First Report of Colletotrichum kahawae Causing Anthracnose on Buckwheat (Fagopyrum tataricum) in China and Biological Characterization of the Pathogen
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation and Purification of Pathogens
2.2. Pathogenicity Tests of Isolate
2.3. Morphological Identification of Fungal Isolates
2.4. Molecular Identification of Fungal Isolates
2.5. Biological Characterization Studies
2.6. Statistical Analysis
3. Results
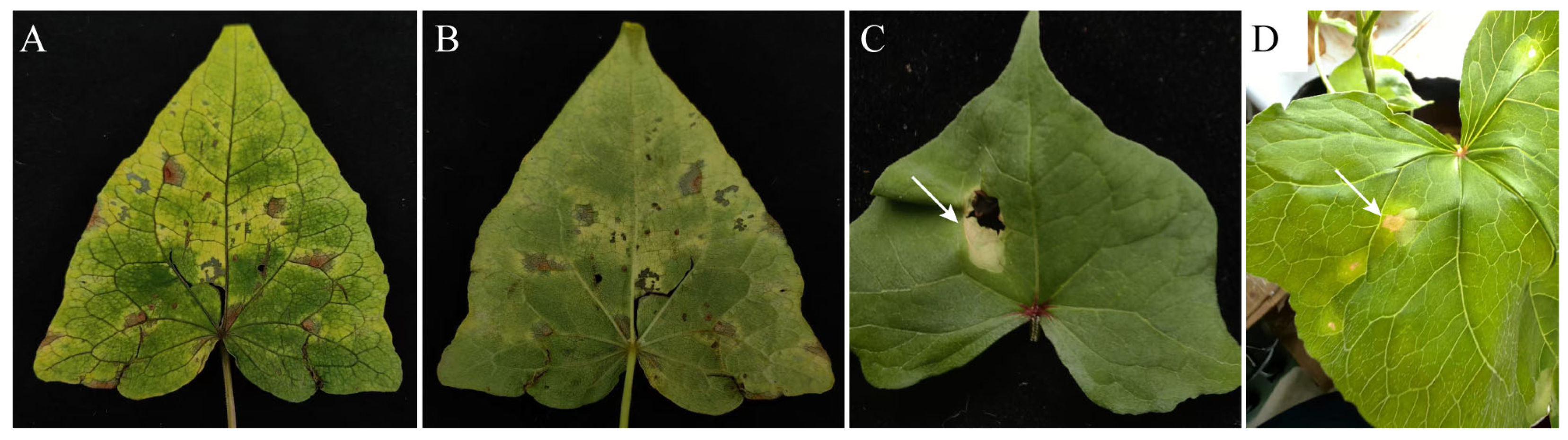
3.1. Natural Symptoms
3.2. Pathogenicity Tests
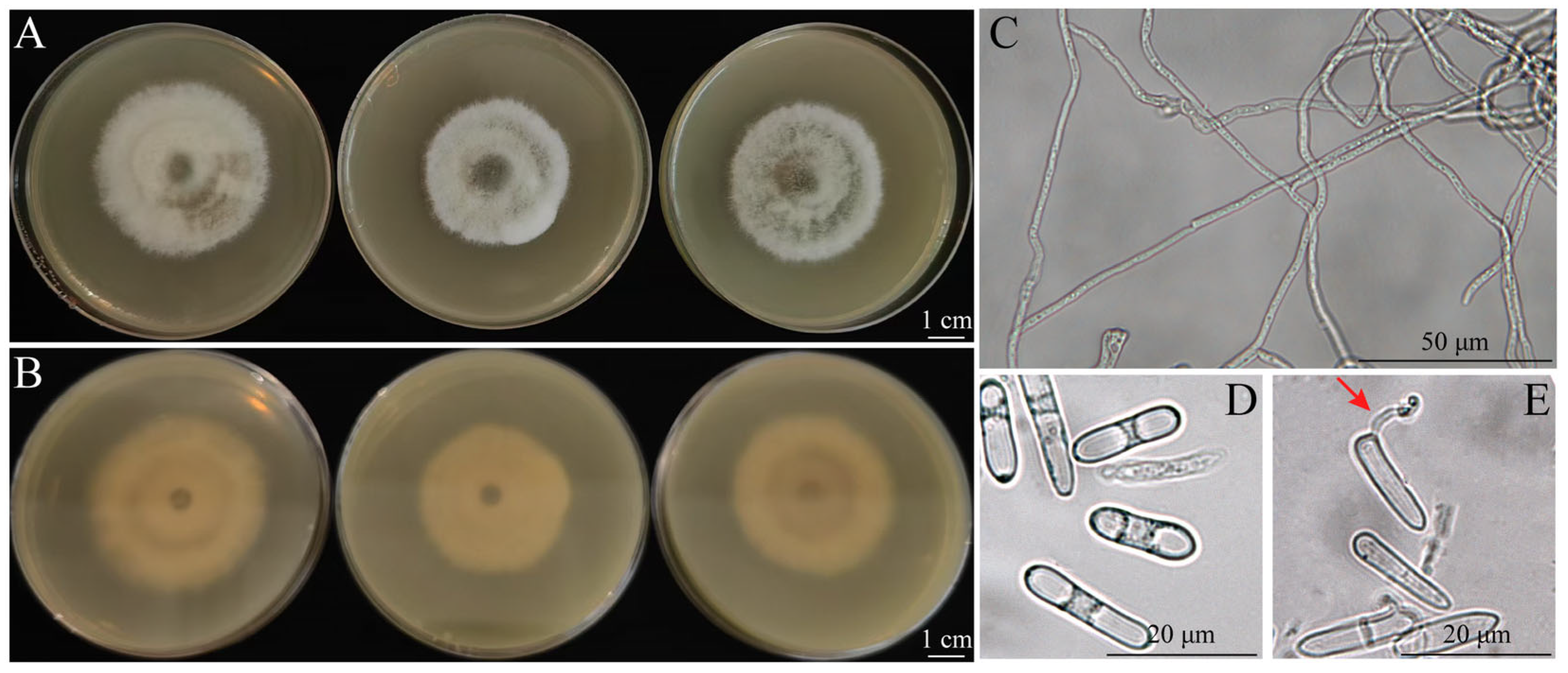
3.3. Morphology of Fungal Isolates
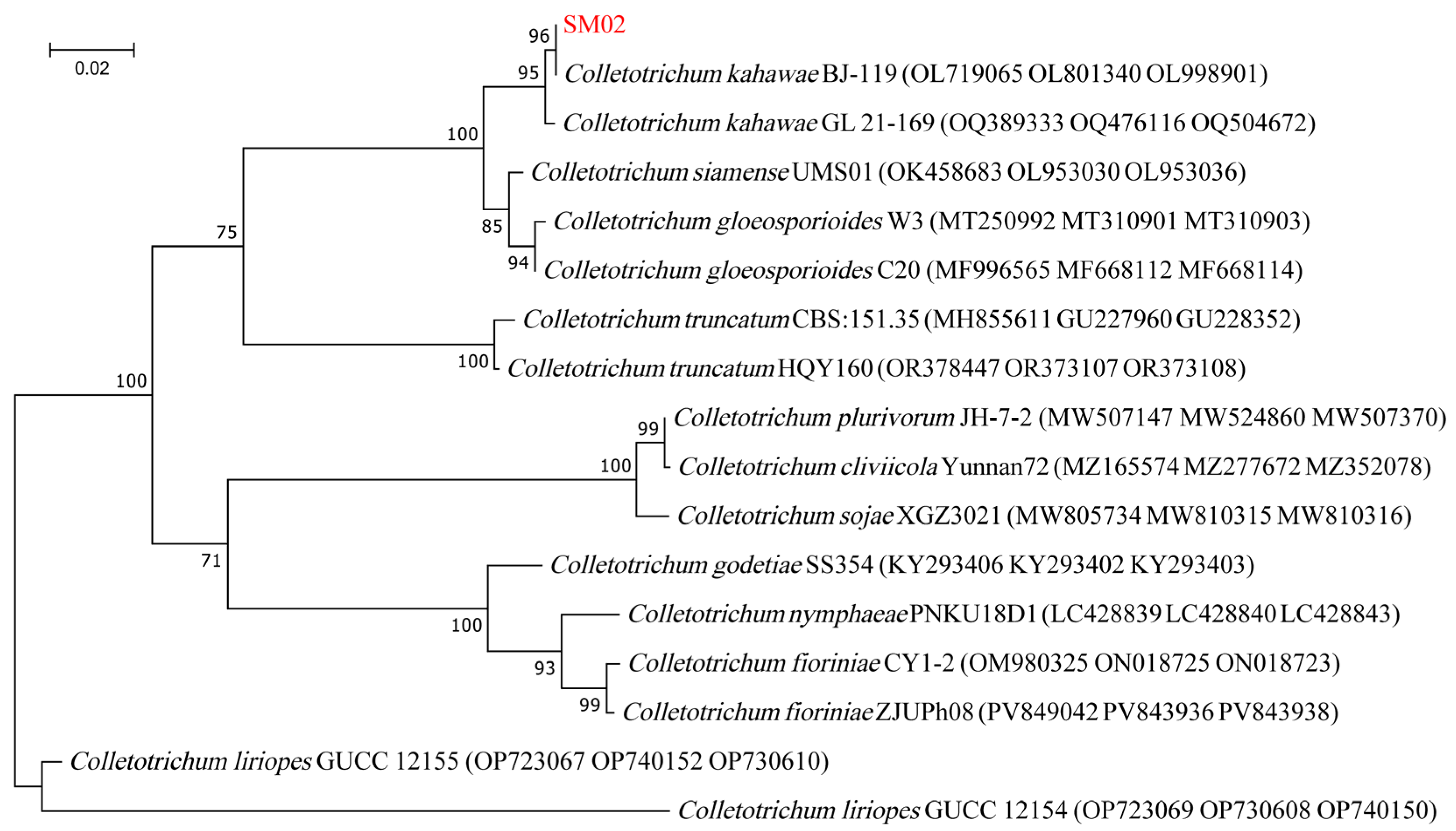
3.4. Phylogenetic Analysis
3.5. Biological Characterizations
3.5.1. Effects of Photoperiod on Growth
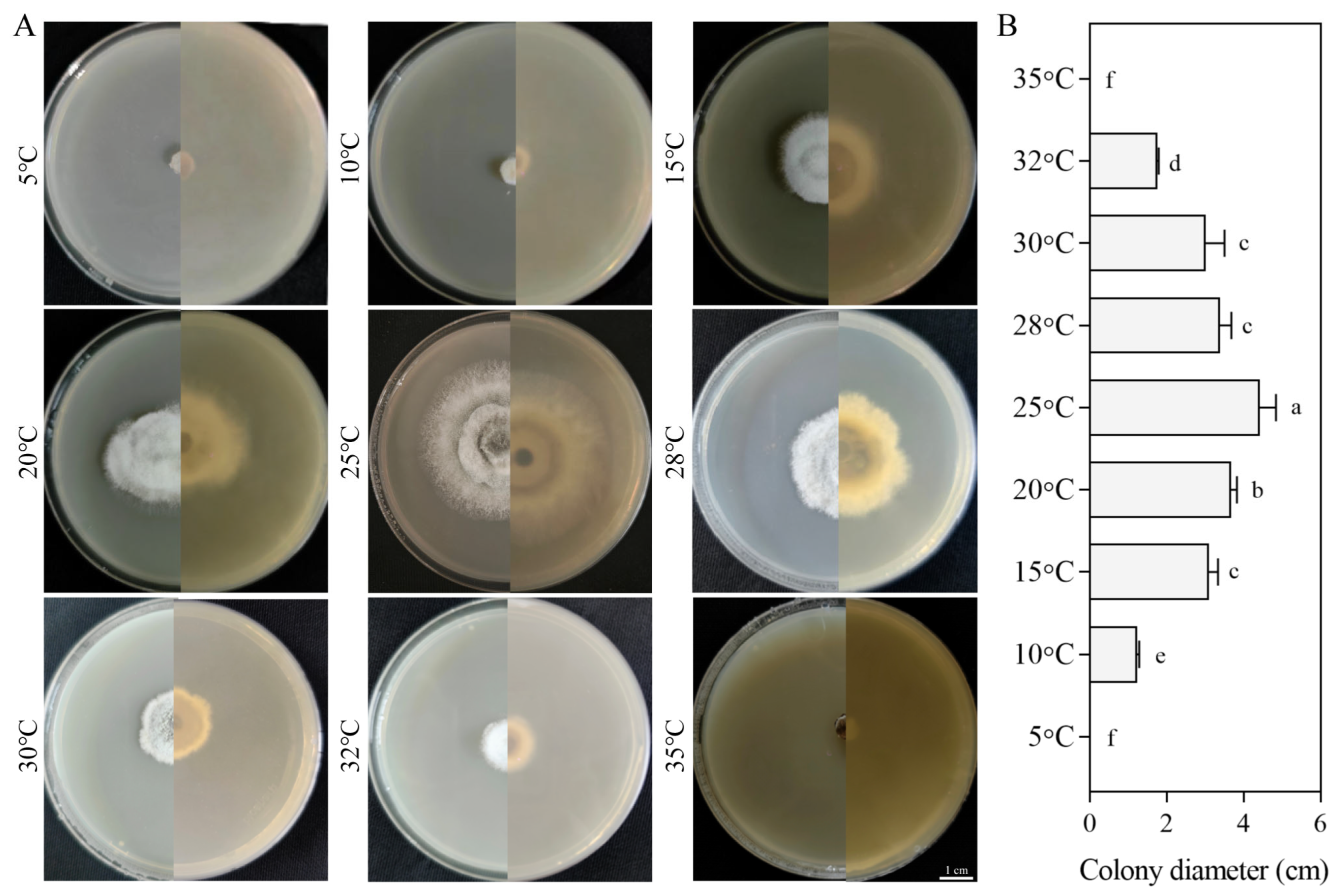
3.5.2. Growth Response to Temperatures
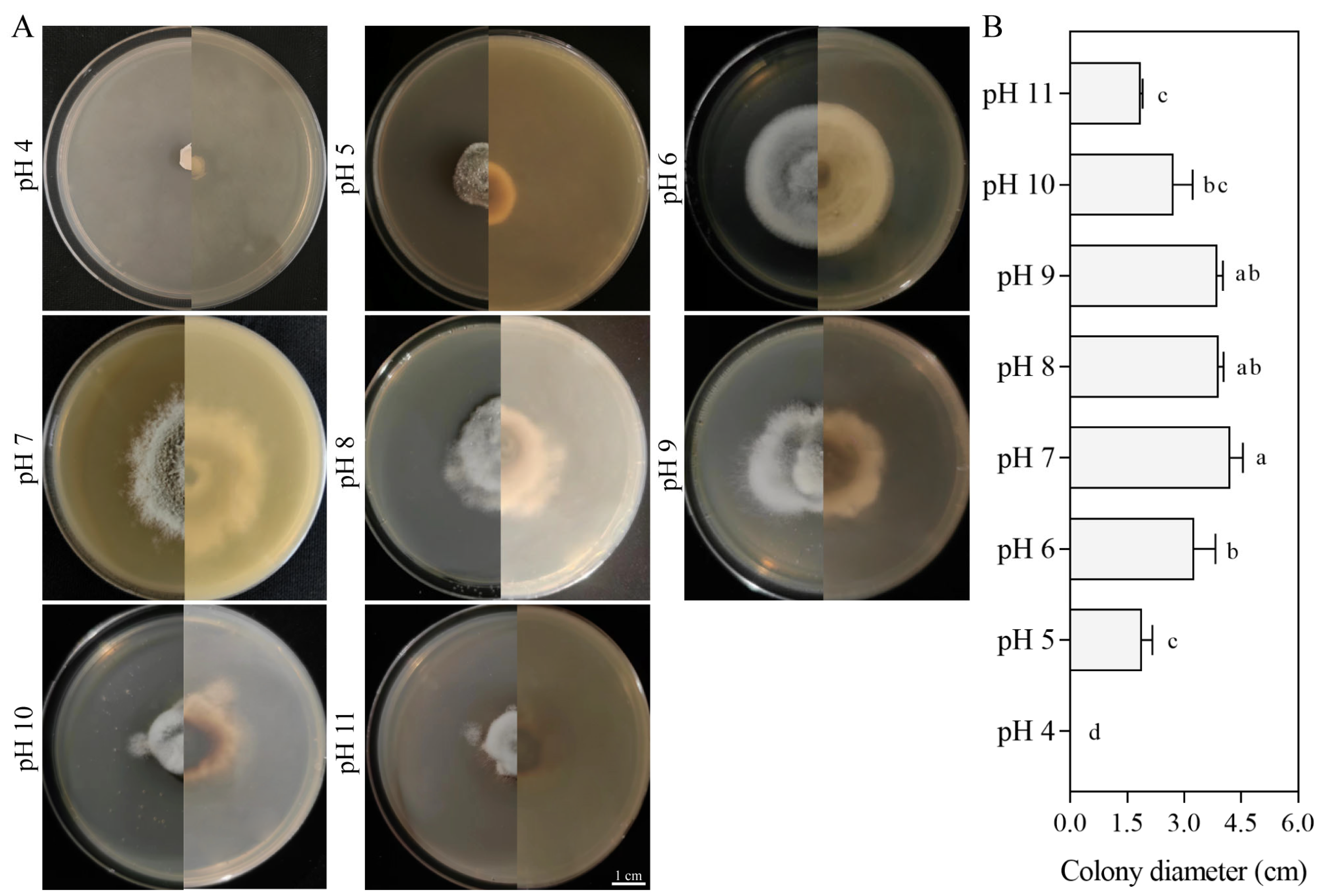
3.5.3. Effect of pH on Growth
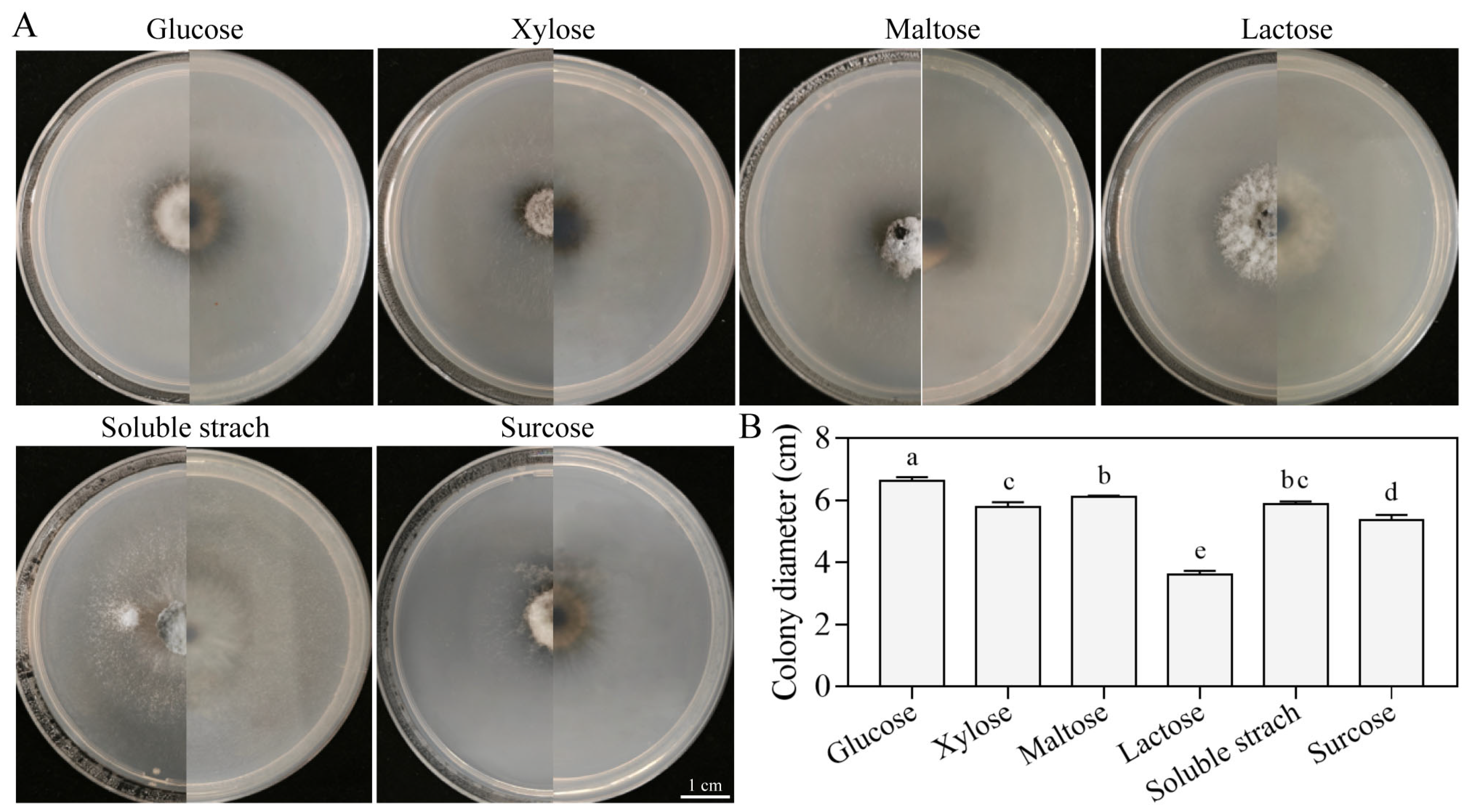
3.5.4. Growth Characteristics on Different Carbon Sources
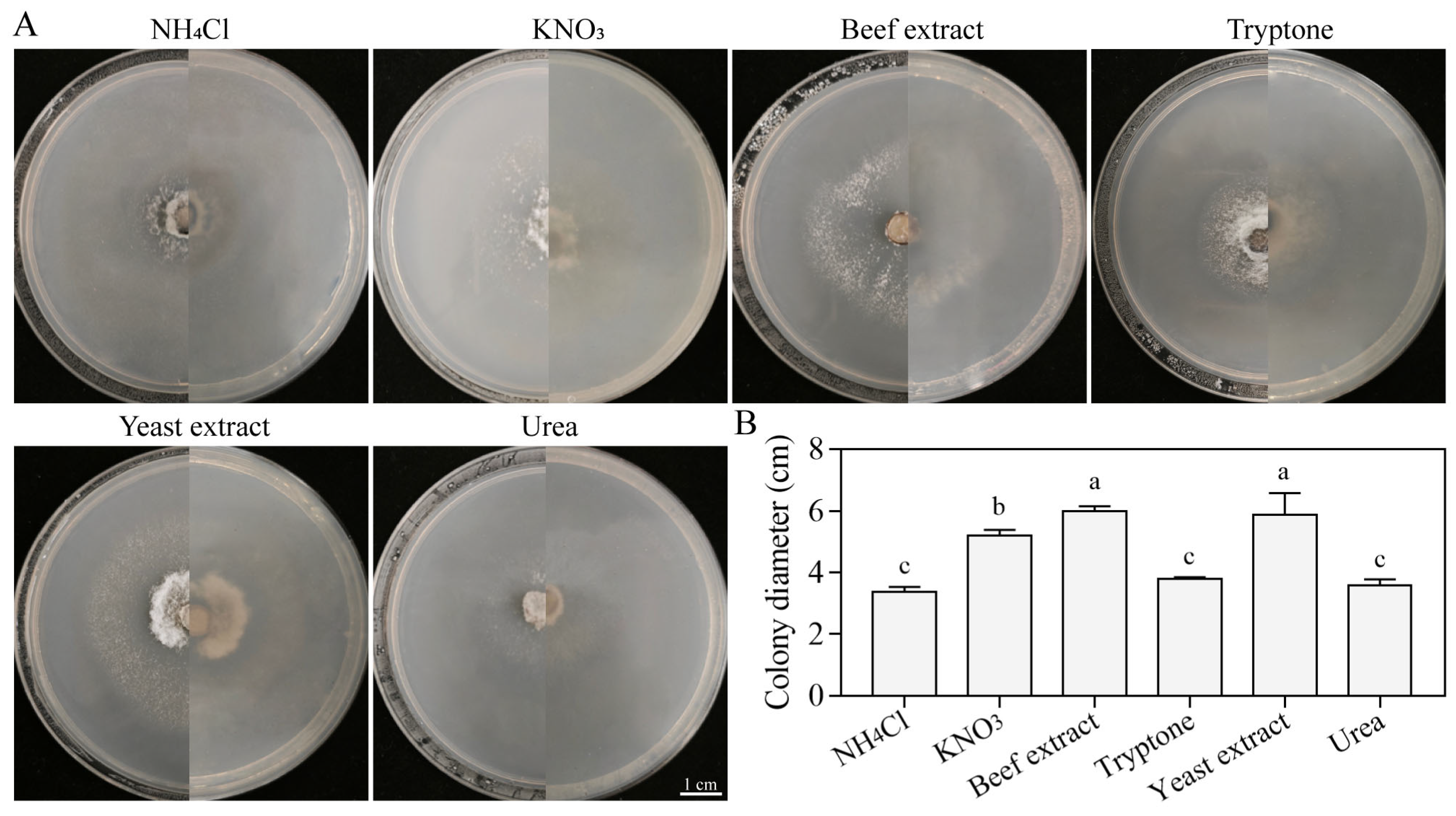
3.5.5. Growth Characteristics on Different Nitrogen Sources
3.6. Pathogenicity Test for Four Crop Plants
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- He, Y.; Zhang, K.; Shi, Y.; Lin, H.; Huang, X.; Lu, X.; Wang, Z.; Li, W.; Feng, X.; Shi, T.; et al. Genomic insight into the origin, domestication, dispersal, diversification and human selection of Tartary buckwheat. Genome Biol. 2024, 25, 61. [Google Scholar] [CrossRef]
- Wen, W.; Li, Z.; Shao, J.; Tang, Y.; Zhao, Z.; Yang, J.; Ding, M.; Zhu, X.; Zhou, M. The distribution and sustainable utilization of buckwheat resources under climate change in China. Plants 2021, 10, 2081. [Google Scholar] [CrossRef]
- Sharma, S.; Kumar, S.; Singh, R.K. A recent advance on phytochemicals, nutraceutical and pharmacological activities of buckwheat. Comb. Chem. High Throughput Screen. 2024, 27, 2654–2666. [Google Scholar] [CrossRef]
- Chen, T.X.; Wang, T.; Gou, Y.D.; Wang, L.H.; Li, C.J. Identification of Colletotrichum liriopes as the causative agent of anthracnose in buckwheat (Fagopyrum esculentum) in China. Plant Dis. 2021, 105, 3741. [Google Scholar] [CrossRef]
- Zhao, S.-Y.; Wang, Q.-T.; Guo, M.-J.; Hou, C.-L. Colletotrichum xishanense as a new etiological anthracnose agent on Fagopyrum gracilipes in Yunnan Province, China. Phytotaxa 2024, 644, 201–210. [Google Scholar] [CrossRef]
- Taixiang, C.; Qi, Y.; Wang, L.; White, J.; Li, C. First report of leaf spot disease caused by Stemphylium vesicarium on Fagopyrum esculentum in China. Plant Dis. 2021, 105, 2242. [Google Scholar] [CrossRef]
- Hsiao, H.-Y.; Ariyawansa, H.; Hsu, C.-C.; Wang, C.-J.; Shen, Y.-M. New records of powdery mildews from Taiwan: Erysiphe ipomoeae comb. nov., E. aff. betae on Buckwheat, and E. neolycopersici comb. nov. on Cardiospermum halicacabum. Diversity 2022, 14, 204. [Google Scholar] [CrossRef]
- Dean, R.; Van Kan, J.A.; Pretorius, Z.A.; Hammond-Kosack, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar] [CrossRef]
- Talhinhas, P.; Baroncelli, R. Colletotrichum species and complexes: Geographic distribution, host range and conservation status. Fungal Divers. 2021, 110, 109–198. [Google Scholar] [CrossRef]
- da Silva, L.L.; Moreno, H.L.A.; Correia, H.L.N.; Santana, M.F.; de Queiroz, M.V. Colletotrichum: Species complexes, lifestyle, and peculiarities of some sources of genetic variability. Appl. Microbiol. Biotechnol. 2020, 104, 1891–1904. [Google Scholar] [CrossRef] [PubMed]
- Garrido, C.; Carbú, M.; Fernández-Acero, F.; Vallejo, I.; Cantoral, J. Phylogenetic relationships and genome organisation of Colletotrichum acutatum causing anthracnose in strawberry. Eur. J. Plant Pathol. 2009, 125, 397–411. [Google Scholar] [CrossRef]
- Xiao, C.; Mackenzie, S.; Legard, D. Genetic and pathogenic analyses of Colletotrichum gloeosporioides isolates from strawberry and noncultivated hosts. Phytopathology 2004, 94, 446–453. [Google Scholar] [CrossRef]
- Peres, N.A.; Mackenzie, S.J.; Peever, T.L.; Timmer, L.W. Postbloom fruit drop of citrus and key lime anthracnose are caused by distinct phylogenetic lineages of Colletotrichum acutatum. Phytopathology 2008, 98, 345–352. [Google Scholar] [CrossRef]
- Talhinhas, P.; Mota Capitão, C.; Martins, S.; Ramos, A.; Martins, J.; Guerra-Guimaraes, L.; Várzea, V.; Silva, M.d.C.; Sreenivasaprasad, S.; Oliveira, H. Epidemiology, histopathology and aetiology of olive anthracnose caused by Colletotrichum acutatum and C. gloeosporioides in Portugal. Plant Pathol. 2011, 60, 483–495. [Google Scholar] [CrossRef]
- Freeman, S.; Katan, T.; Shabi, E. Characterization of Colletotrichum species responsible for anthracnose diseases of various fruits. Plant Dis. 1998, 82, 596–605. [Google Scholar] [CrossRef]
- Timmer, L.W.; Brown, G.E.; Zitko, S.E. The role of Colletotrichum spp. in postharvest anthracnose of citrus and survival of C. acutatum on fruit. Plant Dis. 1998, 82, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Ding, Z.; Zhou, M.; Shang, Y.; Li, C.; Li, Q.; Bu, T.; Tang, Z.; Chen, H. The diversity of endophytic fungi in Tartary buckwheat (Fagopyrum tataricum) and its correlation with flavonoids and phenotypic traits. Front. Microbiol. 2024, 15, 1360988. [Google Scholar] [CrossRef]
- Gomdola, D.; Jeewon, R.; McKenzie, E.H.C.; Jayawardena, R.S.; Al-Otibi, F.; Tang, X.; Wang, Y.; Hyde, K.D.; Fu, L. Phylogenetic diversity of Colletotrichum species (Sordariomycetes, Glomerellales, Glomerellaceae) associated with plant diseases in Thailand. MycoKeys 2025, 119, 137–195. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Hyde, K.; Taylor, P.; Weir, B.; Waller, J.; Abang, M.; Zhang, J.Z.; Yang, Y.L.; Phoulivong, S.; Liu, Z.Y. A polyphasic approach for studying Colletotrichum. Fungal Divers. 2009, 39, 183–204. [Google Scholar]
- Jayawardena, R.; Hyde, K.; Damm, U.; Cai, L.; Liu, M.; Li, X.H.; Zhang, W.; Zhao, W.; Yan, J. Notes on currently accepted species of Colletotrichum. Mycosphere 2016, 7, 1192–1260. [Google Scholar] [CrossRef]
- Liu, F.; Ma, Z.Y.; Hou, L.W.; Diao, Y.Z.; Wu, W.P.; Damm, U.; Song, S.; Cai, L. Updating species diversity of Colletotrichum, with a phylogenomic overview. Stud. Mycol. 2022, 101, 1–56. [Google Scholar] [CrossRef]
- Weir, B.S.; Johnston, P.R.; Damm, U. The Colletotrichum gloeosporioides species complex. Stud. Mycol. 2012, 73, 115–180. [Google Scholar] [CrossRef] [PubMed]
- Sreenivasaprasad, S.; Brown, A.; Mills, P. Coffee berry disease pathogen in Africa: Genetic structure and relationship to the group species Colletotrichum gloeosporioides. Mycol. Res. 1993, 97, 995–1000. [Google Scholar] [CrossRef]
- Ferrucho, R.; Marin Ramirez, G.; Ochoa Corona, F.; Angel-Calle, C. PCR-Based detection for the quarantine fungus Colletotrichum kahawae, a biosecurity threat to the coffee (Coffea arabica) industry worldwide. Plant Dis. 2024, 108, 2615–2624. [Google Scholar] [CrossRef]
- Sun, W.; Lei, T.; Yuan, H.; Chen, S. First report of anthracnose caused by Colletotrichum kahawae and Colletotrichum horii on tea-oil tree in China. Plant Dis. 2022, 107, 1944. [Google Scholar] [CrossRef]
- Ding, J.; Shen, H.; Yao, L.; Gao, X.; Zhang, M.; Wang, Z.; Li, Y.; Yang, X. First report of anthracnose caused by Colletotrichum kahawae on Hypericum chinensis in China. Plant Dis. 2023, 108, 795. [Google Scholar] [CrossRef]
- Dai, X.; Zhang, M.; Liu, T.; Chen, X.; Zhu, T. Brown leaf spot of Cunninghamia lanceolata caused by Colletotrichum kahawae in Sichuan Province, China. Plant Dis. 2023, 107, 2548. [Google Scholar] [CrossRef]
- Batista, D.; Silva, D.N.; Vieira, A.; Cabral, A.; Pires, A.S.; Loureiro, A.; Guerra-Guimarães, L.; Pereira, A.P.; Azinheira, H.; Talhinhas, P.; et al. Legitimacy and implications of reducing Colletotrichum kahawae to subspecies in plant pathology. Front. Plant Sci. 2016, 7, 2051. [Google Scholar] [CrossRef] [PubMed]
- Cabral, A.; Azinheira, H.G.; Talhinhas, P.; Batista, D.; Ramos, A.P.; Silva, M.D.C.; Oliveira, H.; Várzea, V. Pathological, morphological, cytogenomic, biochemical and molecular data support the distinction between Colletotrichum cigarro comb. et stat. nov. and Colletotrichum kahawae. Plants 2020, 9, 502. [Google Scholar] [CrossRef]
- Kebati, R.; Nyangeri, J.; Omondi, C.; Kubochi, J. Effect of artificial shading on severity of coffee berry disease in Kiambu County, Kenya. Annu. Res. Rev. Biol. 2016, 9, 1–11. [Google Scholar] [CrossRef]
- Chen, Z.; Liang, J.; Zhang, C.; Rodrigues, C.J., Jr. Epicatechin and catechin may prevent coffee berry disease by inhibition of appressorial melanization of Colletotrichum kahawae. Biotechnol. Lett. 2006, 28, 1637–1640. [Google Scholar] [CrossRef]
- Zhonghou, L.; Jingna, L.; Xiang, L.; Ping, L.; Hua, Y. Isolation and identification of pathogens causing dragon stripe disease on Kadsura coccinea. J. King Saud Univ. Sci. 2022, 35, 102516. [Google Scholar] [CrossRef]
- Chen, W.; Fan, X.; Tian, C.; Li, F.; Chou, G. Identification and characterization of Neocosmospora silvicola causing canker disease on Pinus armandii in China. Plant Dis. 2023, 107, 3026–3036. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, H.; Zhang, Q.; Xi, J.; Jiang, K.; Li, J.; Xue, H.; Bi, Y. Isolation, identification, and analyzing the biological characteristics of pathogens causing stem rot of Lanzhou onion during postharvest storage and studying the influence of pathogen infection on the active components of Lanzhou onion. J. Fungi 2024, 10, 789. [Google Scholar] [CrossRef]
- Wang, G.; Zhang, X.; Chen, B.; Peng, Y. Construction of an efficient Agrobacterium tumefaciens-based transformation system in the entomopathogenic fungus Metahizium rileyi. J. Agric. Food Chem. 2024, 72, 21711–21719. [Google Scholar] [CrossRef]
- Silva, D.N.; Talhinhas, P.; Cai, L.; Manuel, L.; Gichuru, E.K.; Loureiro, A.; Várzea, V.; Paulo, O.S.; Batista, D. Host-jump drives rapid and recent ecological speciation of the emergent fungal pathogen Colletotrichum kahawae. Mol. Ecol. 2012, 21, 2655–2670. [Google Scholar] [CrossRef] [PubMed]
- Vieira, A.; Diniz, I.; Loureiro, A.; Pereira, A.; Silva, M.d.C.; Várzea, V.; Batista, D. Aggressiveness profiling of the coffee pathogen Colletotrichum kahawae. Plant Pathol. 2018, 68, 358–368. [Google Scholar] [CrossRef]
- Bridge, P.; Waller, J.; Davies, D.; Buddie, A. Variability of Colletotrichum kahawae in relation to other Colletotrichum species from tropical perennial crops and the development of diagnostic techniques. J. Phytopathol. 2008, 156, 274–280. [Google Scholar] [CrossRef]
- Loureiro, A.; Guerra-Guimaraes, L.; Lidon, F.; Bertrand, B.; Silva, M.d.C.; Várzea, V. Isoenzymatic characterization of Colletotrichum kahawae isolates with different levels of aggressiveness. Trop. Plant Pathol. 2011, 36, 287–293. [Google Scholar] [CrossRef]
- Pires, A.; Azinheira, H.; Cabral, A.; Tavares, S.; Tavares, D.; Castro, M.; Várzea, V.; Silva, M.d.C.; Abranches, R.; Loureiro, J.; et al. Cytogenomic characterisation of Colletotrichum kahawae, the causal agent of Coffee Berry Disease, reveals diversity in minichromosome profiles and genome size expansion. Plant Pathol. 2015, 65, 968–977. [Google Scholar] [CrossRef]
- Damm, U.; O’Connell, R.J.; Groenewald, J.Z.; Crous, P.W. The Colletotrichum destructivum species complex—Hemibiotrophic pathogens of forage and field crops. Stud. Mycol. 2014, 79, 49–84. [Google Scholar] [CrossRef]
- Chen, Z.; Liang, J.; Rodrigues, C.J., Jr. Colletotrichum gloeosporioides can overgrow Colletotrichum kahawae on green coffee berries first inoculated with C. kahawae. Biotechnol. Lett. 2005, 27, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Bardas, G.; Koutita, O.; Tzavella-Klonari, K. Molecular diversity and assessment of biological characteristics of greek Colletotrichum lindemuthianum populations. J. Phytopathol. 2008, 157, 311–318. [Google Scholar] [CrossRef]
- Li, P.; Dai, X.; Wang, S.; Luo, Q.; Tang, Q.; Xu, Z.; Zhao, W.; Wu, F. Biological characteristics and fungicide screening of Colletotrichum fructicola causing mulberry anthracnose. Microorganisms 2024, 12, 2386. [Google Scholar] [CrossRef]
- Tao, R.; Yang, B.; Lin, L.; Munir, S.; Li, Y.; Wang, X.; Huang, M. Biological characterization of emerging fungal pathogen Colletotrichum associated with mango (Mangifera indica L.) post-harvest anthracnose from Vietnam. Mol. Biol. Rep. 2024, 51, 557. [Google Scholar] [CrossRef]
- Salotti, I.; Ji, T.; Rossi, V. Temperature requirements of Colletotrichum spp. belonging to different clades. Front. Plant Sci. 2022, 13, 953760. [Google Scholar] [CrossRef]
- Khodadadi, F.; Santander, R.D.; McHenry, D.J.; Jurick, W.M., 2nd; Aćimović, S.G. A Bitter, Complex problem: Causal Colletotrichum species in virginia orchards and apple fruit susceptibility. Plant Dis. 2023, 107, 3164–3175. [Google Scholar] [CrossRef]
- Boeckman, N.J.; Borba, M.C.; Bernal, V.V.; Khodadadi, F.; Jurick, W.M., 2nd; Aćimović, S.G. Apple bitter rot: Biology, ecology, omics, virulence factors, and management of causal Colletotrichum species. Mol. Plant Pathol. 2025, 26, e70050. [Google Scholar] [CrossRef] [PubMed]
- Hassan, O.; Jeon, J.Y.; Chang, T.; Shin, J.S.; Oh, N.K.; Lee, Y.S. Molecular and morphological characterization of Colletotrichum species in the Colletotrichum gloeosporioides complex associated with persimmon anthracnose in South Korea. Plant Dis. 2018, 102, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Aragaw, G.; Chala, A.; Terefe, H. Cultural and morphological characteristics of Colletotrichum sublineolum isolates infecting sorghum in eastern Ethiopia. Heliyon 2023, 9, e13057. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.W.; Shim, S.H. The fungus Colletotrichum as a source for bioactive secondary metabolites. Arch. Pharmacal Res. 2019, 42, 735–753. [Google Scholar] [CrossRef] [PubMed]
- McCulloch, M.; Gauthier, N.; Vaillancourt, L. First report of bitter rot of apple caused by a Colletotrichum sp. in the C. kahawae clade in Kentucky. Plant Dis. 2019, 104, 289. [Google Scholar] [CrossRef]
- Adugna, G. Coffee berry disease: A century-old anthracnose of green berries of Arabica coffee (Coffea arabica L.) in Africa. J. Plant Dis. Prot. 2023, 131, 315–328. [Google Scholar] [CrossRef]
- Nguyen, P.; Vinnere Pettersson, O.; Olsson, P.; Liljeroth, E. Identification of Colletotrichum species associated with anthracnose disease of coffee in Vietnam. Eur. J. Plant Pathol. 2009, 127, 73–87. [Google Scholar] [CrossRef][Green Version]
- Vieira, A.; Silva, D.N.; Várzea, V.; Paulo, O.S.; Batista, D. Genome-wide signatures of selection in Colletotrichum kahawae reveal candidate genes potentially involved in pathogenicity and aggressiveness. Front. Microbiol. 2019, 10, 1374. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Wang, G.; Sun, D.; Tan, J.; Xie, J.; Zhai, B.; Huang, C.; Lu, W.; Wang, L. First Report of Colletotrichum kahawae Causing Anthracnose on Buckwheat (Fagopyrum tataricum) in China and Biological Characterization of the Pathogen. J. Fungi 2025, 11, 633. https://doi.org/10.3390/jof11090633
Liu X, Wang G, Sun D, Tan J, Xie J, Zhai B, Huang C, Lu W, Wang L. First Report of Colletotrichum kahawae Causing Anthracnose on Buckwheat (Fagopyrum tataricum) in China and Biological Characterization of the Pathogen. Journal of Fungi. 2025; 11(9):633. https://doi.org/10.3390/jof11090633
Chicago/Turabian StyleLiu, Xin, Guang Wang, Daowang Sun, Jing Tan, Jiaxing Xie, Binxin Zhai, Chunyan Huang, Wenjie Lu, and Lihua Wang. 2025. "First Report of Colletotrichum kahawae Causing Anthracnose on Buckwheat (Fagopyrum tataricum) in China and Biological Characterization of the Pathogen" Journal of Fungi 11, no. 9: 633. https://doi.org/10.3390/jof11090633
APA StyleLiu, X., Wang, G., Sun, D., Tan, J., Xie, J., Zhai, B., Huang, C., Lu, W., & Wang, L. (2025). First Report of Colletotrichum kahawae Causing Anthracnose on Buckwheat (Fagopyrum tataricum) in China and Biological Characterization of the Pathogen. Journal of Fungi, 11(9), 633. https://doi.org/10.3390/jof11090633





