Integrating Network Pharmacology and Molecular Docking to Analyse the Potential Mechanism of action of Macleaya cordata (Willd.) R. Br. in the Treatment of Bovine Hoof Disease
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data Collection
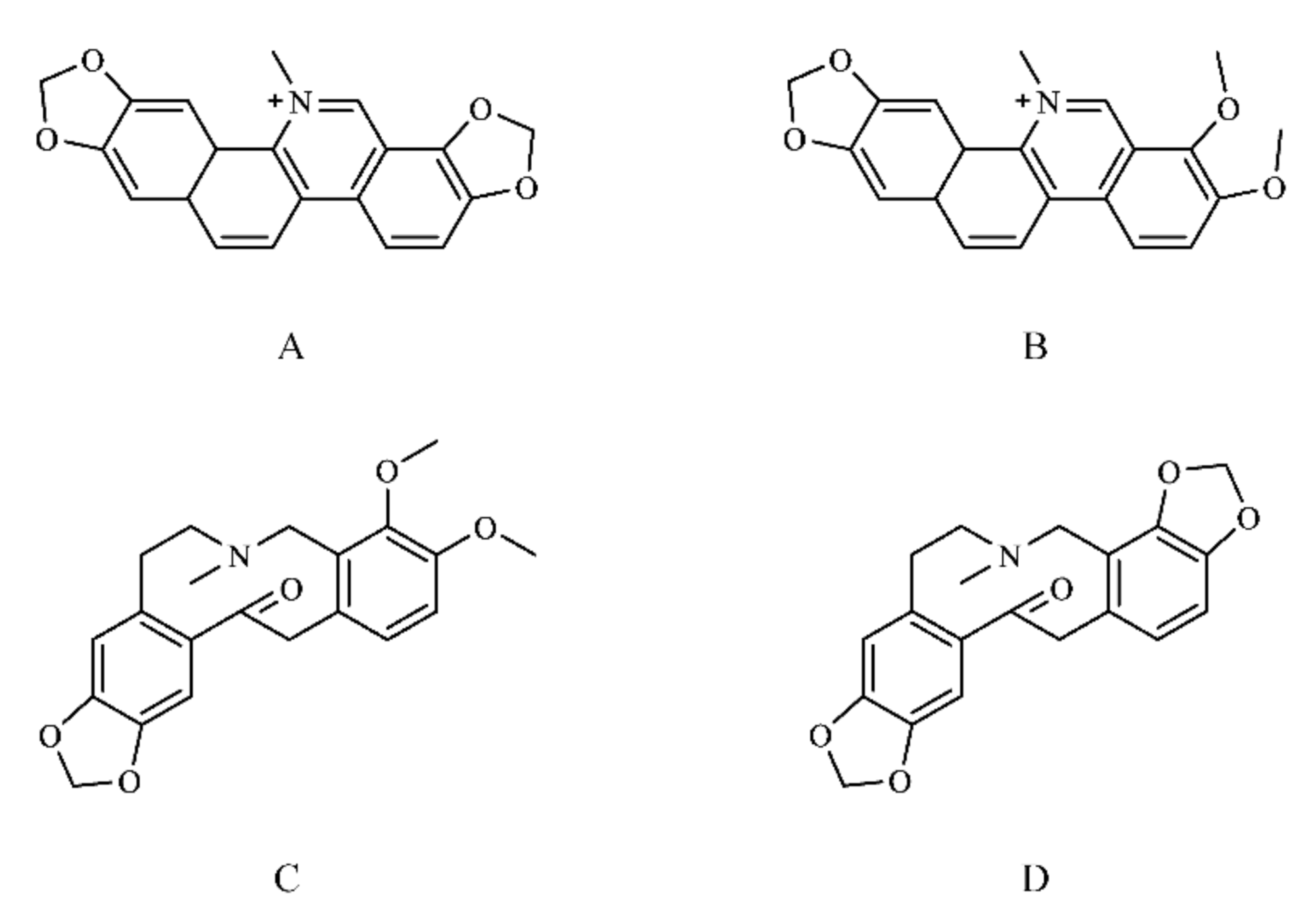
2.1.1. Chemical Composition of M. cordata
2.1.2. ADME Pharmacokinetic Evaluation
2.1.3. Target Gene Prediction for the Selected Compounds
2.1.4. Target Gene Screening for Disease
2.1.5. Venn Analysis
2.2. Network Topology Analysis and Protein Interactions Network Construction
2.2.1. Network Topology Analysis
2.2.2. Protein Interaction Network Construction
2.2.3. Enrichment Analysis
2.2.4. Molecular Docking
3. Results
3.1. Data Collection
3.1.1. Screening of Compounds in the M. cordata
3.1.2. ADME Screening and Associated Target Gene Prediction
3.1.3. Disease Target Gene Acquisition
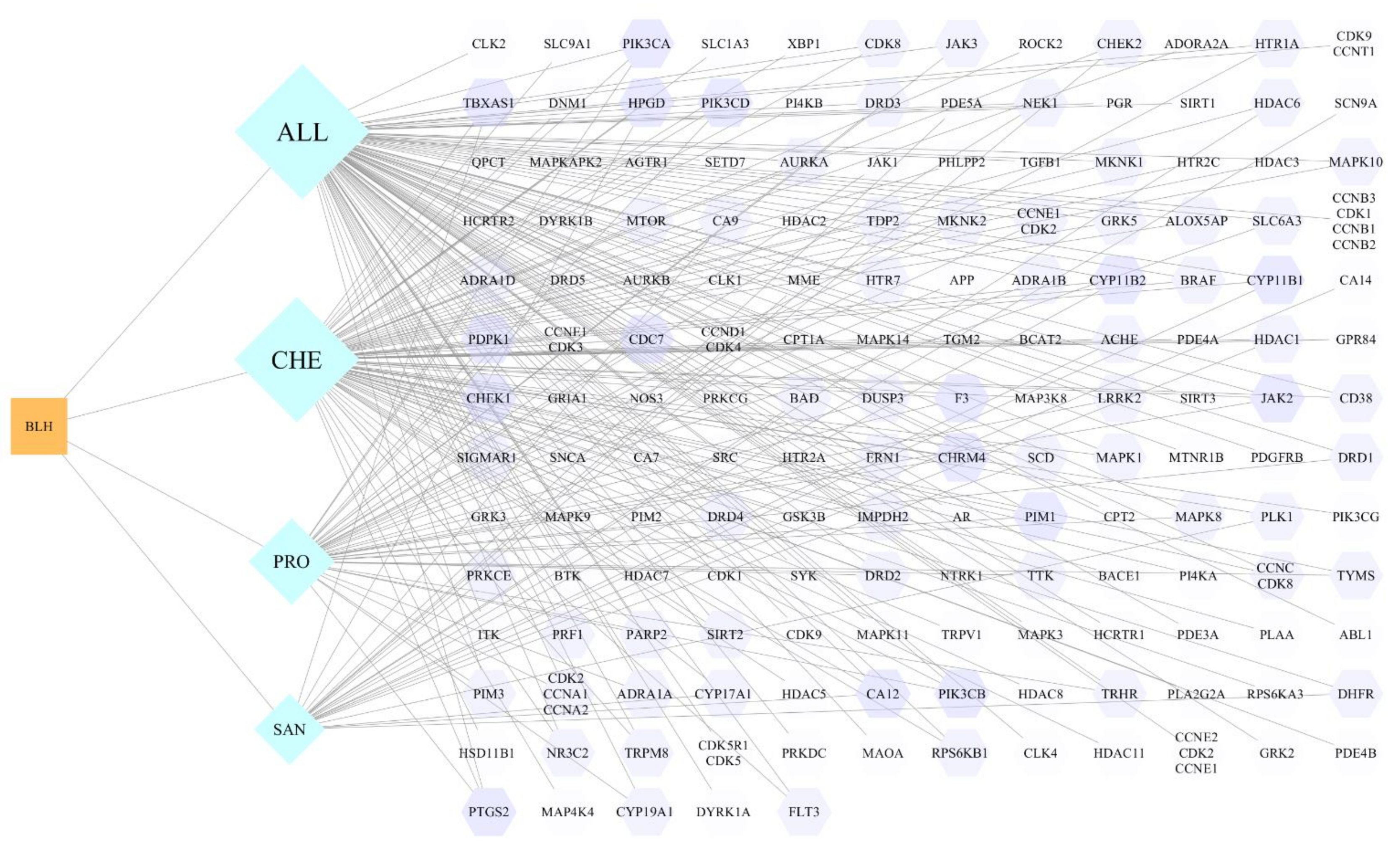
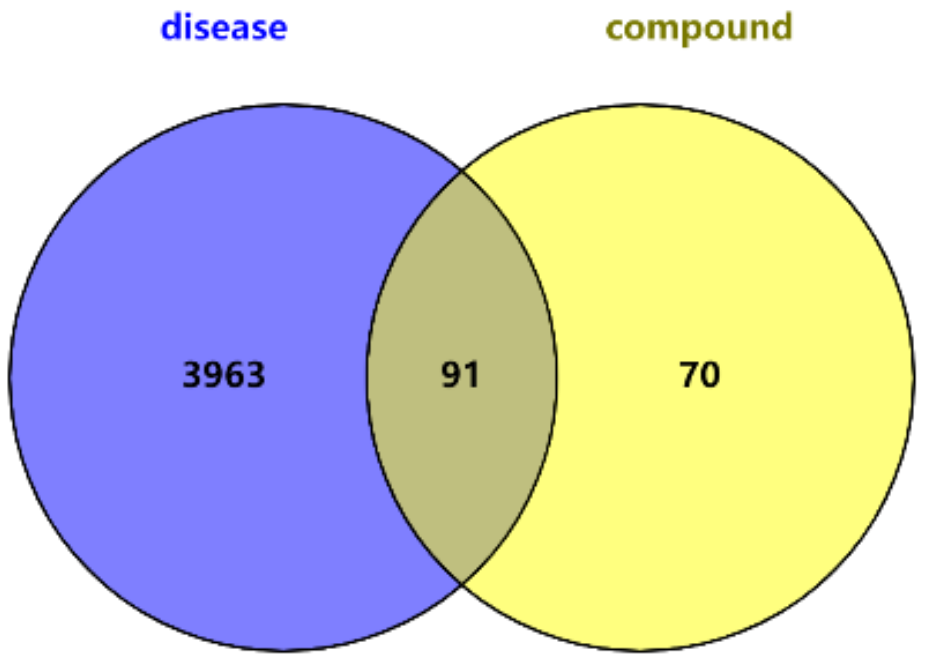
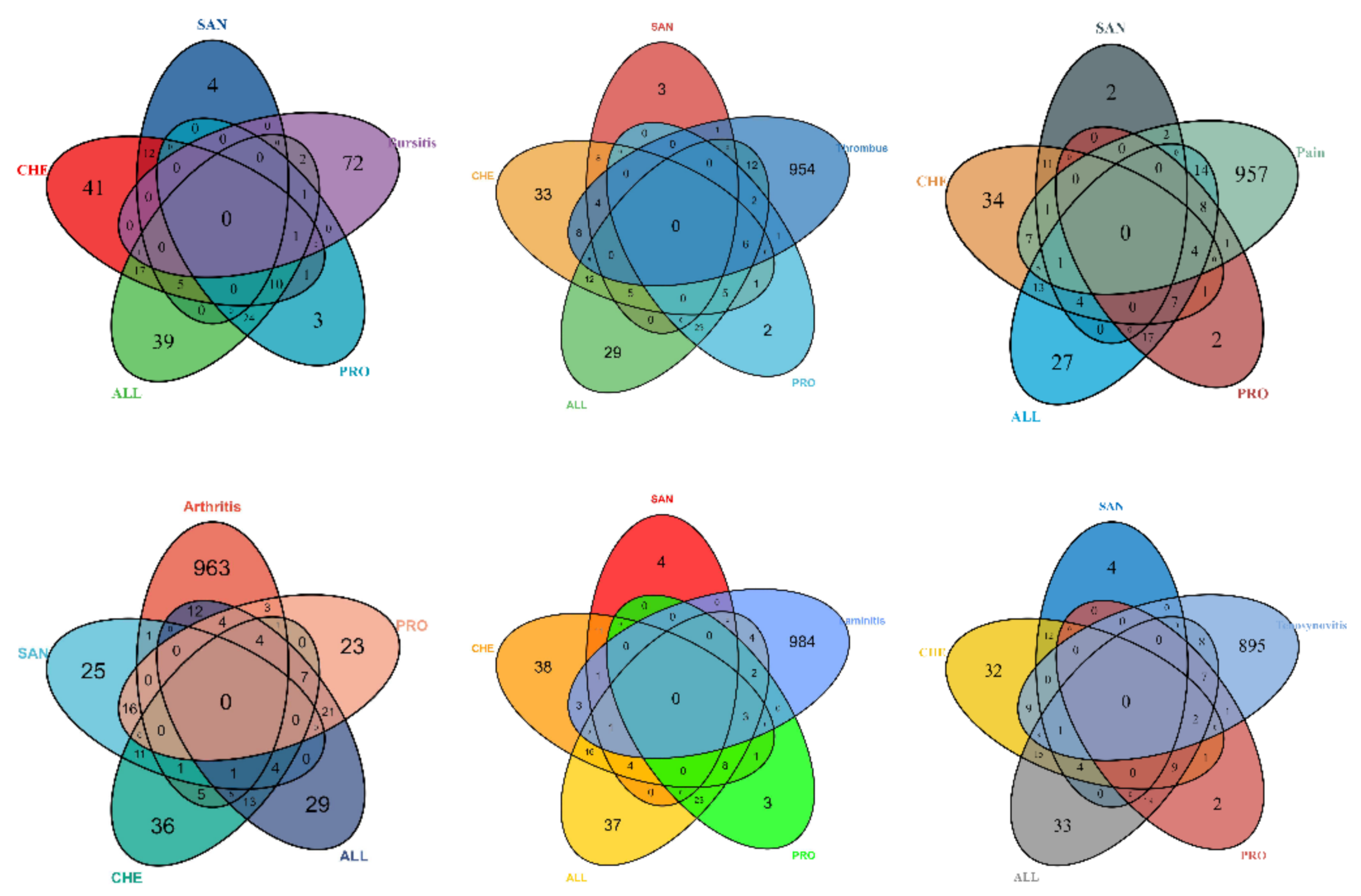
3.1.4. Genetic Intersection of Compounds and Disease Targets
3.2. Network Topology Analysis and PPI Network Construction
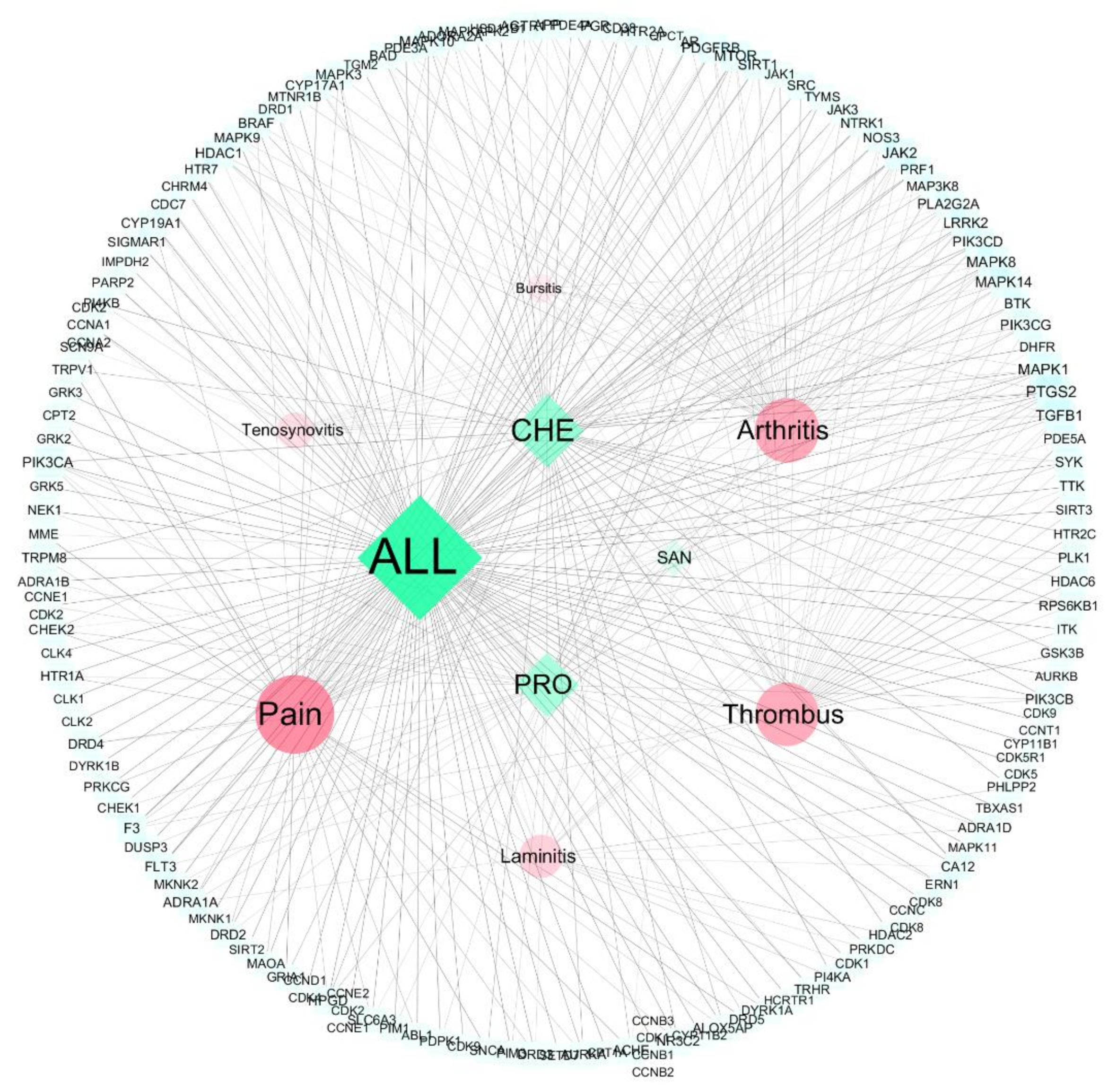
3.2.1. Network Topology Analysis
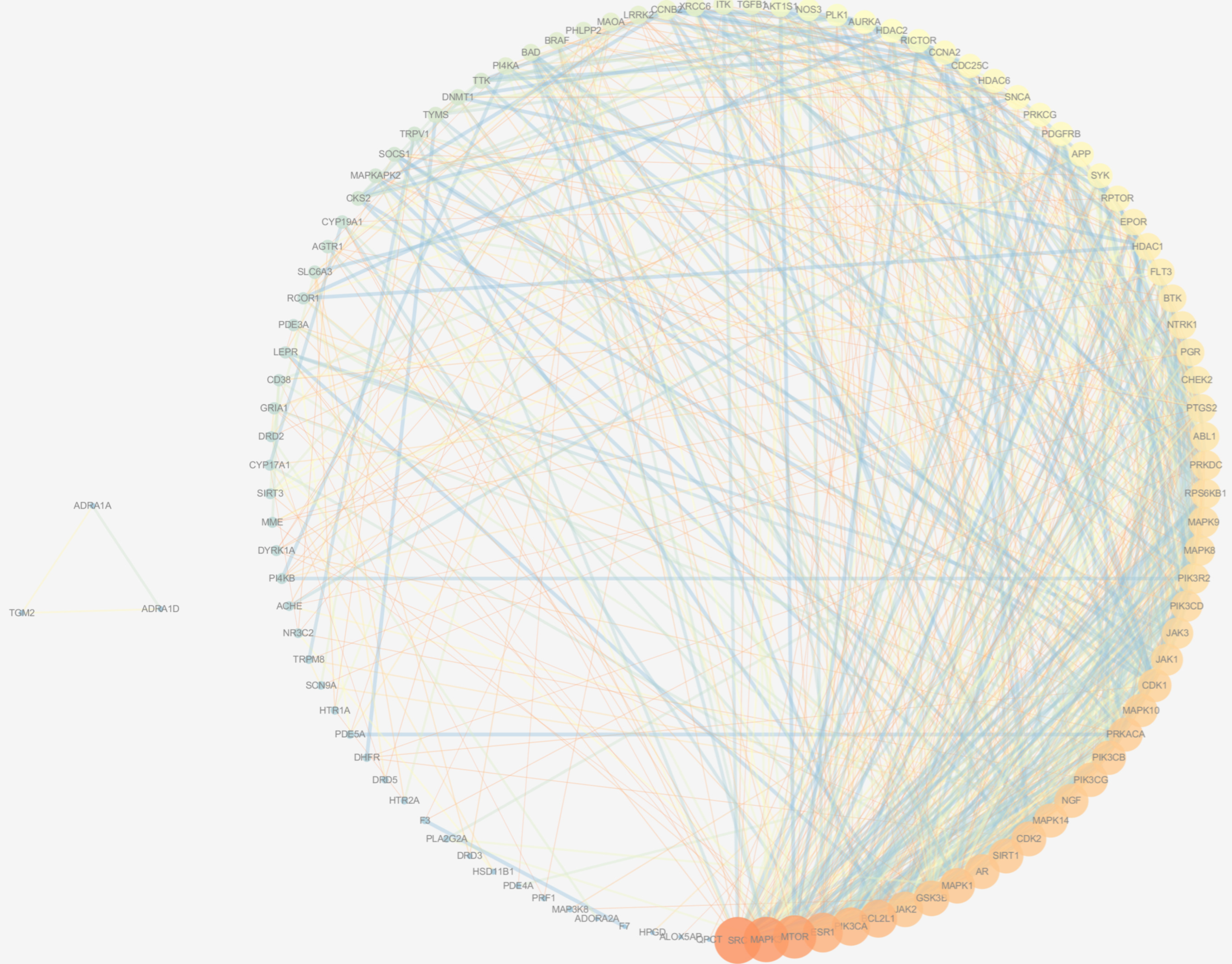
3.2.2. PPI Network
3.3. Enrichment Analysis
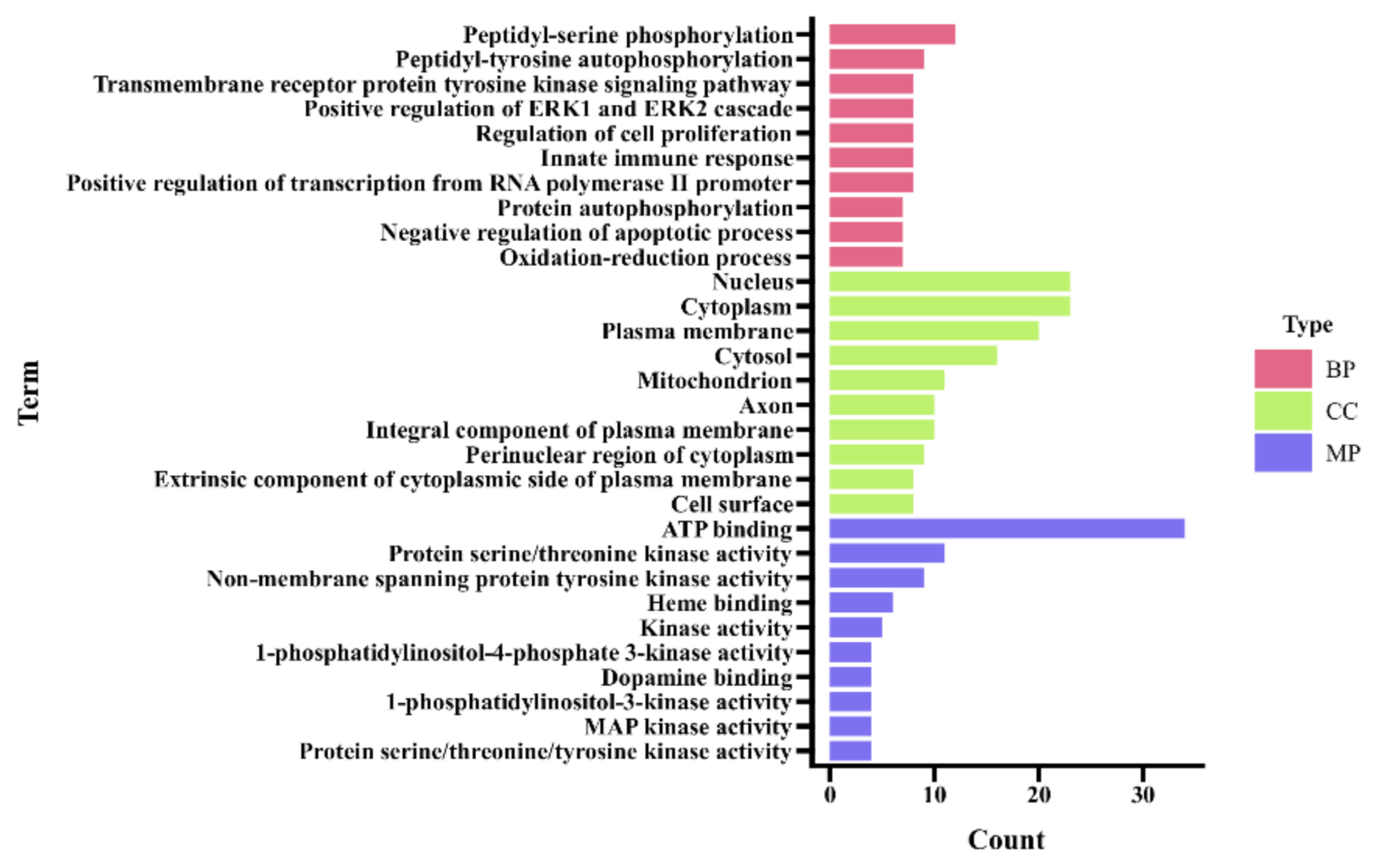
3.3.1. GO Enrichment Analysis
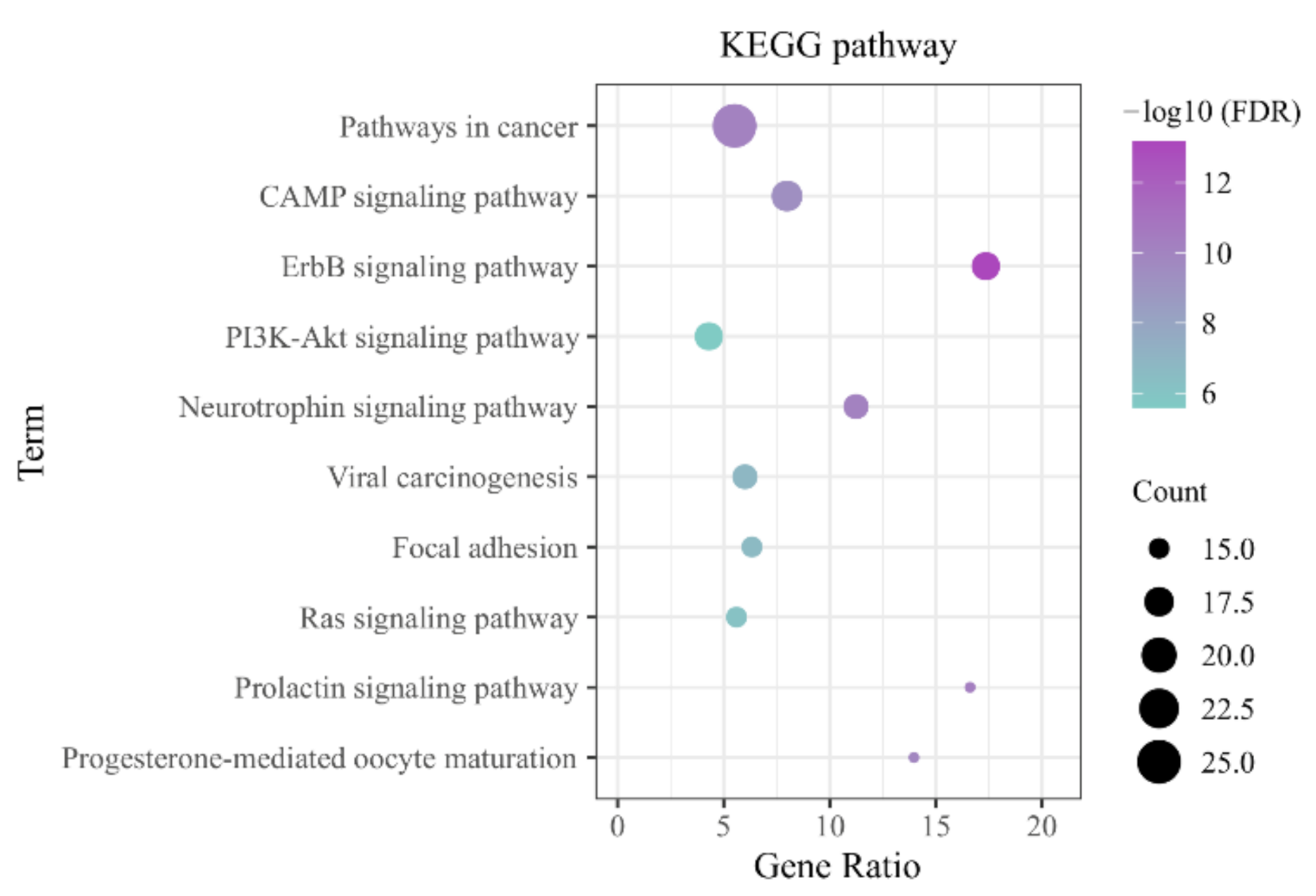
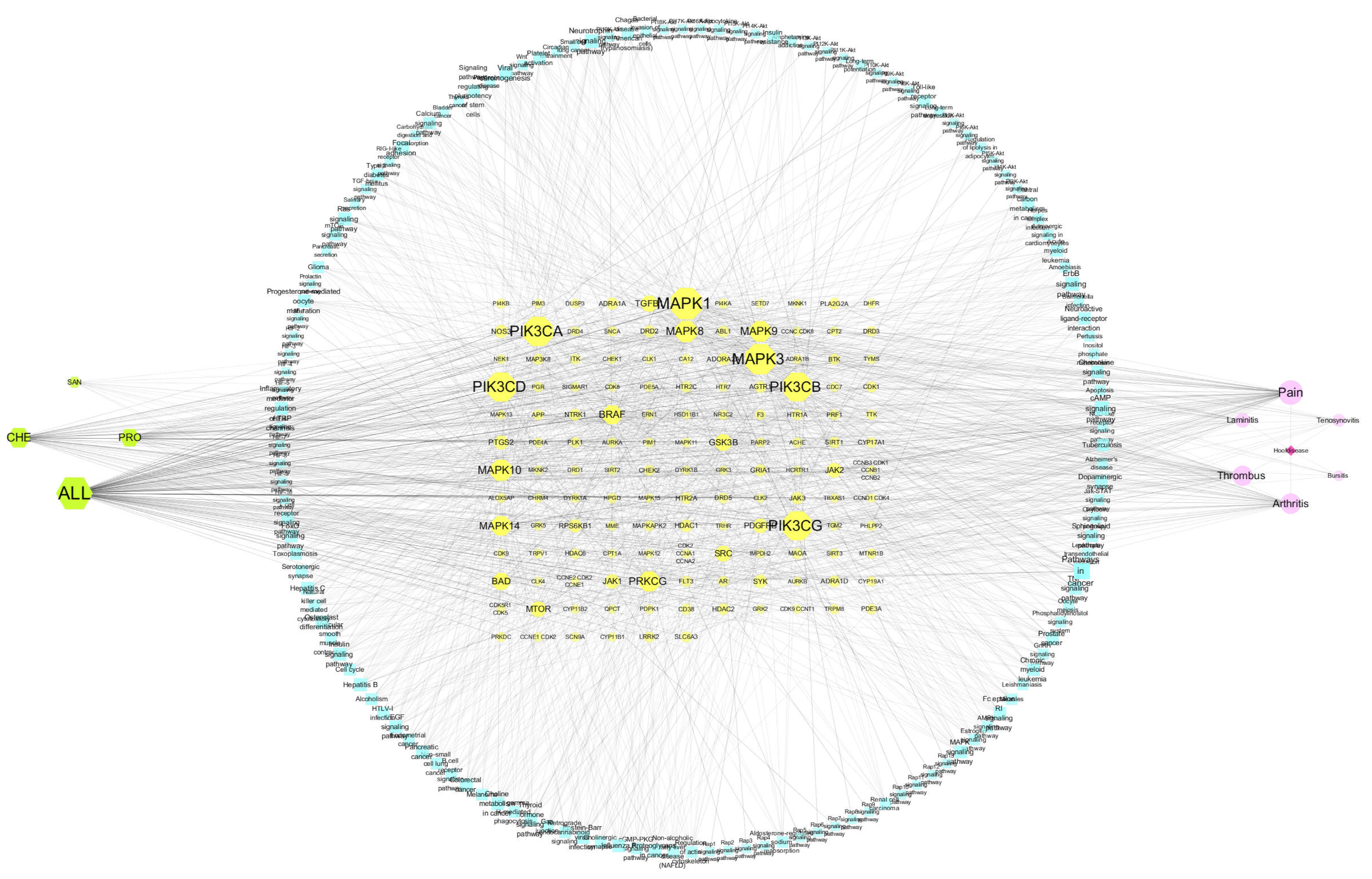
3.3.2. KEGG Pathway Enrichment
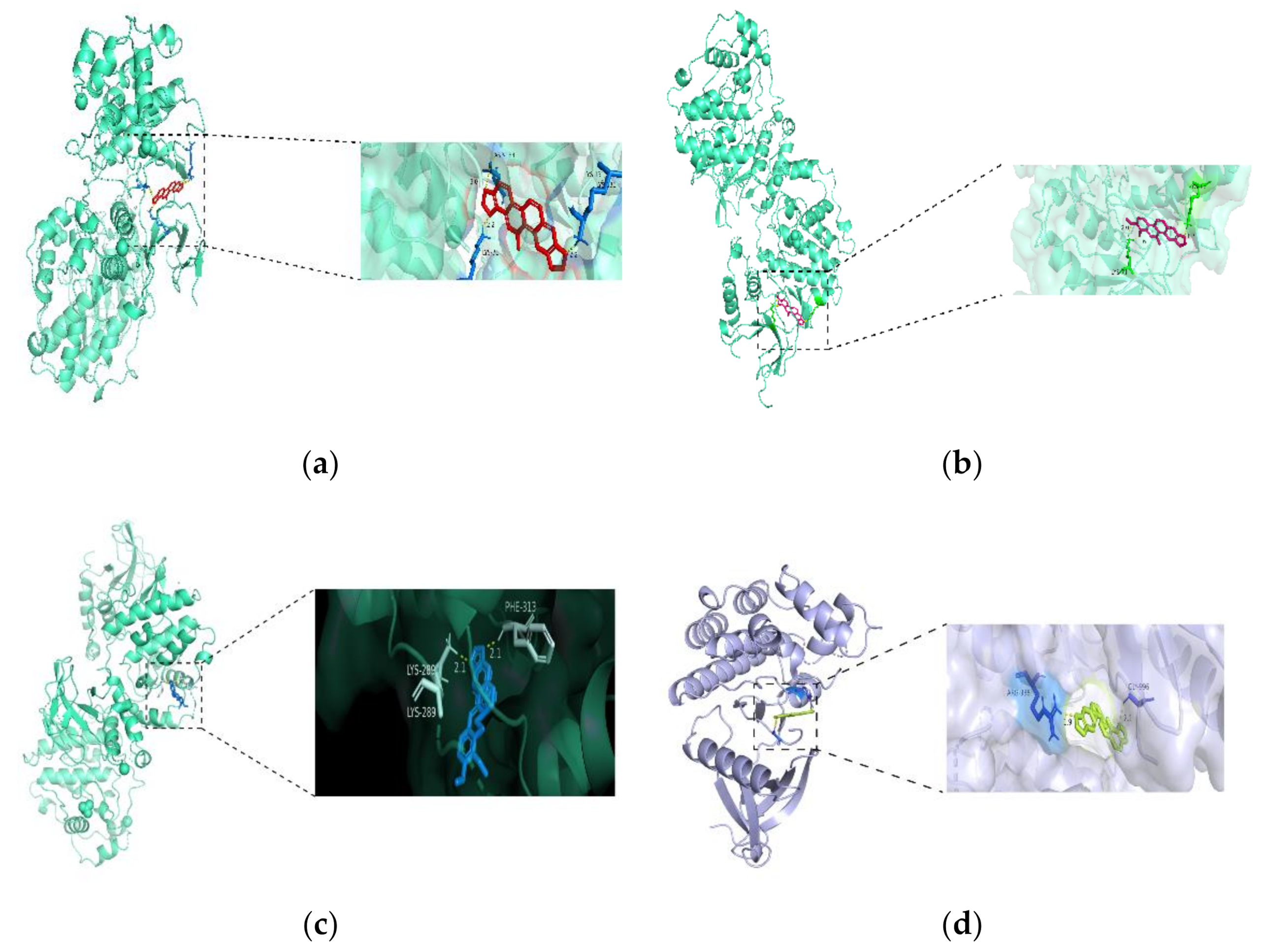
3.4. Molecular Docking
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Beaver, A.; Weary, D.M.; von Keyserlingk, M.A.G. Invited Review: The Welfare of Dairy Cattle Housed in Tiestalls Compared to Less-Restrictive Housing Types: A Systematic Review. J. Dairy Sci. 2021, 104, 9383–9417. [Google Scholar] [CrossRef] [PubMed]
- Goetz, T.E. Anatomic, Hoof, and Shoeing Considerations for the Treatment of Laminitis in Horses. J. Am. Vet. Med. Assoc. 1987, 190, 1323–1332. [Google Scholar] [PubMed]
- Al-Agele, R.; Paul, E.; Taylor, S.; Watson, C.; Sturrock, C.; Drakopoulos, M.; Atwood, R.C.; Rutland, C.S.; Menzies-Gow, N.; Knowles, E.; et al. Physics of Animal Health: On the Mechano-Biology of Hoof Growth and Form. J. R. Soc. Interface 2019, 16, 20190214. [Google Scholar] [CrossRef] [PubMed]
- Bergsten, C. Effects of Conformation and Management System on Hoof and Leg Diseases and Lameness in Dairy Cows. Vet. Clin. N. Am. Food Anim. Pract. 2001, 17, 1–23. [Google Scholar] [CrossRef]
- Bindu, S.; Mazumder, S.; Bandyopadhyay, U. Non-Steroidal Anti-Inflammatory Drugs (NSAIDs) and Organ Damage: A Current Perspective. Biochem. Pharmacol. 2020, 180, 114147. [Google Scholar] [CrossRef]
- Williamson, D.A.; Carter, G.P.; Howden, B.P. Current and Emerging Topical Antibacterials and Antiseptics: Agents, Action, and Resistance Patterns. Clin. Microbiol. Rev. 2017, 30, 827–860. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, P.; Zhang, Y.; Xiao, K.; Jiang, F.; Wang, H.; Tang, D.; Liu, D.; Liu, B.; Liu, Y.; He, X.; et al. The Chicken Gut Metagenome and the Modulatory Effects of Plant-Derived Benzylisoquinoline Alkaloids. Microbiome 2018, 6, 211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Kang, B.; Zhao, Y.; Yao, K.; Fu, C. Effects of Natural Dietary Supplementation with Macleaya Cordata Extract Containing Sanguinarine on Growth Performance and Gut Health of Early-Weaned Piglets. J. Anim. Physiol. Anim. Nutr. 2018, 102, 1666–1674. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhang, J.Q.; Han, X.G.; Wang, Z.L.; Xu, Y.Y.; Miao, J.F. Macleaya Cordata Helps Improve the Growth-Promoting Effect of Chlortetracycline on Broiler Chickens. J. Zhejiang Univ. Sci. B 2018, 19, 776–784. [Google Scholar] [CrossRef] [PubMed]
- Khadem, A.; Soler, L.; Everaert, N.; Niewold, T.A. Growth Promotion in Broilers by Both Oxytetracycline and Macleaya Cordata Extract Is Based on Their Anti-Inflammatory Properties. Br. J. Nutr. 2014, 112, 1110–1118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Q.; Zhang, Z.; Zhou, S.; Jin, M.; Lu, T.; Cui, L.; Qian, H. Macleaya Cordata Extract, an Antibiotic Alternative, Does Not Contribute to Antibiotic Resistance Gene Dissemination. J. Hazard. Mater. 2021, 412, 125272. [Google Scholar] [CrossRef]
- Editorial Committee of the Chinese Materia Medica. State Administration of Traditional Chinese Medicine Chinese Materia Madica-3; Shanghai Science and Technology Press: Shanghai, China, 1999. [Google Scholar]
- van Hasselt, J.; Iyengar, R. Systems Pharmacology: Defining the Interactions of Drug Combinations. Annu. Rev. Pharmacol. Toxicol. 2019, 59, 21–40. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Li, Y. Systems Pharmacology; Dalian University of Technology Press: Dalian, China, 2016. [Google Scholar]
- Wu, Q.; Yang, F.; Tang, H. Based on Network Pharmacology Method to Discovered the Targets and Therapeutic Mechanism of Paederia Scandens against Nonalcoholic Fatty Liver Disease in Chicken. Poult. Sci. 2021, 100, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Bai, D.D.; Liu, L.L.; Bai, Y.B.; Wang, W.W.; Zhang, J.Y.; Zhou, X.Z. The Mechanism of Gui Qi Yimu Decoction Powder in Treating Cow Qi and Blood Two Deficiency Syndrome Based on Network Pharmacology. Acta Vet. Et Zootech. Sin. 2018, 49, 2733–2744. [Google Scholar]
- Shanghai Institute of Organic Chemistry of CAS Chemistry Database [DB/OL]. Available online: http://www.organchem.csdb.cn (accessed on 12 October 2021).
- MacCoss, M.; Baillie, T.A. Organic Chemistry in Drug Discovery. Science 2004, 303, 1810–1813. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound Databases. Nucleic Acids Res. 2016, 44, D1202-13. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated Data and New Features for Efficient Prediction of Protein Targets of Small Molecules. Nucleic Acids Res. 2019, 47, W357–W364. [Google Scholar] [CrossRef] [Green Version]
- UniProt: The Universal Protein Knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D480–D489. [CrossRef] [PubMed]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Stein, T.I.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr. Protoc. Bioinform. 2016, 54, 1.30.1–1.30.33. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING Database in 2021: Customizable Protein-Protein Networks, and Functional Characterization of User-Uploaded Gene/Measurement Sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics Enrichment Tools: Paths toward the Comprehensive Functional Analysis of Large Gene Lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and Integrative Analysis of Large Gene Lists Using DAVID Bioinformatics Resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Burley, S.K.; Bhikadiya, C.; Bi, C.; Bittrich, S.; Chen, L.; Crichlow, G.V.; Christie, C.H.; Dalenberg, K.; Di Costanzo, L.; Duarte, J.M.; et al. RCSB Protein Data Bank: Powerful New Tools for Exploring 3D Structures of Biological Macromolecules for Basic and Applied Research and Education in Fundamental Biology, Biomedicine, Biotechnology, Bioengineering and Energy Sciences. Nucleic Acids Res. 2021, 49, D437–D451. [Google Scholar] [CrossRef] [PubMed]
- Seeliger, D.; de Groot, B.L. Ligand Docking and Binding Site Analysis with PyMOL and Autodock/Vina. J. Comput. Aided. Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef] [Green Version]
- Pěnčíková, K.; Urbanová, J.; Musil, P.; Táborská, E.; Gregorová, J. Seasonal Variation of Bioactive Alkaloid Contents in Macleaya Microcarpa (Maxim.) Fedde. Molecules 2011, 16, 3391–3401. [Google Scholar] [CrossRef]
- Liu, X.; Liu, Y.; Huang, P.; Ma, Y.; Qing, Z.; Tang, Q.; Cao, H.; Cheng, P.; Zheng, Y.; Yuan, Z.; et al. The Genome of Medicinal Plant Macleaya Cordata Provides New Insights into Benzylisoquinoline Alkaloids Metabolism. Mol. Plant 2017, 10, 975–989. [Google Scholar] [CrossRef] [Green Version]
- Udrescu, L.; Bogdan, P.; Chis, A.; Sirbu, I.O.; Topirceanu, A.; Varut, R.M.; Udrescu, M. Uncovering New Drug Properties in Target-Based Drug-Drug Similarity Networks. Pharmaceutics 2020, 12, 879. [Google Scholar] [CrossRef]
- Wang, X.; Wang, Z.Y.; Zheng, J.H.; Li, S. TCM Network Pharmacology: A New Trend towards Combining Computational, Experimental and Clinical Approaches. Chin. J. Nat. Med. 2021, 19, 1–11. [Google Scholar] [CrossRef]
- Liao, S.; Han, L.; Zheng, X.; Wang, X.; Zhang, P.; Wu, J.; Liu, R.; Fu, Y.; Sun, J.; Kang, X.; et al. Tanshinol Borneol Ester, a Novel Synthetic Small Molecule Angiogenesis Stimulator Inspired by Botanical Formulations for Angina Pectoris. Br. J. Pharmacol. 2019, 176, 3143–3160. [Google Scholar] [CrossRef]
- Alam, M.B.; Ju, M.-K.; Kwon, Y.-G.; Lee, S.H. Protopine Attenuates Inflammation Stimulated by Carrageenan and LPS via the MAPK/NF-ΚB Pathway. Food Chem. Toxicol. 2019, 131, 110583. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Liu, W.; Liu, M.; Zhang, J.; Yang, M.; Wang, T.; Qian, W. In Vitro Anti-Biofilm Efficacy of Sanguinarine against Carbapenem-Resistant Serratia Marcescens. Biofouling 2021, 37, 341–351. [Google Scholar] [CrossRef] [PubMed]
- Wong-Deyrup, S.W.; Song, X.; Ng, T.-W.; Liu, X.-B.; Zeng, J.-G.; Qing, Z.-X.; Deyrup, S.T.; He, Z.-D.; Zhang, H.-J. Plant-Derived Isoquinoline Alkaloids That Target Ergosterol Biosynthesis Discovered by Using a Novel Antifungal Screening Tool. Biomed. Pharmacother. 2021, 137, 111348. [Google Scholar] [CrossRef] [PubMed]
- Jampilek, J.; Brychtova, K. Azone Analogues: Classification, Design, and Transdermal Penetration Principles. Med. Res. Rev. 2012, 32, 907–947. [Google Scholar] [CrossRef]
- Supe, S.; Takudage, P. Methods for Evaluating Penetration of Drug into the Skin: A Review. Skin Res. Technol. 2021, 27, 299–308. [Google Scholar] [CrossRef]
- Wu, Y.; Zhao, N.-J.; Cao, Y.; Sun, Z.; Wang, Q.; Liu, Z.-Y.; Sun, Z.-L. Sanguinarine Metabolism and Pharmacokinetics Study in Vitro and in Vivo. J. Vet. Pharmacol. Ther. 2020, 43, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Hu, N.-X.; Chen, M.; Liu, Y.-S.; Shi, Q.; Yang, B.; Zhang, H.-C.; Cheng, P.; Tang, Q.; Liu, Z.-Y.; Zeng, J.-G. Pharmacokinetics of Sanguinarine, Chelerythrine, and Their Metabolites in Broiler Chickens Following Oral and Intravenous Administration. J. Vet. Pharmacol. Ther. 2019, 42, 197–206. [Google Scholar] [CrossRef]
- Orland, A.; Knapp, K.; König, G.M.; Ulrich-Merzenich, G.; Knöß, W. Combining Metabolomic Analysis and Microarray Gene Expression Analysis in the Characterization of the Medicinal Plant Chelidonium Majus L. Phytomedicine 2014, 21, 1587–1596. [Google Scholar] [CrossRef]
- Gao, L.; Schmitz, H.-J.; Merz, K.-H.; Schrenk, D. Characterization of the Cytotoxicity of Selected Chelidonium Alkaloids in Rat Hepatocytes. Toxicol. Lett. 2019, 311, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Mazzanti, G.; Di Sotto, A.; Franchitto, A.; Mammola, C.L.; Mariani, P.; Mastrangelo, S.; Menniti-Ippolito, F.; Vitalone, A. Chelidonium Majus Is Not Hepatotoxic in Wistar Rats, in a 4 Weeks Feeding Experiment. J. Ethnopharmacol. 2009, 126, 518–524. [Google Scholar] [CrossRef]
- De Jong, W.H.; Borm, P.J.A. Drug Delivery and Nanoparticles:Applications and Hazards. Int. J. Nanomed. 2008, 3, 133–149. [Google Scholar] [CrossRef] [Green Version]
- European Medicines Agency. Assessment Report on Chelidonium Majus L., Herba; Committee on Herbal Medicinal Products (HMPC): London, UK, 2011. [Google Scholar]
- Zielińska, S.; Jezierska-Domaradzka, A.; Wójciak-Kosior, M.; Sowa, I.; Junka, A.; Matkowski, A.M. Greater Celandine’s Ups and Downs−21 Centuries of Medicinal Uses of Chelidonium Majus From the Viewpoint of Today’s Pharmacology. Front. Pharmacol. 2018, 9, 299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fravor, L.; Khachemoune, A. Dermatologic Uses of Bloodroot: A Review and Reappraisal. Int. J. Dermatol. 2021, 60, 1070–1075. [Google Scholar] [CrossRef]
- Miyazaki, T.; Neff, L.; Tanaka, S.; Horne, W.C.; Baron, R. Regulation of Cytochrome c Oxidase Activity by C-Src in Osteoclasts. J. Cell Biol. 2003, 160, 709–718. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taniguchi, K.; Wu, L.W.; Grivennikov, S.I.; de Jong, P.R.; Lian, I.; Yu, F.X.; Wang, K.; Ho, S.B.; Boland, B.S.; Chang, J.T.; et al. A Gp130-Src-YAP Module Links Inflammation to Epithelial Regeneration. Nature 2015, 519, 57–62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harbi, M.H.; Smith, C.W.; Nicolson, P.L.R.; Watson, S.P.; Thomas, M.R. Novel Antiplatelet Strategies Targeting GPVI, CLEC-2 and Tyrosine Kinases. Platelets 2021, 32, 29–41. [Google Scholar] [CrossRef]
- Song, J.; Ye, B.; Liu, H.; Bi, R.; Zhang, N.; Hu, J.; Luo, E. Fak-Mapk, Hippo and Wnt Signalling Pathway Expression and Regulation in Distraction Osteogenesis. Cell Prolif. 2018, 51, e12453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Boehm, J.; Lee, J.C. P38 MAP Kinases: Key Signalling Molecules as Therapeutic Targets for Inflammatory Diseases. Nat. Rev. Drug Discov. 2003, 2, 717–726. [Google Scholar] [CrossRef]
- Yu, X.; Quan, J.; Long, W.; Chen, H.; Wang, R.; Guo, J.; Lin, X.; Mai, S. LL-37 Inhibits LPS-Induced Inflammation and Stimulates the Osteogenic Differentiation of BMSCs via P2X7 Receptor and MAPK Signaling Pathway. Exp. Cell Res. 2018, 372, 178–187. [Google Scholar] [CrossRef]
- Dai, Q.; Zhou, D.; Xu, L.; Song, X. Curcumin Alleviates Rheumatoid Arthritis-Induced Inflammation and Synovial Hyperplasia by Targeting MTOR Pathway in Rats. Drug Des. Dev. Ther. 2018, 12, 4095–4105. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Deng, Z.; Ma, Y.; Jin, J.; Qi, F.; Li, S.; Liu, C.; Lyu, F.J.; Zheng, Q. The Role of Autophagy and Mitophagy in Bone Metabolic Disorders. Int. J. Biol. Sci. 2020, 16, 2675–2691. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Wu, M.; Lu, S.; Zhang, H.; Shen, Y.; Shen, C.; Liang, H.; Ge, H.; Ding, X.; Wang, Z. Fgf9 Negatively Regulates Bone Mass by Inhibiting Osteogenesis and Promoting Osteoclastogenesis Via MAPK and PI3K/AKT Signaling. J. Bone Miner. Res. 2021, 36, 779–791. [Google Scholar] [CrossRef] [PubMed]
- Coryell, P.R.; Diekman, B.O.; Loeser, R.F. Mechanisms and Therapeutic Implications of Cellular Senescence in Osteoarthritis. Nat. Rev. Rheumatol. 2021, 17, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Bruey, J.M.; Bruey-Sedano, N.; Luciano, F.; Zhai, D.; Balpai, R.; Xu, C.; Kress, C.L.; Bailly-Maitre, B.; Li, X.; Osterman, A.; et al. Bcl-2 and Bcl-XL Regulate Proinflammatory Caspase-1 Activation by Interaction with NALP1. Cell 2007, 129, 45–56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buttgereit, F.; Dejaco, C.; Matteson, E.L.; Dasgupta, B. Polymyalgia Rheumatica and Giant Cell Arteritis: A Systematic Review. JAMA 2016, 315, 2442–2458. [Google Scholar] [CrossRef]
- Erdogan, S.; Aslantas, O.; Celik, S.; Atik, E. The Effects of Increased CAMP Content on Inflammation, Oxidative Stress and PDE4 Transcripts during Brucella Melitensis Infection. Res. Vet. Sci. 2008, 84, 18–25. [Google Scholar] [CrossRef]
- Chen, C.C.; Stairs, D.B.; Boxer, R.B.; Belka, G.K.; Horseman, N.D.; Alvarez, J.V.; Chodosh, L.A. Autocrine Prolactin Induced by the Pten-Akt Pathway Is Required for Lactation Initiation and Provides a Direct Link between the Akt and Stat5 Pathways. Genes Dev. 2012, 26, 2154–2168. [Google Scholar] [CrossRef] [Green Version]
- Khan, M.Z.; Khan, A.; Xiao, J.; Ma, Y.; Ma, J.; Gao, J.; Cao, Z. Role of the JAK-STAT Pathway in Bovine Mastitis and Milk Production. Animals 2020, 10, 2107. [Google Scholar] [CrossRef] [PubMed]
- de Dios, N.; Orrillo, S.; Irizarri, M.; Theas, M.S.; Boutillon, F.; Candolfi, M.; Seilicovich, A.; Goffin, V.; Pisera, D.; Ferraris, J. JAK2/STAT5 Pathway Mediates Prolactin-Induced Apoptosis of Lactotropes. Neuroendocrinology 2019, 108, 84–97. [Google Scholar] [CrossRef]
- Mor, A.; Philips, M.R. Compartmentalized Ras/MAPK Signaling. Annu. Rev. Immunol. 2006, 24, 771–800. [Google Scholar] [CrossRef]
- Simanshu, D.K.; Nissley, D.V.; McCormick, F. RAS Proteins and Their Regulators in Human Disease. Cell 2017, 170, 17–33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newbern, J.; Birchmeier, C. Nrg1/ErbB Signaling Networks in Schwann Cell Development and Myelination. Semin. Cell Dev. Biol. 2010, 21, 922–928. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karar, J.; Maity, A. PI3K/AKT/MTOR Pathway in Angiogenesis. Front. Mol. Neurosci. 2011, 4, 51. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hughes, G.C. Progesterone and Autoimmune Disease. Autoimmun Rev. 2012, 11, A502–A514. [Google Scholar] [CrossRef] [Green Version]
- Cutini, P.H.; Massheimer, V.L. In Vitro Effects of Progesterone and the Synthetic Progestin Medroxyprogesterone Acetate on Vascular Remodeling. Mol. Cell Endocrinol. 2019, 498, 110543. [Google Scholar] [CrossRef]
- Shu, D.; Zhu, Y.; Lu, M.; He, A.-D.; Chen, J.-B.; Ye, D.-S.; Liu, Y.; Zeng, X.-B.; Ma, R.; Ming, Z.-Y. Sanguinarine Attenuates Collagen-Induced Platelet Activation and Thrombus Formation. Biomedicines 2021, 9, 444. [Google Scholar] [CrossRef]
- Basini, G.; Santini, S.E.; Bussolati, S.; Grasselli, F. Sanguinarine Inhibits VEGF-Induced Akt Phosphorylation. Ann. N. Y. Acad. Sci. 2007, 1095, 371–376. [Google Scholar] [CrossRef]
- Zhang, J.; Mao, K.; Gu, Q.; Wu, X. The Antiangiogenic Effect of Sanguinarine Chloride on Experimental Choroidal Neovacularization in Mice via Inhibiting Vascular Endothelial Growth Factor. Front Pharmacol. 2021, 12, 638215. [Google Scholar] [CrossRef]
- Elsaid, M.B.; Elnaggar, D.M.; Owis, A.I.; AbouZid, S.F.; Eldahmy, S. Production of Isoquinoline Alkaloids from the in Vitro Conserved Fumaria Parviflora and Their in Vitro Wound Healing Activity. Nat. Prod. Res. 2021, 1–8. [Google Scholar] [CrossRef]
- Li, P.; Wang, Y.-X.; Yang, G.; Zheng, Z.-C.; Yu, C. Sanguinarine Attenuates Neuropathic Pain in a Rat Model of Chronic Constriction Injury. Biomed. Res. Int. 2021, 2021, 3689829. [Google Scholar] [CrossRef]
- Zhu, Q.; Jiang, M.; Liu, Q.; Yan, S.; Feng, L.; Lan, Y.; Shan, G.; Xue, W.; Guo, R. Enhanced Healing Activity of Burn Wound Infection by a Dextran-HA Hydrogel Enriched with Sanguinarine. Biomater. Sci. 2018, 6, 2472–2486. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Sun, X.; Huang, K.; Shen, S.; Lin, X.; Xie, Z.; Wang, J.; Fan, S.; Ma, J.; Zhao, X. Sanguinarine Protects against Osteoarthritis by Suppressing the Expression of Catabolic Proteases. Oncotarget 2017, 8, 62900–62913. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, F.; Xie, J.; Wang, G.; Zhang, G.; Yang, H. Anti-Osteoporosis Activity of Sanguinarine in Preosteoblast MC3T3-E1 Cells and an Ovariectomized Rat Model. J. Cell. Physiol. 2018, 233, 4626–4633. [Google Scholar] [CrossRef]
- Ardura, J.A.; Portal-Núñez, S.; Castelbón-Calvo, I.; Martínez de Toda, I.; De la Fuente, M.; Esbrit, P. Parathyroid Hormone-Related Protein Protects Osteoblastic Cells From Oxidative Stress by Activation of MKP1 Phosphatase. J. Cell. Physiol. 2017, 232, 785–796. [Google Scholar] [CrossRef] [PubMed]

| Compounds | Molecular Weight | Water-Oil Distribution Factor | Estimated SOLubility (ESOL) Class | Skin Penetration | Drug-Like Properties (Number of Yes/5) | Bioavailability |
|---|---|---|---|---|---|---|
| Sanguinarine (SAN) | 332.33 | 2.88 | Moderately soluble | −5.17 | 5/5 | 0.55 |
| Chelerythrine (CHE) | 348.37 | 3.02 | Moderately soluble | −5.17 | 5/5 | 0.55 |
| Allocryptopine (ALL) | 369.41 | 2.81 | Moderately soluble | −6.48 | 5/5 | 0.55 |
| Protopine (PRO) | 353.37 | 2.67 | Moderately soluble | −6.47 | 5/5 | 0.55 |
| Compounds | Degree | Betweenness |
|---|---|---|
| SAN | 22 | 16,525.8 |
| CHE | 89 | 15,077.4 |
| ALL | 101 | 2489.3 |
| PRO | 41 | 1679.5 |
| UniProt Entry | Gene Symbol | Protein Name | Degree | Betweenness |
|---|---|---|---|---|
| P12931 | SRC | SRC proto-oncogene, non-receptor tyrosine kinase | 49 | 712.8 |
| P27361 | MAPK3 | mitogen-activated protein kinase 3 | 47 | 620.4 |
| P42345 | MTOR | mechanistic target of rapamycin kinase | 44 | 478.5 |
| P03372 | ESR1 | Estrogen receptor | 39 | 459.0 |
| P42336 | PIK3CA | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform | 37 | 203.3 |
| Q07817 | BCL2L1 | Bcl-2-like protein 1 | 36 | 266.0 |
| P28482 | MAPK1 | Mitogen-activated protein kinase 1 | 33 | 169.5 |
| P49841 | GSK3B | Glycogen synthase kinase-3 beta | 33 | 438.6 |
| O60674 | JAK2 | Tyrosine-protein kinase JAK2 | 33 | 154.9 |
| P10275 | AR | Androgen receptor | 32 | 271.3 |
| Q96EB6 | SIRT1 | NAD-dependent protein deacetylase sirtuin-1 | 32 | 419.3 |
| Combination of Energy (kcal/mol) | Number of Hydrogen Bonds | |
|---|---|---|
| SAN-MAPK3 | −6.98 | 3 |
| CHE-MAPK3 | −7.50 | 3 |
| ALL-MAPK3 | −8.62 | 2 |
| PRO-JAK2 | −6.58 | 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, Z.; Liu, M.; Zou, X.; Sun, W.; Liu, X.; Zeng, J.; Yang, Z. Integrating Network Pharmacology and Molecular Docking to Analyse the Potential Mechanism of action of Macleaya cordata (Willd.) R. Br. in the Treatment of Bovine Hoof Disease. Vet. Sci. 2022, 9, 11. https://doi.org/10.3390/vetsci9010011
Dong Z, Liu M, Zou X, Sun W, Liu X, Zeng J, Yang Z. Integrating Network Pharmacology and Molecular Docking to Analyse the Potential Mechanism of action of Macleaya cordata (Willd.) R. Br. in the Treatment of Bovine Hoof Disease. Veterinary Sciences. 2022; 9(1):11. https://doi.org/10.3390/vetsci9010011
Chicago/Turabian StyleDong, Zhen, Mengting Liu, Xianglin Zou, Wenqing Sun, Xiubin Liu, Jianguo Zeng, and Zihui Yang. 2022. "Integrating Network Pharmacology and Molecular Docking to Analyse the Potential Mechanism of action of Macleaya cordata (Willd.) R. Br. in the Treatment of Bovine Hoof Disease" Veterinary Sciences 9, no. 1: 11. https://doi.org/10.3390/vetsci9010011
APA StyleDong, Z., Liu, M., Zou, X., Sun, W., Liu, X., Zeng, J., & Yang, Z. (2022). Integrating Network Pharmacology and Molecular Docking to Analyse the Potential Mechanism of action of Macleaya cordata (Willd.) R. Br. in the Treatment of Bovine Hoof Disease. Veterinary Sciences, 9(1), 11. https://doi.org/10.3390/vetsci9010011







