Molecular Mechanisms of Lipopolysaccharide (LPS) Induced Inflammation in an Immortalized Ovine Luteal Endothelial Cell Line (OLENDO)
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Cultures with Ovine Luteal Endothelial (OLENDO) Cells
2.2. RNA Isolation, Reverse Transcription and Qualitative RT-PCR
2.3. Semi-Quantitative RT-PCR and Data Evaluation
2.4. Protein Preparation and Western Blot Analysis
2.5. Statistical Analysis
3. Results
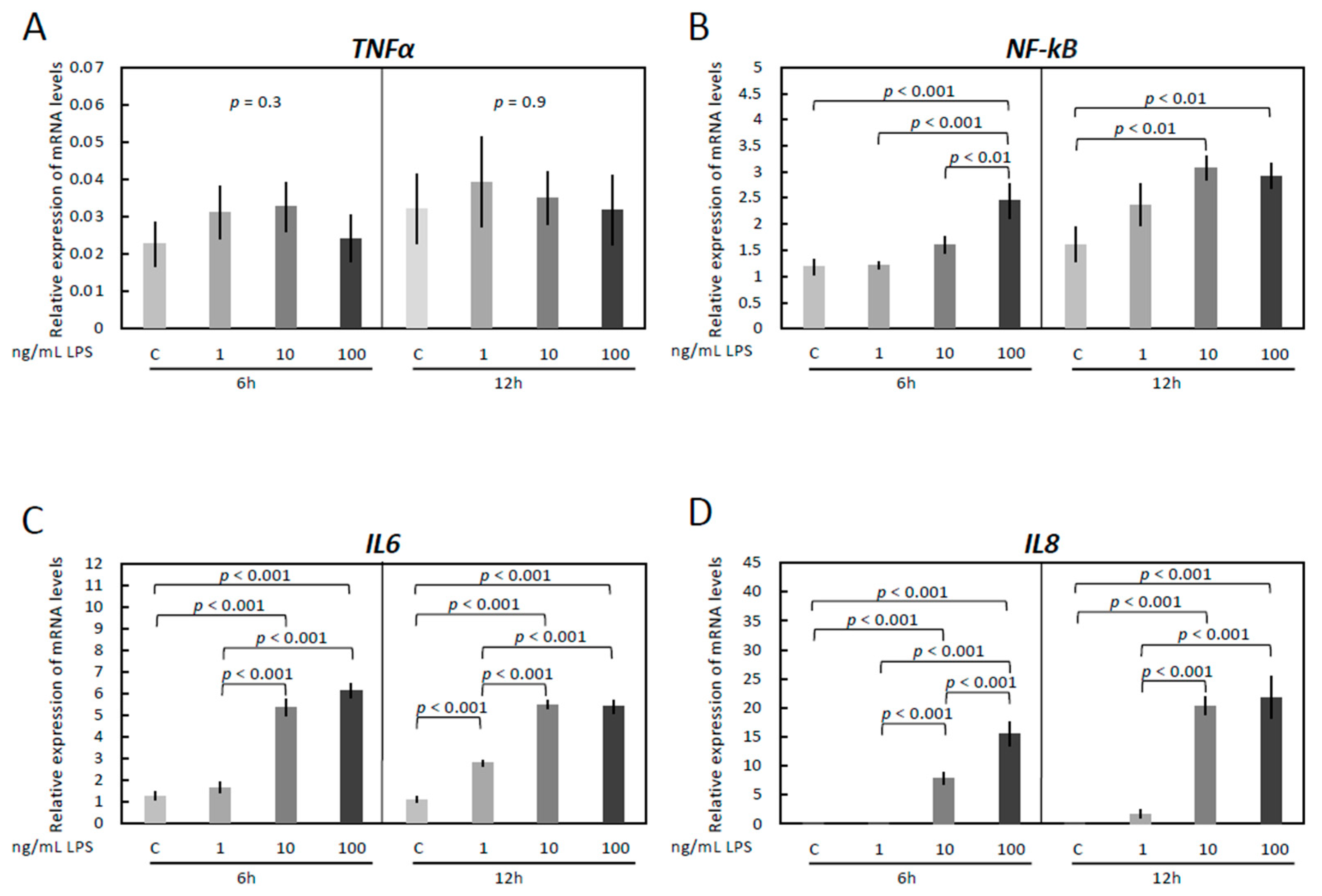
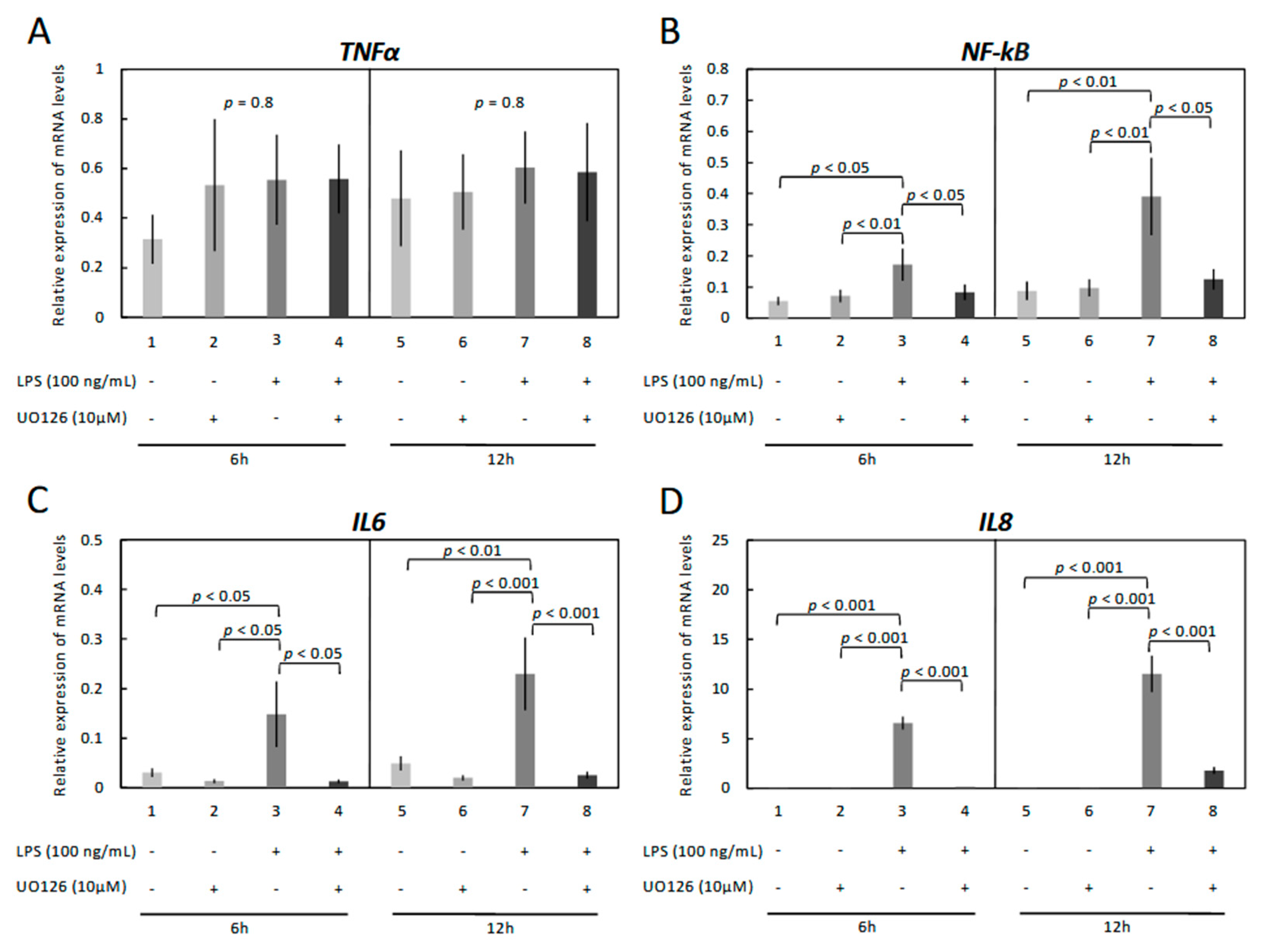
3.1. LPS Treatment Induces Inflammatory Reaction in OLENDO Cells
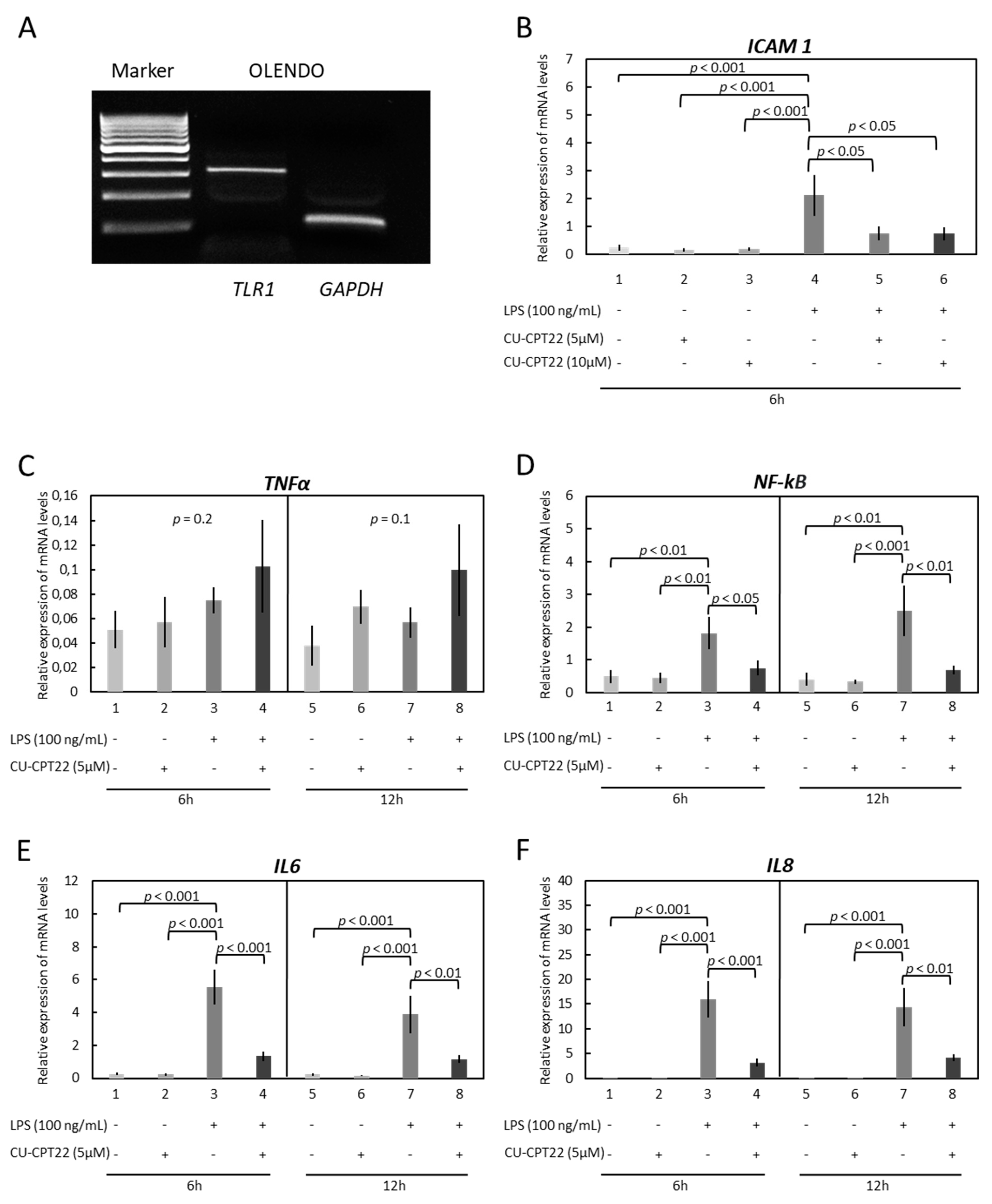
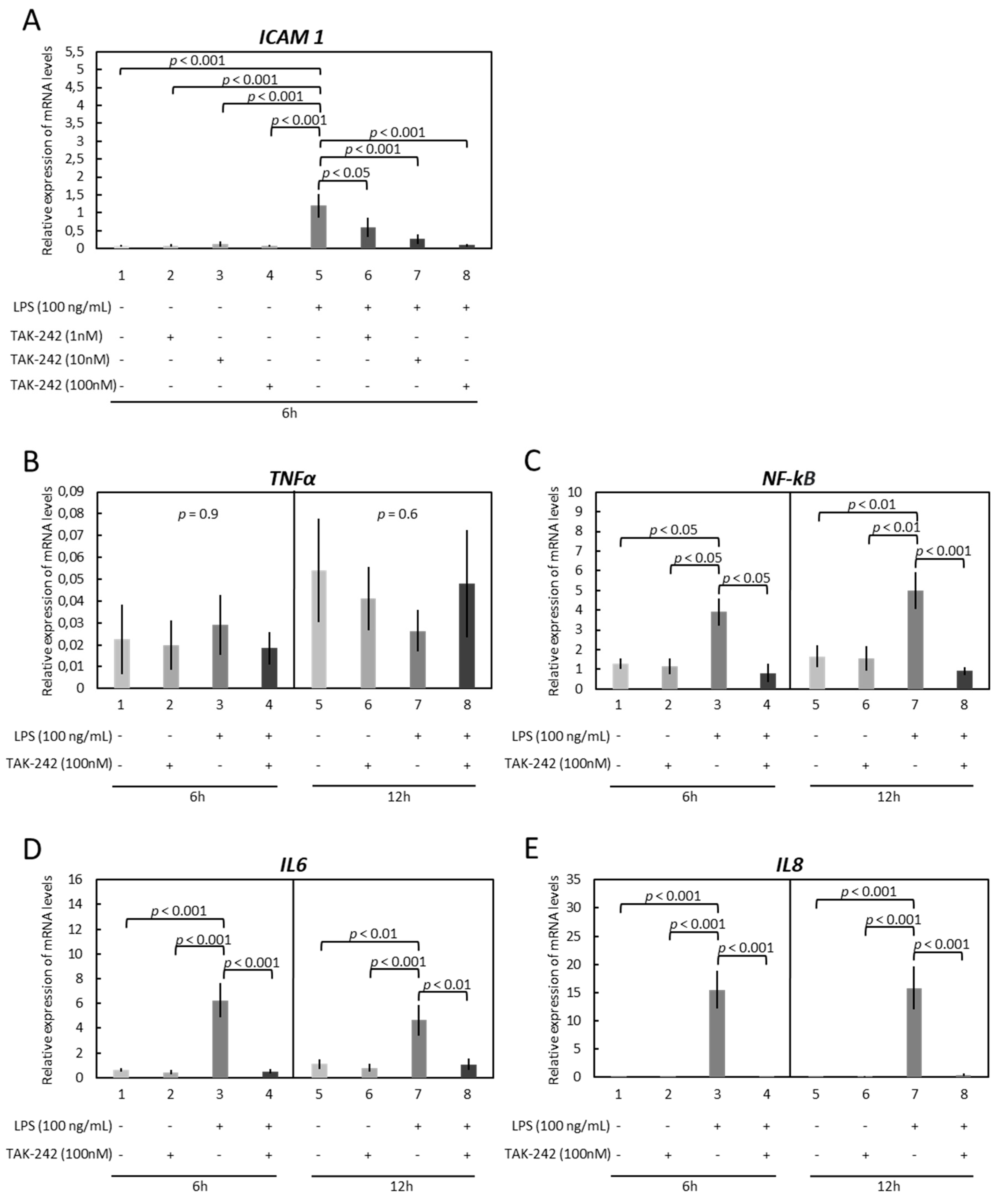
3.2. OLENDO Cells Respond to E. coli LPS through TLR1/TL2 and TLR4 Pathways
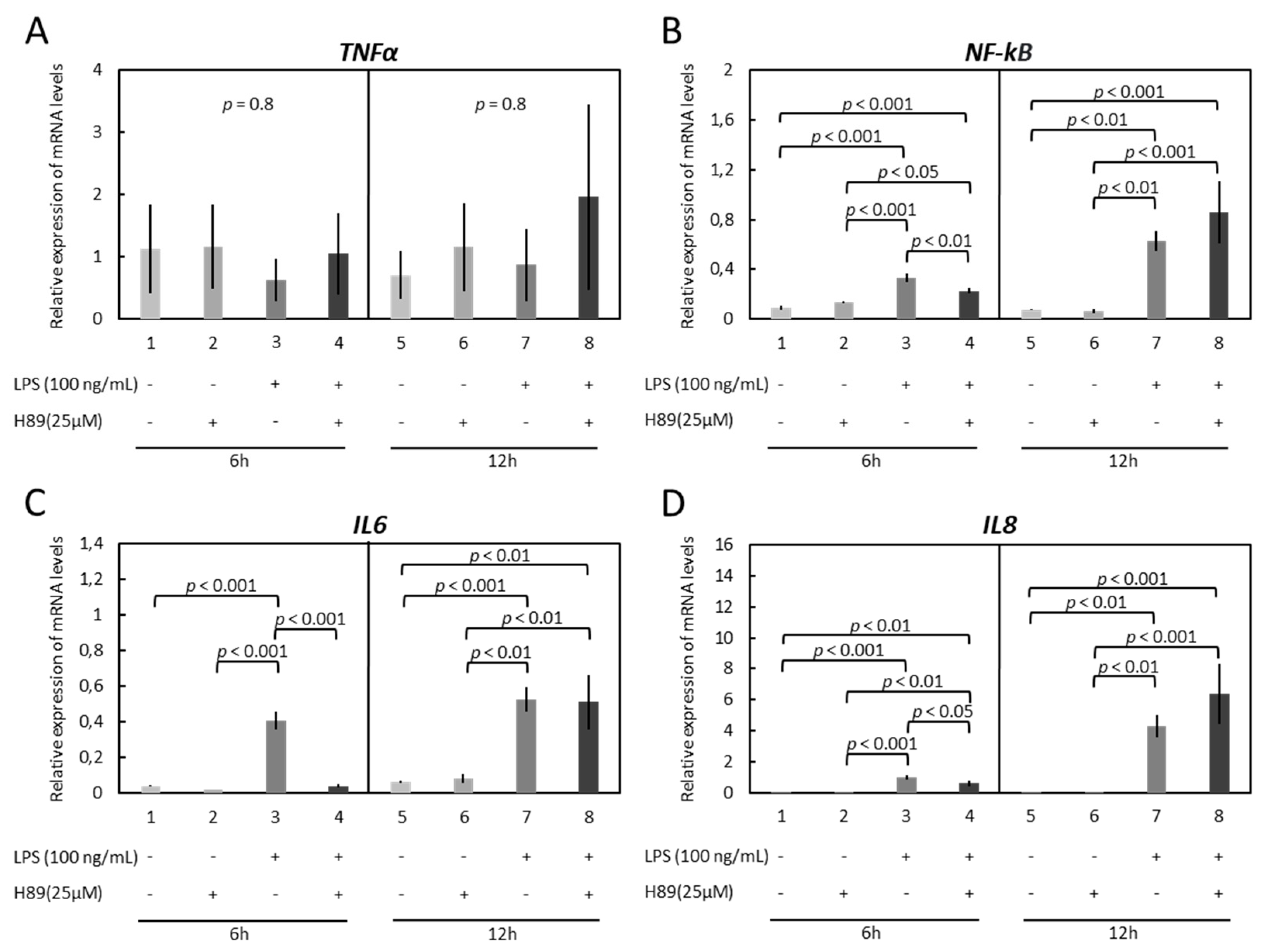
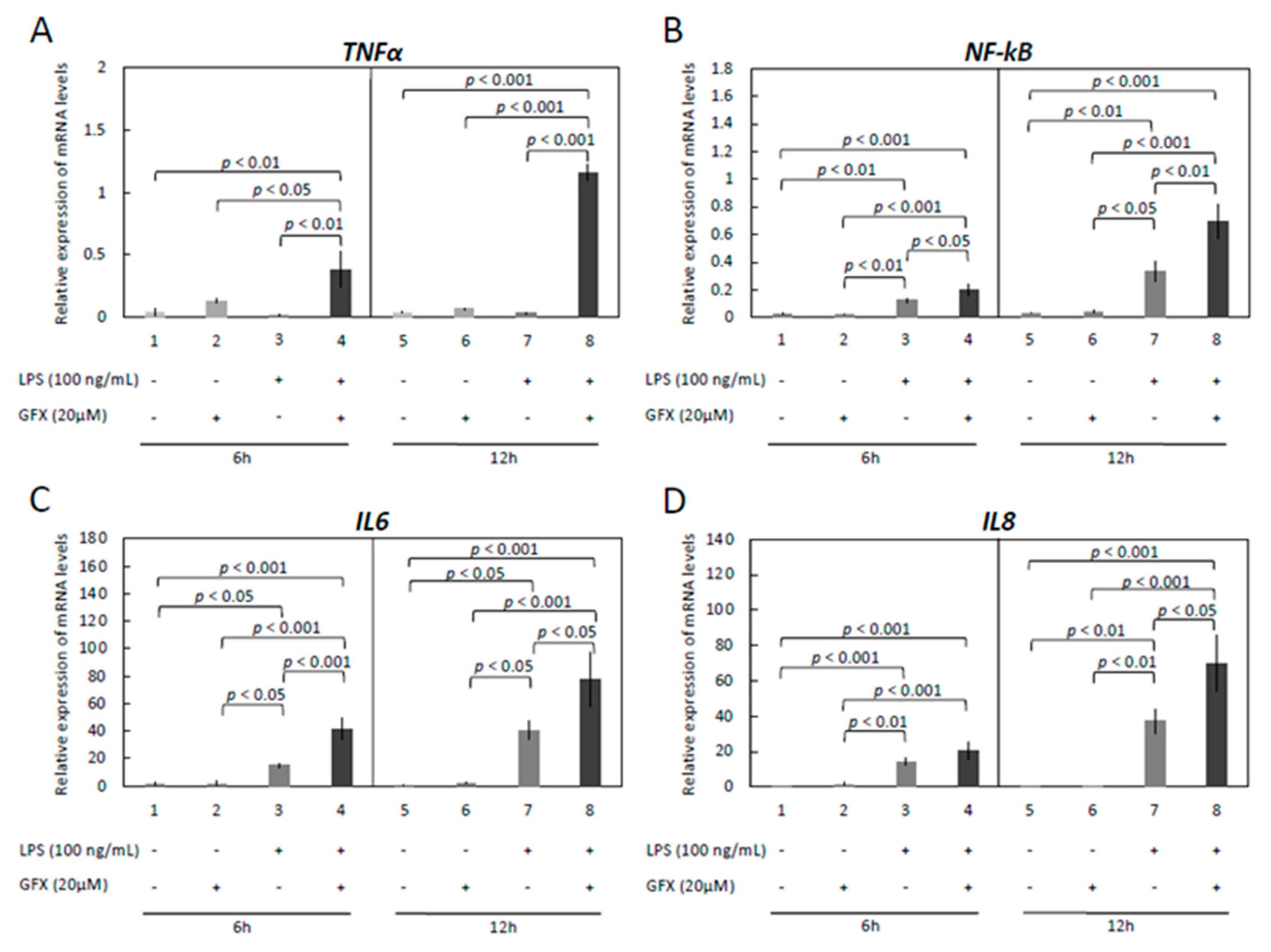
3.3. Modulation of Functionality of Kinases (PKA, PKC, MAPKs) Alters the Expression of Inflammatory Markers in OLENDO Cells
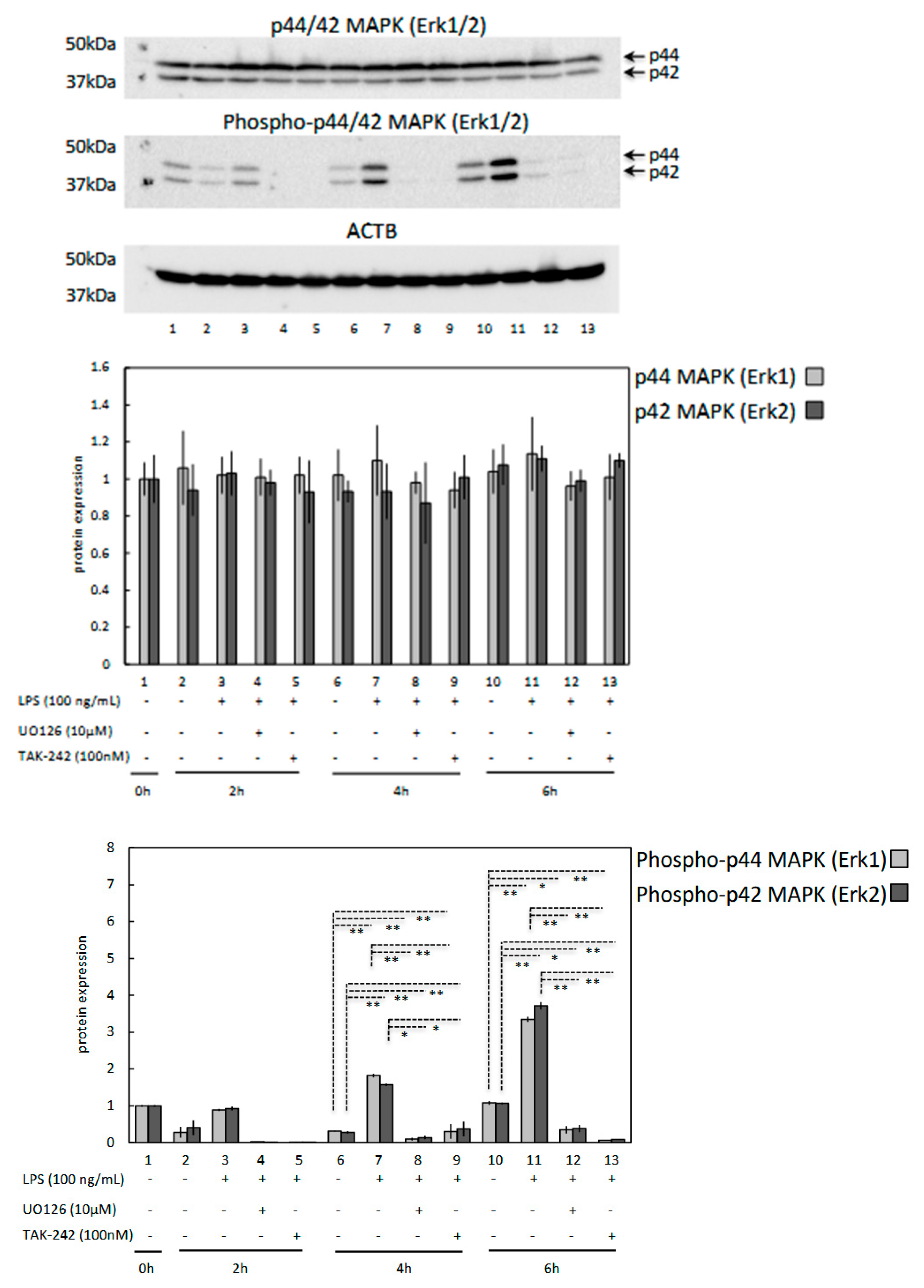
3.4. LPS Induces Phosphorylation of p44/42 MAPK (Erk1/2) in OLENDO Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sheldon, I.M.; Rycroft, A.N.; Dogan, B.; Craven, M.; Bromfield, J.J.; Chandler, A.; Roberts, M.H.; Price, S.B.; Gilbert, R.O.; Simpson, K.W. Specific strains of Escherichia coli are pathogenic for the endometrium of cattle and cause pelvic inflammatory disease in cattle and mice. PLoS ONE 2010, 5, e9192. [Google Scholar] [CrossRef] [PubMed]
- Hertl, J.A.; Grohn, Y.T.; Leach, J.D.; Bar, D.; Bennett, G.J.; Gonzalez, R.N.; Rauch, B.J.; Welcome, F.L.; Tauer, L.W.; Schukken, Y.H. Effects of clinical mastitis caused by gram-positive and gram-negative bacteria and other organisms on the probability of conception in New York State Holstein dairy cows. J. Dairy Sci. 2010, 93, 1551–1560. [Google Scholar] [CrossRef] [PubMed]
- Lavon, Y.; Leitner, G.; Klipper, E.; Moallem, U.; Meidan, R.; Wolfenson, D. Subclinical, chronic intramammary infection lowers steroid concentrations and gene expression in bovine preovulatory follicles. Domest. Anim. Endocrinol. 2011, 40, 98–109. [Google Scholar] [CrossRef] [PubMed]
- Herath, S.; Williams, E.J.; Lilly, S.T.; Gilbert, R.O.; Dobson, H.; Bryant, C.E.; Sheldon, I.M. Ovarian follicular cells have innate immune capabilities that modulate their endocrine function. Reproduction 2007, 134, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, I.M.; Noakes, D.E.; Rycroft, A.N.; Pfeiffer, D.U.; Dobson, H. Influence of uterine bacterial contamination after parturition on ovarian dominant follicle selection and follicle growth and function in cattle. Reproduction 2002, 123, 837–845. [Google Scholar] [CrossRef] [PubMed]
- Williams, E.J.; Sibley, K.; Miller, A.N.; Lane, E.A.; Fishwick, J.; Nash, D.M.; Herath, S.; England, G.C.; Dobson, H.; Sheldon, I.M. The effect of Escherichia coli lipopolysaccharide and tumour necrosis factor alpha on ovarian function. Am. J. Reprod. Immunol. 2008, 60, 462–473. [Google Scholar] [CrossRef]
- Haziak, K.; Herman, A.P.; Wojtulewicz, K.; Pawlina, B.; Paczesna, K.; Bochenek, J.; Tomaszewska-Zaremba, D. Effect of CD14/TLR4 antagonist on GnRH/LH secretion in ewe during central inflammation induced by intracerebroventricular administration of LPS. J. Anim. Sci. Biotechnol. 2018, 9, 52. [Google Scholar] [CrossRef]
- Battaglia, D.F.; Bowen, J.M.; Krasa, H.B.; Thrun, L.A.; Viguie, C.; Karsch, F.J. Endotoxin inhibits the reproductive neuroendocrine axis while stimulating adrenal steroids: A simultaneous view from hypophyseal portal and peripheral blood. Endocrinology 1997, 138, 4273–4281. [Google Scholar] [CrossRef]
- Williams, C.Y.; Harris, T.G.; Battaglia, D.F.; Viguie, C.; Karsch, F.J. Endotoxin inhibits pituitary responsiveness to gonadotropin-releasing hormone. Endocrinology 2001, 142, 1915–1922. [Google Scholar] [CrossRef]
- Bromfield, J.J.; Sheldon, I.M. Lipopolysaccharide initiates inflammation in bovine granulosa cells via the TLR4 pathway and perturbs oocyte meiotic progression in vitro. Endocrinology 2011, 152, 5029–5040. [Google Scholar] [CrossRef]
- Magata, F.; Horiuchi, M.; Miyamoto, A.; Shimizu, T. Lipopolysaccharide (LPS) inhibits steroid production in theca cells of bovine follicles in vitro: Distinct effect of LPS on theca cell function in pre- and post-selection follicles. J. Reprod. Dev. 2014, 60, 280–287. [Google Scholar] [CrossRef] [PubMed]
- Magata, F.; Ishida, Y.; Miyamoto, A.; Furuoka, H.; Inokuma, H.; Shimizu, T. Comparison of bacterial endotoxin lipopolysaccharide concentrations in the blood, ovarian follicular fluid and uterine fluid: A clinical case of bovine metritis. J. Vet. Med. Sci. 2015, 77, 81–84. [Google Scholar] [CrossRef]
- Shimizu, T.; Miyauchi, K.; Shirasuna, K.; Bollwein, H.; Magata, F.; Murayama, C.; Miyamoto, A. Effects of lipopolysaccharide (LPS) and peptidoglycan (PGN) on estradiol production in bovine granulosa cells from small and large follicles. Toxicol. In Vitro 2012, 26, 1134–1142. [Google Scholar] [CrossRef] [PubMed]
- Taylor, C.C.; Terranova, P.F. Lipopolysaccharide inhibits rat ovarian thecal-interstitial cell steroid secretion in vitro. Endocrinology 1995, 136, 5527–5532. [Google Scholar] [CrossRef] [PubMed]
- Luttgenau, J.; Herzog, K.; Struve, K.; Latter, S.; Boos, A.; Bruckmaier, R.M.; Bollwein, H.; Kowalewski, M.P. LPS-mediated effects and spatio-temporal expression of TLR2 and TLR4 in the bovine corpus luteum. Reproduction 2016, 151, 391–399. [Google Scholar] [CrossRef] [PubMed]
- Herzog, K.; Struve, K.; Kastelic, J.P.; Piechotta, M.; Ulbrich, S.E.; Pfarrer, C.; Shirasuna, K.; Shimizu, T.; Miyamoto, A.; Bollwein, H. Escherichia coli lipopolysaccharide administration transiently suppresses luteal structure and function in diestrous cows. Reproduction 2012, 144, 467–476. [Google Scholar] [CrossRef]
- Luttgenau, J.; Moller, B.; Kradolfer, D.; Wellnitz, O.; Bruckmaier, R.M.; Miyamoto, A.; Ulbrich, S.E.; Bollwein, H. Lipopolysaccharide enhances apoptosis of corpus luteum in isolated perfused bovine ovaries in vitro. Reproduction 2016, 151, 17–28. [Google Scholar] [CrossRef]
- Rietschel, E.T.; Kirikae, T.; Schade, F.U.; Mamat, U.; Schmidt, G.; Loppnow, H.; Ulmer, A.J.; Zahringer, U.; Seydel, U.; Di Padova, F.; et al. Bacterial endotoxin: Molecular relationships of structure to activity and function. FASEB J. 1994, 8, 217–225. [Google Scholar] [CrossRef]
- Armant, M.A.; Fenton, M.J. Toll-like receptors: A family of pattern-recognition receptors in mammals. Genome Biol. 2002, 3, REVIEWS3011. [Google Scholar] [CrossRef]
- Beutler, B. Inferences, questions and possibilities in Toll-like receptor signalling. Nature 2004, 430, 257–263. [Google Scholar] [CrossRef]
- Takeuchi, O.; Sato, S.; Horiuchi, T.; Hoshino, K.; Takeda, K.; Dong, Z.; Modlin, R.L.; Akira, S. Cutting edge: Role of Toll-like receptor 1 in mediating immune response to microbial lipoproteins. J. Immunol. 2002, 169, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Gioannini, T.L.; Weiss, J.P. Regulation of interactions of Gram-negative bacterial endotoxins with mammalian cells. Immunol. Res. 2007, 39, 249–260. [Google Scholar] [CrossRef] [PubMed]
- Miyake, K. Innate immune sensing of pathogens and danger signals by cell surface Toll-like receptors. Semin. Immunol. 2007, 19, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, L.A.; Kramer, J.M.; Williams, R.S.; Bromfield, J.J. Human granulosa-luteal cells initiate an innate immune response to pathogen-associated molecules. Reproduction 2016, 152, 261–270. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Price, J.C.; Bromfield, J.J.; Sheldon, I.M. Pathogen-associated molecular patterns initiate inflammation and perturb the endocrine function of bovine granulosa cells from ovarian dominant follicles via TLR2 and TLR4 pathways. Endocrinology 2013, 154, 3377–3386. [Google Scholar] [CrossRef] [PubMed]
- Oda, K.; Kitano, H. A comprehensive map of the toll-like receptor signaling network. Mol. Syst. Biol. 2006, 2, 0015. [Google Scholar] [CrossRef]
- Hortelano, S.; Lopez-Fontal, R.; Traves, P.G.; Villa, N.; Grashoff, C.; Bosca, L.; Luque, A. ILK mediates LPS-induced vascular adhesion receptor expression and subsequent leucocyte trans-endothelial migration. Cardiovasc. Res. 2010, 86, 283–292. [Google Scholar] [CrossRef]
- Takeuchi, S.; Kawashima, S.; Rikitake, Y.; Ueyama, T.; Inoue, N.; Hirata, K.; Yokoyama, M. Cerivastatin suppresses lipopolysaccharide-induced ICAM-1 expression through inhibition of Rho GTPase in BAEC. Biochem. Biophys. Res. Commun. 2000, 269, 97–102. [Google Scholar] [CrossRef]
- Sawa, Y.; Ueki, T.; Hata, M.; Iwasawa, K.; Tsuruga, E.; Kojima, H.; Ishikawa, H.; Yoshida, S. LPS-induced IL-6, IL-8, VCAM-1, and ICAM-1 expression in human lymphatic endothelium. J. Histochem. Cytochem. 2008, 56, 97–109. [Google Scholar] [CrossRef]
- Yan, W.; Zhao, K.; Jiang, Y.; Huang, Q.; Wang, J.; Kan, W.; Wang, S. Role of p38 MAPK in ICAM-1 expression of vascular endothelial cells induced by lipopolysaccharide. Shock 2002, 17, 433–438. [Google Scholar] [CrossRef]
- Gram, A.; Grazul-Bilska, A.T.; Boos, A.; Rahman, N.A.; Kowalewski, M.P. Lipopolysaccharide disrupts gap junctional intercellular communication in an immortalized ovine luteal endothelial cell line. Toxicol. In Vitro 2019, 60, 437–449. [Google Scholar] [CrossRef]
- Kowalewski, M.P.; Dyson, M.T.; Boos, A.; Stocco, D.M. Vasoactive intestinal peptide (VIP)-mediated expression and function of steroidogenic acute regulatory protein (StAR) in granulosa cells. Mol. Cell. Endocrinol. 2010, 328, 93–103. [Google Scholar] [CrossRef]
- Manna, P.R.; Chandrala, S.P.; King, S.R.; Jo, Y.; Counis, R.; Huhtaniemi, I.T.; Stocco, D.M. Molecular mechanisms of insulin-like growth factor-I mediated regulation of the steroidogenic acute regulatory protein in mouse leydig cells. Mol. Endocrinol. 2006, 20, 362–378. [Google Scholar] [CrossRef] [PubMed]
- Gram, A.; Buchler, U.; Boos, A.; Hoffmann, B.; Kowalewski, M.P. Biosynthesis and degradation of canine placental prostaglandins: Prepartum changes in expression and function of prostaglandin F2alpha-synthase (PGFS, AKR1C3) and 15-hydroxyprostaglandin dehydrogenase (HPGD). Biol. Reprod. 2013, 89, 2. [Google Scholar] [CrossRef] [PubMed]
- Kowalewski, M.P.; Beceriklisoy, H.B.; Aslan, S.; Agaoglu, A.R.; Hoffmann, B. Time related changes in luteal prostaglandin synthesis and steroidogenic capacity during pregnancy, normal and antiprogestin induced luteolysis in the bitch. Anim. Reprod. Sci. 2009, 116, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Gram, A.; Boos, A.; Kowalewski, M.P. Uterine and placental expression of canine oxytocin receptor during pregnancy and normal and induced parturition. Reprod. Domest. Anim. 2014, 49 (Suppl. 2), 41–49. [Google Scholar] [CrossRef]
- Mako, V.; Czucz, J.; Weiszhar, Z.; Herczenik, E.; Matko, J.; Prohaszka, Z.; Cervenak, L. Proinflammatory activation pattern of human umbilical vein endothelial cells induced by IL-1beta, TNF-alpha, and LPS. Cytom. Part A 2010, 77, 962–970. [Google Scholar] [CrossRef]
- Wong, E.; Xu, F.; Joffre, J.; Nguyen, N.; Wilhelmsen, K.; Hellman, J. ERK1/2 has Divergent Roles in LPS-Induced Microvascular Endothelial Cell Cytokine Production and Permeability. Shock 2021, 55, 349. [Google Scholar] [CrossRef]
- Zeuke, S.; Ulmer, A.J.; Kusumoto, S.; Katus, H.A.; Heine, H. TLR4-mediated inflammatory activation of human coronary artery endothelial cells by LPS. Cardiovasc. Res. 2002, 56, 126–134. [Google Scholar] [CrossRef]
- Kim, T.H.; Ku, S.K.; Lee, I.C.; Bae, J.S. Anti-inflammatory functions of purpurogallin in LPS-activated human endothelial cells. BMB Rep. 2012, 45, 200–205. [Google Scholar] [CrossRef]
- Hoareau, L.; Bencharif, K.; Rondeau, P.; Murumalla, R.; Ravanan, P.; Tallet, F.; Delarue, P.; Cesari, M.; Roche, R.; Festy, F. Signaling pathways involved in LPS induced TNFalpha production in human adipocytes. J. Inflamm. 2010, 7, 1. [Google Scholar] [CrossRef] [PubMed]
- Dentener, M.A.; Louis, R.; Cloots, R.H.; Henket, M.; Wouters, E.F. Differences in local versus systemic TNFalpha production in COPD: Inhibitory effect of hyaluronan on LPS induced blood cell TNFalpha release. Thorax 2006, 61, 478–484. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dragoni, S.; Hudson, N.; Kenny, B.A.; Burgoyne, T.; McKenzie, J.A.; Gill, Y.; Blaber, R.; Futter, C.E.; Adamson, P.; Greenwood, J.; et al. Endothelial MAPKs Direct ICAM-1 Signaling to Divergent Inflammatory Functions. J. Immunol. 2017, 198, 4074–4085. [Google Scholar] [CrossRef] [PubMed]
- Sano, H.; Nakagawa, N.; Chiba, R.; Kurasawa, K.; Saito, Y.; Iwamoto, I. Cross-linking of intercellular adhesion molecule-1 induces interleukin-8 and RANTES production through the activation of MAP kinases in human vascular endothelial cells. Biochem. Biophys. Res. Commun. 1998, 250, 694–698. [Google Scholar] [CrossRef]
- de Souza, A.P.; Vale, V.L.; Silva Mda, C.; Araujo, I.B.; Trindade, S.C.; de Moura-Costa, L.F.; Rodrigues, G.C.; Sales, T.S.; dos Santos, H.A.; de Carvalho-Filho, P.C.; et al. MAPK involvement in cytokine production in response to Corynebacterium pseudotuberculosis infection. BMC Microbiol. 2014, 14, 230. [Google Scholar] [CrossRef]
- Leonard, M.; Ryan, M.P.; Watson, A.J.; Schramek, H.; Healy, E. Role of MAP kinase pathways in mediating IL-6 production in human primary mesangial and proximal tubular cells. Kidney Int. 1999, 56, 1366–1377. [Google Scholar] [CrossRef]
- Schuh, K.; Pahl, A. Inhibition of the MAP kinase ERK protects from lipopolysaccharide-induced lung injury. Biochem. Pharmacol. 2009, 77, 1827–1834. [Google Scholar] [CrossRef]
- Newton, K.; Dixit, V.M. Signaling in innate immunity and inflammation. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef]
- Guha, M.; O’Connell, M.A.; Pawlinski, R.; Hollis, A.; McGovern, P.; Yan, S.F.; Stern, D.; Mackman, N. Lipopolysaccharide activation of the MEK-ERK1/2 pathway in human monocytic cells mediates tissue factor and tumor necrosis factor alpha expression by inducing Elk-1 phosphorylation and Egr-1 expression. Blood 2001, 98, 1429–1439. [Google Scholar] [CrossRef]
- Ara, T.; Fujinami, Y.; Urano, H.; Hirai, K.; Hatori, T.; Miyazawa, H. Protein kinase A enhances lipopolysaccharide-induced IL-6, IL-8, and PGE(2) production by human gingival fibroblasts. J. Negat. Results Biomed. 2012, 11, 10. [Google Scholar] [CrossRef]
- Geng, Y.; Zhang, B.; Lotz, M. Protein tyrosine kinase activation is required for lipopolysaccharide induction of cytokines in human blood monocytes. J. Immunol. 1993, 151, 6692–6700. [Google Scholar] [PubMed]
- Lohrer, P.; Gloddek, J.; Nagashima, A.C.; Korali, Z.; Hopfner, U.; Pereda, M.P.; Arzt, E.; Stalla, G.K.; Renner, U. Lipopolysaccharide directly stimulates the intrapituitary interleukin-6 production by folliculostellate cells via specific receptors and the p38alpha mitogen-activated protein kinase/nuclear factor-kappaB pathway. Endocrinology 2000, 141, 4457–4465. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Duncan, M.J.; Li, G.; Chan, C.; Grady, R.; Stapleton, A.; Abraham, S.N. A novel TLR4-mediated signaling pathway leading to IL-6 responses in human bladder epithelial cells. PLoS Pathog. 2007, 3, e60. [Google Scholar] [CrossRef] [PubMed]
- Fronhofer, V.; Lennartz, M.R.; Loegering, D.J. Role of PKC isoforms in the Fc(gamma)R-mediated inhibition of LPS-stimulated IL-12 secretion by macrophages. J. Leukoc. Biol. 2006, 79, 408–415. [Google Scholar] [CrossRef]
- McGettrick, A.F.; Brint, E.K.; Palsson-McDermott, E.M.; Rowe, D.C.; Golenbock, D.T.; Gay, N.J.; Fitzgerald, K.A.; O’Neill, L.A. Trif-related adapter molecule is phosphorylated by PKC{epsilon} during Toll-like receptor 4 signaling. Proc. Natl. Acad. Sci. USA 2006, 103, 9196–9201. [Google Scholar] [CrossRef]
- Kontny, E.; Kurowska, M.; Szczepanska, K.; Maslinski, W. Rottlerin, a PKC isozyme-selective inhibitor, affects signaling events and cytokine production in human monocytes. J. Leukoc. Biol. 2000, 67, 249–258. [Google Scholar] [CrossRef]
- Zhou, X.; Yang, W.; Li, J. Ca2+- and protein kinase C-dependent signaling pathway for nuclear factor-kappaB activation, inducible nitric-oxide synthase expression, and tumor necrosis factor-alpha production in lipopolysaccharide-stimulated rat peritoneal macrophages. J. Biol. Chem. 2006, 281, 31337–31347. [Google Scholar] [CrossRef]
| Gene | Primer Sequence | Product Length (bp) | Accession Number |
|---|---|---|---|
| TLR1 | Forward: 5′-GCC ACC CTA CTC TGA ACC TC-3′ Reverse: 5′-ACT CAC TGT GGT GCT GAC TG-3′ | 317 | NM_001135060.2 |
| TNFα | Forward: 5′-GCC CTG GTA CGA ACC CAT CT-3′ Reverse: 5′-CAG GTA TTC CGG CAG GTT GA-3′ TaqMan probe: 5′-CCA GCT GGA GAA GGG AGA TCG CCT C-3′ | 91 | NM_001024860.1 |
| NF-kB | Forward: 5′-CAT CGA GGT TCG GTT CTA CGA-3′ Reverse: 5′-GGA GGT GTC CGG AAC ACA AT-3′ TaqMan probe: 5′-TGA GAA TGG ATG GCA AGC CTT TGG G-3′ | 108 | AF283892.1 |
| IL6 | Forward: 5′-CCA CTG CTG GTC TTC TGG AGT ATC-3′ Reverse: 5′-CTC TGC AAC TCC ATG ACA GTT TCC-3′ TaqMan probe: 5′-ACC TGG ACT TCC TCC AGA ACG AGT TTG AG-3′ | 91 | NM_001009392.1 |
| IL8 | Forward: 5′-CTC TGT GTG AAG CTG CAG TTC TG-3′ Reverse: 5′-GCA GTG TGG CCC ACT CTC AA-3′ TaqMan probe: 5′-ACA CAT TCC ACA CCT TTC CAC CCC A-3′ | 131 | NM_001009401.2 |
| GAPDH | Forward: 5′-GGC ACA GTC AAG GCA GAG AAC-3′ Reverse: 5′-CAC GTA CTC AGC ACC AGC ATC A-3′ TaqMan probe: 5′-AAG GCC ATC ACC ATC TTC CAG GAG C-3′ | 114 | NM_001190390.1 |
| ACTB | Forward: 5′-AGA GGC ATC CTG ACC CTC AA-3′ Reverse: 5′-GTT GTA GAA GGT GTG GTG CCA GAT-3′ TaqMan probe: 5′-TAC CCC ATT GAG CAC GGC ATT GTC A -3′ | 93 | U39357.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gram, A.; Kowalewski, M.P. Molecular Mechanisms of Lipopolysaccharide (LPS) Induced Inflammation in an Immortalized Ovine Luteal Endothelial Cell Line (OLENDO). Vet. Sci. 2022, 9, 99. https://doi.org/10.3390/vetsci9030099
Gram A, Kowalewski MP. Molecular Mechanisms of Lipopolysaccharide (LPS) Induced Inflammation in an Immortalized Ovine Luteal Endothelial Cell Line (OLENDO). Veterinary Sciences. 2022; 9(3):99. https://doi.org/10.3390/vetsci9030099
Chicago/Turabian StyleGram, Aykut, and Mariusz P. Kowalewski. 2022. "Molecular Mechanisms of Lipopolysaccharide (LPS) Induced Inflammation in an Immortalized Ovine Luteal Endothelial Cell Line (OLENDO)" Veterinary Sciences 9, no. 3: 99. https://doi.org/10.3390/vetsci9030099
APA StyleGram, A., & Kowalewski, M. P. (2022). Molecular Mechanisms of Lipopolysaccharide (LPS) Induced Inflammation in an Immortalized Ovine Luteal Endothelial Cell Line (OLENDO). Veterinary Sciences, 9(3), 99. https://doi.org/10.3390/vetsci9030099







