Seroprevalences of Newly Discovered Porcine Pestiviruses in German Pig Farms
Abstract
1. Introduction
2. Materials and Methods
2.1. Diagnostic Samples
2.2. Construction of the Plasmid pA/CP7_E1E2_APPV
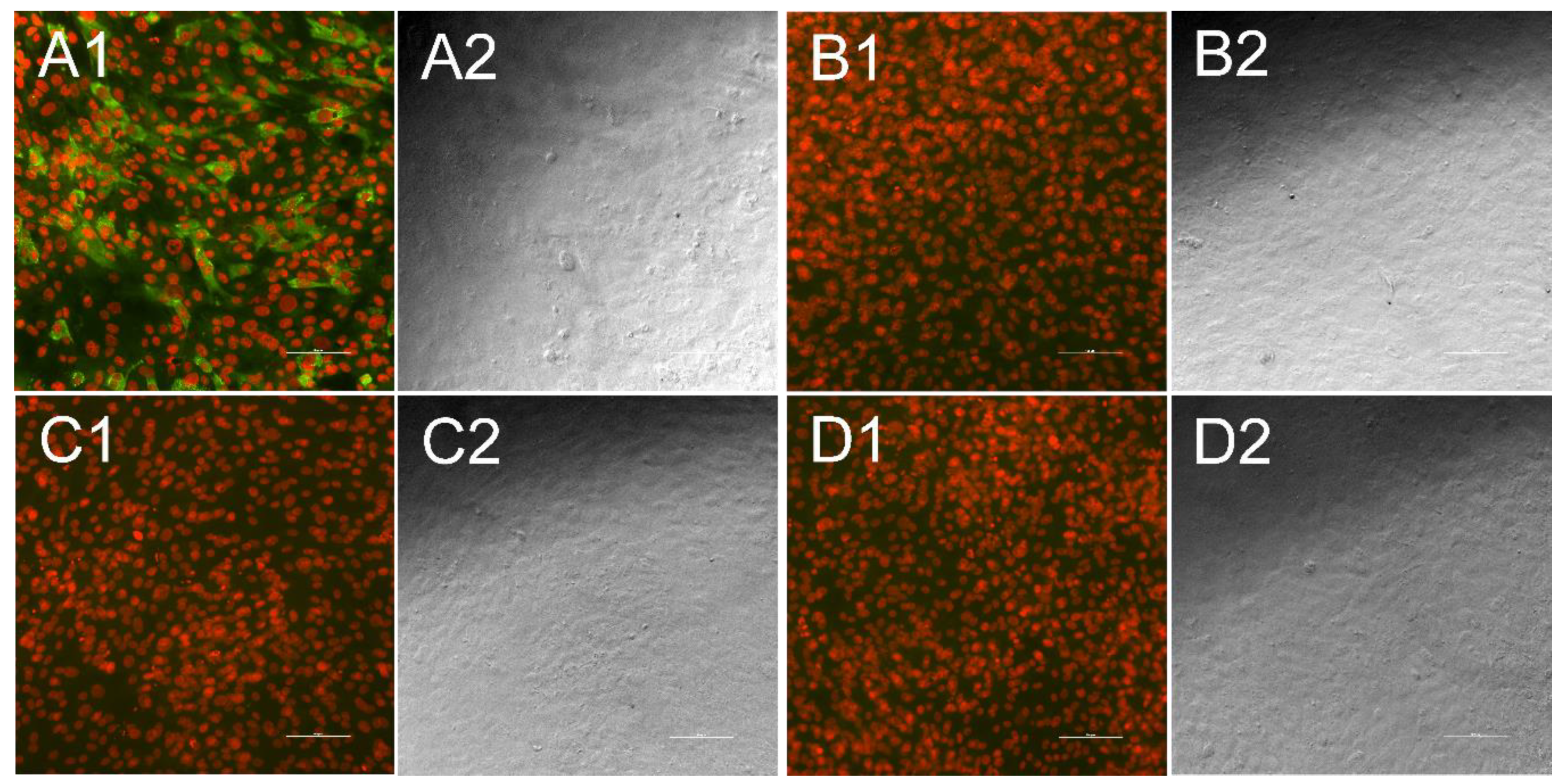
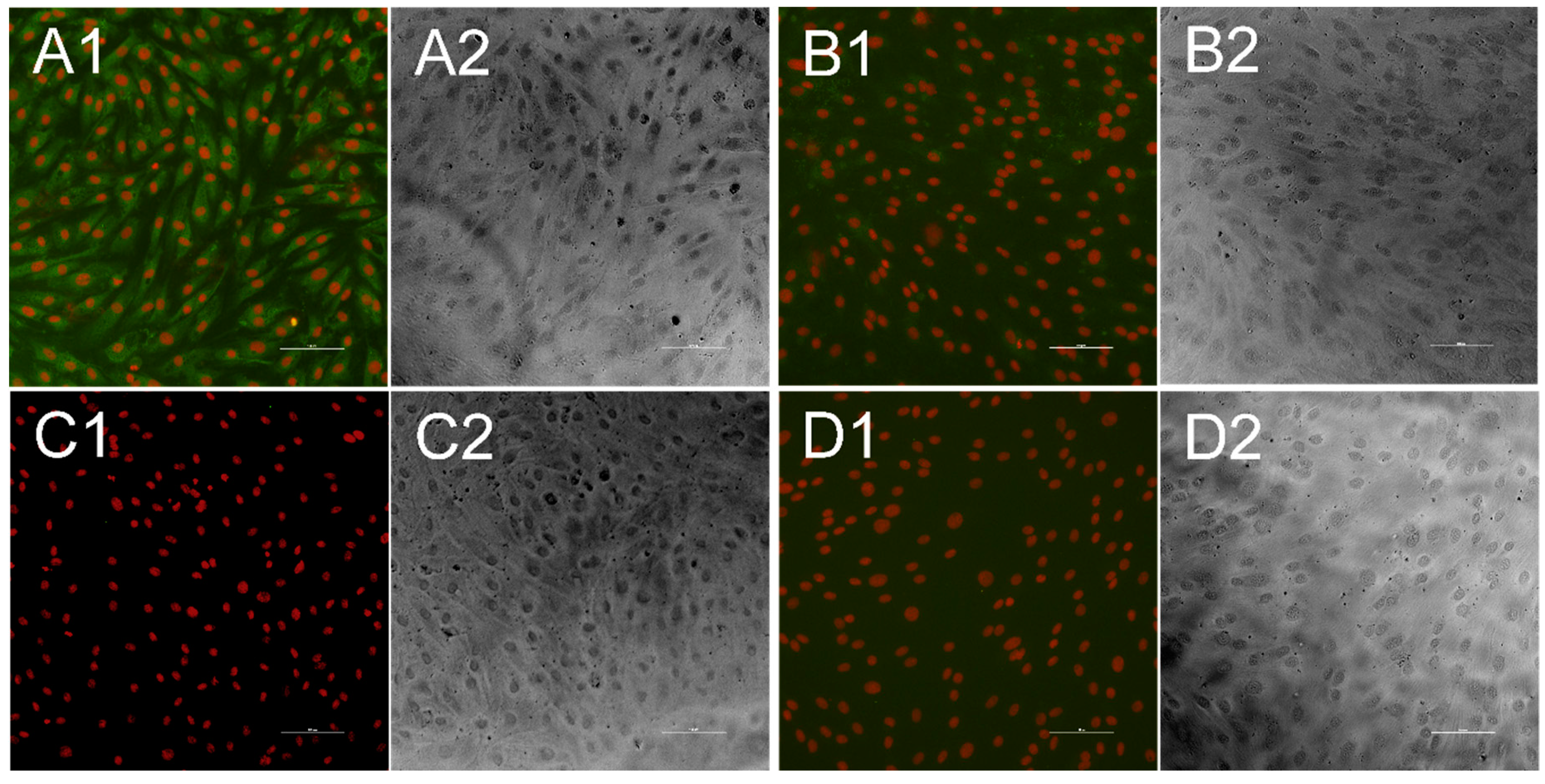
2.3. Indirect Immunofluorescence Test
2.4. RNA Extraction and RT-PCR
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Blome, S.; Beer, M.; Wernike, K. New leaves in the growing tree of pestiviruses. Adv. Virus Res. 2017, 99, 139–160. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.Q.; Lefkowitz, E.J.; Mushegian, A.R.; Adams, M.J.; Dutilh, B.E.; Gorbalenya, A.E.; Harrach, B.; Harrison, R.L.; Junglen, S.; Knowles, N.J.; et al. Changes to taxonomy and the international code of virus classification and nomenclature ratified by the international committee on taxonomy of viruses (2018). Arch. Virol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.B.; Meyers, G.; Bukh, J.; Gould, E.A.; Monath, T.; Scott Muerhoff, A.; Pletnev, A.; Rico-Hesse, R.; Stapleton, J.T.; Simmonds, P.; et al. Proposed revision to the taxonomy of the genus pestivirus, family Flaviviridae. J. Gen. Virol. 2017, 98, 2106–2112. [Google Scholar] [CrossRef] [PubMed]
- Becher, P.; Orlich, M.; Shannon, A.D.; Horner, G.; König, M.; Thiel, H.J. Phylogenetic analysis of pestiviruses from domestic and wild ruminants. J. Gen. Virol. 1997, 78, 1357–1366. [Google Scholar] [CrossRef]
- Vilcek, S.; Ridpath, J.F.; Van Campen, H.; Cavender, J.L.; Warg, J. Characterization of a novel pestivirus originating from a pronghorn antelope. Virus Res. 2005, 108, 187–193. [Google Scholar] [CrossRef]
- Wu, Z.; Ren, X.; Yang, L.; Hu, Y.; Yang, J.; He, G.; Zhang, J.; Dong, J.; Sun, L.; Du, J.; et al. Virome analysis for identification of novel mammalian viruses in bat species from Chinese provinces. J. Virol. 2012, 86, 10999–11012. [Google Scholar] [CrossRef]
- Firth, C.; Bhat, M.; Firth, M.A.; Williams, S.H.; Frye, M.J.; Simmonds, P.; Conte, J.M.; Ng, J.; Garcia, J.; Bhuva, N.P.; et al. Detection of zoonotic pathogens and characterization of novel viruses carried by commensal Rattus norvegicus in New York City. MBio 2014, 5, e01933-14. [Google Scholar] [CrossRef]
- Kirkland, P.D.; Frost, M.J.; Finlaison, D.S.; King, K.R.; Ridpath, J.F.; Gu, X. Identification of a novel virus in pigs—Bungowannah virus: A possible new species of pestivirus. Virus Res. 2007, 129, 26–34. [Google Scholar] [CrossRef]
- Finlaison, D.S.; King, K.R.; Frost, M.J.; Kirkland, P.D. Field and laboratory evidence that Bungowannah virus, a recently recognised pestivirus, is the causative agent of the porcine myocarditis syndrome (PMC). Vet. Microbiol. 2009, 136, 259–265. [Google Scholar] [CrossRef]
- Finlaison, D.S.; King, K.R.; Gabor, M.; Kirkland, P.D. An experimental study of Bungowannah virus infection in weaner aged pigs. Vet. Microbiol. 2012, 160, 245–250. [Google Scholar] [CrossRef]
- Abrahante, J.E.; Zhang, J.W.; Rossow, K.; Zimmerman, J.J.; Murtaugh, M.P. Surveillance of bungowannah pestivirus in the upper midwestern USA. Transbound. Emerg. Dis. 2012. [Google Scholar] [CrossRef] [PubMed]
- Hause, B.M.; Collin, E.A.; Peddireddi, L.; Yuan, F.; Chen, Z.; Hesse, R.A.; Gauger, P.C.; Clement, T.; Fang, Y.; Anderson, G. Discovery of a novel putative atypical porcine pestivirus in pigs in the USA. J. Gen. Virol. 2015, 96, 2994–2998. [Google Scholar] [CrossRef]
- Beer, M.; Wernike, K.; Dräger, C.; Höper, D.; Pohlmann, A.; Bergermann, C.; Schröder, C.; Klinkhammer, S.; Blome, S.; Hoffmann, B. High prevalence of highly variable atypical porcine pestiviruses found in Germany. Transbound. Emerg. Dis. 2017, 64, e22–e26. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Han, Z.; Li, J.; Huang, Y.; Yang, J.; Ding, H.; Zhang, J.; Zhu, M.; Zhang, Y.; Liao, J.; et al. Atypical porcine pestivirus as a novel type of pestivirus in Pigs in China. Front. Microbiol. 2017, 8, 862. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-González, S.; Canturri, A.; Peréz-Simó, M.; Bohórquez, J.A.; Rosell, R.; Cabezón, O.; Segalés, J.; Domingo, M.; Ganges, L. First report of the novel atypical porcine pestivirus in Spain and a retrospective study. Transbound. Emerg. Dis. 2017, 64, 1645–1649. [Google Scholar] [CrossRef] [PubMed]
- Gatto, I.R.H.; Harmon, K.; Bradner, L.; Silva, P.; Linhares, D.C.L.; Arruda, P.H.; de Oliveira, L.G.; Arruda, B.L. Detection of atypical porcine pestivirus in Brazil in the central nervous system of suckling piglets with congenital tremor. Transbound. Emerg. Dis. 2018. [Google Scholar] [CrossRef] [PubMed]
- Postel, A.; Meyer, D.; Cagatay, G.N.; Feliziani, F.; De Mia, G.M.; Fischer, N.; Grundhoff, A.; Milićević, V.; Deng, M.C.; Chang, C.Y.; et al. High abundance and genetic variability of atypical porcine pestivirus in pgs from Europe and Asia. Emerg. Infect. Dis. 2017, 23, 2104–2107. [Google Scholar] [CrossRef] [PubMed]
- Dénes, L.; Biksi, I.; Albert, M.; Szeredi, L.; Knapp, D.G.; Szilasi, A.; Bálint, A.; Balka, G. Detection and phylogenetic characterization of atypical porcine pestivirus strains in Hungary. Transbound. Emerg. Dis. 2018. [Google Scholar] [CrossRef]
- Dessureault, F.G.; Choinière, M.; Provost, C.; Gagnon, C.A. First report of atypical porcine pestivirus in piglets with congenital tremor in Canada. Can. Vet. J. 2018, 59, 429–432. [Google Scholar]
- de Groof, A.; Deijs, M.; Guelen, L.; van Grinsven, L.; van Os-Galdos, L.; Vogels, W.; Derks, C.; Cruijsen, T.; Geurts, V.; Vrijenhoek, M.; et al. Atypical porcine pestivirus: A possible cause of congenital tremor type A-II in newborn piglets. Viruses 2016, 8, 271. [Google Scholar] [CrossRef]
- Arruda, B.L.; Arruda, P.H.; Magstadt, D.R.; Schwartz, K.J.; Dohlman, T.; Schleining, J.A.; Patterson, A.R.; Visek, C.A.; Victoria, J.G. Identification of a divergent lineage porcine pestivirus in nursing piglets with congenital tremors and reproduction of disease following experimental inoculation. PLoS ONE 2016, 11, e0150104. [Google Scholar] [CrossRef] [PubMed]
- Lamp, B.; Schwarz, L.; Hogler, S.; Riedel, C.; Sinn, L.; Rebel-Bauder, B.; Weissenböck, H.; Ladinig, A.; Rümenapf, T. Novel pestivirus species in pigs, Austria, 2015. Emerg. Infect. Dis. 2017, 23, 1176–1179. [Google Scholar] [CrossRef] [PubMed]
- Postel, A.; Meyer, D.; Petrov, A.; Becher, P. Recent emergence of a novel porcine pestivirus: Interference with classical swine fever diagnosis? Emerg. Microbes Infect. 2017, 6, e19. [Google Scholar] [CrossRef] [PubMed]
- Meyers, G.; Tautz, N.; Becher, P.; Thiel, H.-J.; Kümmerer, B. Recovery of cytopathogenic and noncytopathogenic bovine viral diarrhea viruses from cDNA constructs. J. Virol. 1996, 70, 8606–8613. [Google Scholar]
- Richter, M.; Reimann, I.; Wegelt, A.; Kirkland, P.D.; Beer, M. Complementation studies with the novel “Bungowannah” virus provide new insights in the compatibility of pestivirus proteins. Virology 2011, 418, 113–122. [Google Scholar] [CrossRef]
- Postel, A.; Hansmann, F.; Baechlein, C.; Fischer, N.; Alawi, M.; Grundhoff, A.; Derking, S.; Tenhundfeld, J.; Pfankuche, V.M.; Herder, V.; et al. Presence of atypical porcine pestivirus (APPV) genomes in newborn piglets correlates with congenital tremor. Sci. Rep. 2016, 6, 27735. [Google Scholar] [CrossRef]
- Dalmann, A.; Reimann, I.; Wernike, K.; Beer, M. Autonomously replicating RNAs of Bungowannah pestivirus: ERNS is not essential for the generation of infectious particles. submitted.
- Reimann, I.; Depner, K.; Trapp, S.; Beer, M. An avirulent chimeric Pestivirus with altered cell tropism protects pigs against lethal infection with classical swine fever virus. Virology 2004, 322, 143–157. [Google Scholar] [CrossRef]
- Schwarz, L.; Riedel, C.; Högler, S.; Sinn, L.J.; Voglmayr, T.; Wöchtl, B.; Dinhopl, N.; Rebel-Bauder, B.; Weissenböck, H.; Ladinig, A.; et al. Congenital infection with atypical porcine pestivirus (APPV) is associated with disease and viral persistence. Vet. Res. 2017, 48, 1. [Google Scholar] [CrossRef]
- Cagatay, G.N.; Meyer, D.; Wendt, M.; Becher, P.; Postel, A. Characterization of the humoral immune response induced after infection with Atypical Porcine Pestivirus (APPV). Viruses 2019, 11, 880. [Google Scholar] [CrossRef]
- Cagatay, G.N.; Antos, A.; Meyer, D.; Maistrelli, C.; Keuling, O.; Becher, P.; Postel, A. Frequent infection of wild boar with atypical porcine pestivirus (APPV). Transbound. Emerg. Dis. 2018. [Google Scholar] [CrossRef]
- Kaufmann, C.; Stalder, H.; Sidler, X.; Renzullo, S.; Gurtner, C.; Grahofer, A.; Schweizer, M. Long-term circulation of atypical porcine pestivirus (APPV) within Switzerland. Viruses 2019, 11, 653. [Google Scholar] [CrossRef] [PubMed]
- Gatto, I.R.H.; Sonálio, K.; de Oliveira, L.G. Atypical porcine pestivirus (APPV) as a new species of pestivirus in pig production. Front. Vet. Sci. 2019, 6. [Google Scholar] [CrossRef] [PubMed]


| Federal State | Level | 2009/10 | 2018 | |||||
|---|---|---|---|---|---|---|---|---|
| Number of Positive Samples/Total Number | % pos. | Number of Positive Samples/Total Number | % pos. | Number of Positive Samples/total Number | % pos. | |||
| Mecklenburg-Western Pomerania | farms | 7/10 | 70.0% | 4/9 | 44.4% | Σ | 11/19 | 57.9% |
| individual | 24/99 | 24.2% | 24/90 | 26.7% | 48/189 | 25.4% | ||
| Lower Saxony | farms | 4/13 | 30.8% | 4/5 | 80.0% | Σ | 8/18 | 44.4% |
| individual | 9/199 | 4.5% | 16/59 | 27.1% | 25/258 | 9.7% | ||
| Saxony-Anhalt | farms | 7/9 | 77.8% | 3/12 | 25.0% | Σ | 10/21 | 47.6% |
| individual | 34/145 | 23.4% | 4/55 | 7.3% | 38/200 | 19.0% | ||
| Brandenburg | farms | 4/14 | 28.6% | 3/13 | 23.1% | Σ | 7/27 | 25.9% |
| individual | 9/42 | 21.4% | 9/105 | 8.6% | 18/147 | 12.2% | ||
| Baden-Wuerttemberg | farms | 6/14 | 42.9% | 2/7 | 28.6% | Σ | 8/21 | 38.1% |
| individual | 16/115 | 13.9% | 16/67 | 23.9% | 32/182 | 17.6% | ||
| Bavaria | farms | 5/11 | 45.5% | Σ | 5/11 | 45.5% | ||
| individual | 17/91 | 18.7% | 17/91 | 18.7% | ||||
| North Rhine-Westphalia | farms | 2/5 | 40.0% | Σ | 2/5 | 40.0% | ||
| individual | 4/48 | 8.3% | 4/48 | 8.3% | ||||
| Overall | farms | 28/60 | 46.7% | 23/62 | 37.1% | Σ | 51/122 | 41.8% |
| individual | 92/600 | 15.3% | 90/515 | 17.5% | 182/1115 | 16.3% | ||
| Federal State | Year | Number of Positive Samples/Total Number of Each Farm | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mecklenburg-Western Pomerania | 2009/10 | 9/10 | 5/10 | 3/10 | 3/10 | 1/10 | 1/10 | 0/10 | 0/10 | 0/10 | 2/9 | ||||
| 2018 | 10/10 | 8/10 | 3/10 | 0/10 | 3/10 | 0/10 | 0/10 | 0/10 | 0/10 | ||||||
| Lower Saxony | 2009/10 | 0/80 | 3/10 | 3/10 | 1/10 | 0/10 | 0/10 | 0/10 | 0/10 | 0/10 | 0/10 | 0/10 | 0/10 | 2/9 | |
| 2018 | 0/25 | 3/15 | 7/10 | 3/5 | 3/4 | ||||||||||
| Saxony-Anhalt | 2009/10 | 14/20 | 5/20 | 0/20 | 2/19 | 0/19 | 4/13 | 2/13 | 6/12 | 1/9 | |||||
| 2018 | 2/5 | 1/5 | 0/5 | 0/5 | 0/5 | 0/5 | 0/5 | 0/5 | 0/5 | 0/5 | 1/3 | 0/2 | |||
| Brandenburg | 2009/10 | 3/7 | 2/7 | 0/5 | 1/4 | 3/3 | 0/3 | 0/3 | 0/2 | 0/2 | 0/2 | 0/1 | 0/1 | 0/1 | 0/1 |
| 2018 | 3/12 | 0/12 | 4/11 | 0/12 | 0/11 | 0/11 | 0/10 | 2/8 | 0/5 | 0/5 | 0/5 | 0/2 | 0/1 | ||
| Baden Württemberg | 2009/10 | 3/10 | 1/10 | 0/10 | 6/9 | 2/9 | 1/9 | 0/9 | 3/8 | 0/8 | 0/8 | 0/7 | 0/7 | 0/6 | 0/5 |
| 2018 | 2/25 | 14/15 | 0/19 | 0/2 | 0/2 | 0/2 | 0/2 | ||||||||
| Bavaria | 2018 | 1/9 | 0/9 | 0/9 | 0/9 | 0/9 | 1/8 | 6/8 | 0/8 | 4/8 | 5/7 | 0/7 | |||
| North Rhine-Westphalia | 2018 | 3/14 | 0/14 | 1/10 | 0/9 | 0/1 | |||||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Michelitsch, A.; Dalmann, A.; Wernike, K.; Reimann, I.; Beer, M. Seroprevalences of Newly Discovered Porcine Pestiviruses in German Pig Farms. Vet. Sci. 2019, 6, 86. https://doi.org/10.3390/vetsci6040086
Michelitsch A, Dalmann A, Wernike K, Reimann I, Beer M. Seroprevalences of Newly Discovered Porcine Pestiviruses in German Pig Farms. Veterinary Sciences. 2019; 6(4):86. https://doi.org/10.3390/vetsci6040086
Chicago/Turabian StyleMichelitsch, Anna, Anja Dalmann, Kerstin Wernike, Ilona Reimann, and Martin Beer. 2019. "Seroprevalences of Newly Discovered Porcine Pestiviruses in German Pig Farms" Veterinary Sciences 6, no. 4: 86. https://doi.org/10.3390/vetsci6040086
APA StyleMichelitsch, A., Dalmann, A., Wernike, K., Reimann, I., & Beer, M. (2019). Seroprevalences of Newly Discovered Porcine Pestiviruses in German Pig Farms. Veterinary Sciences, 6(4), 86. https://doi.org/10.3390/vetsci6040086





