Microbiological Characteristics and Pathogenesis of Klebsiella pneumoniae Isolated from Hainan Black Goat
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation and Identification of Bacteria
2.2. Drug Sensitive Test
2.3. Experimental Animals
2.4. Pathogenic Test of K. pneumoniae
2.5. Bacterial Colonization in Mice
2.6. Histopathology Examination of Lungs
2.7. RNA Preparation and Sequencing
2.8. RNA-seq Analysis and DEGs Identification
2.9. qRT-PCR Validation of DEGs
2.10. Statistical Analysis
3. Result
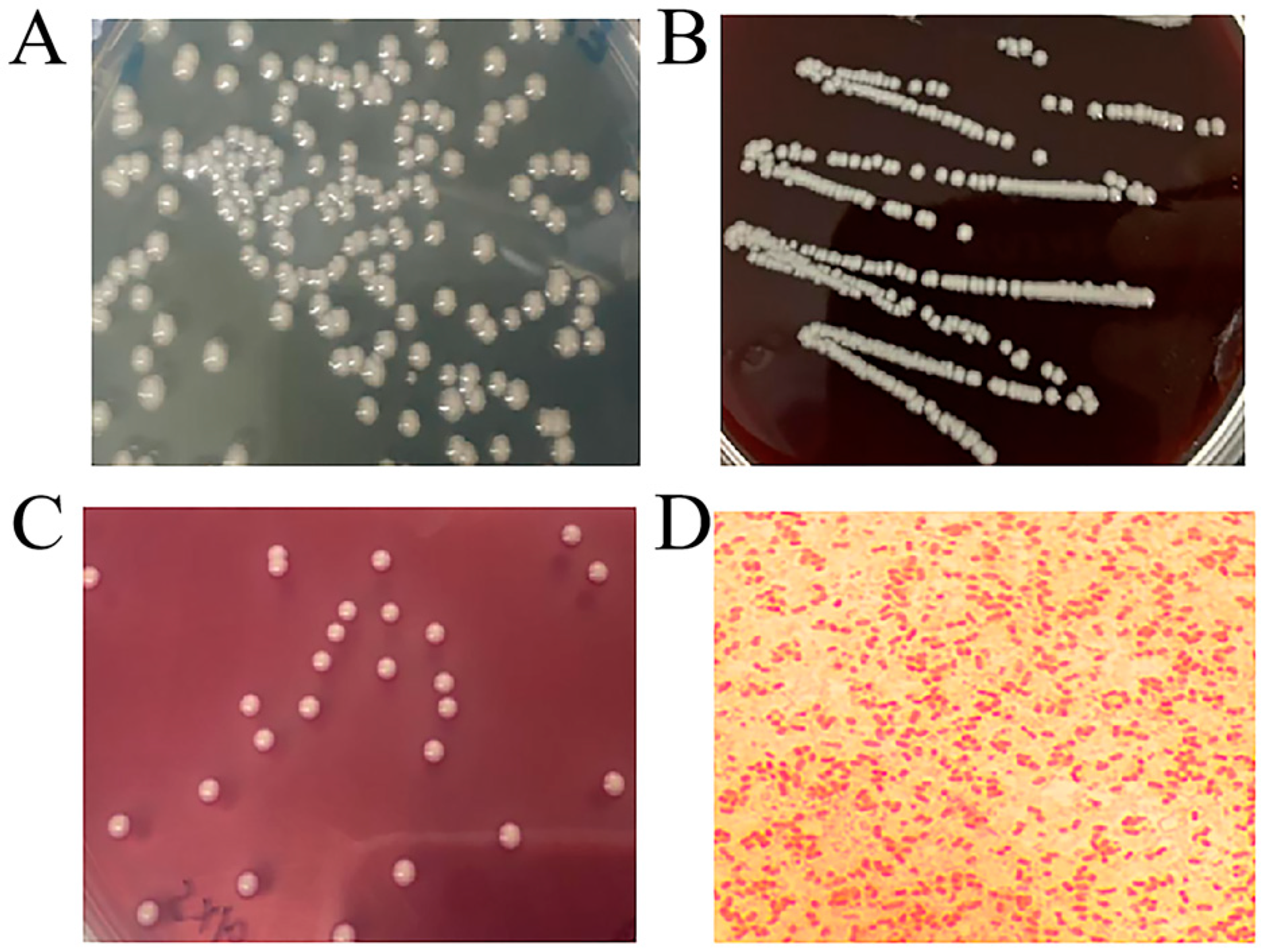
3.1. Colonies and Morphological Characteristics of Isolated Bacteria
3.2. 16S rRNA Identification of Bacteria
3.3. Biochemical Identification of Bacteria
3.4. Drug Sensitive Test of KPHN001
3.5. Mouse Pathogenicity Test
3.6. Bacteria Load in Viscera of Mice
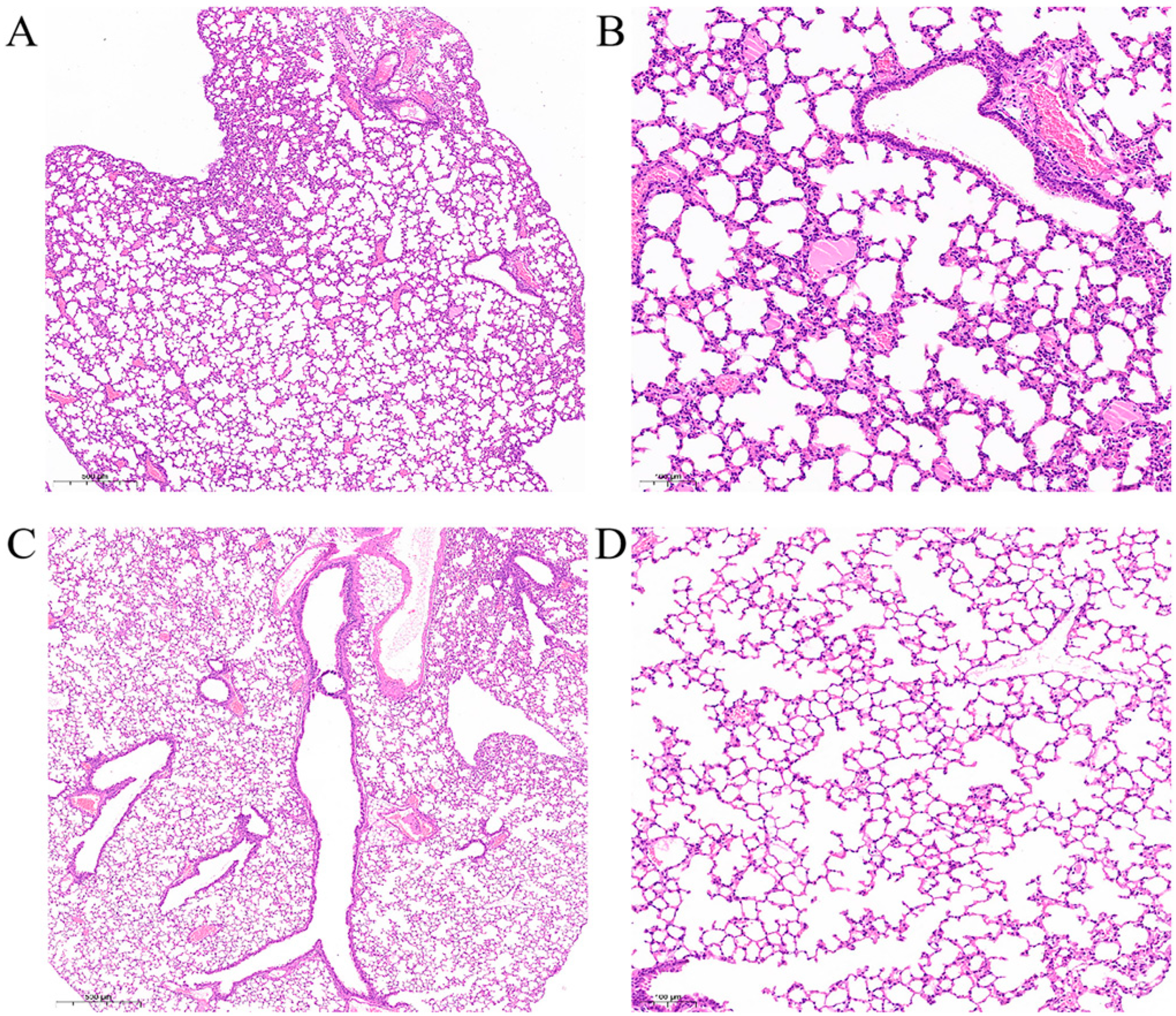
3.7. Histopathology Examination of Mouse Lungs
3.8. RNA-seq Data Analysis
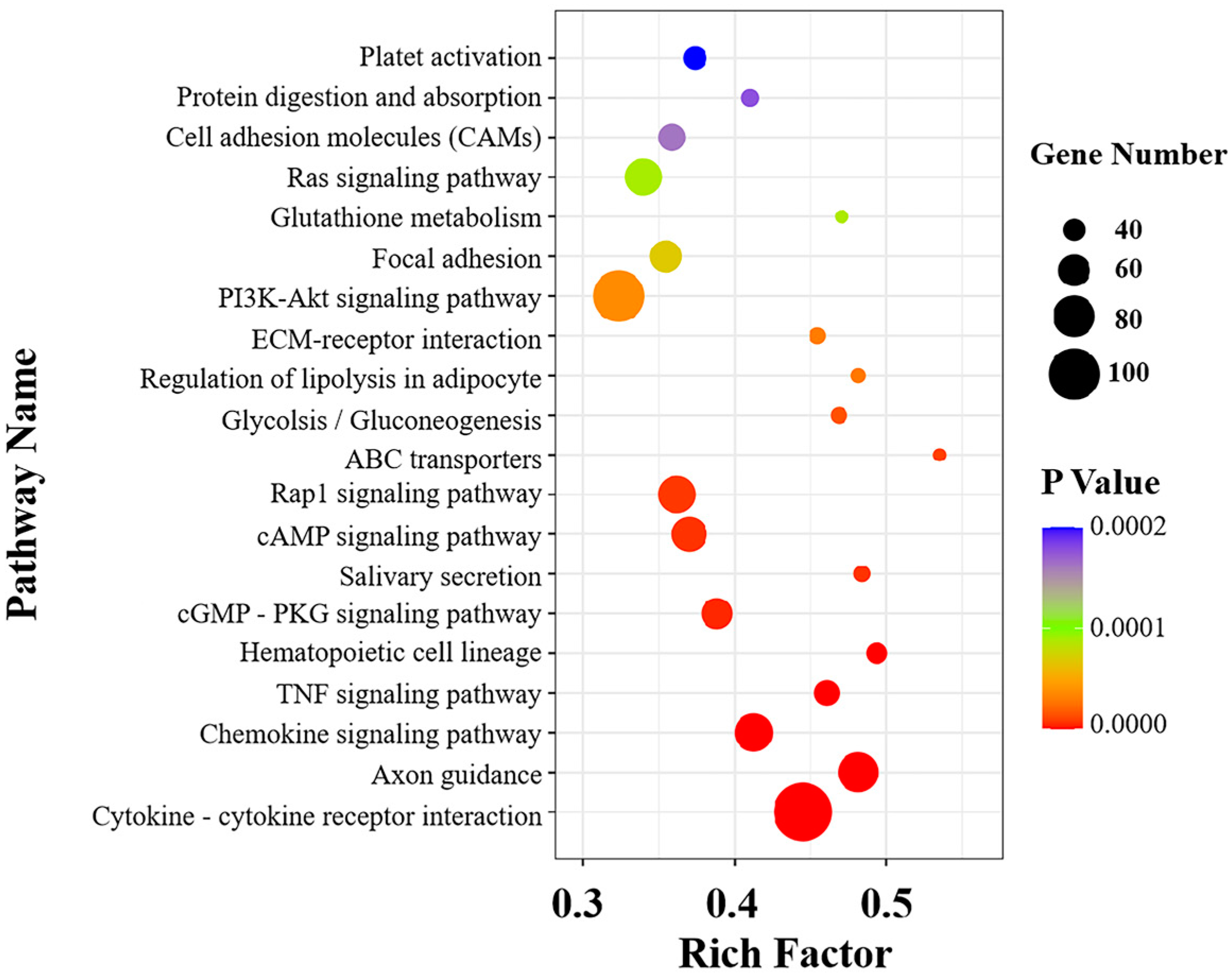
3.9. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Analysis
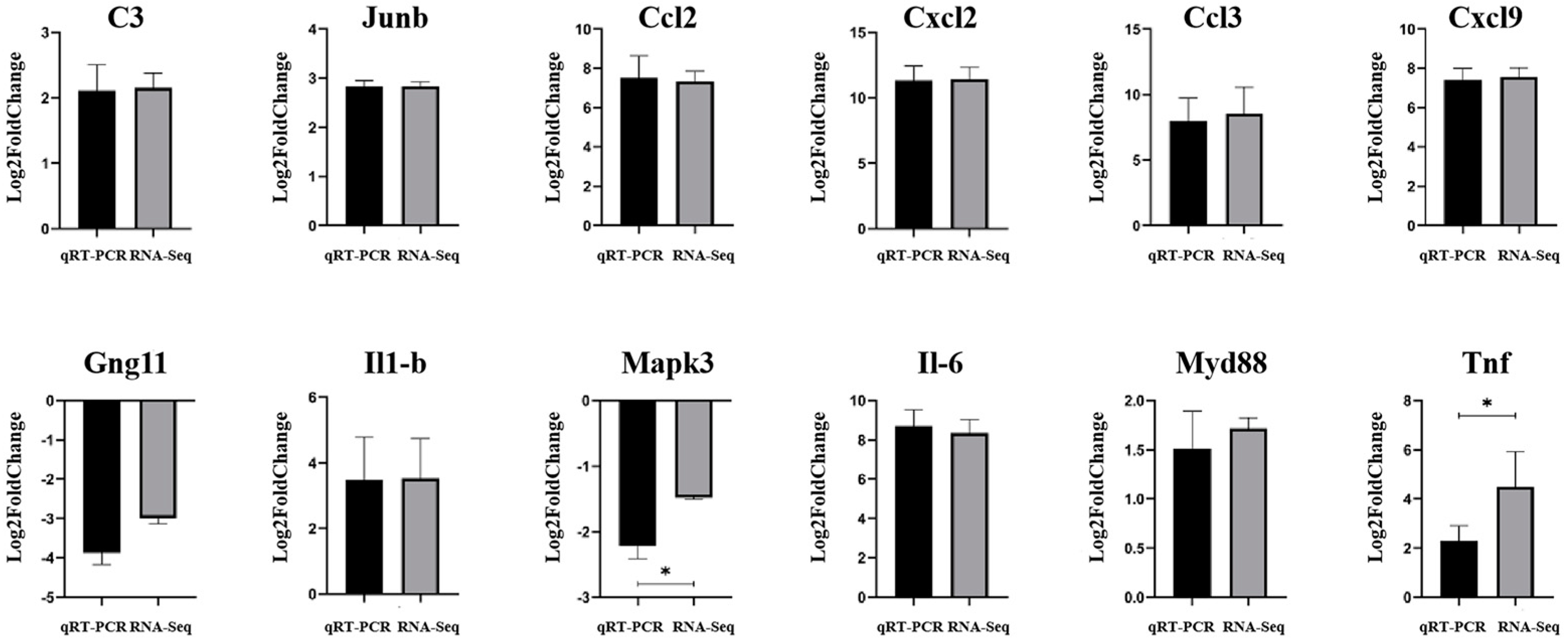
3.10. qRT-PCR Validation of DEGs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Paczosa, M.K.; Mecsas, J. Klebsiella pneumoniae: Going on the offense with a strong defense. Microbiol. Mol. Biol. Rev. 2016, 80, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Kohler, J.E.; Hutchens, M.P. Klebsiella pneumoniae necrotizing fasciitis and septic arthritis: An appearance in the Western hemisphere. Surg. Infect. 2007, 8, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Schelenz, S.; Bramham, K. Septic arthritis due to extended spectrum beta lactamase producing Klebsiella pneumoniae. Jt. Bone Spine 2007, 74, 275–278. [Google Scholar] [CrossRef]
- Chen, R.C.; Shang, H.Q.; Niu, X.Y. Establishment and evaluation of an indirect ELISA for detection of antibodies to goat Klebsiella pneumonia. BMC Vet. Res. 2021, 17, 107. [Google Scholar] [CrossRef] [PubMed]
- Russo, T.A.; Olson, R.; Fang, C.T.; Stoesser, N.; Miller, M.; MacDonald, U.; Hutson, A.; Barker, J.H.; La Hoz, R.M.; Johnson, J.R. Identification of biomarkers for differentiation of hypervirulent Klebsiella pneumoniae from classical K. pneumoniae. J. Clin. Microbiol. 2018, 56, e00776-18. [Google Scholar] [CrossRef]
- Gonzalez-Ferrer, S.; Peñaloza, H.F.; Budnick, J.A. Finding Order in the Chaos: Outstanding Questions in Klebsiella pneumoniae Pathogenesis. Infect. Immun. 2021, 89, e00693-20. [Google Scholar] [CrossRef]
- Sirijatuphat, R.; Sripanidkulchai, K.; Boonyasiri, A.; Rattanaumpawan, P.; Supapueng, O.; Kiratisin, P.; Thamlikitkul, V. Implementation of global antimicrobial resistance surveillance system (GLASS) in patients with bacteremia. PLoS ONE 2018, 13, e0190132. [Google Scholar] [CrossRef]
- Xiong, H.; Keith, J.W.; Samilo, D.W.; Carter, R.A.; Leiner, I.M.; Pamer, E.G. Innate lymphocyte/Ly6C(hi) monocyte crosstalk promotes Klebsiella pneumoniae clearance. Cell 2016, 165, 679–689. [Google Scholar] [CrossRef]
- Sligl, W.I.; Dragan, T.; Smith, S.W. Nosocomial Gram-negative bacteremia in intensive care: Epidemiology, antimicrobial susceptibilities, and outcomes. Int. J. Infect. Dis. 2015, 37, 129–134. [Google Scholar] [CrossRef]
- Holmes, C.L.; Anderson, M.T.; Mobley, H.L.T.; Bachman, M.A. Pathogenesis of Gram-Negative Bacteremia. Clin. Microbiol. Rev. 2021, 34, e00234-20. [Google Scholar] [CrossRef]
- Lawlor, M.S.; Hsu, J.; Rick, P.D.; Miller, V.L. Identification of Klebsiella pneumoniae virulence determinants using an intranasal infection model. Mol. Microbiol. 2005, 58, 1054–1073. [Google Scholar] [CrossRef] [PubMed]
- Fodah, R.A.; Scott, J.B.; Tam, H.H.; Yan, P.; Pfeffer, T.L.; Bundschuh, R.; Warawa, J.M. Correlation of Klebsiella pneumoniae Comparative Genetic Analyses with Virulence Profiles in a Murine Respiratory Disease Model. PLoS ONE 2014, 9, e107394. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.Y.; Zhang, L.L.; Qiu, H.Y.; Jiang, N.; Cheng, P.B.; Lv, M.; Zhou, D.S.; Zhao, Y.E.; Yang, W.H. Establishment and toxicological evaluation of mouse model of hypervirulent Klebsiella pneumoniae infection via aerosolized intratracheal inoculation. Mil. Med. Sci. 2020, 44, 921–925. (In Chinese) [Google Scholar]
- Vornhagen, J.; Sun, Y.; Breen, P.; Forsyth, V.; Zhao, L.; Mobley, H.L.T.; Bachman, M.A. The Klebsiella pneumoniae citrate synthase gene, gltA, influences site specific fitness during infection. PLoS Pathog. 2019, 15, e1008010. [Google Scholar] [CrossRef] [PubMed]
- Carreno, D.; Wanford, J.J.; Jasiunaite, Z.; Hames, R.G.; Chung, W.Y.; Dennison, A.R.; Straatman, K.; Martinez-Pomares, L.; Pareek, M.; Orihuela, C.J.; et al. Splenic macrophages as the source of bacteraemia during pneumococcal pneumonia. EBioMedicine 2021, 72, 103601. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.H.; Yue, C.C.; Zhang, H.; He, L.L.; Dai, Y.Y.; Liu, Y.Y.; Li, J.B. Construction and evaluation of Klebsiella pneumoniae liver abscess mouse model. Acta Univ. Med. Anhui 2021, 56, 1701–1706. (In Chinese) [Google Scholar]
- Anderson, M.T.; Brown, A.N.; Pirani, A.; Smith, S.N.; Photenhauer, A.L.; Sun, Y.; Snitkin, E.S.; Bachman, M.A.; Mobley, H.L.T. Replication Dynamics for Six Gram-Negative Bacterial Species during Bloodstream Infection. mBio 2021, 12, e0111421. [Google Scholar] [CrossRef]
- Yang, Y.B.; Chen, Y.H.; Liu, Y.T.; Song, Y.; Ai, X.H. Klebsiella pneumoniae: A pathogenic bacteria transmitted through Hirudo nipponia that may cause illness in humans. Transbound. Emerg. Dis. 2021, 68, 2051–2058. [Google Scholar] [CrossRef]
- Wu, Y.X.; Yang, Y.B.; Dang, H.F.; Xiao, H.H.; Huang, W.J.; Jia, Z.; Zhao, X.; Chen, K.Y.; Ji, N.; Guo, J.H.; et al. Molecular identification of Klebsiella pneumoniae and expression of immune genes in infected spotted gar Lepisosteus oculatus. Fish Shellfish Immunol. 2021, 119, 220–230. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Coperchini, F.; Chiovato, L.; Croce, L.; Magri, F.; Rotondi, M. The cytokine storm in COVID-19: An overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor Rev. 2020, 53, 25–32. [Google Scholar] [CrossRef]
- Griffith, J.W.; Sokol, C.L.; Luster, A.D. Chemokines and chemokine receptors: Positioning cells for host defense and immunity. Annu. Rev. Immunol. 2014, 32, 659–702. [Google Scholar] [CrossRef]
- Chua, R.L.; Lukassen, S.; Trump, S.; Hennig, B.P.; Wendisch, D.; Pott, F.; Debnath, O.; Thürmann, L.; Kurth, F.; Völker, M.T.; et al. COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 2020, 38, 970–979. [Google Scholar] [CrossRef] [PubMed]
- Darif, D.; Hammi, I.; Kihel, A.; El Idrissi Saik, I.; Guessous, F.; Akarid, K. The pro-inflammatory cytokines in COVID-19 pathogenesis: What goes wrong? Microb. Pathog. 2021, 153, 104799. [Google Scholar] [CrossRef] [PubMed]
- Makwana, R.; Gozzard, N.; Spina, D.; Page, C. TNF-α-induces airway hyperresponsiveness to cholinergic stimulation in Guinea pig airways. Br. J. Pharmacol. 2012, 165, 1978–1991. [Google Scholar] [CrossRef]
- Peiris, J.S.M.; Lai, S.T.; Poon, L.L.M.; Guan, Y.; Yam, L.Y.C.; Lim, W.; Nicholls, J.; Yee, W.K.S.; Yan, W.W.; Cheung, M.T.; et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 2003, 361, 1319–1325. [Google Scholar] [CrossRef]
- Sironi, M.; Breviario, F.; Proserpio, P.; Biondi, A.; Vecchi, A.; Van Damme, J.; Dejana, E.; Mantovani, A. IL-1 stimulates IL-6 production in endothelial cells. J. Immunol. 1989, 142, 549–553. [Google Scholar]
- Dinarello, C.A. Interleukin-1beta. Crit. Care Med. 2005, 33, 460–462. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Narazaki, M.; Kishimoto, T. IL-6 in inflammation, immunity, and disease. Cold Spring Harb. Perspect. Biol. 2014, 6, a016295. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, T.; Yamazaki, S.; Fukui, Y.; Aoki, K.; Yagita, H.; Nishina, T.; Mikami, T.; Katagiri, S.; Shiraishi, A.; Kimura, S.; et al. JunB plays a crucial role in development of regulatory T cells by promoting IL-2 signaling. Mucosal. Immunol. 2019, 12, 1104–1117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Tournier, C.; Davis, R.J.; Flavell, R.A. Regulation of IL-4 expression by the transcription factor JunB during T helper cell differentiation. EMBO J. 1999, 18, 420–432. [Google Scholar] [CrossRef]
- Thomsen, M.K.; Bakiri, L.; Hasenfuss, S.C.; Hamacher, R.; Martinez, L.; Wagner, E.F. JUNB/AP-1 controls IFN-γ during inflammatory liver disease. J. Clin. Investig. 2013, 123, 5258–5268. [Google Scholar] [CrossRef] [Green Version]





| Name of Genes | Primer Sequence (5′-3′) | Product Length (bp) |
|---|---|---|
| Cxcl2 | F: CACTCTCAAGGGCGGTCAA | 96 |
| R: AGTTAGCCTTGCCTTTGTTCAG | ||
| Ccl3 | F: TCCCAGCCAGGTGTCATTTTC | 105 |
| R: GGCATTCAGTTCCAGGTCAGT | ||
| Il1β | F: AATGCCACCTTTTGACAGTGATG | 139 |
| R: AGCTTCTCCACAGCCACAAT | ||
| Il6 | F: TGATGGATGCTACCAAACTGGA | 199 |
| R: CTGTGACTCCAGCTTATCTCTTG | ||
| C3 | F: ACTTCTTCATTGACCTGCGGC | 197 |
| R: CGAGGACTTGGGAGGGATTT | ||
| Tnf | F: CCCTCACACTCACAAACCAC | 134 |
| R: ACAAGGTACAACCCATCGGC | ||
| Ccl2 | F: CACTCACCTGCTGCTACTCA | 117 |
| R: GCTTGGTGACAAAAACTACAGC | ||
| Cxcl9 | F: GTGTGGAGTTCGAGGAACCCT | 173 |
| R: GGCAGGTTTGATCTCCGTTC | ||
| Myd88 | F: AAGCAGCAGAACCAGGAGTC | 150 |
| R: GCAGTAGCAGATAAAGGCATCG | ||
| Junb | F: CAGCCTTTCTATCACGACGAC | 96 |
| R: GGTGGGTTTCAGGAGTTTGTAG | ||
| Gng11 | F: CAAGTTGCAGAGACAACAGGTATC | 130 |
| R: GCTGCCCTTTTCTTTGAAGGG | ||
| Mapk3 | F: CAACACCACCTGCGACCTTA | 154 |
| R: GGATTTGGTGTAGCCCTTGGAA | ||
| β-actin | F: CCTCTATGCCAACACAGT | 146 |
| R: TAGGAGCCAGAGCAGTAA |
| Biochemical Project | Result | Biochemical Project | Result |
|---|---|---|---|
| Glucose | + | Sorbitol | + |
| Sucrose | + | Urea | + |
| Maltose | + | Malonate | + |
| Mannose | + | Citrate | + |
| Arabinose | + | Ornithine | − |
| Raffinose | + | Lysine | + |
| Rhamnose | + | Indole | − |
| Xylose | + | Methyl red test (MR) | − |
| Mannitol | + | Voges–Proskauer test | + |
| Salicin | + | Hydrogen sulfide | − |
| Inositol | + | Nitrate reduction | + |
| Antimicrobial Agents | Inhibitory Zone Diameter/mm | Drug Resistance | Antimicrobial Agents | Inhibitory Zone Diameter/mm | Drug Resistance |
|---|---|---|---|---|---|
| Penicillin | 0 | R | Ciprofloxacin | 14 | R |
| Oxacillin | 0 | R | Vancomycin | 0 | R |
| Ampicillinum | 0 | R | Polymyxin B | 17 | S |
| Carbenicillin | 0 | R | Trimethoprim | 0 | R |
| Piperacillin | 10 | R | Furazolidone | 19 | S |
| Cefalexin | 0 | R | Chloramphenicol | 0 | R |
| Cefazolin | 0 | R | Amikacin | 0 | R |
| Cefradine | 0 | R | Gentamicin | 0 | R |
| Cefuroxime | 0 | R | Kanamycin | 0 | R |
| Ceftazidime | 15 | R | Neomycin | 0 | R |
| Ceftriaxone | 9 | R | Tetracycline | 7 | R |
| Cefoperazone | 12 | R | Doxycycline | 0 | R |
| Midecamycin | 0 | R | Minocycline | 10 | R |
| Norfloxacin | 10 | R | Erythromycin | 0 | R |
| Ofloxacin | 12 | R | Clindamycin | 0 | R |
| Serial Number | Heart Bacteria Load (CFU/g) | Liver Bacteria Load (CFU/g) | Spleen Bacteria Load (CFU/g) | Lung Bacteria Load (CFU/g) | Kidney Bacteria Load (CFU/g) |
|---|---|---|---|---|---|
| E1 | 7.34 × 107 | 3.07 × 108 | 6.16 × 108 | 1.79 × 108 | 1.59 × 108 |
| E2 | 2.25 × 107 | 1.27 × 108 | 9.88 × 108 | 3.27 × 107 | 7.60 × 107 |
| E3 | 4.58 × 107 | 2.41 × 109 | 5.04 × 109 | 1.15 × 109 | 5.25 × 108 |
| Serial Number | Total Reads | Q10 Bases Ratio(%) | Q20 Bases Ratio(%) | Q30 Bases Ratio(%) | GC Bases Ratio(%) | N Bases Ratio(%) |
|---|---|---|---|---|---|---|
| C1 | 71,187,134 | 100.00% | 98.98% | 96.12% | 51.62% | 0.00% |
| C2 | 55,851,916 | 100.00% | 99.01% | 96.21% | 51.69% | 0.00% |
| C3 | 55,792,564 | 100.00% | 98.99% | 96.17% | 51.72% | 0.00% |
| E1 | 63,352,576 | 100.00% | 99.03% | 96.27% | 51.38% | 0.00% |
| E2 | 61,343,784 | 100.00% | 98.98% | 96.1% | 51.69% | 0.00% |
| E3 | 58,422,366 | 100.00% | 98.98% | 96.1% | 51.11% | 0.00% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, M.; Li, H.; Zhang, Z.; Jiang, J.; Li, H.; Yang, W.; Cheng, Y.; Gao, H.; Chen, Q.; Du, L.; et al. Microbiological Characteristics and Pathogenesis of Klebsiella pneumoniae Isolated from Hainan Black Goat. Vet. Sci. 2022, 9, 471. https://doi.org/10.3390/vetsci9090471
He M, Li H, Zhang Z, Jiang J, Li H, Yang W, Cheng Y, Gao H, Chen Q, Du L, et al. Microbiological Characteristics and Pathogenesis of Klebsiella pneumoniae Isolated from Hainan Black Goat. Veterinary Sciences. 2022; 9(9):471. https://doi.org/10.3390/vetsci9090471
Chicago/Turabian StyleHe, Meirong, Haoyang Li, Zhenxing Zhang, Junming Jiang, Hong Li, Weijie Yang, Yiwen Cheng, Hongyan Gao, Qiaoling Chen, Li Du, and et al. 2022. "Microbiological Characteristics and Pathogenesis of Klebsiella pneumoniae Isolated from Hainan Black Goat" Veterinary Sciences 9, no. 9: 471. https://doi.org/10.3390/vetsci9090471
APA StyleHe, M., Li, H., Zhang, Z., Jiang, J., Li, H., Yang, W., Cheng, Y., Gao, H., Chen, Q., Du, L., Chen, S., Man, C., & Wang, F. (2022). Microbiological Characteristics and Pathogenesis of Klebsiella pneumoniae Isolated from Hainan Black Goat. Veterinary Sciences, 9(9), 471. https://doi.org/10.3390/vetsci9090471






