Evaluation of Candida spp. and Other Fungi in Feces from Dogs with Naturally Occurring Diabetes Mellitus
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Criteria for Selection of Cases and Study Design
2.2. Data and Sample Collection
2.3. Fungal Culture, Isolation, Quantification Methods
2.4. Identification Methods
2.5. Statistical Analysis
3. Results
3.1. Animal Population
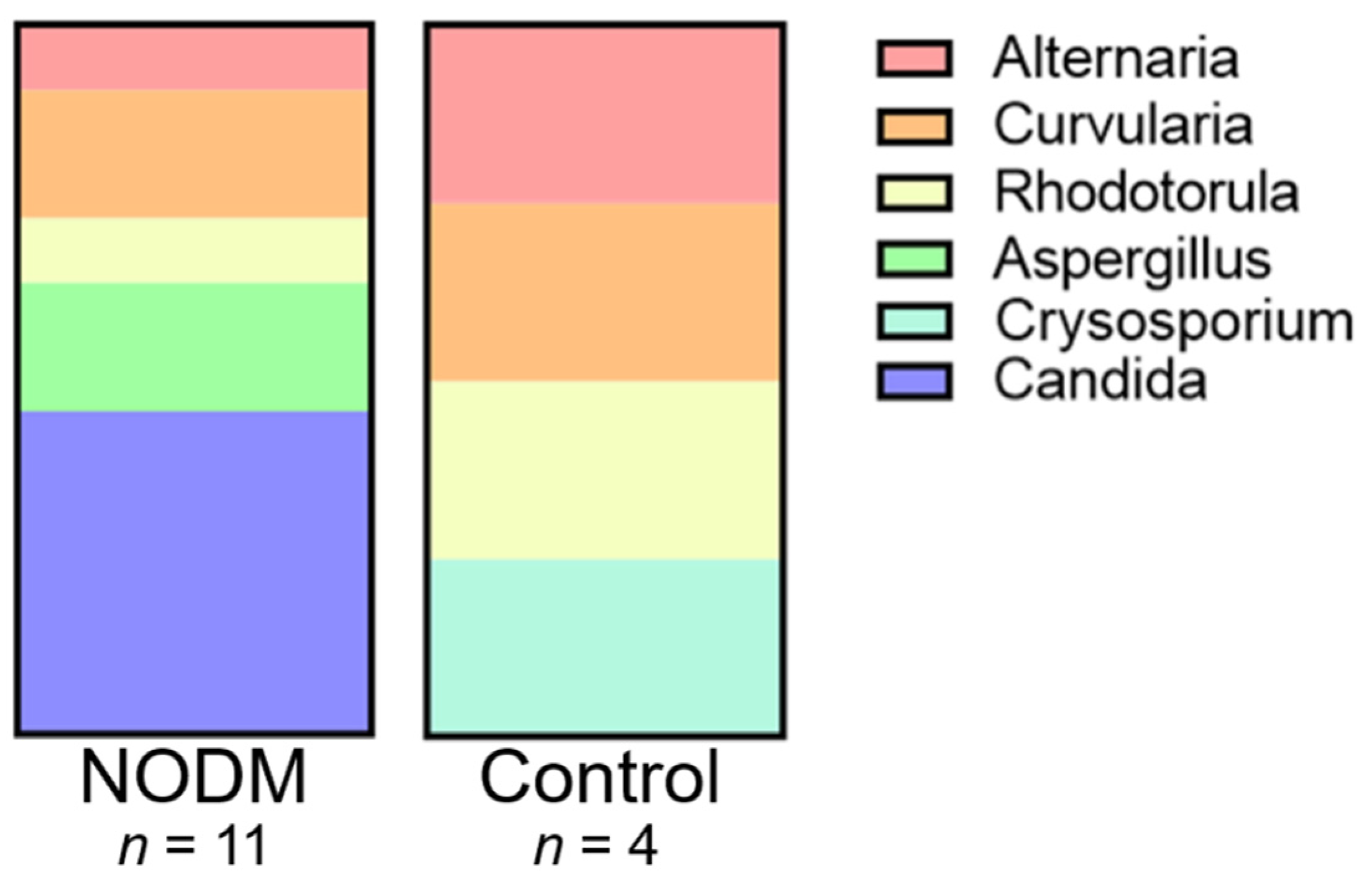
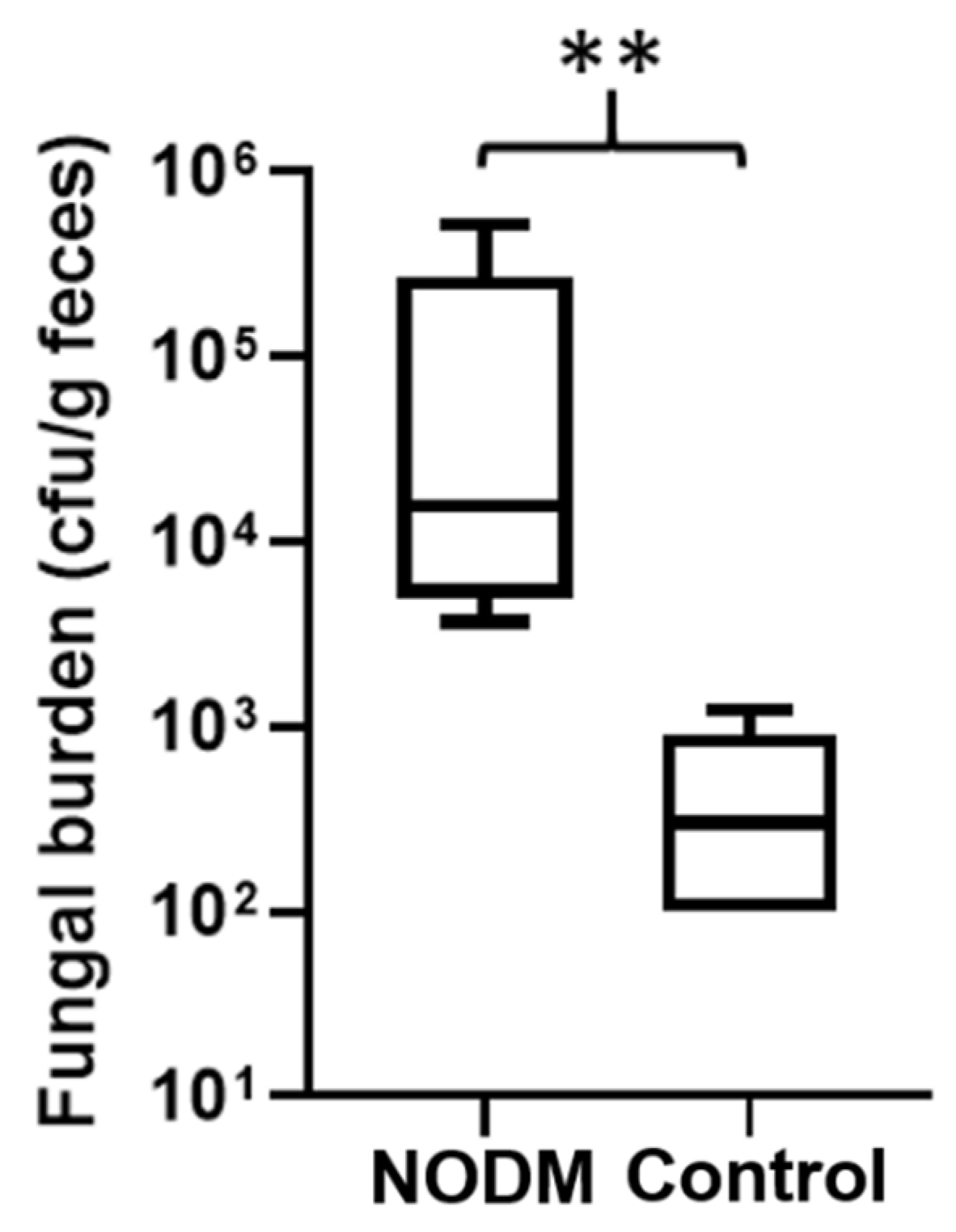
3.2. Fungi Characterization and Distribution
3.3. Risk Factors for Fungi Colonization
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hess, R.S.; Kass, P.H.; Ward, C.R. Breed distribution of dogs with diabetes mellitus admitted to a tertiary care facility. J. Am. Vet. Med. Assoc. 2000, 216, 1414–1417. [Google Scholar] [CrossRef] [PubMed]
- Guptill, L.; Glickman, L.; Glickman, N. Time trends and risk factors for diabetes mellitus in dogs: Analysis of veterinary medical data base records (1970–1999). Vet. J. 2003, 165, 240–247. [Google Scholar] [CrossRef]
- Fracassi, F.; Pietra, M.; Boari, A.; Aste, G.; Giunti, M.; Famigli-Bergamini, P. Breed distribution of canine diabetes mellitus in Italy. Vet. Res. Commun. 2004, 28 (Suppl. S1), 339–342. [Google Scholar] [CrossRef] [PubMed]
- Davison, L.J.; Herrtage, M.E.; Catchpole, B. Study of 253 dogs in the United Kingdom with diabetes mellitus. Vet. Rec. 2005, 156, 467–471. [Google Scholar] [CrossRef]
- Fall, T.; Hamlin, H.H.; Hedhammar, A.; Kampe, O.; Egenvall, A. Diabetes mellitus in a population of 180,000 insured dogs: Incidence, survival, and breed distribution. J. Vet. Intern. Med. 2007, 21, 1209–1216. [Google Scholar] [CrossRef]
- Catchpole, B.; Adams, J.P.; Holder, A.L.; Short, A.D.; Ollier, W.E.; Kennedy, L.J. Genetics of canine diabetes mellitus: Are the diabetes susceptibility genes identified in humans involved in breed susceptibility to diabetes mellitus in dogs? Vet. J. 2013, 195, 139–147. [Google Scholar] [CrossRef]
- Denyer, A.L.; Massey, J.P.; Davison, L.J.; Ollier, W.E.R.; Catchpole, B.; Kennedy, L.J. Dog leucocyte antigen (DLA) class II haplotypes and risk of canine diabetes mellitus in specific dog breeds. Canine Med. Genet. 2020, 7, 15. [Google Scholar] [CrossRef]
- Yoon, S.; Fleeman, L.M.; Wilson, B.J.; Mansfield, C.S.; McGreevy, P. Epidemiological study of dogs with diabetes mellitus attending primary care veterinary clinics in Australia. Vet. Rec. 2020, 187, e22. [Google Scholar] [CrossRef]
- Brito-Casillas, Y.; Melian, C.; Holder, A.; Wiebe, J.C.; Navarro, A.; Quesada-Canales, O.; Exposito-Montesdeoca, A.B.; Catchpole, B.; Wagner, A.M. Studying the heterogeneous pathogenesis of canine diabetes: Observational characterization of an island population. Vet. Med. Sci. 2021, 7, 1071–1081. [Google Scholar] [CrossRef]
- Moshref, M.; Tangey, B.; Gilor, C.; Papas, K.K.; Williamson, P.; Loomba-Albrecht, L.; Sheehy, P.; Kol, A. Concise review: Canine diabetes mellitus as a translational model for innovative regenerative medicine approaches. Stem Cells Transl. Med. 2019, 8, 450–455. [Google Scholar] [CrossRef]
- Gilor, C.; Niessen, S.J.; Furrow, E.; DiBartola, S.P. What’s in a name? Classification of diabetes mellitus in veterinary medicine and why it matters. J. Vet. Intern. Med. 2016, 30, 927–940. [Google Scholar] [CrossRef] [PubMed]
- Cleff, M.B.; Lima, A.P.d.; Faria, R.O.d.; Meinerz, A.R.M.; Antunes, T.d.Á.; Araújo, F.B.d.; Nascente, P.d.S.; Nobre, M.d.O.; Meireles, M.C.A. Isolation of Candida spp. from vaginal microbiota of healthy canine females during estrous cycle. Braz. J. Microbiol. 2005, 36, 201–204. [Google Scholar] [CrossRef]
- Brito, E.H.; Fontenelle, R.O.; Brilhante, R.S.; Cordeiro, R.A.; Monteiro, A.J.; Sidrim, J.J.; Rocha, M.F. The anatomical distribution and antimicrobial susceptibility of yeast species isolated from healthy dogs. Vet. J. 2009, 182, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Foster, M.L.; Dowd, S.E.; Stephenson, C.; Steiner, J.M.; Suchodolski, J.S. Characterization of the fungal microbiome (mycobiome) in fecal samples from dogs. Vet. Med. Int. 2013, 2013, 658373. [Google Scholar] [CrossRef] [PubMed]
- Kumamoto, C.A.; Gresnigt, M.S.; Hube, B. The gut, the bad and the harmless: Candida albicans as a commensal and opportunistic pathogen in the intestine. Curr. Opin. 2020, 56, 7–15. [Google Scholar] [CrossRef]
- Kuwamura, M.; Ide, M.; Yamate, J.; Shiraishi, Y.; Kotani, T. Systemic candidiasis in a dog, developing spondylitis. J. Vet. Med. Sci. 2006, 68, 1117–1119. [Google Scholar] [CrossRef]
- Rogers, C.L.; Gibson, C.; Mitchell, S.L.; Keating, J.H.; Rozanski, E.A. Disseminated candidiasis secondary to fungal and bacterial peritonitis in a young dog. J. Vet. Emerg. Crit. Care 2009, 19, 193–198. [Google Scholar] [CrossRef]
- Ong, R.K.; Raisis, A.L.; Swindells, K.L. Candida albicans peritonitis in a dog. J. Vet. Emerg. Crit. Care 2010, 20, 143–147. [Google Scholar] [CrossRef]
- Bradford, K.; Meinkoth, J.; McKeirnen, K.; Love, B. Candida peritonitis in dogs: Report of 5 cases. Vet. Clin. Pathol. 2013, 42, 227–233. [Google Scholar] [CrossRef]
- Reagan, K.L.; Dear, J.D.; Kass, P.H.; Sykes, J.E. Risk factors for Candida urinary tract infections in dogs and cats. J. Vet. Intern. Med. 2019, 33, 648–653. [Google Scholar] [CrossRef]
- Stamler, E.F.; Cruz, M.L.; Mimouni, F.; Rosenn, B.; Siddiqi, T.; Khoury, J.; Miodovnik, M. High infectious morbidity in pregnant women with insulin-dependent diabetes: An understated complication. Am. J. Obstet. Gynecol. 1990, 163, 1217–1221. [Google Scholar] [CrossRef]
- Michalopoulos, A.; Kriaras, J.; Geroulanos, S. Systemic candidiasis in cardiac surgery patients. Eur. J. Cardiothorac. Surg. 1997, 11, 728–731. [Google Scholar] [CrossRef]
- Nowakowska, D.; Kurnatowska, A.; Stray-Pedersen, B.; Wilczynski, J. Species distribution and influence of glycemic control on fungal infections in pregnant women with diabetes. J. Infect. 2004, 48, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Yismaw, G.; Asrat, D.; Woldeamanuel, Y.; Unakal, C. Prevalence of candiduria in diabetic patients attending Gondar University Hospital, Gondar, Ethiopia. Iran. J. Kidney Dis. 2013, 7, 102–107. [Google Scholar] [PubMed]
- Gosiewski, T.; Salamon, D.; Szopa, M.; Sroka, A.; Malecki, M.T.; Bulanda, M. Quantitative evaluation of fungi of the genus Candida in the feces of adult patients with type 1 and 2 diabetes-a pilot study. Gut Pathog. 2014, 6, 43. [Google Scholar] [CrossRef] [PubMed]
- Soyucen, E.; Gulcan, A.; Aktuglu-Zeybek, A.C.; Onal, H.; Kiykim, E.; Aydin, A. Differences in the gut microbiota of healthy children and those with type 1 diabetes. Pediatr. Int. 2014, 56, 336–343. [Google Scholar] [CrossRef]
- Falahati, M.; Farahyar, S.; Akhlaghi, L.; Mahmoudi, S.; Sabzian, K.; Yarahmadi, M.; Aslani, R. Characterization and identification of candiduria due to Candida species in diabetic patients. Curr. Med. Mycol. 2016, 2, 10–14. [Google Scholar] [CrossRef]
- Segal, E.; Soroka, A.; Lehrer, N. Attachment of Candida to mammalian tissues—Clinical and experimental studies. Zent. Bakteriol. Mikrobiol. Hyg. A 1984, 257, 257–265. [Google Scholar] [CrossRef]
- Wilson, R.M.; Reeves, W.G. Neutrophil phagocytosis and killing in insulin-dependent diabetes. Clin. Exp. Immunol. 1986, 63, 478–484. [Google Scholar]
- Manns, J.M.; Mosser, D.M.; Buckley, H.R. Production of a hemolytic factor by Candida albicans. Infect. Immun. 1994, 62, 5154–5156. [Google Scholar] [CrossRef]
- Manfredi, M.; McCullough, M.; Al-Karaawi, Z.; Vescovi, P.; Porter, S. In vitro evaluation of virulence attributes of Candida spp. isolated from patients affected by diabetes mellitus. Oral Microbiol. Immunol. 2006, 21, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Fatahinia, M.; Poormohamadi, F.; Zarei Mahmoudabadi, A. Comparative study of esterase and hemolytic activities in clinically Important Candida species, isolated from oral cavity of diabetic and non-diabetic individuals. Jundishapur J. Microbiol. 2015, 8, e20893. [Google Scholar] [CrossRef] [PubMed]
- Guinan, J.; Thangamani, S. Antibiotic-induced alterations in taurocholic acid levels promote gastrointestinal colonization of Candida albicans. FEMS Microbiol. Lett. 2018, 365, fny196. [Google Scholar] [CrossRef] [PubMed]
- Guinan, J.; Wang, S.; Hazbun, T.R.; Yadav, H.; Thangamani, S. Antibiotic-induced decreases in the levels of microbial-derived short-chain fatty acids correlate with increased gastrointestinal colonization of Candida albicans. Sci. Rep. 2019, 9, 8872. [Google Scholar] [CrossRef] [PubMed]
- Finkel, J.S.; Xu, W.; Huang, D.; Hill, E.M.; Desai, J.V.; Woolford, C.A.; Nett, J.E.; Taff, H.; Norice, C.T.; Andes, D.R. Portrait of Candida albicans adherence regulators. PLoS Pathog. 2012, 8, e1002525. [Google Scholar] [CrossRef] [PubMed]
- Trama, B.; Fernandes, J.D.S.; Labuto, G.; de Oliveira, J.C.F.; Viana-Niero, C.; Pascon, R.C.; Vallim, M.A. The evaluation of bioremediation potential of a yeast collection isolated from composting. Adv. Microbiol. 2014, 4, 796. [Google Scholar] [CrossRef][Green Version]
- Hallen-Adams, H.E.; Kachman, S.D.; Kim, J.; Legge, R.M.; Martínez, I. Fungi inhabiting the healthy human gastrointestinal tract: A diverse and dynamic community. Fungal Ecol. 2015, 15, 9–17. [Google Scholar] [CrossRef]
- Gursoy, S.; Kockar, T.; Atik, S.U.; Onal, Z.; Onal, H.; Adal, E. Autoimmunity and intestinal colonization by Candida albicans in patients with type 1 diabetes at the time of the diagnosis. Korean J. Pediatr. 2018, 61, 217–220. [Google Scholar] [CrossRef]
- Handl, S.; Dowd, S.E.; Garcia-Mazcorro, J.F.; Steiner, J.M.; Suchodolski, J.S. Massive parallel 16S rRNA gene pyrosequencing reveals highly diverse fecal bacterial and fungal communities in healthy dogs and cats. FEMS Microbiol. Ecol. 2011, 76, 301–310. [Google Scholar] [CrossRef]
- Swanson, K.S.; Dowd, S.E.; Suchodolski, J.S.; Middelbos, I.S.; Vester, B.M.; Barry, K.A.; Nelson, K.E.; Torralba, M.; Henrissat, B.; Coutinho, P.M.; et al. Phylogenetic and gene-centric metagenomics of the canine intestinal microbiome reveals similarities with humans and mice. ISME J. 2011, 5, 639–649. [Google Scholar] [CrossRef]
- Suchodolski, J.S.; Morris, E.K.; Allenspach, K.; Jergens, A.E.; Harmoinen, J.A.; Westermarck, E.; Steiner, J.M. Prevalence and identification of fungal DNA in the small intestine of healthy dogs and dogs with chronic enteropathies. Vet. Microbiol. 2008, 132, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, M.; Kaneko, J.; Heusner, A. Relation of fructosamine to serum protein, albumin and glucose concentrations in healthy and diabetic dogs. Am. J. Vet. Res. 1992, 53, 851–855. [Google Scholar] [PubMed]
- Reusch, C.; Liehs, M.; Hoyer, M. Fructosamine. A new parameter for diagnosis and metabolic control in diabetic dogs and cats. J. Vet. Intern. Med. 1993, 7, 177–182. [Google Scholar] [CrossRef]
- Goswami, R.; Dadhwal, V.; Tejaswi, S.; Datta, K.; Paul, A.; Haricharan, R.N.; Banerjee, U.; Kochupillai, N.P. Species-specific prevalence of vaginal candidiasis among patients with diabetes mellitus and its relation to their glycaemic status. J. Infect. 2000, 41, 162–166. [Google Scholar] [CrossRef]
- Atabek, M.E.; Akyurek, N.; Eklioglu, B.S. Frequency of vaginal candida colonization and relationship between metabolic parameters in children with type 1 diabetes mellitus. J. Pediatr. Adolesc. Gynecol. 2013, 26, 257–260. [Google Scholar] [CrossRef] [PubMed]
- Hammad, M.M.; Darwazeh, A.M.; Idrees, M.M. The effect of glycemic control on Candida colonization of the tongue and the subgingival plaque in patients with type II diabetes and periodontitis. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2013, 116, 321–326. [Google Scholar] [CrossRef]
- Shenoy, M.P.; Puranik, R.S.; Vanaki, S.S.; Puranik, S.R.; Shetty, P.; Shenoy, R. A comparative study of oral candidal species carriage in patients with type1 and type2 diabetes mellitus. J. Oral Maxillofac. Pathol. 2014, 18, S60–S65. [Google Scholar] [CrossRef]
- Rodrigues, C.F.; Rodrigues, M.E.; Henriques, M. Candida sp. infections in patients with diabetes mellitus. J. Clin. Med. 2019, 8, 76. [Google Scholar] [CrossRef]
- Elderman, M.; Hugenholtz, F.; Belzer, C.; Boekschoten, M.; van Beek, A.; de Haan, B.; Savelkoul, H.; de Vos, P.; Faas, M. Sex and strain dependent differences in mucosal immunology and microbiota composition in mice. Biol. Sex Differ. 2018, 9, 26. [Google Scholar] [CrossRef]
- Yuan, X.; Chen, R.; Zhang, Y.; Lin, X.; Yang, X. Sexual dimorphism of gut microbiota at different pubertal status. Microb. Cell Fact. 2020, 19, 152. [Google Scholar] [CrossRef]
- Lacour, M.; Zunder, T.; Huber, R.; Sander, A.; Daschner, F.; Frank, U. The pathogenetic significance of intestinal Candida colonization—A systematic review from an interdisciplinary and environmental medical point of view. Int. J. Hyg. Environ. Health 2002, 205, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Mentula, S.; Harmoinen, J.; Heikkila, M.; Westermarck, E.; Rautio, M.; Huovinen, P.; Kononen, E. Comparison between cultured small-intestinal and fecal microbiotas in beagle dogs. Appl. Environ. Microbiol. 2005, 71, 4169–4175. [Google Scholar] [CrossRef] [PubMed]
| Variable | NODM | Control | p-Value |
|---|---|---|---|
| Number of dogs | 14 | 14 | |
| Age (years) a | 9.4 (2.2) | 9.5 (2.2) | 0.94 c |
| Weight (kgs) b | 8.8 (19.2) | 9.8 (21.8) | 0.58 d |
| BCS b | 5 (1) | 5 (1) | 0.71 d |
| Sex (MN, MI, FS, FI) | 8, 0, 5, 1 | 8, 0, 5, 1 | 1.0 e |
| Breeds | Rottweiler, Pomeranian, Labrador retriever, Chihuahua, Havanese, Australian cattle dog, Bichon frise, Miniature pinscher, Yorkshire terrier, Poodle-mix, Labrador retriever-mix, Maltese-mix, Miniature pinscher-mix, Australian shepherd-mix | ||
| Variable | NODM | Control | Reference Range | p-Value |
|---|---|---|---|---|
| Number | 14 | 14 | ||
| Glucose (mg/dL) | 277.5 (311.8) | 94.5 (18.7) | 70–138 | 0.01 |
| Fructosamine (µmol/L) | 519.5 (175.5) | 258.5 (65.5) | 136–350 | <0.001 |
| Triglyceride (mg/dL) | 408.5 (792.8) | 178.0 (243.3) | 29–291 | 0.04 |
| Cholesterol (mg/dL) | 510.0 (276.8) | 249.0 (134.3) | 92–324 | <0.001 |
| Fungal Species | NODM | Control |
|---|---|---|
| Candida glabrata | 4 | 0 |
| Candida albicans | 1 | 0 |
| Alternaria alternata | 1 | 1 |
| Curvularia lunata | 1 | 1 |
| Curvularia pallascens | 1 | 0 |
| Crysosporium shanxiense | 0 | 1 |
| Rhodotorula mucilaginosa | 1 | 1 |
| Aspergillus versicolor | 1 | 0 |
| Aspergillus nidulans | 1 | 0 |
| Variable | Fungi-Positive | Fungi-Negative | p-Value |
|---|---|---|---|
| Number | 16 | 12 | |
| Age (years) a | 9.5 (2.5) | 9.4 (1.8) | 0.94 c |
| Weight (kgs) b | 8.8 (15.7) | 9.3 (39.6) | 0.69 d |
| Sex (male/female) | 6, 10 | 10, 2 | 0.02 e |
| Glucose (mg/dL) b | 157.0 (241.8) | 97.0 (120.5) | 0.28 d |
| Fructosamine (µmol/L) b | 412.5 (307.5) | 287.0 (231) | 0.30 d |
| Triglyceride (mg/dL) b | 267.5 (725.3) | 198.5 (493.3) | 0.80 d |
| Cholesterol (mg/dL) b | 372.0 (367) | 340.5 (256) | 0.76 d |
| Variable | Candida spp.-Positive | Candida spp.-Negative | p-Value |
|---|---|---|---|
| Number | 5 | 23 | |
| Age (years) a | 10.6 (1.0) | 9.2 (2.3) | 0.19 c |
| Weight (kgs) b | 23.4 (25.2) | 8.8 (11.6) | 0.47 d |
| Sex (male/female) | 3, 2 | 13, 10 | 1.00 e |
| Glucose (mg/dL) b | 348.0 (226.5) | 100 (123.0) | 0.01 d |
| Fructosamine (µmol/L) b | 594.0 (67.0) | 279.0 (190.0) | 0.002 d |
| Triglyceride (mg/dL) b | 513.0 (684.0) | 179.0 (487.0) | 0.07 d |
| Cholesterol (mg/dL) b | 608.0 (475.5) | 301.0 (205.0) | 0.01 d |
| Variable | Candida spp.-Positive | Candida spp.-Negative | p-Value |
|---|---|---|---|
| Number | 5 | 9 | |
| Age (years) a | 10.6 (1.0) | 8.8 (2.4) | 0.13 c |
| Weight (kgs) b | 23.4 (25.2) | 8.4 (6.7) | 0.29 d |
| Sex (male/female) | 3, 2 | 5, 4 | 1.00 e |
| Glucose (mg/dL) b | 372.2 (114.6) | 271.6 (216.5) | 0.36 c |
| Fructosamine (µmol/L) a | 588.8 (34.4) | 428.7 (141.0) | 0.03 c |
| Triglyceride (mg/dL) b | 513.0 (684.0) | 211.0 (807.5) | 0.42 d |
| Cholesterol (mg/dL) a | 672.2 (309.2) | 479.7 (194.7) | 0.17 c |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jaffey, J.A.; Okwumabua, O.; Graves, T.K.; Al-Nakkash, L.; Monasky, R.; Wilson, A.; Thangamani, S. Evaluation of Candida spp. and Other Fungi in Feces from Dogs with Naturally Occurring Diabetes Mellitus. Vet. Sci. 2022, 9, 567. https://doi.org/10.3390/vetsci9100567
Jaffey JA, Okwumabua O, Graves TK, Al-Nakkash L, Monasky R, Wilson A, Thangamani S. Evaluation of Candida spp. and Other Fungi in Feces from Dogs with Naturally Occurring Diabetes Mellitus. Veterinary Sciences. 2022; 9(10):567. https://doi.org/10.3390/vetsci9100567
Chicago/Turabian StyleJaffey, Jared A., Ogi Okwumabua, Thomas K. Graves, Layla Al-Nakkash, Ross Monasky, Alec Wilson, and Shankar Thangamani. 2022. "Evaluation of Candida spp. and Other Fungi in Feces from Dogs with Naturally Occurring Diabetes Mellitus" Veterinary Sciences 9, no. 10: 567. https://doi.org/10.3390/vetsci9100567
APA StyleJaffey, J. A., Okwumabua, O., Graves, T. K., Al-Nakkash, L., Monasky, R., Wilson, A., & Thangamani, S. (2022). Evaluation of Candida spp. and Other Fungi in Feces from Dogs with Naturally Occurring Diabetes Mellitus. Veterinary Sciences, 9(10), 567. https://doi.org/10.3390/vetsci9100567





