains with reduced drug susceptibility. This drives the search for novel drug targets and the development of new antifungal therapies [27].
To identify potential drug targets, it is crucial to determine which molecules are essential for C. albicans to survive and proliferate but are absent in humans. The genome of C. albicans has been known and annotated for a long time, allowing for the identification of essential genes at the whole-genome level. This search has been conducted using various approaches. One of the latest studies employed transposon mutagenesis in a haploid C. albicans isolate [27]. Among the genes identified as essential in this study, there is a subset that lacks orthologs not only in humans but also in several other yeast species. These genes are mostly uncharacterized, and their lack of orthologs suggests that they may be considered orphan genes, which are relatively young from an evolutionary perspective [28]. Surprisingly, some of these genes contain one or more CUG codons, indicating that C. albicans can tolerate the dual-meaning codon even in essential genes.
We used AlphaFold2 predictions to examine how serine-to-leucine changes affect the structure of proteins encoded by these essential genes lacking orthologs. Since AlphaFold2 relies on known homologous protein structures for its predictions, the analysis of this particular protein set is less reliable and should be interpreted with caution. Nevertheless, we believe it is valuable to highlight these uncharacterized and potentially interesting proteins in an important human pathogen and to discuss structural changes in these proteins, despite some uncertainty.
2. Materials and Methods
The gene sequences were obtained from the Candida Genome Database (www.candidagenome.org) and translated into amino acid sequences using the Translate Tool available at the ExPASy Bioinformatics Resource Portal (https://web.expasy.org/translate/, accessed on 4 November 2024). Positions of Ser/Leu residues encoded by the CUG codons were found manually. Intrinsic-disorder prediction was performed using NetSurfP-2.0 [29]. The structural prediction of the uncharacterized proteins and their mutant variants was carried out using a combination of computational tools. The wild-type (WT) protein structure was retrieved from the AlphaFold Protein Structure Database, which offers highly accurate structural models based on deep learning algorithms [30]. To assess the potential structural effects of specific point mutations, selected serine residues in the WT sequence were substituted with leucine residues at positions corresponding to CTG (CUG) codons. These targeted mutations were introduced using a custom Python 3 script executed in the Google Colab environment, enabling rapid in silico modification and prediction of the mutant sequence [31]. Following model generation, the three-dimensional structures of both the WT and mutant proteins were visualized and analyzed using UCSF ChimeraX (v. 1.10) [32]. Structural alignment and comparison were conducted to identify conformational changes, potential disruption of secondary structure elements, or alterations in predicted functional regions caused by the serine-to-leucine substitutions.
3. Results
The selection of the open reading frames included in this study stems from the work of Segal et al. (2018) [27], who identified 1610 C. albicans genes essential for the growth of a haploid C. albicans strain under standard laboratory conditions. Among them, 17 genes were identified as having no detectable orthologs in the human genome, no orthologs in the two model yeasts Saccharomyces cerevisiae and Schizosaccharomyces pombe, and no orthologs in three major fungal pathogens Aspergillus fumigatus, Cryptococcus neoformans, and Histoplasma capsulatum. Twelve of these open reading frames contain at least one CUG codon and became the subject of this study (Table 1).

Table 1.
List of the genes included in this study. Systematic gene names are according to the Candida Genome Database, and protein accession numbers are according to Uniprot.
Due to limited homology to known protein sequences, AlphaFold2 predicts the structures of these proteins with low confidence. With this limitation in mind, we categorized them according to the predicted influence that an ambiguous CUG codon reading may have on the resulting proteins.
3.1. Proteins with Insignificant Ser-Leu Exchange Effect
AlphaFold2 predictions suggested that in eight out of twelve ORFs, the Ser-to-Leu substitution would not significantly affect the overall protein structure (Figure 1). This finding was further supported by NetSurfP-2.0 analysis, which predicts secondary structure elements and regions of disorder. However, in several cases, Ser-Leu exchange may lead to a rearrangement in the relative positions of α-helices.
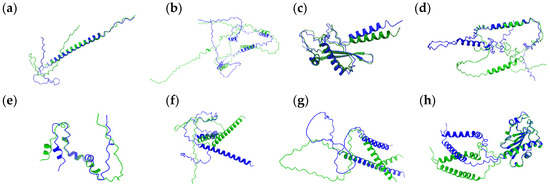
Figure 1.
Structural alignment of proteins with no statistically significant changes in their structure. Superimposed variants are shown in green and blue. Residue identities are not labeled due to a lack of statistically relevant substitutions. Panels correspond to the following proteins: (a) Q5AB58, (b) Q5A299, (c) Q5A3K8, (d) A0A1D8PG02, (e) A0A1D8PF20 (TLO4), (f) Q5ALW9, (g) A0A1D8PLD5, and (h) A0A1D8PPD0. Green (WT—predicted structure, AlphaFold2) and blue (predicted structure with S ↔ L substitution, CoLab) indicate two protein variants aligned for comparison.
The gene C1_00020C_A has been identified as haploinsufficient for cell size, with its heterozygous deletion leading to smaller cells [33]. Although transcription of C1_00020C_A has been confirmed [34], the corresponding gene product has not yet been experimentally validated. The predicted protein (Q5AB58) is 103 amino acids long and contains three Ser-Leu exchange sites, one within a predicted helical region and two within the C-terminal unstructured region. According to NetSurfP-2.0, when leucine occupies positions 92 and 97, the segment spanning residues 89–98 may adopt a helical conformation. In contrast, AlphaFold2 predicts that the C-terminal region remains disordered regardless of the Ser-Leu substitutions (Figure 1a).
Although the gene C1_05250W_A has orthologs in Candida lusitaniae and Candida guilliermondii, it remains uncharacterized. It is predicted to encode a protein of 204 amino acids (Q5A299), 4 of which are encoded by the CUG codon. These residues are at positions 71, 100, 123, and 180, located in loops or unstructured regions of the protein (Figure 1b). While they may influence the relative positioning of the three predicted helices, their overall impact on the protein’s architecture is likely to be minimal.
The protein Q5A3K8, a potential product of the C1_12040W_A locus, is a 143-amino-acid uncharacterized protein with a conserved ortholog in Candida dubliniensis. Two Ser-Leu exchange sites are present at positions 94 and 101. AlphaFold modeling (Figure 1c) indicates that residue 94 lies in a surface-exposed loop, while residue 101 is located at the C-terminal end of a short α-helix spanning residues 99–101. The Ser-Leu substitution is predicted to have no significant effect on the protein’s overall structure.
The locus C1_14580C_A is annotated in the Candida Genome Database as an uncharacterized gene associated with a transposable element. The corresponding protein sequence matches the nucleocapsid protein of the Zorro2 retrotransposon, a member of the Zorro family of non-LTR retrotransposons in Candida albicans that are known to be transcriptionally active and capable of transposition [35]. The protein (A0A1D8PG02) is composed of 175 amino acids and contains a single Ser-Leu exchange site at position 10, which is located on a loop. According to AlphaFold2 (Figure 1d), this substitution is not expected to affect the overall protein structure.
TLO4 (telomere-linked open reading frame) is one of the telomere-proximal genes of unknown function. The corresponding protein (A0A1D8PFZ0) listed in the Candida Genome Database consists of 66 amino acids, with the Ser-Leu exchange sites at positions 24 and 35. While transcription of the TLO4 gene was found to be induced during oral candidiasis and regulated by Hap43p transcription factor, the existence of the Tlo4p protein has not been experimentally verified. According to the AlphaFold2 model (Figure 1e), residues 24 and 35 are both on the helix in the central part of the protein, and Ser-Leu exchange would not cause structural changes.
C2_01500W_A is an uncharacterized open reading frame predicted to encode a 144-amino-acid protein (Q5ALW9). To date, no studies have reported on this gene. A potential Ser-Leu substitution at position 102 may slightly affect the positioning of the C-terminal helix (Figure 1f). However, the overall protein structure is likely to remain unchanged.
The gene C4_01710C_A encodes a 142-amino-acid protein (A0A1D8PLD5) that has been identified as part of the transcriptional network regulating biofilm formation in C. albicans [36]. Among the proteins analyzed in this study, it contains the highest number of residues encoded by the ambiguous CUG codon, with seven potential Ser-Leu exchange sites. Notably, this includes a continuous stretch of five residues (positions 124–128), each of which can be either serine or leucine. Both AlphaFold2 and NetSurfP-2.0 predict this stretch to be located between the two helices in the C-terminal part of the protein, with no significant effect on the protein structure. Residues 18 and 132 are predicted by AlphaFold2 to be part of short N- and C-terminal α-helices, respectively. However, NetSurfP-2.0 classifies the N-terminal region as disordered, without a defined helical structure. In either case, Ser-Leu exchanges near the termini are also predicted to have minimal structural consequences (Figure 1g). The amino acid composition of this protein is unusual, with asparagine accounting for nearly 30% of the residues and glutamine for an additional 9%. Reflecting this low sequence complexity, the central region of the protein is predicted to be intrinsically disordered by both AlphaFold2 and NetSurfP-2.0. Such an N/Q-rich region may be capable of switching between a prion-like state and a more defined fold upon binding to an interaction partner. Although the Ser-Leu exchange sites are not located within this unstructured region, CUG codon translation may still influence overall structural alterations if the disordered segment adopts specific folds.
Although the gene C6_00270W_A remains uncharacterized, it has conserved orthologs in Candida dubliniensis and Candida tropicalis. It encodes a relatively long protein of 230 amino acids (A0A1D8PPD0), making it the longest protein analyzed in this study. AlphaFold2 predictions suggest that the protein is relatively well structured. Two potential Ser-Leu exchange sites are present at positions 100 and 107, both located within a surface-exposed loop, and are not expected to affect the overall protein structure (Figure 1h).
3.2. Proteins with Predicted Serine Phosphorylation
Leucine-serine exchange can have more significant consequences if the serine is phosphorylated. Sárkány et al. suggested that some CUG-encoded serines, particularly within signal transduction proteins, could be post-translationally modified [37]. However, to the best of our knowledge, there are no confirmed examples of a CUG-encoded serine in Candida albicans that have been shown to be directly phosphorylated in vivo or in vitro. Despite this, we used Prosite to find serine residues in our sequences encoded by CUG codons, which might be phosphorylated. Potential phosphorylation sites were identified in five proteins, one site in each of them: Q5AB58, Q5A3K8, A0A1D8PG02, A0A1D8PFZ0, and A0A1D8PLD5. The usage of these sites is questionable; however, at least in the case of A0A1D8PG02, the nucleocapsid protein of the Zorro-2 retrotransposon, serine phosphorylation would not be surprising.
3.3. Proteins Affected by Ser-Leu Exchange
In four proteins, AlphaFold2 prediction indicated that Ser-Leu exchange may induce more profound changes. C1_04440W_A is an uncharacterized ORF with no references in the literature, to the best of our knowledge. It is predicted to encode a 122-amino-acid protein (Q59KZ4) with a Ser-Leu site at position 13. The AlphaFold2 models show notable differences: the Ser13 variant appears partially disordered and includes β-sheets, while the Leu13 variant adopts a more helical conformation (Figure 2a). Of note, the NetSurfP-2.0 prediction suggests that both variants are structurally similar, containing β-sheets and disordered regions.
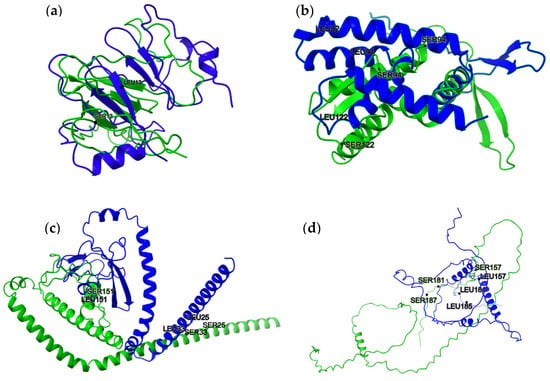
Figure 2.
Structural alignment of proteins with statistically significant differences at the indicated residue positions. Superimposed structures highlight substitutions between serine (SER) and leucine (LEU) residues in positions (a) Q59KZ4 (SER13 ↔ LEU13), (b) Q5ACW4 (SER82 ↔ LEU82, SER94 ↔ LEU94 and SER122 ↔ LEU122), (c) Q5A7K2 (SER25 ↔ LEU25, SER33 ↔ LEU33 and SER151 ↔ LEU151), and (d) A0A1D8PNC2 (SER157 ↔ LEU157, SER181 ↔ LEU181 and SER185 ↔ LEU185). Green (WT—predicted structure, Alphafold2) and blue (predicted structure with Ser ↔ Leu substitution, CoLab) indicate two protein variants aligned for comparison. Labels indicate residue identity and position.
C2_00310W_A encodes a 148-amino-acid protein (Q5ACW4) that contains three Ser-Leu exchange sites. This protein remains uncharacterized, with no reports in the literature to date. According to AlphaFold2 predictions (Figure 2b), both the serine- and leucine-containing variants are relatively well-structured, featuring a central helical core along with some β-structures. While the overall fold is largely preserved between the two versions, local topological differences are evident. The variant containing serines at all three CUG-encoded positions appears more compact. In contrast, the leucine-containing version displays extended β-strands and longer, more disordered loops. NetSurfP-2.0 also predicts the serine variant to be more compact; however, it does not identify any β-structures in either version of the protein.
C3_00120W_A encodes an uncharacterized protein consisting of 208 amino acids (Q5A7K2) and contains three Ser-Leu exchange sites. Both the serine- and leucine-containing variants are characterized by a prominent extended α-helix. In the serine-containing version, this helix is bent, resulting in a more compact overall structure. While both variants include β-structures, the serine version appears to have more tightly packed strands. Likewise, the loop regions in the serine-containing variant are more compact, exhibiting tighter turns and shorter loops compared to the leucine variant (Figure 2c).
The final protein in the analyzed panel, A0A1D8PNC2 (192 amino acids), encoded by the C5_02130W_A locus, is uncharacterized and contains three Ser-Leu exchange sites. Both the serine- and leucine-containing variants feature a prominent helical structure along with a substantial proportion of disordered regions. In this case, the leucine-containing variant appears more structured and compact compared to the more extended and disordered serine-containing version (Figure 2d).
Figure 3 illustrates the confidence levels of the protein models in which the Ser-Leu exchange is predicted to cause structural changes.
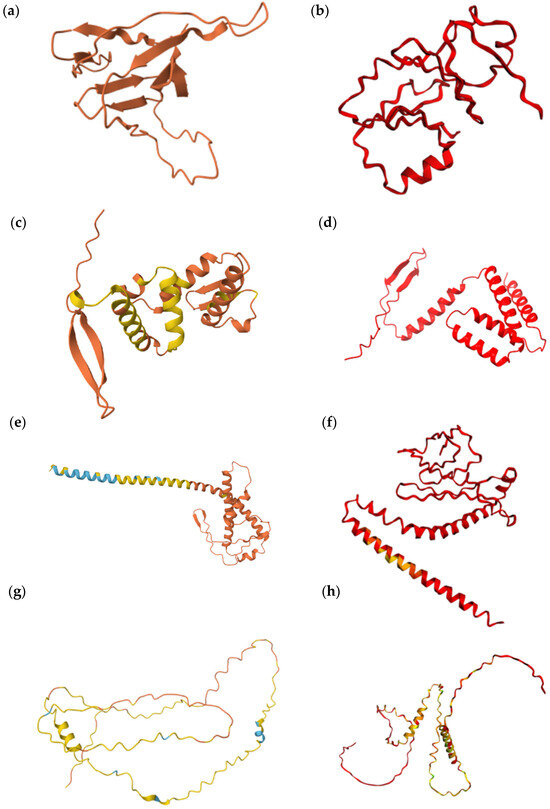
Figure 3.
Predicted protein structures. Wild-type proteins were predicted with AlphaFold2, whereas S–L exchange variants were modeled using Colab: (a) Q59KZ4 WT, (b) Q59KZ4 (Ser13 ↔ Leu13), (c) Q5ACW4 WT, (d) Q5ACW4 (Ser82 ↔ Leu82, Ser94 ↔ Leu94, and Ser122 ↔ Leu122), (e) Q5A7K2 WT, (f) Q5A7K2 (Ser25 ↔ Leu25, Ser33 ↔ Leu33, and Ser151 ↔ Leu151), (g) A0A1D8PNC2 WT, (h) A0A1D8PNC2 (Ser157 ↔ Leu157, Ser181 ↔ Leu181, and Ser185 ↔ Leu185). Model confidence levels differed between AlphaFold2 and Colab predictions. Model Confidence AlphaFold2: very high (pLDDT > 90, royal blue), high (pLDDT > 70; cyan), low (pLDDT > 50, yellow), and very low (pLDDT < 50, orange); Model Confidence CoLab: very high (pLDDT > 90, royal blue), confident pLDDT ~80, cyan), OK (pLDDT ~70, green), low (pLDDT ~60, yellow), and very low (pLDDT < 50, red).
4. Discussion
The existence of essential orphan genes in C. albicans that retain CUG codons highlights a paradox at the intersection of genetic code evolution, protein stability, and species-specific adaptation. Genes lacking homologs in other organisms are often considered to be evolutionarily young. This raises the following question: How could C. albicans have entrusted essential functions to genes that are both taxonomically unique and potentially unstable due to the dual meaning of the CUG codon?
One possibility is that the impact of the CUG codon is negligible in these proteins due to their structural robustness, particularly if the Ser-Leu substitutions occur in disordered or functionally tolerant regions. Alternatively, the dual meaning of CUG may confer a functional advantage, introducing a subtle form of adaptive flexibility that allows proteins to respond to changing conditions. Indeed, both scenarios have been documented in the literature. For example, the C. albicans translation initiation factor 4E contains a CUG site where decoding as serine or leucine results in altered protein properties, supporting the idea that codon ambiguity plays a regulatory role [38]. Conversely, the exo-β-1,3-glucanase from C. albicans showed no detectable structural differences upon Ser-Leu substitution [39], nor did the secreted aspartic protease from C. parapsilosis, another CTG-clade species [40]. These contrasting outcomes reinforce the view that the structural and functional consequences of CUG codon usage are context-dependent.
However, it remains unclear whether these codons have persisted in the essential orphan genes analyzed in this study, simply because these genes are evolutionarily young and have not yet undergone purging, or whether the CUGs were introduced after the genes became essential. The latter scenario, de novo insertion of ambiguous codons into already essential genes, seems less likely. Nonetheless, the retention of CUG codons in these genes suggests that their presence is at least selectively neutral, if not advantageous.
To address whether the presence of CUG codons in these essential orphan genes reflects protein robustness or adds a potential functional advantage, we employed AlphaFold2 and other structural prediction tools to compare the serine- and leucine-containing variants of selected proteins. These analyses aimed to distinguish between cases where the Ser-Leu substitution has minimal structural impact—suggesting robustness—and those where the exchange may induce conformational differences that could confer functional versatility. However, it is important to note that all predictions must be interpreted with caution. Due to the lack of sequence homology and structural templates for orphan proteins, the confidence of these models is inherently limited, particularly in disordered regions or unconventional folds. It is also important to emphasize that most of these proteins have not been experimentally confirmed at the protein level; their existence remains predicted based on gene models and computational annotation.
In eight cases, AlphaFold2 modeling suggested that the Ser-Leu exchange would not lead to significant structural changes. These proteins typically contained extensive unstructured regions, which may contribute to their apparent tolerance to amino acid variation at the CUG-encoded sites. The flexibility of intrinsically disordered regions likely buffers the structural impact of Ser-Leu substitution, supporting the idea of protein robustness. On the other hand, unstructured regions have the potential to adopt defined conformations upon interaction with other proteins, nucleic acids, or other types of ligands. An open question is whether the translation of the CUG codon can contribute to such structural changes. In this study, this issue is particularly relevant for proteins Q5AB58, A0A1D8PLD5, and A0A1D8PNC2, which contain large disordered regions with Ser-Leu exchange sites located either within these regions or in their proximity.
In four of the analyzed proteins, AlphaFold2 modeling predicted structural alterations resulting from Ser-Leu exchange at CUG-encoded positions. The case of Q59KZ4 is particularly striking. It contains only a single CUG site, yet the predicted structural difference between the serine and leucine variants was substantial. This observation is difficult to interpret, as it would imply that a single amino acid substitution exerts a disproportionately large effect on the overall fold. This outcome, while possible in theory, remains speculative and without experimental validation. Two other proteins, Q5ACW4 and Q5A7K2, each harbor three CUG sites. In both cases, the serine-containing variants appeared more compact compared to their leucine counterparts. This shift in conformation may point to differences in protein flexibility or stability and raises the possibility that the dual encoding could confer functional or regulatory adaptability under varying cellular conditions. A similar observation was made for the last protein, A0A1D8PNC2, which also contains three CUG sites. In this case, however, the leucine-containing variant appeared more structured and compact than the serine variant. This inverse pattern further supports the notion that Ser-Leu exchange can influence protein conformation in a context-dependent manner, potentially contributing to functional diversity.
Another layer of complexity arises from the potential post-translational modification of CUG-encoded serines through phosphorylation. While phosphorylation of serines resulting from CUG translation has not yet been experimentally confirmed in C. albicans, it remains a plausible regulatory mechanism—particularly in proteins exposed to dynamic cellular processes. One notable candidate is the predicted Zorro-2 nucleocapsid protein A0A1D8PG02, where the CUG-encoded serine is located in a region likely to be accessible and conformationally flexible. Given the role of nucleocapsid proteins in retrotransposon packaging and regulation, phosphorylation at this site could modulate protein–protein or protein–RNA interactions, influence assembly dynamics, or serve as a regulatory switch in retrotransposition [41].
In this study, we aimed to draw attention to a subset of potentially interesting and under-studied genes, essential orphan genes in C. albicans that retain CUG codons. These genes occupy a unique intersection of evolutionary novelty, genetic code ambiguity, and functional indispensability. They have not been studied from the perspective of virulence, but their essentiality makes them relevant as potential drug targets. While our structural predictions provide initial insights, a central question remains unresolved: How can C. albicans rely on proteins encoded by genes that are both unique and subject to the dual interpretation of one of the codons?
Author Contributions
O.H. conceived the study, selected the ORFs for analysis, and drafted the manuscript. M.Č. performed the majority of the structural modeling and contributed to writing the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
The data presented in this study are available in Laboratory of Pathogenic Yeasts at https://natur.cuni.cz/chemie/katedry-a-pracoviste/katedra-biochemie/veda-a-vyzkum/laborator-biochem accessed on 4 November 2024.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| AA | Amino Acid |
| ORF | Open Reading Frame |
| P | Phosphorylation |
References
- Chen, W.; Geng, Y.; Zhang, B.; Yan, Y.; Zhao, F.; Miao, M. Stop or Not: Genome-Wide Profiling of Reassigned Stop Codons in Ciliates. Mol. Biol. Evol. 2023, 40, msad064. [Google Scholar] [CrossRef]
- Yamao, F.; Muto, A.; Kawauchi, Y.; Iwami, M.; Iwagami, S.; Azumi, Y.; Osawa, S. UGA Is Read as Tryptophan in Mycoplasma Capricolum. Proc. Natl. Acad. Sci. USA 1985, 82, 2306–2309. [Google Scholar] [CrossRef] [PubMed]
- Knight, R.D.; Landweber, L.F.; Yarus, M. How Mitochondria Redefine the Code. J. Mol. Evol. 2001, 53, 299–313. [Google Scholar] [CrossRef]
- Miranda, I.; Silva, R.; Santos, M.A.S. Evolution of the Genetic Code in Yeasts. Yeast 2006, 23, 203–213. [Google Scholar] [CrossRef]
- Záhonová, K.; Kostygov, A.Y.; Ševčíková, T.; Yurchenko, V.; Eliáš, M. An Unprecedented Non-Canonical Nuclear Genetic Code with All Three Termination Codons Reassigned as Sense Codons. Curr. Biol. 2016, 26, 2364–2369. [Google Scholar] [CrossRef] [PubMed]
- Swart, E.C.; Serra, V.; Petroni, G.; Nowacki, M. Genetic Codes with No Dedicated Stop Codon: Context-Dependent Translation Termination. Cell 2016, 166, 691–702. [Google Scholar] [CrossRef]
- Kachale, A.; Pavlíková, Z.; Nenarokova, A.; Roithová, A.; Durante, I.M.; Miletínová, P.; Záhonová, K.; Nenarokov, S.; Votýpka, J.; Horáková, E.; et al. Short tRNA Anticodon Stem and Mutant eRF1 Allow Stop Codon Reassignment. Nature 2023, 613, 751–758. [Google Scholar] [CrossRef]
- Kollmar, M.; Mühlhausen, S. Nuclear Codon Reassignments in the Genomics Era and Mechanisms behind Their Evolution. BioEssays 2017, 39, 1600221. [Google Scholar] [CrossRef]
- Ohama, T.; Suzuki, T.; Mori, M.; Osawa, S.; Ueda, T.; Watanabe, K.; Nakase, T. Non-Universal Decoding of the Leucine Codon CUG in Several Candida Species. Nucleic Acids Res. 1993, 21, 4039–4045. [Google Scholar] [CrossRef]
- Ueda, T.; Suzuki, T.; Tokogawa, T.; Nishikawa, K.; Watanabe, K. Unique Structure of New Serine tRNAs Responsible for Decoding Leucine Codon CUG in Various Candida Species and Their Putative Ancestral tRNA Genes. Biochimie 1994, 76, 1217–1222. [Google Scholar] [CrossRef] [PubMed]
- Mühlhausen, S.; Findeisen, P.; Plessmann, U.; Urlaub, H.; Kollmar, M. A Novel Nuclear Genetic Code Alteration in Yeasts and the Evolution of Codon Reassignment in Eukaryotes. Genome Res. 2016, 26, 945–955. [Google Scholar] [CrossRef]
- Saini, J.K.; Saini, R.; Tewari, L. Lignocellulosic Agriculture Wastes as Biomass Feedstocks for Second-Generation Bioethanol Production: Concepts and Recent Developments. 3 Biotech. 2015, 5, 337–353. [Google Scholar] [CrossRef] [PubMed]
- Krassowski, T.; Coughlan, A.Y.; Shen, X.-X.; Zhou, X.; Kominek, J.; Opulente, D.A.; Riley, R.; Grigoriev, I.V.; Maheshwari, N.; Shields, D.C.; et al. Evolutionary Instability of CUG-Leu in the Genetic Code of Budding Yeasts. Nat. Commun. 2018, 9, 1887. [Google Scholar] [CrossRef]
- Suzuki, T. The ‘polysemous’ Codon_a Codon with Multiple Amino Acid Assignment Caused by Dual Specificity of tRNA Identity. EMBO J. 1997, 16, 1122–1134. [Google Scholar] [CrossRef]
- Knight, R.D.; Freeland, S.J.; Landweber, L.F. Rewiring the Keyboard: Evolvability of the Genetic Code. Nat. Rev. Genet. 2001, 2, 49–58. [Google Scholar] [CrossRef]
- Massey, S.E.; Moura, G.; Beltrão, P.; Almeida, R.; Garey, J.R.; Tuite, M.F.; Santos, M.A.S. Comparative Evolutionary Genomics Unveils the Molecular Mechanism of Reassignment of the CTG Codon in Candida spp. Genome Res. 2003, 13, 544–557. [Google Scholar] [CrossRef]
- Butler, G.; Rasmussen, M.D.; Lin, M.F.; Santos, M.A.S.; Sakthikumar, S.; Munro, C.A.; Rheinbay, E.; Grabherr, M.; Forche, A.; Reedy, J.L.; et al. Evolution of Pathogenicity and Sexual Reproduction in Eight Candida Genomes. Nature 2009, 459, 657–662. [Google Scholar] [CrossRef]
- Santos, M.A.; Perreau, V.M.; Tuite, M.F. Transfer RNA Structural Change Is a Key Element in the Reassignment of the CUG Codon in Candida albicans. EMBO J. 1996, 15, 5060–5068. [Google Scholar] [CrossRef] [PubMed]
- Santos, M.A.S.; Cheesman, C.; Costa, V.; Moradas-Ferreira, P.; Tuite, M.F. Selective Advantages Created by Codon Ambiguity Allowed for the Evolution of an Alternative Genetic Code in Candida spp. Mol. Microbiol. 1999, 31, 937–947. [Google Scholar] [CrossRef] [PubMed]
- Bezerra, A.R.; Simões, J.; Lee, W.; Rung, J.; Weil, T.; Gut, I.G.; Gut, M.; Bayés, M.; Rizzetto, L.; Cavalieri, D.; et al. Reversion of a Fungal Genetic Code Alteration Links Proteome Instability with Genomic and Phenotypic Diversification. Proc. Natl. Acad. Sci. USA 2013, 110, 11079–11084. [Google Scholar] [CrossRef]
- Brown, A.J.P.; Budge, S.; Kaloriti, D.; Tillmann, A.; Jacobsen, M.D.; Yin, Z.; Ene, I.V.; Bohovych, I.; Sandai, D.; Kastora, S.; et al. Stress Adaptation in a Pathogenic Fungus. J. Exp. Biol. 2014, 217, 144–155. [Google Scholar] [CrossRef]
- Katsipoulaki, M.; Stappers, M.H.T.; Malavia-Jones, D.; Brunke, S.; Hube, B.; Gow, N.A.R. Candida albicans and Candida glabrata: Global Priority Pathogens. Microbiol. Mol. Biol. Rev. 2024, 88, e00021-23. [Google Scholar] [CrossRef]
- Rai, L.S.; Wijlick, L.V.; Bougnoux, M.; Bachellier-Bassi, S.; d’Enfert, C. Regulators of Commensal and Pathogenic Life-styles of an Opportunistic Fungus—Candida albicans. Yeast 2021, 38, 243–250. [Google Scholar] [CrossRef]
- Lass-Flörl, C.; Steixner, S. The Changing Epidemiology of Fungal Infections. Mol. Aspects Med. 2023, 94, 101215. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Lionakis, M.S.; Arendrup, M.C.; Ostrosky-Zeichner, L.; Kullberg, B.J. Invasive Candidiasis. Nat. Rev. Dis. Primer 2018, 4, 18026. [Google Scholar] [CrossRef]
- Wall, G.; Lopez-Ribot, J.L. Current Antimycotics, New Prospects, and Future Approaches to Antifungal Therapy. Antibiotics 2020, 9, 445. [Google Scholar] [CrossRef]
- Segal, E.S.; Gritsenko, V.; Levitan, A.; Yadav, B.; Dror, N.; Steenwyk, J.L.; Silberberg, Y.; Mielich, K.; Rokas, A.; Gow, N.A.R.; et al. Gene Essentiality Analyzed by In Vivo Transposon Mutagenesis and Machine Learning in a Stable Haploid Isolate of Candida albicans. mBio 2018, 9, e02048-18. [Google Scholar] [CrossRef] [PubMed]
- Tautz, D.; Domazet-Lošo, T. The Evolutionary Origin of Orphan Genes. Nat. Rev. Genet. 2011, 12, 692–702. [Google Scholar] [CrossRef]
- Klausen, M.S.; Jespersen, M.C.; Nielsen, H.; Jensen, K.K.; Jurtz, V.I.; Sønderby, C.K.; Sommer, M.O.A.; Winther, O.; Nielsen, M.; Petersen, B.; et al. NetSurfP-2.0: Improved Prediction of Protein Structural Features by Integrated Deep Learning. Proteins Struct. Funct. Bioinforma. 2019, 87, 520–527. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Bisong, E. Google Colaboratory. In Building Machine Learning and Deep Learning Models on Google Cloud Platform; Apress: Berkeley, CA, USA, 2019; pp. 59–64. ISBN 978-1-4842-4469-2. [Google Scholar]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure Visualization for Researchers, Educators, and Developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Chaillot, J.; Cook, M.A.; Corbeil, J.; Sellam, A. Genome-Wide Screen for Haploinsufficient Cell Size Genes in the Opportunistic Yeast Candida albicans. G3 Genes|Genomes|Genetics 2017, 7, 355–360. [Google Scholar] [CrossRef]
- Sellam, A.; Hogues, H.; Askew, C.; Tebbji, F.; Van Het Hoog, M.; Lavoie, H.; Kumamoto, C.A.; Whiteway, M.; Nantel, A. Experimental Annotation of the Human Pathogen Candida albicans Coding and Noncoding Transcribed Regions Using High-Resolution Tiling Arrays. Genome Biol. 2010, 11, R71. [Google Scholar] [CrossRef]
- Goodwin, T.J.D.; Ormandy, J.E.; Poulter, R.T.M. L1-like Non-LTR Retrotransposons in the Yeast Candida albicans. Curr. Genet. 2001, 39, 83–91. [Google Scholar] [CrossRef]
- Nobile, C.J.; Fox, E.P.; Nett, J.E.; Sorrells, T.R.; Mitrovich, Q.M.; Hernday, A.D.; Tuch, B.B.; Andes, D.R.; Johnson, A.D. A Recently Evolved Transcriptional Network Controls Biofilm Development in Candida albicans. Cell 2012, 148, 126–138. [Google Scholar] [CrossRef]
- Sárkány, Z.; Silva, A.; Pereira, P.J.B.; Macedo-Ribeiro, S. Ser or Leu: Structural Snapshots of Mistranslation in Candida albicans. Front. Mol. Biosci. 2014, 1, 27. [Google Scholar] [CrossRef]
- Feketová, Z.; Mašek, T.; Vopálenský, V.; Pospíšek, M. Ambiguous Decoding of the CUG Codon Alters the Functionality of the Candida albicans Translation Initiation Factor 4E: Candida albicans eIF4E. FEMS Yeast Res. 2010, 10, 558–569. [Google Scholar] [CrossRef][Green Version]
- Cutfield, J.F.; Sullivan, P.A.; Cutfield, S.M. Minor Structural Consequences of Alternative CUG Codon Usage (Ser for Leu) in Candida albicans Exoglucanase. Protein Eng. Des. Sel. 2000, 13, 735–738. [Google Scholar] [CrossRef] [PubMed]
- Dostál, J.; Brynda, J.; Hrušková-Heidingsfeldová, O.; Sieglová, I.; Pichová, I.; Řezáčová, P. The Crystal Structure of the Secreted Aspartic Protease 1 from Candida parapsilosis in Complex with Pepstatin A. J. Struct. Biol. 2009, 167, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Botova, M.; Camacho-Zarco, A.R.; Tognetti, J.; Bessa, L.M.; Guseva, S.; Mikkola, E.; Salvi, N.; Maurin, D.; Herrmann, T.; Blackledge, M. A Specific Phosphorylation-Dependent Conformational Switch in SARS-CoV-2 Nucleocapsid Protein Inhibits RNA Binding. Sci. Adv. 2024, 10, eaax2323. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).