Abstract
The emergence of human immunodeficiency virus (HIV) causing acquired immunodeficiency syndrome (AIDS) in infected humans has resulted in a global pandemic that has killed millions. HIV-1 and HIV-2 belong to the lentivirus genus of the Retroviridae family. This genus also includes viruses that infect other vertebrate animals, among them caprine arthritis-encephalitis virus (CAEV) and Maedi-Visna virus (MVV), the prototypes of a heterogeneous group of viruses known as small ruminant lentiviruses (SRLVs), affecting both goat and sheep worldwide. Despite their long host-SRLV natural history, SRLVs were never found to be responsible for immunodeficiency in contrast to primate lentiviruses. SRLVs only replicate productively in monocytes/macrophages in infected animals but not in CD4+ T cells. The focus of this review is to examine and compare the biological and pathological properties of SRLVs as prototypic Tat-independent lentiviruses with HIV-1 as prototypic Tat-dependent lentiviruses. Results from this analysis will help to improve the understanding of why and how these two prototypic lentiviruses evolved in opposite directions in term of virulence and pathogenicity. Results may also help develop new strategies based on the attenuation of SRLVs to control the highly pathogenic HIV-1 in humans.
1. Background
HIV type 1 and 2, like small ruminant lentiviruses SRLVs, belong to the lentivirus genus of the Retroviridae family. They are small single-stranded enveloped RNA viruses characterized by reverse transcription of their RNA into double-stranded DNA for their replication. Additional members of this genus are the simian immunodeficiency viruses (SIVs) which infect various species of monkeys, bovine immunodeficiency virus (BIV) which infects cattle, feline immunodeficiency (FIV) which infects domestic cats and a variety of wild felids, equine infectious anemia virus (EIAV) which infects horses, and the small ruminant lentiviruses (SRLVs) with the prototypic caprine arthritis encephalitis virus (CAEV) and Maedi Visna Virus (MVV) which infect mainly goats and sheep, respectively. It has been well established now that HIV-1 and HIV-2 arose in humans following recent zoonosis of SIVcpz from chimpanzees (Pan troglodyte), SIVgor from gorillas, and SIVsmm from Sooty mangabey macaques, respectively [1,2,3]. The viruses have repeatedly crossed the species barrier, caused infections, and adapted into human cells to become highly replication-competent and transmissible from human to human, resulting in increased pathogenesis in their new host.
Since the discovery of HIV-1 by Montagnier and Barré-Sinoussi’s laboratory [4] over three decades ago, there have been nearly 80 million individuals infected and, prior to the development of efficient therapy, more than 35 million worldwide have progressed to AIDS and died from disease. Today there are over 35 million individuals living infected with HIV-1.
AIDS is characterized by low CD4+ T cell counts (below 200 cells/µL), hemogram abnormalities, chronic immune activation, and the occurrence of opportunistic infections. AIDS is often preceded by the co-receptor usage shift of the virus from CCR5 (macrophage-tropic) expressed predominantly at the surface of memory CD4+ T cells to CXCR4 (lymphocyte-tropic) at the surface of activated naïve CD4+ T cells, leading to severe depletion of these cells and impaired regeneration of the immune system [5]. The molecular mechanisms responsible for the CCR5 to CXCR4 switch remain still to be determined.
MVV causing disease was intensely studied in Icelandic sheep following the import of Karakul Asian breed sheep from Germany in 1933 to genetically enrich local breeds. The sheep were quarantined and distributed to different places in Iceland. Within a span of few years, Icelandic sheep started showing signs of diseases, mostly lung (maedi) and CNS (visna) diseases [6,7]. CAEV causing disease was identified in the 1970s when caprine flocks showed neurological symptoms in kids and synovial arthritis in adults. In cell culture, MVV shows markedly more cytopathogenicity in infected monolayer cells than CAEV. The major tropism of MVV and CAEV is for macrophages and dendritic cells but not CD4+ T cells in vivo [8,9]. Although HIV-1 also infects these cells, the virus replication occurs predominantly in CD4+ T lymphocytes in infected humans. Thus, SRLVs only cause productive infection, inflammation, and chronic degenerative diseases in 20–50% of naturally infected animals; this affects the central nervous system, lungs, udder, and joints. In contrast, HIV-1 infection in humans is associated with impairment of the immune system in nearly 100% of infected individuals. In the absence of therapy, infected patients undergo progressive destruction of CD4+ T lymphocytes, leading to AIDS and death due to opportunistic infections (Figure 1).
Here we will examine and compare the biological, pathological, and host/pathogen interactions in SRLVs of sheep and goats versus HIV-1/SIV of human and non-human primates.
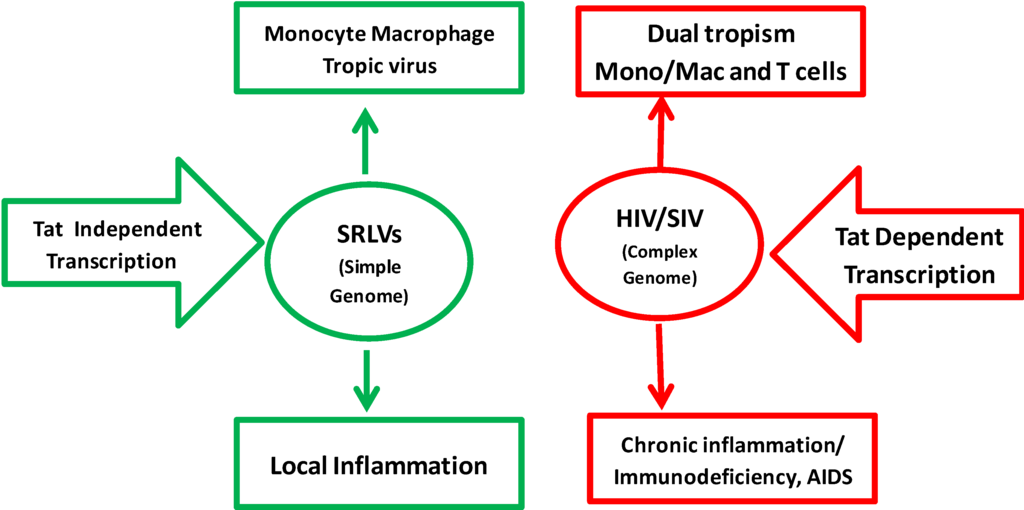
Figure 1.
Comparison of SRLV and HIV-1 properties. SRLV replication is restricted to monocyte/macrophage cell lineage. HIV-1 can target both monocytes/macrophages and CD4+ T cells using CD4 as the main receptor and either CCR5 or CXCR4 as the co-receptor. The replication of HIV/SIV is highly dependent on Tat transactivation, while replication of SRLVs is fully independent and constitutive. The pathogenesis of SRLVs is characterized by local inflammatory diseases in target organs while HIV-1 induces chronic systemic inflammation and immunodeficiency leading to AIDS and multi-organ degenerative diseases.
Figure 1.
Comparison of SRLV and HIV-1 properties. SRLV replication is restricted to monocyte/macrophage cell lineage. HIV-1 can target both monocytes/macrophages and CD4+ T cells using CD4 as the main receptor and either CCR5 or CXCR4 as the co-receptor. The replication of HIV/SIV is highly dependent on Tat transactivation, while replication of SRLVs is fully independent and constitutive. The pathogenesis of SRLVs is characterized by local inflammatory diseases in target organs while HIV-1 induces chronic systemic inflammation and immunodeficiency leading to AIDS and multi-organ degenerative diseases.
2. Genome Organization of SRLVs and HIV-1
The HIV-1 genome contains the classical gag, pol, and env genes found in all replication-competent retroviruses. The genome has also three regulatory genes, tat, rev and vif, whose products are necessary for efficient replication, and three auxiliary genes, nef, vpr and vpu, whose products are involved in virus/host interactions and pathogenesis. The HIV-1 genome with its nine major genes is more complex than SRLV genomes which, in addition to gag, pol and env, contain only two regulatory vif and rev genes and one auxiliary gene called “tat”. The latter from here on will be referred to as vpr-like gene [10].The low pathogenicity of SRLVs compared to HIV-1 can potentially be linked to the absence of proteins encoded by nef, vpu, and tat genes (Figure 2).
The HIV-1 gag gene produces a 55 kilodalton (kD) Gag precursor protein (Gag Pr-55) that is subsequently cleaved to generate the matrix p17, the capsid p24, the nucleocapsid p7 proteins, and the p6, which is critical for virus budding. The 160 kD Env glycoprotein precursor (gp160) is expressed from singly spliced viral mRNA and then is matured by cellular protease cleavage to generate the surface gp120 and transmembrane gp41 mature Env glycoproteins. The catalytic proteins are produced from a precursor protein encoded by the pol gene which, following cleavage, generates the protease (Pro p10), reverse transcriptase (RT p50), and integrase (IN p31). Proteases are known to play essential roles in many biological processes. They catalyze the hydrolysis of peptide bonds with high sequence selectivity and catalytic proficiency. HIV-1 protease, a member of the aspartic protease family, is a symmetrically assembled homodimer consisting of two identical subunits of 99 amino acids. Both subunits are involved in the catalytic activity (through an aspartic acid at codon 25). It is responsible for the cleavage of Gag-Pol, where Gag and Pol can be released during budding. In the absence of functional protease the viral assembly is not impaired, but the resulting particles are non-infectious [11,12]. RT has two enzymatic activities, a DNA polymerase that can copy either a DNA or a RNA template, and a RNase H activity that removes RNA from the RNA/DNA intermediate. Two different domains of RT are implicated in these two activities and they cooperate to convert the genomic RNA into a double-stranded linear DNA. The resulting double-stranded viral DNA is then transported into the nucleus and integrated in the host genome by IN [14].
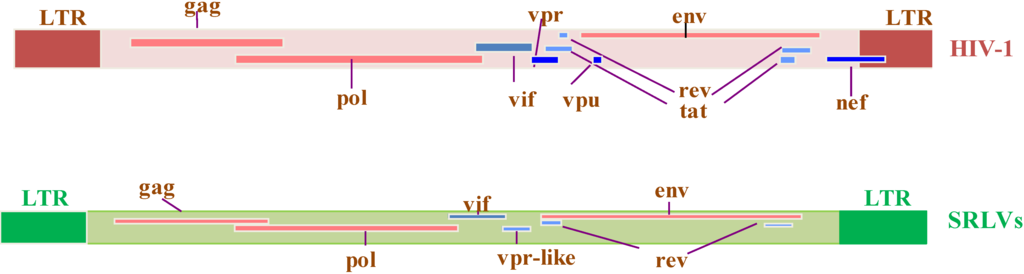
Figure 2.
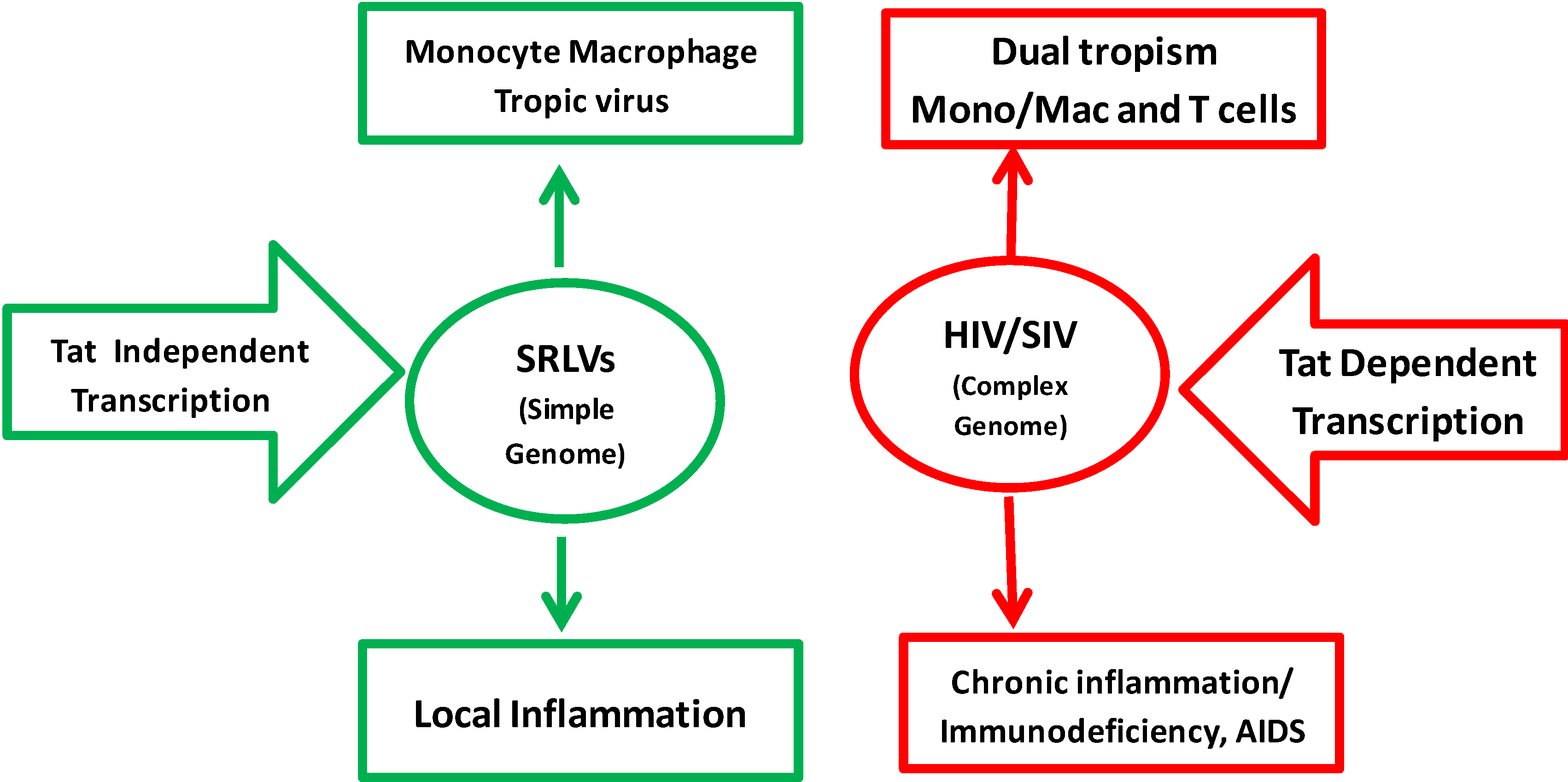
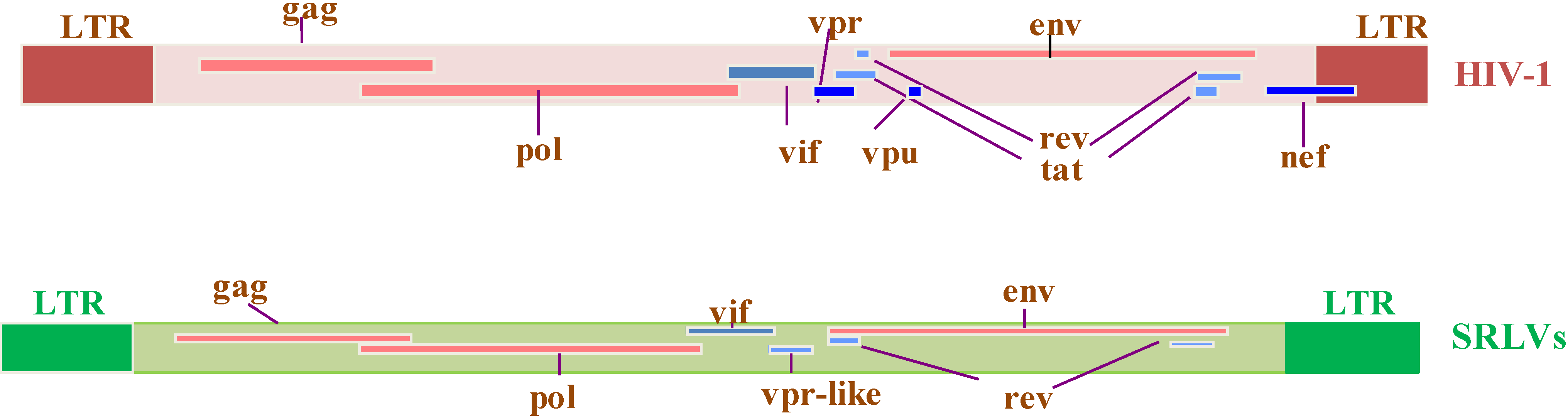
Genomic organization of HIV-1 and SRLV proviruses. Both genomes have the structural and enzyme gag, pol, and env genes (solid red bars), with LTRs at the 5’ and 3’ends (solid brown and green rectangles in HIV-1 and SRLVs, respectively) common to all retroviruses. The six additional open reading frames (vif, vpr, vpu, tat, rev, and nef) that encode regulatory and accessory proteins are indicated (solid blue bars), while the SRLV genomes have only three additional open reading frames (vif, vpr-like, and rev) [13].
Figure 2.
Genomic organization of HIV-1 and SRLV proviruses. Both genomes have the structural and enzyme gag, pol, and env genes (solid red bars), with LTRs at the 5’ and 3’ends (solid brown and green rectangles in HIV-1 and SRLVs, respectively) common to all retroviruses. The six additional open reading frames (vif, vpr, vpu, tat, rev, and nef) that encode regulatory and accessory proteins are indicated (solid blue bars), while the SRLV genomes have only three additional open reading frames (vif, vpr-like, and rev) [13].
2.1. Virus-Encoded Regulatory Proteins
2.1.1. Regulatory Tat
Tat is a key protein that regulates positively the expression of all genome encoded proteins in some of the lentiviruses. Detailed description of Tat and Tat functions are discussed in Section 2.3 and Section 2.4.
2.1.2. Regulatory Rev
Rev and Tat are expressed early during the initial phases of the replication cycle from multi-spliced RNA templates [15,16]. HIV-1 Rev is a 116-amino-acid phosphoprotein translated from fully spliced viral mRNA and encodes a protein that has four functional motifs. They include (i) a nuclear localization signal (NLS) directly interacting with importin-β for the import of Rev to the nucleus; (ii) a RNA-binding domain (RBD) that specifically directs the interaction of Rev with the Rev response elements (RRE) sequence; (iii) sequences that mediate self-interactions of Rev-Rev ending with a complex formation with the RRE; and (iv) a C-terminal leucine-rich nuclear export signal (NES) that binds to the chromosomal region maintenance 1 (CRM1) for the nuclear export of unspliced and unispliced RNA [17,18,19,20].
2.1.3. Regulatory Vif
Viral infectivity factor (Vif) is a 23 kD basic protein that is expressed in a Rev-dependent manner, largely localized in the cytoplasm of infected cells, and is essential for HIV-1 replication in lymphocytes and macrophages. Many studies using Δvif mutant genomes have demonstrated that these genotypes produce virions that are non-infections, yet there is no difference in the protein or RNA contents of Δvif compared to the wild type [21,22,23]. The main role of Vif is to suppress the host antiviral defense mechanism involving the DNA editing enzyme APOBEC3G. Studies have demonstrated that Vif neutralizes the potent intracellular defense pathway of APOBEC3G that protects host cells from a retrovirus. Several members of the APOBEC family act on single-stranded DNA or RNA and they alter the nucleotide sequence through cytidine deamination, converting cytidine to uridine and thus providing an intrinsic immunity to the host [24], which is an edit-dependent process [25,26]. APOBEC inhibits the retrovirus by several mechanisms which are edit-independent processes as well [27].
2.2. Accessory Virus Gene Encoded Proteins
2.2.1 Accessory Nef
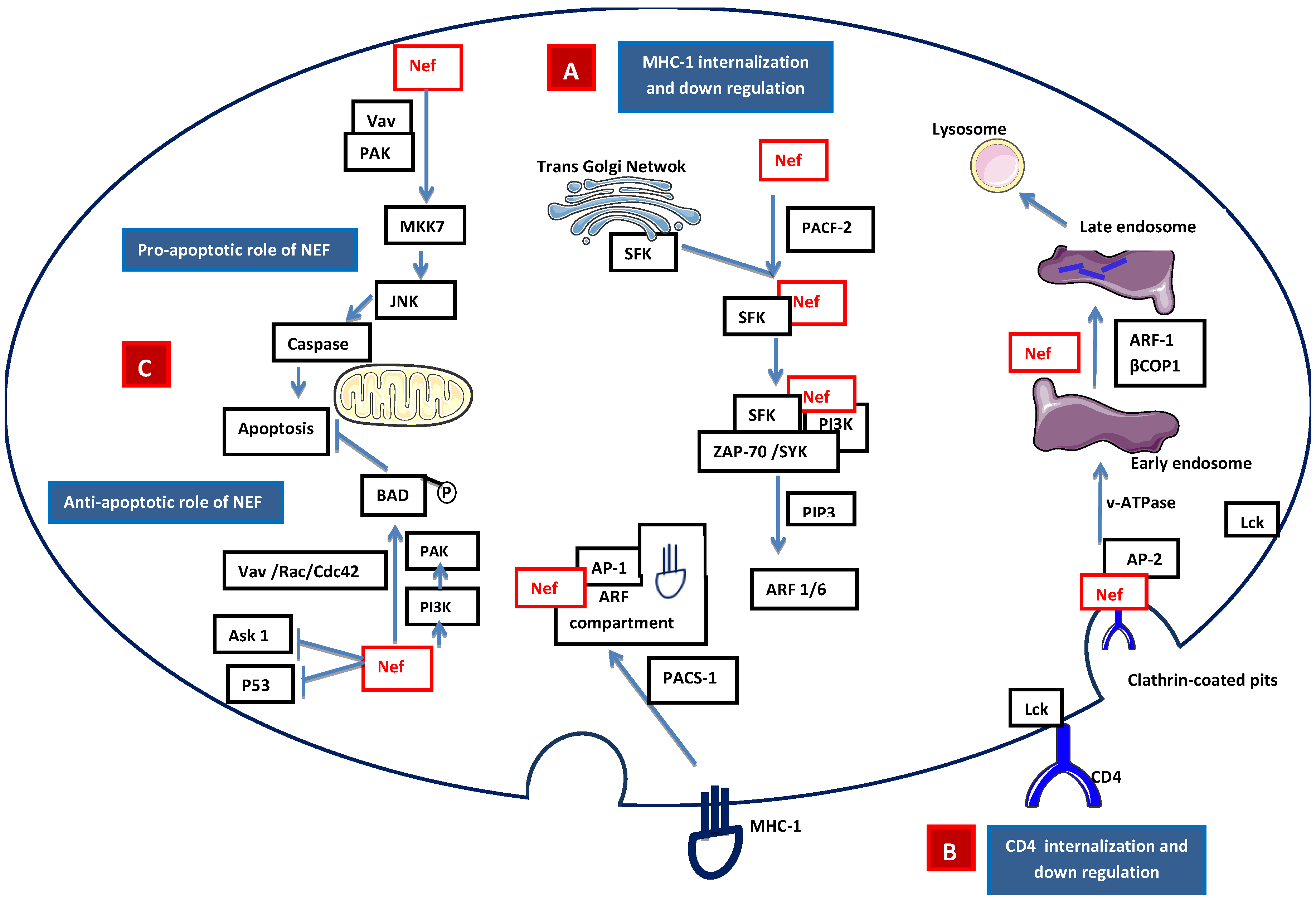
HIV-1 Nef is a 27 kD myristoylated protein that was initially thought to be a negative factor for virus replication and presents only in the intracellular compartment of infected cells. Later, Nef was found to have a positive effect on viral replication [28]. During HIV-1 infection, Nef down-regulates the surface expression of the CD4 and selective MHC class I receptors, thereby avoiding lytic HIV-1 superinfections and early elimination of HIV-1-infected cells by natural killer cells and cytotoxic T cells [29]. Nef induces internalization of MHC-I molecules that accumulate in the endosomal vesicles and are subsequently degraded. Nef also induces relocalization of internalized MHC-I molecules from the cell surface to the transgolgi network (TGN) [30]. MHC-I down-regulation may involve clathrin adapter protein complex 1 (AP-1) or Src family kinase-ZAP70/Syk-PI3K cascade recruited by phosphofurin acidic cluster sorting protein 1 (PACS2) [31,32,33] (Figure 3A). Nef interacts with several downstream cascades and complexes, including the adaptor complex of clathrin-coated pits and the beta subunits of COP-I coatomer, to down-regulate CD4 [34]. The CD4-p56lck complex is disrupted by Nef, allowing for the internalization of CD4 and the diversion of internalized CD4 to the lysosomal pathway which results in its destruction through the phosphatidylinositol 3-kinase (PI3K) pathway [35] (Figure 3B). Nef is also found to be in the extracellular compartment, leading to interference with hematopoiesis, and its excretion may be associated with exosomes [36,37]. Nef also has an anti-apoptotic effect that promotes efficient viral replication [38] and pathogenesis. Nef activates NAK (PAK2) through guanine nucleotide exchange factor Vav and the small GTPase Rac1 and Cdc42. Nef-mediated activation of PAK involves PI3K, which acts upstream of PAK, and the Nef-associated PI3-PAK complex phosphorylates the pro-apoptotic Bad protein, thus blocking apoptosis [39,40] (Figure 3C).
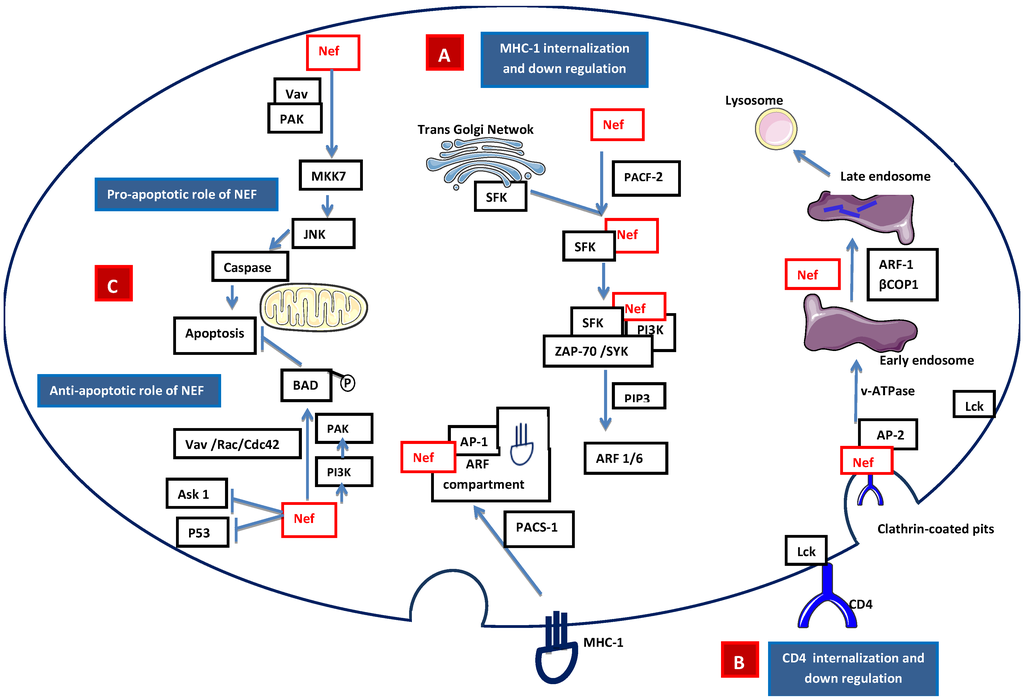
Figure 3.
Signaling model of Nef-mediated (A) apoptosis/anti-apoptosis, (B) down-regulation of expression of MHC class I molecules, and (C) down-regulation of the expression of CD4 molecules. A. Internalization of MHC-I: Nef accelerates the endocytosis of MHC class I molecules through PI3K-dependent activation of ADP ribosylation factor 6 (ARF6)-mediated endocytosis. First, the Nef binds PACS-2 and targets to the late Golgi/TGN; at the TGN, Nef binds and activates a TGN-localized SFK. The activated Nef–SFK complex, which leads to the signal transduction pathway recruiting and activating ZAP-70/Syk, then binds to a class I PI3K. The activated PI3K causes the accumulation of phosphatidylinositol-3,4,5-triphosphate (PtdInsP3) (PIP3) on the inner leaflet of the plasma membrane. PIP3 recruits the ARF6 guanosine exchange factor (ARF6-GEF) to the plasma membrane. Nef mediates the internalization of MHC-I from the plasma membrane to an ARF-6 endosomal compartment. The endocytosed MHC-I forms a complex with Nef and AP-1 via phosphofurin acidic cluster sorting protein 1 (PACS1). B. Internalization of CD4: Lck disassociates from the cytoplasmic tail of CD4 and Nef is attached to the cytoplasmic tail of CD4 within clathrin-coated pits through the interaction of adaptor protein 2 (AP-2) and vacuolar ATPase (v-ATPase). This leads to the internalization of CD4 into the early endosome. During the formation of the late endosome, Nef intacts with βCOP1 and through ARF-1, which targets CD4 to the lysosomal degradation. C. Apoptotic role of Nef: Intrinsic Nef signals localized on plasma membrane activate MAP kinase kinase 7 (MKK7) and c-Jun N-terminal kinase (JNK) to induce caspase-dependent apoptosis, while the anti-apoptotic pathway is mediated by Fas cell surface death receptor (FAS) and tumor-necrosis factor receptor (TNFR) by inhibiting the apoptosis signal-regulating kinase (ASK1) and P53. Nef-mediated p21-activated kinase (PAK) activation involves PI3K, which acts upstream of PAK. Nef-associated PI3K and PAK phosphorylate the pro-apoptotic Bad protein to inactivate Bad, thus blocking the apoptosis mediated by BCL-2.
Figure 3.
Signaling model of Nef-mediated (A) apoptosis/anti-apoptosis, (B) down-regulation of expression of MHC class I molecules, and (C) down-regulation of the expression of CD4 molecules. A. Internalization of MHC-I: Nef accelerates the endocytosis of MHC class I molecules through PI3K-dependent activation of ADP ribosylation factor 6 (ARF6)-mediated endocytosis. First, the Nef binds PACS-2 and targets to the late Golgi/TGN; at the TGN, Nef binds and activates a TGN-localized SFK. The activated Nef–SFK complex, which leads to the signal transduction pathway recruiting and activating ZAP-70/Syk, then binds to a class I PI3K. The activated PI3K causes the accumulation of phosphatidylinositol-3,4,5-triphosphate (PtdInsP3) (PIP3) on the inner leaflet of the plasma membrane. PIP3 recruits the ARF6 guanosine exchange factor (ARF6-GEF) to the plasma membrane. Nef mediates the internalization of MHC-I from the plasma membrane to an ARF-6 endosomal compartment. The endocytosed MHC-I forms a complex with Nef and AP-1 via phosphofurin acidic cluster sorting protein 1 (PACS1). B. Internalization of CD4: Lck disassociates from the cytoplasmic tail of CD4 and Nef is attached to the cytoplasmic tail of CD4 within clathrin-coated pits through the interaction of adaptor protein 2 (AP-2) and vacuolar ATPase (v-ATPase). This leads to the internalization of CD4 into the early endosome. During the formation of the late endosome, Nef intacts with βCOP1 and through ARF-1, which targets CD4 to the lysosomal degradation. C. Apoptotic role of Nef: Intrinsic Nef signals localized on plasma membrane activate MAP kinase kinase 7 (MKK7) and c-Jun N-terminal kinase (JNK) to induce caspase-dependent apoptosis, while the anti-apoptotic pathway is mediated by Fas cell surface death receptor (FAS) and tumor-necrosis factor receptor (TNFR) by inhibiting the apoptosis signal-regulating kinase (ASK1) and P53. Nef-mediated p21-activated kinase (PAK) activation involves PI3K, which acts upstream of PAK. Nef-associated PI3K and PAK phosphorylate the pro-apoptotic Bad protein to inactivate Bad, thus blocking the apoptosis mediated by BCL-2.
2.2.2. Accessory Vpr
Viral protein R (Vpr), a small basic protein (14 kDa) of 96 amino acids, is conserved in human (HIV-1 and HIV-2) and non-human (SIVs) lentiviruses. Vpr is mainly involved in the G2/M arrest of dividing cells [41,42,43], but it is also one of the virus protein members of the pre-integration complex that targets neo-synthesized double-stranded viral DNA to the nucleus, and it induces apoptosis and has some transactivation activity of HIV-1 long terminal repeats (LTR) [44]. The innate and cellular immunity of infected individuals is affected by the action of Vpr via the inhibition of IL-2 production and enhancement of glucocorticoid activity [45]. Vpr also triggers virus production by inducing TNF which activates HIV-1 expression and replication via the NFκB pathway. In the absence of Vpr, there is a delay in viral replication and disease progression as shown experimentally with SIVmac with mutated vpr [46,47].
Previously, several chimeric lentiviruses have been generated in our laboratory [48,49,50] by inserting the SIV nef and vpr/vpx genes separately or together in the infectious molecular genome of CAEV. All chimeras were found to be replication competent and showed increased cytopathicity in cell culture systems. The chimeric virus that has both nef and vpx/vpr genes was used along with the parental CAEV to inoculate newborn kids which were monitored for six months. The results showed that kids infected with this chimera have an increased persistence of viral replication in the blood mononuclear cells compared to kids infected with the wild-type CAEV. In addition, a persistent decrease in the proportion of circulating T cells was observed only in kids infected with the chimeric virus. Altogether, these data clearly provide the demonstration that the increased complexity of the CAEV viral genome is associated with the increased virulence of this virus.
2.3. HIV Tat Protein
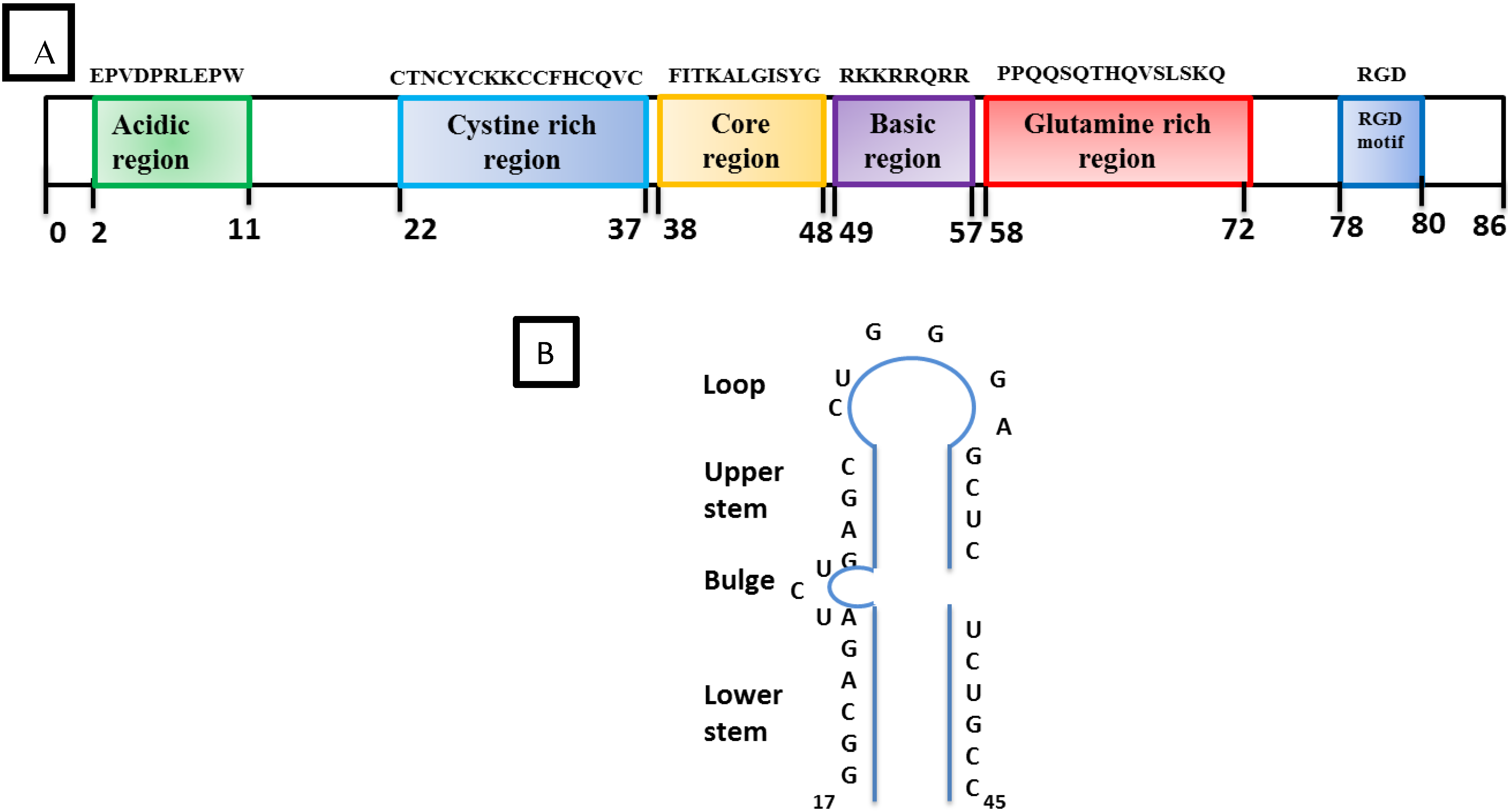
HIV-1 Tat is a 9–14 KD protein containing 86 amino acids encoded by two exons. It is one of the HIV-1 highly conserved proteins that is produced early in the HIV-1 infection. The first exon encodes the first 72 amino acids. Tat is subdivided into six functional domains (Figure 4A) including a proline and cysteine-rich N-terminal domain, a hydrophobic core, a basic region followed by a glutamine-rich region, and a C-terminal domain that contains a tripeptide RGD (Arg, Gly, Asp) [51,52,53,54,55]. In addition, there is a basic region called the protein transduction domain (PTD) which facilitates the trafficking of Tat through the plasma cell membrane. Thus, Tat can be found in the nuclear and cytosolic compartment of the cell as well as in the extracellular compartment. The primary function of Tat is to transactivate the promoter in the HIV-1 5’LTR for increased transcription of the proviral genome into full-length viral RNA [56].
2.4. Absence of SRLV Tat
Earlier studies suggested that SRLV genomes have tat-coding sequences upstream of the env gene. Furthermore, the encoded protein was found to be associated with high transactivation activity of the LTR promoter of SRLVs [57,58,59]. These data were not confirmed in the later studies and, in contrast, recent findings from our laboratory using more sophisticated and novel tools have proven that these proteins were associated with weak, if any, transactivation activities on their respective or heterologous LTRs [60]. In addition, the nucleotide sequence of the open reading frame called tat does not produce a protein structurally or functionally comparable to primate lentivirus Tat [61]. Historically, this open reading frame located upstream of the env coding sequences in the genome of SRLVs has been named tat because of its position rather than its structure or function; as such, it appears a misnomer. Recent studies from our laboratory clearly demonstrated that this protein was associated with activities ascribed to HIV-1 Vpr and, therefore, this SRLV open reading frame was renamed Vpr-like [60,61,62,63].
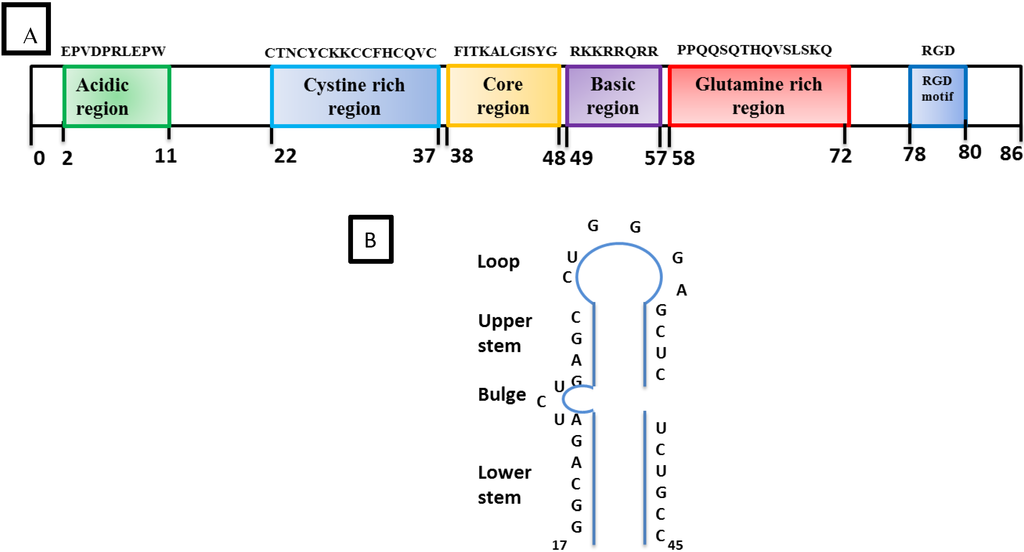
Figure 4.
Structural organization of Tat and TAR: (A) The boxes in different colors indicate the position of the six different domains in the amino acid sequence of HIV-1 Tat. The Basic region from 49–57 is required for RNA binding. (B) Schematic representation of the trans-activation response (TAR) stem-loop structure with its different regions and the bulge structure which is essential for the high-affinity binding of Tat. The minimal sequence element required for Tat-responsiveness is from residues 19–42.
Figure 4.
Structural organization of Tat and TAR: (A) The boxes in different colors indicate the position of the six different domains in the amino acid sequence of HIV-1 Tat. The Basic region from 49–57 is required for RNA binding. (B) Schematic representation of the trans-activation response (TAR) stem-loop structure with its different regions and the bulge structure which is essential for the high-affinity binding of Tat. The minimal sequence element required for Tat-responsiveness is from residues 19–42.
3. Natural History of HIV-1 and SRLV
3.1. Discovery of HIV-1
In 1981, AIDS was first recognized as a disease when homosexual men were reported with unusual opportunistic infections and malignancies. The principle causative agent of AIDS was a retrovirus now termed HIV-1, which was first discovered by Barré-Sinoussi et al. in the laboratory of Montagnier at the Pasteur Institute in Paris [4].
3.2. Cross-Species Infection from Monkeys to Human
HIV-1 is closely related to SIVs which have, on many occasions, jumped the species barrier from non-human primates to humans and spread successfully in the human population [64,65,66,67]. There are four phylogenetic lineage groups of HIV-1 (M, N, O, P) that were all thought to result from independent cross-species transmissions. It is now clearly established that cross-species transmission of the M and N groups of HIV-1 occurred from SIVcpz from naturally infected chimpanzees (Pan troglodytes) in South Cameroon, while a recent report indicated that O group originated following cross-species transmission from western lowland gorillas [68].
3.3. HIV Transmission in Humans
HIV-1 is generally transmitted through: (1) exchange of bodily fluids such as blood, semen, rectal fluids, vaginal fluids, breast colostrum, and milk from an infected person to recipient persons; (2) vertical transmission from mother to child either in utero, during delivery, or during post-natal breast feeding of colostrum or milk containing infected cells; (3) contaminated needle exchange during drug use or accidental iatrogenic exposure (reviewed in detail in [69]).
3.4. HIV Disease Progression
The rate of disease progression upon HIV-1 infection is variable and depends on host and viral factor interactions. The regular pattern of progression of HIV-1 infection has been found to be punctuated in vivo in three phases: 1) an acute primary infection associated with seroconversion and transient illness; 2) an asymptomatic phase during which the controlled virus continues to replicate, leading to a chronic immune activation; 3) a symptomatic phase during which the virus replicates to a higher titer. At this stage, the immune system is impaired and allows the proliferation of opportunistic infections [70].
3.5. First Descriptions of SRLV Infection
A disease that is now known to be associated with lentivirus infection in sheep was first described in South Africa by Mitchell in 1915 and termed as progressive pneumonia of sheep [71,72]. The next description was reported in sheep from Montana in 1923 with severe chronic interstitial pneumonia that resulted in wasting and, eventually, death of the affected animals [73]. In Iceland, about a decade later, “maedi”, a chronic form of pneumonia, emerged in a local breed of Icelandic sheep, progressing to a fatal form associated with important weight loss and shortness of breath [74]. There was also a neurological form of disease, “visna”, associated with paralysis and wasting that affected the Icelandic sheep during the next decade [72]. Epidemiological studies showed that the appearance of Maedi/Visna diseases began in a local breed of sheep after the import of asymptomatic Karakul rams from Germany in 1933. These lambs were imported for introducing genetic diversity because of the valuable Persian lamb skins of Karakul sheep for the Icelandic industry. An eradication program that lasted nearly a decade and in which all the sheep (over 100,000) in the affected areas were slaughtered was the only way to stop the fatal loss of the local breed [75]. During the following years, intensive studies were conducted for better understanding the Maedi and Visna diseases. Virological and pathological studies under Dr. Bjorn Sigurdsson helped to conclude that a lentivirus was the etiologic agent responsible for both Maedi and Visna [6,7]. This was described as a “slow/low” virus that induces disease in infected sheep only years post-infection. Since then, the virus causing this type of infection in sheep was widely called “Maedi Visna Virus” (MVV) or ovine progressive pneumonia (OPPV) or ovine lentivirus (OvLV).
The pathological changes that are known to be attributed to the goat lentivirus infection CAEV were reported in the late 1950s in adult goats in Switzerland [76] and about a decade later in Germany [77]. Later, Linda Cork in the early 1970s described in detail the anatomic-pathological changes associated with this disease [78]. The first isolation of the etiologic agent was simultaneously reported from the joints of an arthritic goat [79] and from the CNS of an encephalitic kid [80]. Later, the virus was isolated from goats all over the world. All lentivirus isolates from sheep and goats are referred to now as Small Ruminant Lentiviruses (SRLVs).
3.6. Cellular Tropism in Vivo and in Vitro
The main cell types that support SRLV replication in vivo are those of the monocyte/macrophage cell lineage [8]. Unlike primate lentiviruses, SRLVs do not replicate productively in CD4+ T lymphocytes [9]. The mechanisms underlying this restriction are still not well defined. In vivo in circulating monocytes, the viral DNA remains silent, not producing viral proteins and, thus, latently infected monocytes largely escape immune surveillance, providing for a Trojan horse mechanism [81]. Virus replication becomes productive only upon their differentiation into macrophages following their homing in tissues [82,83,84] in different organs [85]. SRLVs can infect cells other than monocytes/macrophages, including microglia, astrocytes, and oligodendrocytes [86,87]. In cell culture, SRLVs productively infect the goat synovial membrane cells, sheep choroid plexus cells [88,89], fibroblasts [90], and epithelial and endothelial cells [91]. Unlike primate lentiviruses, SRLVs do not use CD4, CCR5, and CXCR4 molecules as receptors/coreceptors for target cell infection. However, the receptor/coreceptor used by SRLVs are still not well identified.
HIV-1 infects predominantly the CD4+ T cells following attachment to the principal CD4 receptor and either the CCR5 or CXCR4 co-receptor. Following the entry of HIV-1 in the human body, there is a bottle neck selection of a founder virus that predominantly uses the CCR5 as a co-receptor [92,93]. This is primarily expressed on the surface of memory CD4+ T cells. The infection of such cells may lead either to productive HIV-1 replication and cell death via cytopathic effects or non-productive HIV-1 latently infected cells, depending on the activation dynamic of the host cells [94]. In addition to CD4+ T lymphocytes, HIV-1 has the ability to infect myeloid cell lineages such as dendritic cells and monocytes/macrophages [95] both in vitro in cultured cells and in vivo in infected humans [96]. In blood, HIV-1 proviral DNA is detectable in less than 1% of the circulating monocytes [97,98,99,100]. However, monocytes disseminate to all tissues of the organism where they rapidly differentiate into specialized tissue-specific macrophages [101] and persist as infected cells for a long period of time. In addition to monocytes/macrophages, epithelial and endothelial cells and fibroblasts are susceptible to in vitro infection with varying levels of viral replication [10,91,102,103]. The other cell types susceptible to HIV-1 infection are dendritic cells (myeloid, Langerhans, and plasmacytoid) that are antigen-presenting cells which can be targets of HIV-1 infection [104,105,106]. Dendritic cells were found to capture the virus through lectin-type receptors such as DC-specific ICAM-3-grabbing non-integrin (DC-SIGN) or sialic acid-binding Ig-like lectin 1 (SIGLEC-1), and act primarily as intermediates for virus transmission to T cells since HIV-1 replication in these lineages is limited [107,108,109].
3.7. Cross-Species Infection
Earlier experimental infections of sheep with the goat lentivirus (CAEV) and goats with ovine lentivirus (MVV) have provided the demonstration of the lack of species barrier for these viruses in sheep and goats [110]. Later, viruses genetically closer to CAEV were found in naturally infected sheep [111,112] and others genetically closer to MVV were found in naturally infected goats [113]. The experimental infection of mouflons [114] and calves [115] clearly demonstrated that the virus is capable of causing infection in a large variety of ruminants. The natural cross-species transmission of SRLVs in domestic and wild ruminants has been recently documented [116]; there is also evidence of mixed infections in field conditions with two closely related SRLVs. In addition, there have been descriptions of goats persistently infected with both CAEV and MVV and vice versa [117].
3.8. Natural SRLV Transmission
In adult sheep the virus is mainly transmitted by the respiratory route through the exchange of droplets when animals are housed in high density in closed facilities during the winter [118]. In newborn lambs and kids, the virus is transmitted mainly by consumption of virus-infected cells in the colostrum and milk from infected dams and female goats [119,120,121]. In early stages of small ruminant development, the gastrointestinal tract is porous, allowing infected cells to cross the epithelium barrier and invade lymph nodes and blood where they then cause infection of the peripheral blood mononuclear cells [122].
3.9. SRLV Viral Dynamics in Vivo
Unlike primate lentivirus infection, there are no defined phases of infection with SRLV [118,119,123]. Most infected animals remain lifelong seropositive following seroconversion and the virus is thought to cause lifelong persistence in the host monocytes where it can stay in a latent form. Following the differentiation of monocytes into macrophages in tissues, latent proviruses become transcriptionally active and productive of new infectious virus. The time between infection and seroconversion is greatly variable and can take more than two years [124]. Only a fraction of persistently seropositive animals remain productively infected and capable of transmitting the virus without showing any clinical symptoms of disease [79,118,125].
3.10. Acute Primary Infection, Disease Progression, and Characterization
The initial phase of HIV-1 infection in humans includes high activity of viral replication, virus genome mutagenesis, and cell death that lasts for two to eight weeks. During this stage, viral replication can produce up to 10 billion virions/day and the concentration of HIV-1 RNA in the plasma can exceed 107 copies/mL [126,127]. As mentioned earlier in section 3.6, the initial phase of infection of both SRLVs and HIV-1 targets monocyte/macrophage cell lineages but, unlike SRLVs, HIV-1 also infects the CCR5+ CD4+ T cells where it mainly replicates in the gut-associated lymphoid tissues (GALT) early post-infection [128]. This productive replication is associated with the spread of the circulating virus throughout the body to establish the main reservoirs/sanctuaries of viruses in the lymphoid tissues, the central nervous system, and the genital tract [129]. The clinical symptoms during this stage can include fever, pharyngitis, lymphadenopathy, cough, rashes, myalgia, diarrhea, headache, nausea [130,131,132].
Following two to six months post-infection with HIV-1 in humans, a balance is reached between the rate of viral replication and induced antiviral immune defenses, leading to a steady level of viremia evaluated by HIV-1 RNA in the plasma [133,134,135]. The initial drop of CD4+ T lymphocytes stabilizes around 350–800 cells/µL, and this phase can be maintained or show progressive decline for years. This was initially considered as the asymptomatic phase. The progression towards symptomatic stages and AIDS varies greatly between infected individuals and is linked to host genetic factors and the reactivation of latent infections of other pathogens or new co-infections, but there are other potential environmental factors [136,137,138,139]. Disease progression in the absence of therapy can evolve in three distinct patterns; the normal progressors with a progressive loss of CD4+ T cells over six to eight years before developing AIDS, the rapid progressors whose loss of CD4+ T cells occurs within less than two years, and the long-term non-progressors in whom the CD4 count remains stable and the viral load is less than 50 copies/mL [140,141]. Although there is a tiny proportion of individuals, the elite controllers who, in absence of antiviral treatment, progress very slowly or do not progress, the vast majority of infected individuals (>95%) develop progressively typical HIV-1-associated pathogenesis [142,143]. The CD4+ T cell count progressively declines during approximately the eight years post-infection. Clinical manifestations are those associated with CDC category B symptomatic conditions such as Herpes zoster, oral Hairy leukoplakia, Listeriosis, Idiopathic thrombocytopenic purpura, Peripheral neuropathy, oropharyngeal Candidiasis, Bacillary angiomatosis, Kaposi sarcoma, HIV-associated idiopathic thrombocytopenic purpura, cervical intraepithelial neoplasia II-III, and lymphoid interstitial pneumonitis. With further disease progression and the decline of the CD4+ cell count below 200 cells/µL, the immune system further weakens and the individual shows susceptibility for CDC category C disease conditions such as Pneumocystis jiroveci pneumonia (pcp), mucocutaneous Herpes simplex, cryptosporidial/microsporidia diarrhea, esophageal candidiasis, extrapulmonary/military tuberculosis, HIV-associated wasting, and peripheral neuropathy [144,145,146]. The end-stage disease is generally associated with CD4 counts below 100 cells/µL and with systemic fungal diseases, symptomatic cytomegalovirus infection, and HIV encephalopathy, ending with the death of the infected patient. Such disease progression is, however, unlikely to occur in patients undergoing highly active antiretroviral therapy (HAART), which markedly controls viral replication to a barely detectable level and substantially prolongs the lifespan of infected patients. There is an increase of the CD4+ T cell count to around 800 cells/µL and there are no common opportunistic infections, but some patients develop degenerative diseases associated with persistent chronic inflammation in addition to the risk of virus rebound and the emergence of drug-resistant variants [114,147].
For SLRVs, after a long preclinical period lasting many years, animals may develop inflammatory symptoms characterized by arthritis and mastitis in adult goats and, in rare conditions, encephalitis in kids [80]. In some unique circumstances, CAEV-infected kids can also undergo CNS disease characterized by leukoencephalomyelitis in animals that are one to four months old [148]. This might result from the synergistic effects of coinfections of CAEV with other pathogens, inducing local inflammation leading to permeabilization of the blood barrier and infiltration of CAEV-infected monocytes in the CNS. Animals can also develop chronic progressive arthritis with lympho-plasmocytic synovitis predominantly in adults [149].
In sheep, the symptoms are mainly in the lungs as a result of interstitial pneumonia, in the udder (mastitis), and the CNS (encephalitis) [6,116]. The clinical symptoms described by Sigurdsson et al. in 1957 appear mainly in adults, including weight loss, shortness of breath with progressive respiratory distress, and the enlargement of lungs. The progression of the disease to Visna, on the other hand, causes massive demyelination in the central nervous system associated with neuronal death, including in the spinal cord, leading to progressive paralysis of infected animals [7,150].
As mentioned previously in section 3.6, SRLVs do not replicate productively in T lymphocytes and both in early and late phases of infection there is no measurable depletion of CD4+ T lymphocytes in the periphery; therefore, infected animals do not undergo immunodeficiency. Moreover, unlike the cross-species infection of SIVs that has originated pathogenic persistent HIV-1 in humans, the experimental cross-species infection of adult mouflons and newborn calves with CAEV resulted in the productive infection of animals but did not result in the increased pathogenicity of virus. In mouflons, CAEV caused productive persistent infection [114] while, in contrast, in calves, it resulted in the diffusion of CAEV in target organs and transient replication, though, after four months, there was a total spontaneous clearance of all evidence of CAEV infection [115]. This provided the first demonstration that productive lentivirus experimental infection can be naturally cleared by the host’s defenses. This last finding is promising for the development of new potential vaccine strategies to fight against lentivirus infections.
4. Clinical Pathogenesis
HIV-1 mucosal transmission may occur through several sites including rectal, vaginal, and/or penile and oral mucosa. Rectal transmission is considered to be the easiest mucosal route for viral acquisition [151], where HIV-1 and HIV-infected cells transgress the epithelial layer via small breaks to cause systemic diffusion of the virus following the infection of numerous target CD4+/CCR5+ T lymphocytes. This transmission is thought to be established by one founder virus tropic for CD4+/CCR5+ T cells. This founder has enhanced interaction with dendritic cells and is resistant to the IFN-γ host response [152]. Another mechanism highlighted for this route are M cells which produce mucus and form intraepithelial pockets that have CD4+ memory T cells and dendritic cells and promote the trans-epithelial transport of HIV-1 [153,154].
For vaginal transmission, extensive work is still ongoing to highlight the major target cells throughout the female reproductive tract (FRT). FRT is a strong barrier that effectively prevents the virus from transmission to germinal cells, so for virus entry it is likely that there is more than one site. The vagina and the ectocervix comprise multilayered epithelium, while the endocervix has only a single columnar epithelium layer [155]. In a recent rhesus macaque vaginal transmission model study, using a single round non-replicating SIV-based vector with dual marker genes, it was shown that the entire FRT including the vagina, ecto-, and endocervix, along with the ovaries and local draining lymph nodes, can contain vector transduced cells only 48 hours after inoculation. The results from this study clearly suggest that virions quickly disseminate after the multisite entry of cells throughout the FRT with a preference for infection in squamous vaginal and ectocervical mucous. This study for the first time showed that even if the infection occurs primarily in vaginal and ectocervical tissue, it can spread as far as the ovaries and local draining lymph nodes [156].
Thereafter, the brunt of the viral replication has been found to be in immune cells of the gastrointestinal tract [157,158], named the gut-associated lymphoid tissue (GALT). GALT represents the most important lymphoid organ harboring 40–80% of all T cells in the body [159,160]. Importantly, this site predominantly contains memory CD4+/CCR5+ T cells, expressing the major receptor and co-receptor for HIV-1 and SIV entry [161,162]. Infection in the GALT is extensive, reaching 60% of the mucosal memory CD4+ T cells within days post-infection and the infected cells are rapidly eliminated [163,164].
Intravenous transmission of the virus is faster as there is no selective barrier between virus and target cells, and the virus rapidly disseminates to all lymphoid tissues including thymus, spleen, peripheral lymphoid organs, and mucosal lymphoid tissues, with a peak of viremia at 10–14 days post infection. The productive infection is observed within two weeks in the paracortex of the lymphoid sheath in the spleen, and in the thymus medulla. Like in the mucosal transmission, the main targets of the infection are memory CD4+ T cells expressing the CCR5 receptor (reviewed in [165]).
As alluded above, progressive HIV-1 infection is associated with the progressive decrease of CD4+ T cells both in number and in function, but also, as recently shown, with continuous chronic inflammation, is initiated by a burst of pro-inflammatory cytokines (reviewed in [166]). This is followed by the chronic presence of several pro-inflammatory factors, such as IL-6, TNF, and others, leading to early immune senescence and remodeling of lymphoid tissue [167].
In the CNS, the late stages of HIV-1 infection can be complicated by AIDS dementia complex or HIV-associated Dementia (HAD), which is a neurological syndrome characterized by abnormalities in cognition, motor performance, and behavior [168]. Invasion of the CNS occurs both by cell-free and cell-associated virus very early post-infection [169,170,171]. This occurs following virus and virus-infected cells crossing the blood-brain barrier using complex and multiple strategies [172]. Viral replication in the CNS relies predominantly on macrophages and microglia, often leading to multinucleated cells that are due to virus-induced cell fusion. The dementia is partially if not totally due to the inflammatory effect of the virus in the brain [173]. There is absence of productive infection in neurons and oligodendrocytes, while astrocytes may under select circumstances appear infected with limited contribution to viral replication. In the era of HAART, AIDS dementia complex has been largely eliminated but is replaced by more subtle motor and cognitive impairments regrouped under the term HIV-1 associated neurocognitive disorder (HAND), which is hypothesized to be caused by the low-grade chronic inflammation and neurotoxicity of select areas of the brain. The issue is often complicated by the poor penetration of many anti-retroviral drugs across the blood-brain barrier [174].
In SRLVs, the multiple routes of infection might influence the outcome of pathogenesis. Although the main route of SRLV transmission is thought to be through ingesting contaminated milk and colostrum containing infected cells, studies have shown that various cells in the female as well as male genital tracts are susceptible to permissive infection [175]. The main clinical disease is characterized by inflammatory lesions including infiltrates of lymphocytes, plasma cells, and macrophages into various regions of the central nervous system, joints, lungs, and the mammary glands [176,177,178]. The inflammation is fueled by infected cells expressing viral antigens promoting the local activation of immune cells. This sustained activation/inflammation causes tissue damage by cytokine-mediated amplification of antiviral immune responses leading to the damage of uninfected cells (bystander effect) in the affected tissue. In the CNS, the lesions are limited to the white matter and appear as focal discolored areas, which are prominent in the spinal cord [78,148,178]. Microscopic changes are observed in the white matter with perivascular cuffing and the accumulation of mononuclear cells. In all infected kids undergoing CNS disease, the spinal cord, especially the segments in the cervical and lumbosacral parts, is involved. The CNS inflammatory disease also comprises meningoencephalitis, microgliosis, and astrocytosis in different parts of the CNS, including the spinal cord [78,179]. Lesions are generally not seen in spinal ganglia, nerve roots, and peripheral nerves. In the joints, microscopic lesions include proliferative synovitis of the joints, tendon sheaths, and bursa. As the disease progresses, fibrosis, necrosis, and mineralization of synovial membranes and the enlargement of carpi become more evident [149]. The goats gradually exhibit a poor hair coat and lose weight to become cachectic. The clinical signs include lameness and atrophy of the muscles of the affected limb. In the lungs, pulmonary lesions may vary, depending on the severity of the infection, ranging from mild congestion and interstitial pneumonia to patchy interstitial pneumonia in affected goats [78]. The clinical signs include rapid shallow respiration and increased weight of lungs; tracheobronchial and mediastinal lymph nodes are generally enlarged. In sheep, lung lesions can progress to very severe interstitial pneumonia with massive infiltrates of mononuclear cells in the interstitium causing obstruction of alveoli, which may be associated with smooth muscle hyperplasia and fibrosis of the lungs [180]. This results in the progressive reduction of the respiratory volume and may cause the death of animals [181]. In the mammary glands of infected does and ewes, extensive intralobular infiltration of mostly lymphocytes and marked lymphoid hyperplasia adjacent to lactiferous ducts have been noted [123,182].
5. Receptor/Co-Receptor Usage
As mentioned previously in section 3.6, HIV-1 interacts with the cell surface CD4 receptor and a chemokine co-receptor molecule, which triggers the structural changes in gp120, resulting in movement of the variable loop and exposure of the co-receptor binding site. When the co-receptor binds, further structural changes take place mainly in gp41 which lead to fusion of the viral and cellular membranes and internalization of the virus capsid into the cell cytoplasm. There have been seven transmembrane receptors identified in vitro as co-receptors for HIV/SIV that support the infection of CD4+ cells [183]. However, in vivo, CCR5 and CXCR4 are the major co-receptors supporting the infection [184], and HIV-1 uses either one or both co-receptors for entry.
Receptor usage by SRLVs has been studied but, to date, no receptor/co-receptor mediating entry has been clearly and reproducibly identified. Studies have suggested the mannose receptor (MR) as a potential SRLV receptor either alone or in concert with another receptor. However, cells lacking MR can be infected, suggesting that at least other receptors may be used by the virus [185,186]. In vivo, SRLVs have tropism for and replicate essentially in cells of the monocyte/macrophage lineage and dendritic cells in various tissues [90,187].
Compartmentalization of viral replication and evolution has been observed in both HIV-1 [188,189,190] and the SRLVs [117,191], likely due to site-specific differences in immune selection. Conserved sequences in the hypervariable V3 region of HIV-1 Env have been identified and suggested to be determinant for virus entry of macrophage tropic strains; this region also determines the replication efficiency and cell tropism [192]. In SRLVs, the Env hypervariable region V4 is structurally and functionally similar to the V3 region of HIV-1 [193,194]. The variation in the Env V4 regions gives rise to the subpopulation of viruses that colonize different organs [193]. The diverse compartments of CNS and genital tracts contain unique HIV-1 variants that are clearly distinct from the ones found in blood and lymphatic tissues [195].
6. HAART in HIV-Infected Patients
HAART consists of combinations of drugs like nucleoside/nucleotide reverse transcriptase inhibitors, non-nucleoside reverse transcriptase inhibitors, fusion and entry inhibitors, protease inhibitors, and integrase inhibitors. According to the last UNAIDS 2014 report, there are 35 million HIV-1-infected individuals worldwide and nearly half of them (13.6 million) now have access to antiretroviral therapy. These treatments have significantly increased the life span of HIV-1-infected individuals and virtually eliminated numerous terminal comorbidities such as Kaposi sarcoma and other opportunistic infections. While an important breakthrough in the treatment of HIV-1, these drugs fail to fully eradicate the virus infection from the body of infected, treated individuals since numbers of latently infected CD4+ T cells escape the treatment effects that are highly efficient only on cells productively replicating the provirus [196]. The discontinuation or interruption of HAART invariably results in the rebound of viremia from the latent reservoirs of HIV-1 [197,198]. Reservoirs of HIV-1 (estimated at 105–106 cells) are established within the first few days of infection [199]. The reservoir sites include immune-privileged organs [200] with limited antiviral immune responses (CNS and testes) or organs separated from blood via anatomical barriers that limit the access of antiretroviral therapy [201,202].
7. Latency and Persistence
Lentiviruses have developed multiple complex strategies to persist in the infected hosts, inducing progressive chronic diseases. Among these strategies, there is the post-integration latency of the provirus involving various heterogeneous cellular and viral mechanisms regulating the balance transcriptional active/latent proviruses that are still not fully understood [203,204,205,206]. HIV-1-associated latency has been the most studied, particularly in the context of the HIV-1 cure after HAART treatment [207,208]. After virus entry and reverse transcription, HIV-1 double-stranded DNA can either stay episomal or become irreversibly integrated preferentially in transcriptionally active sites of the host’s cell chromosomal DNA. However, a limited number of proviruses also integrate in transcriptionally inactive sites and remain transcriptionally silent in a state called viral latency. There are two main types of latency. The first is pre-integration latency in which viral DNA remains unintegrated located in the cytoplasm for a few days in the form of preintegration complex (PIC) and will eventually be degraded over time due to the cells' low metabolic rate. However, if the cell is activated before the decay of the PIC, it will then integrate and undergoes productive infection. The second type is post-integration latency whereby proviral DNA integrated into the host genome remains transcriptionally silent. Several factors and mechanisms have been associated with post-integration latency. The most prevalent mechanisms of latency currently recognized are [203,209]:
- Absence of Tat, or non-functional transactivation activity of Tat [210].
- Epigenetic regulations/chromatin remodeling by post-transcriptional modifications (hypoacetylation or trimethylation) [211].
- Insufficient levels or lack of nuclear host transcription factors (SP1, NF-kB) [212,213].
- Expression of transcription factor complexes with negative regulatory activities (YY1, CBF-1/RBP, APOBEC3G) [214,215].
- Influence of integration sites and provirus orientation on HIV-1 transcription efficiency [216].
- Unproductive control of viral RNA splicing, due to absence of Rev and innate host antiviral processes [217].
The viral regulatory protein Tat is thought to be one of the major factors in the regulation of HIV-1 latency. The absence of Tat results in a repressive chromatin environment that blocks the elongation of transcription [218,219].
In SRLVs, the viral DNA can remain in transcriptionally silent until the circulating blood monocytes mature into macrophages after their localization in tissues. This mechanism is independent of Tat, since SRLV genomes lack the gene encoding this protein [60]. In HIV-1, the proviruses in monocytes could be associated with a minimal amount of viral proteins without active expression of the proviral genome, thereby escaping detection and killing by the immune system; however, like SRLV-infected monocytes, upon entry in various tissues their rapid maturation into tissue macrophages is associated with increased expression and the release of fully infectious viral particles [8,220]. In SRLVs, viral persistence is fueled mainly by the lack of or low-grade viral replication in infected circulating monocytes that disseminate productive replication of the virus in various tissues. However, SRLV replication in macrophages can be also restricted by a number of posttranscriptional blocks [103,221,222].
8. Molecular Biology of HIV-1 Latency
8.1. Molecular Mechanisms of the Transcription Involving Tat in HIV/SIV
To date, there is no identified viral negative factor that represses HIV-1 transcription to undergo latency. Factors that were thought to be negative regulators in initial studies were not confirmed in more recent studies. The transcription of HIV-1 is tightly controlled by powerful feedback mechanisms of viral and cellular factors that act like molecular switches to regulate the expression of HIV-1 RNA transcription. HIV-1 transcription comprises two phases: first, an initiation step, and then an elongation step. The core promoter region in the U3 region of the LTRs includes several important motifs for the binding of factors required for the transcription, including a TATA box element, three SP-1 sites, and an initiator element (Figure 5A) [223,224,225,226]. Each of these core promoter elements contributes to the binding of the initiation complex TFIID. During the initiation phase, several factors such as the TATA box binding protein (TBP) and TBP-associated factors (TAFs), which interact with the TATA box and SP-1, are recruited by the promoter region to initiate the transcription complex [227]. A recent study shows the presence on an E-box motif (RBE1) within the core promoter that is implicated in transcriptional activation. RBE1 is a binding site for the RBF-2 transcription factor complex (USF1, USF2, and TFII-I), previously shown to bind an upstream viral element, RBE3. The results indicate that RBE1 is a bona fide RBF-2 binding site and that the RBE1 and RBE3 elements are necessary for mediating proper transcription from the HIV-1 LTR [228]. In another new study, it has been shown that CTGC motifs flanking the HIV-1 TATA box are required for Tat transactivation in living cells and the correct formation of pre-initiation complexes of HIV-1 (PICH); this complex contains the general transcription factor TFIIA that binds the HIV-1 core promoter formation in vitro. The binding of known core promoter binding proteins AP-4 and USF-1 was found to be dispensable for Tat function. The transcriptional response element (TAR) RNA prevented the stable binding of PICH-2. The impact of TAR on PICH-2 specifically required its bulge sequence, which is also known to interact with Tat [229]. A study by Jensen et al. using an in vitro model of a HIV Tat-mediated positive-feedback loop demonstrated that fluctuations in viral Tat-transactivating protein levels generate integration site-dependent, stochastically-driven phenotypes, in which infected cells randomly ‘switch’ between high and low expressing states in a manner that may be related to viral latency. Furthermore, in an extended model, they designed a forward genetic screen that systematically identifies the genetic elements in the HIV LTR promoter that modulate the fraction of genomic integration sites that specify ‘switching’ phenotypes. In this experiment, it has been shown that specific mutations in the core promoter regions, including Sp1 and TATA transcription factor binding sites, can increase the switching fraction several fold. Using the single-cell system experiments with computational modeling, they could further investigate the mechanism of switching-fraction enhancement for a selected Sp1 mutation. Altogether, these experiments demonstrated that mutations in the Sp1 site impaired not only Tat-induced transactivation of expression, but also the basal expression in the absence of Tat. Computational analysis demonstrated that the observed change in basal expression could contribute significantly to the observed increase in viral integrations that specify a switching phenotype, provided that the selected mutations affected Tat-mediated noise amplification differentially across genomic contexts [230].
Upstream of the core promoter region there is the regulatory region of HIV-1 LTR, which has binding sites for the cellular transcription factors nuclear factor of activated T-cells (NFAT), AP-1, p65/p50 nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kB) and Signal Transducer and Activator of Transcription 5 (STAT5), which orchestrate HIV-1 expression. The enhancer region of the regulatory promoter region consists of the NF-κB binding site, which has been studied extensively. Any mutation in one or more core promoter sites or regulatory promoter elements or enhancer regions leads to strong impairment of both basal and Tat-dependent transcription [231,232,233,234,235]. The key factor in the initial stage of the transcription of the HIV-1 LTR promoter by RNA polymerase is the phosphorylation of the carboxyl terminal domain (CTD) [236]. The CTD phosphorylation renders the RNA polymerase highly processive. The phosphorylated polymerase, once cleared from the promoter region, is then able to transcribe through the TAR region [237] (Figure 5B).
The initial phase in the elongation of HIV-1 transcription produces a low-level of viral gene expression which is a result of the hypo-phosphorylated condition of RNA pol II and activity of the negative elongation factor, 5,6-dichloro-1-beta-D-ribofuranosyl-benzimidazole sensitivity-inducing factor (DSIF), and negative elongation factor complex (NELF) [238,239,240]. This stage allows the initial production of small amounts of Tat, which plays an important role during the RNA Polymerase II elongation phase. Recent studies suggest that DSIF binds the elongation complex via association with the nascent transcript and subsequently recruits NELF [241,242]. In the case of the HIV-1 provirus, the recruitment of NELF may be further enhanced by the ability of NELF-E to bind directly to the TAR region of viral RNA [243] (Figure 5C). Unlike other transactivation factors, Tat does not bind to the DNA, but it interacts with the TAR element of the genomic viral RNA. TAR is a stem loop structure containing three nucleotide bulges (23–25 residues) and a loop of six nucleotides (30–35 residues) (Figure 4B). The bulge structure in TAR is the high affinity site for Tat binding [244]. During the elongation phase, which is a Tat dependent phase, Tat binds to TAR via its arginine-rich motif (ARM) and facilitates binding of the positive transcription elongation factor b (P-TEFb) [245,246], which is comprised of cyclin-dependent kinase 9 (Cdk9) and cyclin T1 [247,248,249]. The Bromo-domain protein 4 (Brd4) stimulates the kinase activity of P-TEFb for phosphorylation of the CTD of RNA polymerase II over basal levels. [250]. As a result, the HIV-1 transcription rate is enhanced several-hundred-fold, thereby producing a high level of viral gene transcripts [219,251] because P-TEFb phosphorylates subunits of the NELF and DSIF to release the RNA polymerase II activity preventing its pausing on the HIV-1 promoter [252,253] (Figure 5D,5E). For more details please refer to the following reviews on HIV-1 Tat transcription [206,209,254,255,256,257,258,259,260,261,262,263].
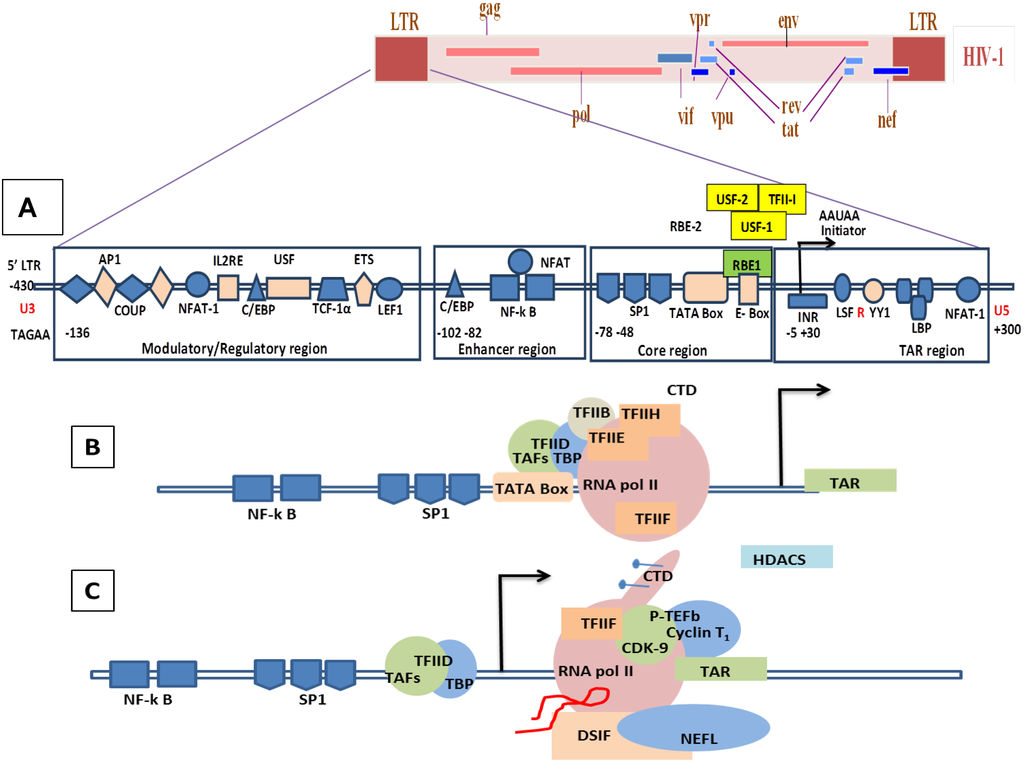
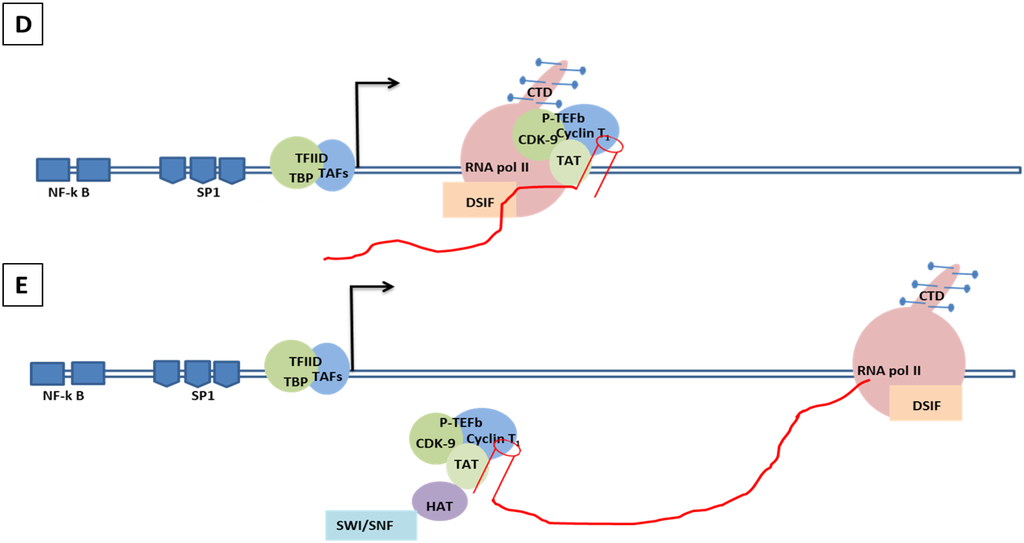
Figure 5.
Genomic organization of HIV-1 and mechanisms of LTR transactivation. (A) The modulatory region consists of transcription factors such as COUP, AP1, NFAT-1, IL2RE, C/EBP, USF, TCF-1α, ETS and LEF1. The enhancer region has the NF-κB and C/EBP, and the core region consists of three tandem SP1 sites and the TATA box and E-box to which RBE1 and RBE2 (USF1, USF2, TFII-I) binds. Downstream to the TATA box is the initiator element from the TAR region. A weak transcription starts at -430 at U3 with TAGAA and is enhanced at the downstream TAR region at the +1 AAUAA polyadenylation site. (Not all the transcription binding sites are shown here). (B) Initiation stage: the cellular transcription factors TAFs and TBP (TAFs + TBP + TFIID) are recruited which, along with the other components of the basal transcription, help in recruiting the RNA polymerase holoenzymes. The CDK7 kinase presents the CTD of the RNA polymerase in the TFIIH phosphorylates. (C) The hypo-phosphorylated state of the CTD correlates with the low processivity of the RNA polymerase enzyme complex. Due to hypo-phosphorylation, and in absence of Tat, there is mainly production of short RNA. Furthermore, DSIF and NELF that bind to the RNA pol II inhibit the transcriptional elongation. (D) CDK9 phosphorylates the CTD and, along with cyclin T1, they produce the complex P-TEFb (positive transcription elongation factor b). TAT is recruited and binds to the bulge sequence of the TAR RNA. The hyper-phosphorylation of CTD is induced by the binding of the P-TEFb to the TAR, resulting in dissociation of DSIF and NELF. HATS and SWI/SNF, which induces acetylation of the nucleosomes, are recruited and remove any blocks due to epigenetic modification and support elongation. The acetylation of Tat creates a binding site for P300/CREB binding protein-associated factors (PCAF) which enhances the transcription elongation.
Figure 5.
Genomic organization of HIV-1 and mechanisms of LTR transactivation. (A) The modulatory region consists of transcription factors such as COUP, AP1, NFAT-1, IL2RE, C/EBP, USF, TCF-1α, ETS and LEF1. The enhancer region has the NF-κB and C/EBP, and the core region consists of three tandem SP1 sites and the TATA box and E-box to which RBE1 and RBE2 (USF1, USF2, TFII-I) binds. Downstream to the TATA box is the initiator element from the TAR region. A weak transcription starts at -430 at U3 with TAGAA and is enhanced at the downstream TAR region at the +1 AAUAA polyadenylation site. (Not all the transcription binding sites are shown here). (B) Initiation stage: the cellular transcription factors TAFs and TBP (TAFs + TBP + TFIID) are recruited which, along with the other components of the basal transcription, help in recruiting the RNA polymerase holoenzymes. The CDK7 kinase presents the CTD of the RNA polymerase in the TFIIH phosphorylates. (C) The hypo-phosphorylated state of the CTD correlates with the low processivity of the RNA polymerase enzyme complex. Due to hypo-phosphorylation, and in absence of Tat, there is mainly production of short RNA. Furthermore, DSIF and NELF that bind to the RNA pol II inhibit the transcriptional elongation. (D) CDK9 phosphorylates the CTD and, along with cyclin T1, they produce the complex P-TEFb (positive transcription elongation factor b). TAT is recruited and binds to the bulge sequence of the TAR RNA. The hyper-phosphorylation of CTD is induced by the binding of the P-TEFb to the TAR, resulting in dissociation of DSIF and NELF. HATS and SWI/SNF, which induces acetylation of the nucleosomes, are recruited and remove any blocks due to epigenetic modification and support elongation. The acetylation of Tat creates a binding site for P300/CREB binding protein-associated factors (PCAF) which enhances the transcription elongation.
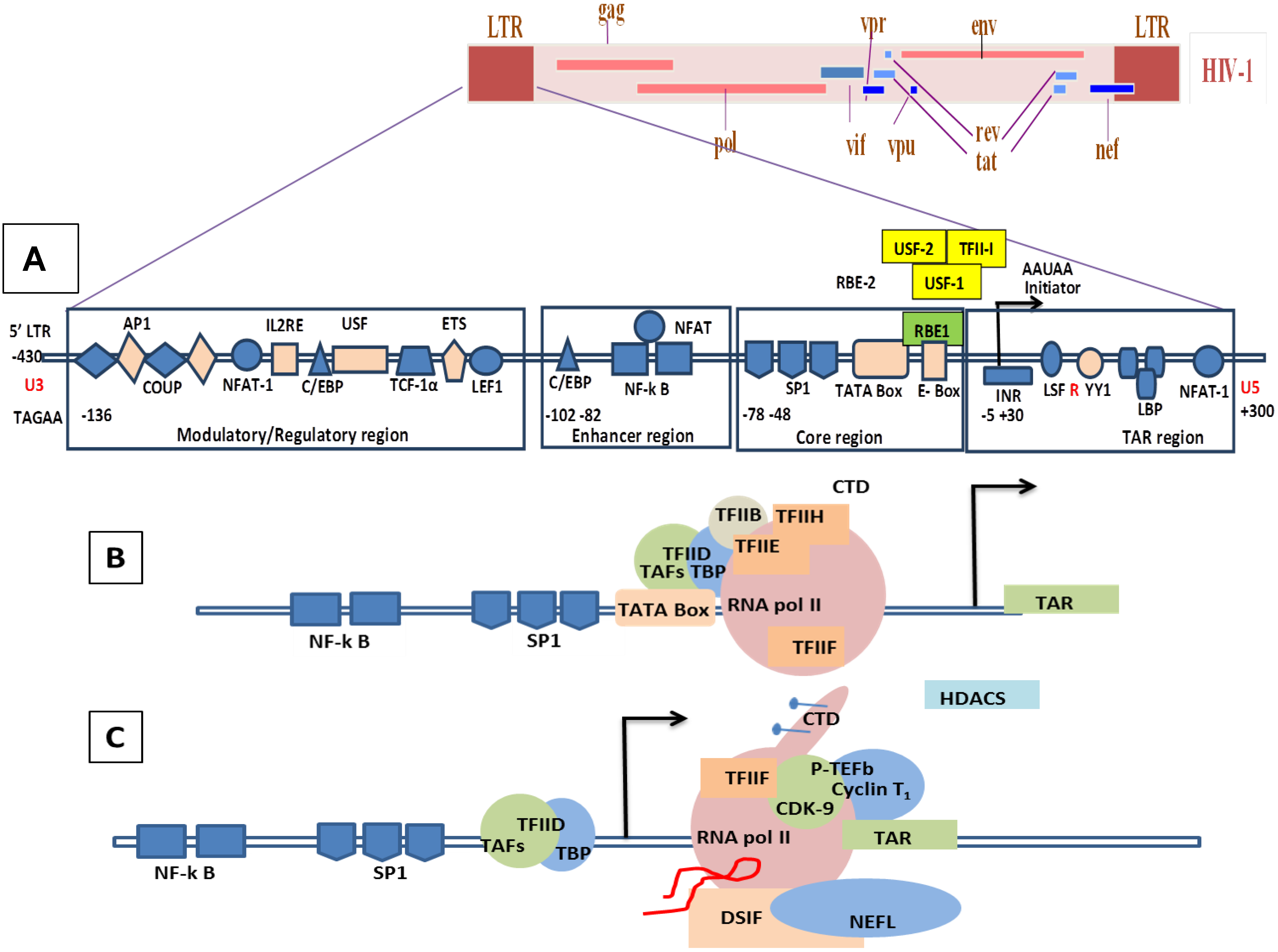
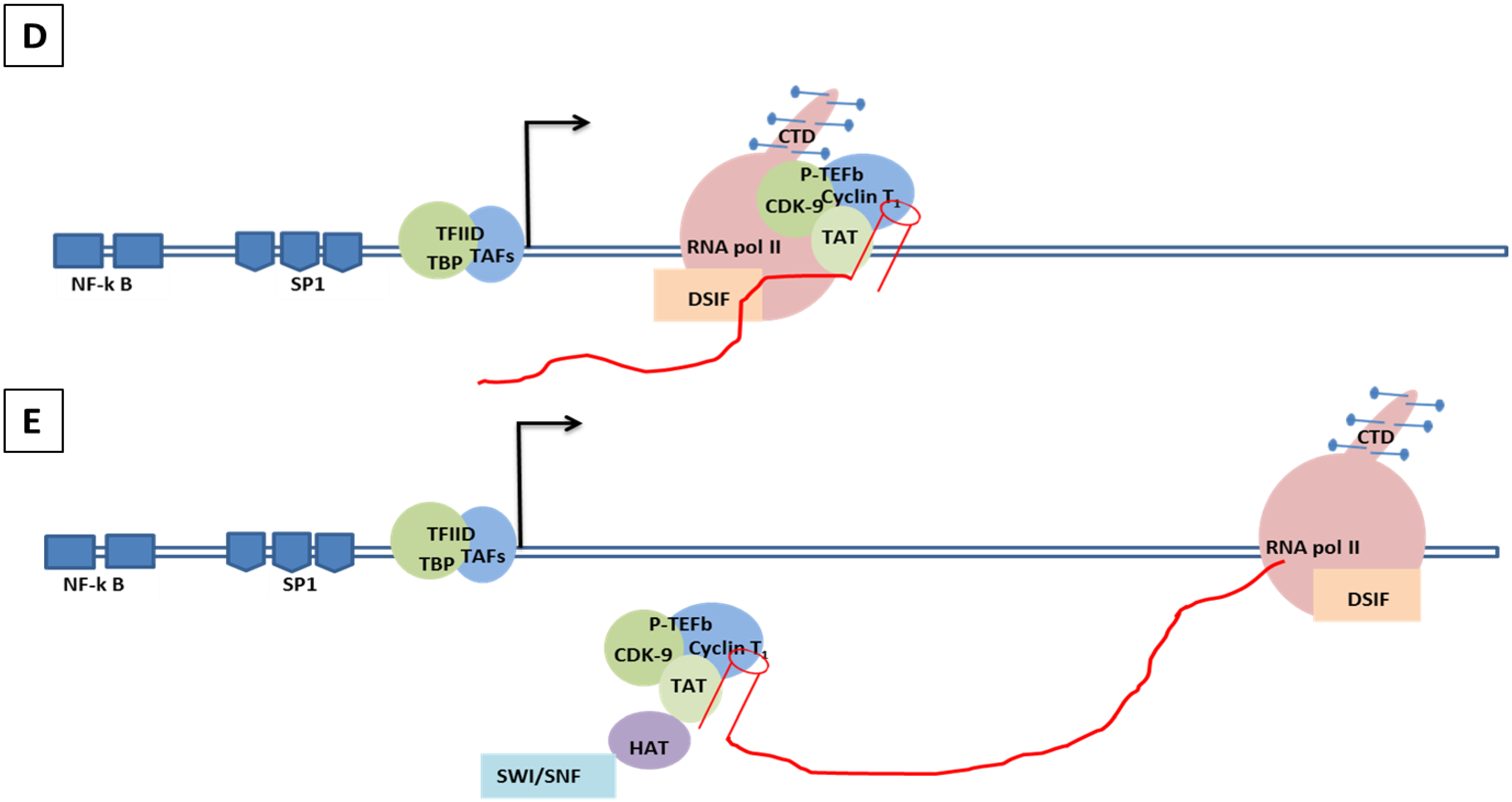
8.2. Molecular Mechanism of Tat-Independent Transcription in SRLVs
SRLV genomes lack both tat-coding sequences and its TAR target motif [60,61]. Earlier publications [58,264,265,266,267] reported interactions of LTR promoters with a protein called “Tat” (Vpr-Like) produced from an open reading frame whose position in SRLV genomes coincided with that of tat-coding sequences of primate lentiviruses. These publications reported the following findings:
- (1).
- Vpr-like protein mediates transcription activation of SRLV genomes and this is dependent on the presence of AP-1 and AP-4 sites located in the U3 region of SRLV LTRs. The proximal AP1 site to the MVV TATA box promoter is the most important for transcription, though SRLV Vpr-like protein does not bind the AP-1 site [264,268].
- (2).
- SRLV Vpr-like protein structure has three domains including: (a) an N-terminal acidic and hydrophobic domain, presumably the activator domain, which interacts with the TATA binding protein (TBP) in vitro [59,266]; (b) a central Leucine-rich domain that can interact with FOS-Jun-specific proteins in the AP-1 sites [266]; (c) a C-terminal Cysteine-rich domain which might direct protein dimerization. An interaction of the SRLV Vpr-like protein with the TBP from the TFIID complex was shown to bind to the TATA box via its activator domain, in addition to the proximal AP-1 binding to the Leucine-rich domain. These interactions allow a stabilization of protein complexes located at the viral promoter, and therefore induce an increase of viral gene transcription as illustrated in Figure 6.
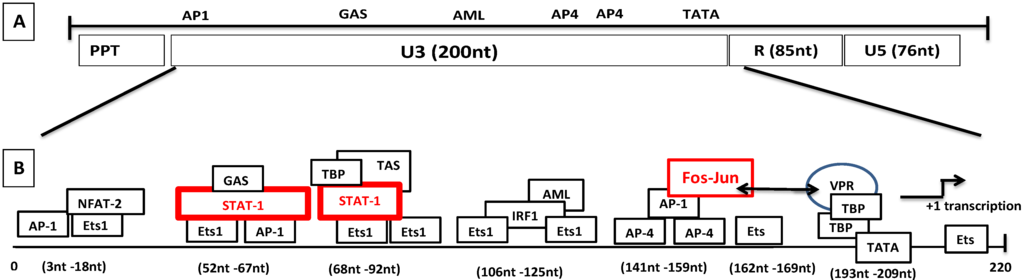
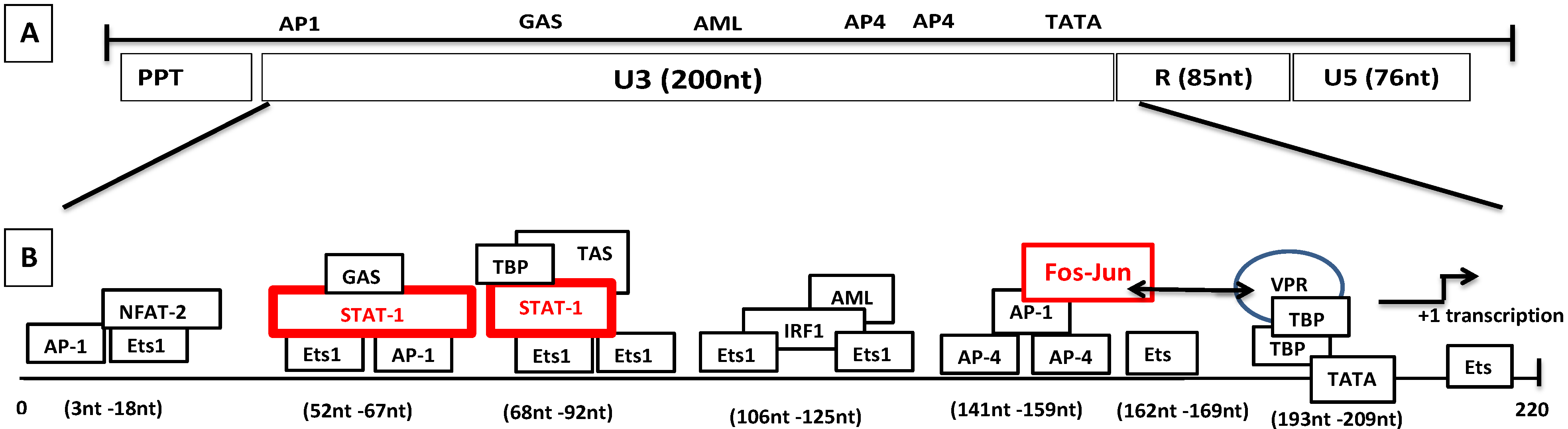
Figure 6.
Genomic organization of SRLVs and mechanism of LTR transactivation independent of Tat. (A) Various regions of the SRLV LTR are distinguished. (B) The transcriptional elements in U3 consist of TATA box, Ets-1, Stat-1, AP4, GAS, AML, NFAT-2, IRF-1, and AP-1 sites. The proposed transactivation pathway suggests that IFN-γ activates STAT-1 phosphorylation. Phosphorylated homodimer binds to the gamma activation site (GAS) while tumor necrosis factor alpha (TNF-α) activates the transcription by binding to the 17-nucleotide TNF-activated site (TAS) within the 70-nucleotide repeats. Vpr interacts with the Jun protein of the Fos-Jun complex bound to the AP-1 site and also interacts with the TATA box-binding proteins TBP, which interact with the TATA box.
Figure 6.
Genomic organization of SRLVs and mechanism of LTR transactivation independent of Tat. (A) Various regions of the SRLV LTR are distinguished. (B) The transcriptional elements in U3 consist of TATA box, Ets-1, Stat-1, AP4, GAS, AML, NFAT-2, IRF-1, and AP-1 sites. The proposed transactivation pathway suggests that IFN-γ activates STAT-1 phosphorylation. Phosphorylated homodimer binds to the gamma activation site (GAS) while tumor necrosis factor alpha (TNF-α) activates the transcription by binding to the 17-nucleotide TNF-activated site (TAS) within the 70-nucleotide repeats. Vpr interacts with the Jun protein of the Fos-Jun complex bound to the AP-1 site and also interacts with the TATA box-binding proteins TBP, which interact with the TATA box.
Studies by Harmache et al. [269] showed that the tat gene of CAEV was fully dispensable for viral replication in vitro and in vivo. Three different infectious molecular clones of CAEV bearing three distinct mutations in the tat gene were used in this study to compare the replication kinetics after transfection or infection of cultured goat synovial membrane cells and infection of blood-derived mononuclear cells or macrophages. The results showed that all mutants replicated at an efficiency similar to that of the wild type. The properties of wild type and tat mutant viruses were also compared in vivo by injecting the infectious molecular clones bearing the proviral DNAs directly into the joints of goats. Animals seroconverted between 27 and 70 days post-inoculation, and the CAEV mutant in tat was isolated from blood-derived macrophages, confirming that tat was dispensable for productive replication of CAEV in vivo as well.
Recent data clearly demonstrated that tat-encoded protein is not only structurally different, but also has no transactivation activity comparable to HIV/SIV Tat [60,61]. Further studies found instead that this protein was associated with functions similar to that of HIV-1 Vpr [61,62,63]. These studies compared the trans-activation activity exerted by MVV, CAEV, and HIV-1 Tat proteins in a variety of cell lines. The sub-cellular localization of MVV and CAEV Vpr-like protein was also investigated. None of the SRLV Vpr-like proteins tested induced the transactivation of SRLV or HIV-1 LTR promoters. Unlike HIV-1 Tat, the SRLV protein was found to be incorporated into the virions. The results also demonstrated that this lack of transactivation was not associated with the inability of this protein to enter the nucleus of expressing cells. The SRLV Vpr-like protein from the MVV strain, however, induced a very modest (two- to three-fold) upregulation of SRLV LTRs. However, in similar conditions, HIV-1 Tat trans-activated HIV-1 LTR over 60-fold [60,61].
The studies suggest that the primary role of the so-called Vpr-like protein in SRLV is not the transactivation of the LTR promoter similar to that of the primate lentiviruses HIV and SIV. Further studies demonstrated that this protein is structurally and functionally related to Vpr. The mechanism of transactivation with SRLV Vpr-like protein involves an interaction with Jun protein from the Fos/Jun complex bound to the AP-1 site located in viral LTR; this mechanism is functionally very close to the HIV-1 Vpr which it interacts with in the Sp1 located in viral LTR [61,62,63].
In the pursuit of exploration of other molecular pathways which lead to the consecutive transcription of the CAEV promoters and transcription factors, various studies were performed by Murphy et al. in the human monocytic U937 cell line expressing the SRLV genome. These studies suggested that the tumor necrosis factor alpha (TNF-α) and granulocyte-macrophage colony stimulating factor (GM-CSF) activate the CAEV transcription. They used SP600125 to block the Jun N-terminal kinase (JNK) and the cells were subsequently treated with cytokines. SP600125 blocked the previously described transactivation pathway through AP-1 [270], yet the SRLV transactivation was observed. Therefore, they proved that the TNF-α and GM-CSF could induce the activation of CAEV promoter independent of AP-1. Furthermore, using a set of sequentially deleted mutants, they found that elements within the 70-bp repeat in the U3 region are involved in the activation by TNF-α [270]. In a similar work performed previously by Sepp et al., they showed that the γ-interferon activation site (GAS) element from the 70-bp motif is adequate for the responsiveness to IFN-γ using a heterologous minimal promoter. They also showed that the binding of the nuclear factor to the GAS element is inhibited by antibodies directed against the signal transducer and activator of transcription STAT1 (p91/84) [271]. In a later work reported by Murphy et al., the authors identified a sequence of 17 nucleotides called TNF-activation site (TAS) which is located within the 70-bp repeat of the U3 region. They also found other sequences within the U3 region required for the IFN-γ activation. However, additional work is needed to establish whether STAT1 binding to the U3 site is involved in the activation of SRLV promoter through TNF-α and GM-CSF regulation.
Data from experiments in these two reports show that the TAS sequence in the SRLV LTRs is essential for TNF-α and IFN-γ to activate STAT-1 phosphorylation and bind to GAS. The STAT-l signaling pathway was shown to play an important role in the regulation of viral promoter activation through IFN-γ [271,272] and GM-CSF [273]. It is also interesting to know that IFN-γ leads to the activation of CAEV LTR through the STAT-1 signaling pathway [271] in monocytes, and STAT1 also plays an important role in the differentiation of monocytes to macrophages [274]. Interestingly, natural deletions in the U3 70-bp, vrp-like accessory gene and dUTPase resulted in highly attenuated genotype E strains of SRLV isolated in Italy [275]. Furthermore, the low pathogenicity and variable virulence can be also due to the deletions and/or mutations of the replication enhancer factors of the LTR, such as AP1, AML, TNF-α, and IFN-γ responsive elements [276,277]. While in SRLV strains from sheep the viral promoter sequences in LTR were found to determine the cell tropism [278], those sequences of strains from goats were not [279,280]
In a serendipitous experiment of nature as quoted by Dr. Murphy, his group isolated SRLV (CAEV-MA) from the mammary glands and from the carpal synovium (CAEV-JT) of the same goat. Sequence analyses in viral promoters revealed that the CAEV MA had a rare loss of GAS that was replaced by 5’-ACTCGAGGGTCTAGGA-3’ [281]. Earlier studies of SRLV-infected goats showed a heterogeneity of molecular clones with different promoters in different anatomical locations [279], but it was the first time that they found the rare mutation in the same goat. Using an assay based on β-galactosidase activity measurement, they found that the mean of basal promoter activity of CAEV-MA was inferior to that of CAEV-JT (mean OD of 0.389 ± 0.083 vs. 007, p < .0001, respectively) [281].
8.3. Role of Epigenetics and Transcriptional Regulation Factors in Modulating HIV-1 Latency
In general, eukaryotic DNA that is not in a transcriptionally active form is tightly packaged around nucleosomes. Each nucleosome is composed of octamers of histone proteins having pairs of H2A, H2B, H3, and H4 molecules densely packed to form the chromatin. Any change in the structure of the chromatin markedly affects gene expression. The complex of DNA/histones undergoes various post-translational modifications like acetylation, methylation, and phosphorylation, leading to modifications of the structure of chromosomes [282,283,284,285,286]. Histone modifications also contribute to the transcriptional regulation by the transcriptionally active euchromatin or transcriptionally repressive region heterochromatin which inhibits the transcription.
The post-integrational event of HIV-1 provirus into the host genome is followed by interactions with two nucleosomes, nuc-1 and nuc-0, on the LTR region. The position of nuc-1 plays an important role in controlling gene expression [287,288]. The cellular transcription factors, such as Yin Yang-1 (YY1), LSF, c-Myc, Sp-1, and AP-4, recruit histone deacetylases HDACs to the HIV-1 promoter, leading to a block of the transcription activity leading to latency [215,289,290,291,292,293,294]. The classical NF-kB complex has two subunits, p50 and p65, and this heterodimer is a potent activator of transcription. However, p50 can also generate homodimers that repress HIV-1 transcription by recruiting the HDAC to the LTR promoter. Knockdown experiments of p50 result in the reinitiation of RNA polymerase recruitment [293,295].
Histone acetyltransferases (HATs) such as E1A binding protein p300/CREB-binding protein (p300/CBP), P300/CBP-associated factor (PCAF), and GCN5 histone acetyltransferase may be recruited to the HIV-1 promoter. These HATs, which are critical for the acetylation of histone tails, interact with Tat, leading to recruitment of the chromatin remodeling complex SWI/SNF/BAF that opens/relaxes the chromatin and enables the transcription elongation by displacing restrictive nucleosomes [296,297,298,299].
There are also post-transcription modifications that are implicated in the regulation of Tat-induced latency [300]. Tat is necessary for sustained transcription of the HIV-1 LTR and suboptimal levels of Tat can lead to latency. Lack or minimal production of Tat can be due to the lack of cellular factors or due to mutations on the Tat/TAR axis. There are post-transcriptional modifications of Tat that control the interactions of Tat with P-TEFb and TAR. The dissociation of Tat from TAR and P-TEFb is due to the demethylation of Lysine 51 by Lysine-specific demethylase 1A (KDM1A/LSD1)/REST corepressor 1 (CoREST) and the acetylation of Lysine 50 mediated by Lysine acetyltransferase p300/KAT3B while, in contrast, Tat activity is enhanced by the methylation of Lysine 51 by Lysine methyltransferase SET7/9/KMT7 [301,302,303,304,305]. The transcriptional activity of pTEFb is reduced by acetylation of CDK9 of pTEFb, by the histone acetyltransferases hGCN5 and PCAF, leading to HIV-1 latency [306]. Moreover, efficient activity of Tat is dependent on the methylation of Lysine 51 by KMT7 that results in enhanced Tat binding to its target TAR in the 5’ extremity of the genomic RNA. Acetylation of Tat by PCAF on Lysine 28 is essential for recruitment of pTEF at the 5’end of viral RNA for an efficient elongation of RNA transcription. Sirtuin-1, a specific Tat deacetylase, increases the amount of unacetylated Tat in number [296].
Histone methylation, another epigenetic modification, also plays a role in regulating the latency of HIV-1. The histone methyltransferases (HMTs) such as Enhancer of zeste homolog 2 (EZH2) and Euchromatic Histone-Lysine N-methyltransferase 2 (EHMT2), also known as G9a and Histone-Lysine N-methyltransferase (SUV39H1), are involved in regulating HIV-1 transcription by inducing H3 methylation at Lysine 9 (H3K9) and Lysine 27 (H3K27). The studies by Karn’s group have demonstrated that EZH2 is a critical enzymatically active compound of the polycomb repressive complex 2 (PRC2), through which H3K27 trimethylation takes place [307]. The knock-out of EZH2 induces the loss of H3K27 trimethylation which leads to latency. SUV39H1 is an H3K9 methyltransferase that initiates the heterochromatin formation by interacting with HP1 and induces latency [204,308,309].
Further details on the epigenetic mechanism and post-translational modifications associated with HIV-1 latency and activation of transcription are reviewed in following [254,262,310,311,312].
8.4. Latency and HIV Cure
Latently infected cells are defined as the cells containing the integrated HIV-1 DNA that are transcriptionally silent but retain the capacity to produce infectious virus upon activation. These latently infected cells reside in viral reservoirs in different anatomical locations. They consist of different cell types that have stable kinetic properties and allow persistent virus replication. In the HAART era, most of the latently infected cells were abundantly found in the sanctuaries where the HAART drug penetration was limited. Theoretically, a cure can be either functional or sterilizing. A functional cure aims at permanent viral suppression following the interruption of therapy to levels that prevent immunodeficiency and transmission, while a sterilizing cure aims at the complete eradication of the viral population from the whole body [313].
Several therapeutic strategies to cure HIV-1 infection have been proposed, targeting the latently infected CD4+ T cells (reviewed in [314,315,316]).
- Reactivation of latent virus while patient is on HAART using HDAC inhibitors, vorinostats (SAHA), valproic acid, [317] “non-oncogenic” phorbol ester (bryostatin, prostratin), and cytokine IL-7 [318]. Induction of viral replication of proviruses in cells from latent reservoirs and clearance is the most promising “Kick and kill” strategy.
- Excision and removal of the proviral DNA [319,320,321],
Strategies that aim to eradicate the virus without targeting latent reservoirs:
- Creation of HIV-1 resistant cells (SB 728-T) [322,323],
- Gene therapy (CCR5Δ32/Δ32) [324,325,326] and Elite suppressors (ESs) model [327,328],
- Reversal of immune exhaustion (PD-1 antibodies) [329],
The Unique Case of Possible Cure
The Berlin patient was infected with HIV-1 and was on ART for more than 11 years before being diagnosed with leukemia at age 40. The curative intervention included an acute myeloid leukemia (AML) intensive conditioning regimen, including chemotherapy, antithymocyte globulin, and total body irradiation, with allogeneic hematopoietic stem cell transplantation (HSCT) from a CCR5 D32 homozygous donor. These interventions have eliminated most of the patient’s immune cells, including latently infected cells, and engrafted cells could not be infected with residual virus if any remained. This patient is established “aviremic”, and even seven years after HSCT and extensive analyses, there is no HIV-1 detected [325,326,330]. With a similar HSCT procedure, two other patients, Boston patient 1 and Boston patient 2, were intervened by reduced intensity conditioning and allogeneic HSCT from CCR5 wild-type donor. They were on ART for 4.3 years and 2.6 years, respectively, after HSCT, but they had detected a rebound after 84 days and 225 days, respectively, after ART interruption [331,332].
A case of “functional cure” has been recently reported in the International AIDS Society’s 8th conference on HIV-1 pathogenesis, treatment, and prevention. Under the French Pediatric Cohort led by Dr.Asier Saez-Cirion, an ART treatment was initiated soon after the birth of a baby girl who continued the treatment for six years and then stopped. The young woman, now 18.5 years old, has been in remission for 12 years. The early antivirals might have limited and reduced the formation of the reservoirs and protected the immune system [333]. This is the only case so far with such a long remission in children. Earlier, the “Mississippi baby” who received an early postnatal ART became aviremic for a period of 24 months. However, after cessation of HAART at 18 months, a rebound was detected, thus suggesting that the early ART treatment in the infected infants did not completely eliminate the reservoirs [334]; however, new studies have proved otherwise. In another long-term successful remission in the “VISCONTI cohort” study in adults, there are 14 post-treatment controllers (PTCs). They were initiated with the ART soon after infection and were on the treatment for at least three years before interruption. The study found that PTCs were able, after therapy interruption, to keep and in some cases further reduce a weak viral reservoir [335].
9. Sites of Latent Reservoirs
9.1. Cell Lineages
The primary cell targets of HIV-1 infections (discussed in detail in section 3.6) include the CD4+ T lymphocytes [336,337] and monocyte/macrophage lineages [338], while other cells were found to be capable of getting infected with HIV-1 including natural killer, CD8+, B and follicular dendritic cells.
9.1.1. CD4+ T Lymphocytes
In normal conditions, the majority of the CD4+ T lymphocytes are not activated (naïve and resting memory cells comprising central memory TCM and transitional memory TM) [339]. Some studies have shown that non-activated CD4+ T cells can be permissive to HIV-1, and target naïve T cells might have a low level of CCR5 expression [340,341,342]. It is estimated that only approximately 1 × 107 resting latently infected CD4+ T cells are present in blood and lymph nodes [343]. During HAART treatment, latent CD4+ reservoirs escape the efficient depletion of infected activated T cells and their progressive depletion occurs only slowly, depending on cell activation and cell death. The half-life of latently infected resting CD4+ T cells is estimated to be around 44 months [344] and, thus, mathematical modeling of eradication, assuming a constant rate of reactivation/elimination of the entire reservoir of 106 cells, would require 73 years [345,346]. Therefore, resting CD4+ T lymphocytes are considered the leading cell type in the HIV-1 reservoir [347]. SRLVs do not infect T lymphocytes productively or latently, therefore this cell lineage is not site of a SRLV reservoir.
9.1.2. Monocytes and Macrophages
As such, monocytes/macrophages can act like a viral reservoir, some of which are immune-privileged sanctuaries such as the CNS where they mature into microglia, perivascular and meningeal macrophages, and choroid plexus [348]. Infected microglia and perivascular macrophages induce clinical pathogenesis of HIV-1-associated dementia [349]. In general, virus production from monocytes/macrophages is markedly lower than from the CD4+ T lymphocytes [101]. Nevertheless, in addition to truly silent reservoirs, these cells may support a continuous low level virus production throughout their life since, unlike T cells, they are less sensitive to cytopathic effects induced by virus infection [101]. Gut-associated macrophages of lamina propria differentiated from blood monocytes could represent an important HIV-1 reservoir [350,351]. The role of monocytes and macrophages is also reviewed in detail in the following reviews [312,352,353,354]. SRLV in sheep and goats would be an excellent model to specifically study the mechanisms involved in the latency associated with these cells in absence of that associated with the CD4+ T lymphocyte reservoir; unfortunately, there are only limited studies.
9.1.3. Dendritic Cells
Their potential implication as players in HIV-1 latency/reservoir was demonstrated by the mature myeloid dendritic cells located in the lymph nodes which can harbor and transmit virus shielded from immune recognition for extended periods of time [355]. There have been reports that SRLVs could also infect and replicate in dendritic cells, which may also play a role in virus dissemination into lymphoid tissues, but it seems to be of low significance [90,356].
9.2. Anatomical Reservoirs
Among the anatomical reservoirs for HIV-1 there are two main categories that deserve special attention: (i) immune-privileged sites in which antiviral effector mechanisms are inhibited and/or (ii) sites that are inaccessible or poorly accessible to drugs used in HAART. Examples of such reservoirs are the CNS, lymphoid organs, the genital tract, and lungs.
9.2.1. CNS
CNS is protected by the blood-brain barrier which is highly selective in the exchange between blood and the brain tissue compartments, thus creating an immunological micro-environment. The tight epithelium of the choroid plexus restricts the flow of the molecules and cells into the cerebrospinal fluid (CSF), thus creating an effective barrier which prevents the passage of most HAART drugs to the CNS. The HIV-1 reservoirs in the CNS are the microglia and, to a lesser extent, the cortical and basal ganglia-derived astrocytes in which HIV-1 DNA has been detected [357,358,359]. Microglia and perivascular macrophages produce neurotoxins and cellular factors that induce clinical pathogenesis of HIV-1-associated dementia [129,170,360,361].
In SRLVs, the endothelial cells of the vascular system of the CNS have been shown to facilitate viral entry into the CNS [10]. The virus in the CNS causes lesions and the obvious signs of meningitis along with the destruction of the myelin sheath [362]. Proviral DNA has been located in macrophages, microglial cells, astrocytes and oligodendrocytes, the ependymal epithelium, and the choroid plexus. Positive areas are also found in the spinal cord, in endothelial cells of small blood vessels of goats and kids naturally infected with SRLVs [87]. Apart from the Icelandic research, MVV is mostly associated with the pulmonary and mammary lesions, but the study done by Benavide’s group described clinical cases in 12 Assaf sheep from Spain in which the CNS lesions were only confined to the spinal cord [363]. In another study on 64 sheep with natural SRLV-associated meningoencephalitis, they found lesions particularly in the cerebellar peduncles (non-suppurative meningoencephalitis), followed by the corpus callosum, hippocampus, and thoracic spinal cord [362]. These studies are cases of CNS alterations in sheep which are not the common pathogenesis currently seen in SRLV-infected flocks.
9.2.2. Lymphoid Organs
Lymphoid organs contain 98% of the lymphocytes in the body, and thus in patients treated with HAART, HIV-1 can be detected in all primary and secondary lymphoid tissues such as tonsils, bone marrow, GALT, spleen, lymph nodes, and thymus. The GALT is the tissue that contains the largest portion (40–60%) of lymphocyte populations and is also one of the largest reservoirs of macrophages [364,365]. In infected individuals, the GALT generally has half- to one-log more HIV-1 RNA molecules than in the peripheral blood mononuclear cells (PBMC) [158,366]. This increase is mainly due to the constant presence of activated CD4+ T cells leading to replication of the viruses even during HAART. Thus, while not immunologically protected or necessarily preventing HAART access, the GALT serves as a reservoir by the sheer number of activated viral targets.
In SRLV-infected animals, provirus DNA was detected in classical peripheral lymphoid tissues (spleen, lymph nodes, thymus, bone marrow, PBMCs, and the lungs) of experimentally and chronically infected goats [102,367,368]. Although lymph nodes are considered an important viral reservoir, they do not appear to be sites of intense viral replication. Indeed, multi-spliced mRNA has not been shown to be present in samples from all goats, whereas un-spliced mRNA has been detected from lymph nodes, spleen, bone marrow, and /or PBMC [369].
9.2.3. Genital Tract
The genital tracts of both males and females are considered to be anatomical reservoirs. The testis in particular represents an immunologically privileged sanctuary of HIV-1, which additionally also restricts the accessibility of HAART drugs [370,371,372,373]. This may result in the presence of HIV-1-infected monocytes and T cells in seminiferous tubules and the interstitium of the testis, as well as in the semen [374]. Earlier works have reported HIV-1 infection of the spermatogonia, spermatocytes, spermatids, and residual germ layer [375,376,377,378,379]. In women, the viral replication is active in the submucosa of the cervix and HIV-1 has also been detected in the epithelial and stromal cells of the uterus, fallopian tubes, and ectocervix. HIV-1 DNA was found in cervicovaginal fluids, with cervical biopsies during HAART suggesting potential limited drug penetration [380,381,382].
Studies in SRLVs have clearly demonstrated that goat uterine epithelial cells are susceptible to SRLV infection in vivo [383]. Also, SRLV proviral DNA has been identified using PCR in tissues of the genital tract (uterus, oviduct, and ovary) [175] and in vitro, in granulosa cells, oviduct epithelial cells [384,385], and early goat embryo cells [386]. Recent studies also demonstrate that cells of the buck genital tract are targets of SRLV infection and are thus a potential reservoir that may shed infectious SRLV into the semen of infected animals [387,388,389]. The work conducted by Fieni’s group demonstrated that the first four washes were sufficient to remove infectious CAEV and infected cells in the semen so that SRLV-free embryos can be produced by in vitro fertilization (IVF) using spermatozoa contaminated in vitro by SRLV [390].
9.2.4. Lungs and Kidney
HIV-1 is positively detected in alveolar macrophages, interstitial macrophages, and lymphocytes. Alveolar macrophages harbor HIV-1 even in patients under HAART with undetectable plasma viral loads, showing that they are a potential reservoir for the virus [391,392]. Additionally, phagocytosis and immune functions are impaired due to HIV-1 replication in macrophages, which increases the risk of lung infection with other pathogens [393]. The generally low levels of replication in alveolar macrophages and the presence of alveolar lymphocytes can be rapidly augmented in lungs due to inflammatory responses to opportunistic infections. In addition, HIV-1-infected individuals are significantly more susceptible to long-term HIV-1-associated complications such as chronic obstructive pulmonary disease (COPD) [394] (reviewed in detail in [395]).
HIV-1 DNA and mRNA have been found in the renal globular and tubular epithelial cells of kidney, suggesting replication in renal tissues. Kidney biopsies have shown the presence of circularized viral DNA, suggesting active replication [396,397]. Recent studies showed increased evidence of the kidney as a potential reservoir where the virus is exchanged between interstitial T cells and renal tubule epithelial cells [398]. Using novel noninvasive tests designed for determining HIV-1 infection of the kidney allografts by measuring HIV-1 DNA and RNA levels in a patient’s urine, it has been demonstrated that HIV-1 can be detected in kidney allografts after transplantation despite undetectable viremia, and this infection might influence the graft outcome [399]. Another study suggested that the HIV-1 persistence is reduced in the transplant recipient following the transplantation of the HIV-1 infected kidney [400].
Studies conducted by Ravazzolo et al. indicated the presence of SRLV provirus transcripts (both un-spliced and multi-spliced) in alveolar macrophages, confirming that the lungs represent an important site of virus replication [369,401,402]. Other studies also confirmed relatively high levels of genomic and sub-genomic viral RNAs in alveolar macrophages of infected goats [403]. In earlier studies, it was found that SRLV-infected sheep alveolar macrophages were able to stimulate CTL activity in vitro and were targets of these activated CTL [404]. In recent studies, the increased expression of chemoattractant chemokines (MCP-1 and IP-10) was linked to the inflamed lungs [50]. Some fatal infections show diffuse interstitial pneumonia characterized histologically by severe lymphoplasmacytic infiltrates with massive alveolar proteinosis, interstitial fibrosis, and type II pneumocyte hyperplasia [405].
10. Can SRLV Help to Develop Strategies to Control HIV-1?
SRLVs in infected sheep and goats do not cause immunodeficiency like HIV-1 does in HIV-1-infected humans. The reason why SRLVs did not evolve towards an immunodeficiency-inducing phenotype is not well understood. It has been established that SRLVs replicate only transiently in the periphery and replicate productively in macrophages but not in CD4+ T cells in target tissues. Our group has demonstrated that SRLVs have Tat-independent constitutive LTR promoters [60]. We have also shown that SRLV infection in newborn calves results in a total clearance of the virus after four months of virus replication and diffusion in target tissues [115].
Taken together, these results helped to conclude that the constitutive expression of viral proteins by SRLV LTRs could have induced antigen-specific T cell responses that were associated with virus clearance. To test the hypothesis, our group has recently generated a chimeric genome, based on the former vaccine integration and replication-defective ∆4-SHIVKU2 by replacing the 5’SIV LTR with that of CAEV (∆4-SHIV-KU2 was derived from SHIV-KU2, from which the rt, int, vif, and the 3’ LTR sequences were deleted). The replication- and integration-deficient lentivector DNA vaccine showed promising results by inducing potent immune responses both in mice and macaques [406].
These encouraging results led to the development of a further enhanced immunogenic chimeric lentivector DNA also based on the SHIV genome in which both 5’ and 3’ LTRs of SIV were replaced with those of CAEV. Additionally, the integrase was deleted, making it a safe, novel, non-integrative one-cycle lentivector DNA vaccine that produces viral antigens and particles to stimulate the immune system in in vivo transfected cells. These viral particles can transduce target cells without integrating vaccine DNA into these cells, thereby expressing further viral proteins and thus amplifying the vaccine antigenicity. This lentivector induced high multifunctional persistent immune responses both in immunized mice and macaque models [406].
These data provide an encouraging perspective of use of CAEV for the development of innovative strategies for the generation of a safe and long-term effective vaccine that could control HIV-1 [407].
11. Conclusions
HIV-1, which causes AIDS, has been responsible for the infection of over 75 million people and the death of nearly 35 million people worldwide. SRLVs infect goats and sheep in flocks all over the world. The reason why these have not evolved to induce AIDS in their natural hosts in contrast to primate lentiviruses remains not well understood. Knowledge from SRLV studies was very helpful to understanding HIV-1 biopathology and host/pathogen interaction early after the discovery of the human lentivirus. However, during the last decades, there was no substantial effort to use the SRLV model for better understanding the complex host/pathogen interaction of primate lentiviruses. Comparative analyses of structural, functional, and pathogenesis differences between SRLVs and primate subgroups of lentiviruses might bring new insights that will help us better understanding the natural history of these viruses and help develop innovative tools to control them.
Acknowledgments
We thank the National Institute of Health, AIDS Research and Reference Reagent Program for providing HIV and SIV overlapping peptides and cell lines.
We also thank the “Institut National de la Recherche Agronomique” (INRA), University Joseph Fourier and the “Agence Nationale de Recherche sur le SIDA” (ANRS) for supporting this work.
Author Contributions
Yahia Chebloune had the original idea and conducted the design of the study. Deepanwita Bose performed the bibliographic study and the writing of the review. Jean Gagnon helped in the writing, discussion and proof reading.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Peeters, M.; Fransen, K.; Delaporte, E.; Van den Haesevelde, M.; Gershy-Damet, G.M.; Kestens, L.; van der Groen, G.; Piot, P. Isolation and characterization of a new chimpanzee lentivirus (simian immunodeficiency virus isolate cpz-ant) from a wild-captured chimpanzee. AIDS 1992, 6, 447–451. [Google Scholar] [CrossRef] [PubMed]
- Peeters, M.; Honore, C.; Huet, T.; Bedjabaga, L.; Ossari, S.; Bussi, P.; Cooper, R.W.; Delaporte, E. Isolation and partial characterization of an HIV-related virus occurring naturally in chimpanzees in Gabon. AIDS 1989, 3, 625–630. [Google Scholar] [CrossRef]
- Van Heuverswyn, F.; Li, Y.; Neel, C.; Bailes, E.; Keele, B.F.; Liu, W.; Loul, S.; Butel, C.; Liegeois, F.; Bienvenue, Y.; et al. Human immunodeficiency viruses: SIV infection in wild gorillas. Nature 2006. [Google Scholar] [CrossRef] [PubMed]
- Barre-Sinoussi, F.; Chermann, J.; Rey, F.; Nugeyre, M.; Chamaret, S.; Gruest, J.; Dauguet, C.; Axler, B.C.; Vezinet- Brun, F.; Rouzioux, C.; et al. Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science 1983, 220, 868–871. [Google Scholar] [CrossRef]
- Terzieva, V. Regulatory T Cells and HIV-1 Infection. Viral Immunol. 2008, 21, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Sigurdsson, B.; Palsson, P.A. Visna of sheep; a slow, demyelinating infection. Br. J. Exp. Pathol. 1958, 39, 519–528. [Google Scholar] [PubMed]
- Sigurdsson, B.; Palsson, P.A.; Tryggvaddottir, A. Transmission experiments with maedi. J. Infect. Dis. 1953, 93, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Gendelman, H.E.; Narayan, O.; Kennedy-Stoskopf, S.; Kennedy, P.G.; Ghotbi, Z.; Clements, J.E.; Stanley, J.; Pezeshkpour, G. Tropism of sheep lentiviruses for monocytes: susceptibility to infection and virus gene expression increase during maturation of monocytes to macrophages. J. Virol. 1986, 58, 67–74. [Google Scholar] [PubMed]
- Gorrell, M.D.; Brandon, M.R.; Sheffer, D.; Adams, R.J.; Narayan, O. Ovine lentivirus is macrophagetropic and does not replicate productively in T lymphocytes. J. Virol. 1992, 66, 2679–2688. [Google Scholar] [PubMed]
- Clements, J.E.; Zink, M.C. Molecular biology and pathogenesis of animal lentivirus infections. Clin. Microbiol. Rev. 1996, 9, 100–117. [Google Scholar] [PubMed]
- Fun, A.; Wensing, A.M.; Verheyen, J.; Nijhuis, M. Human immunodeficiency virus Gag and protease: Partners in resistance. Retrovirology 2012. [Google Scholar] [CrossRef] [PubMed]
- Konvalinka, J.; Krausslich, H.G.; Muller, B. Retroviral proteases and their roles in virion maturation. Virology 2015, 479–480, 403–417. [Google Scholar] [CrossRef]
- Minardi da Cruz, J.; Singh, D.; Lamara, A.; Chebloune, Y. Small ruminant lentiviruses (SRLVs) break the species barrier to acquire new host range. Viruses 2013, 5, 1867–1884. [Google Scholar] [CrossRef] [PubMed]
- Sarafianos, S.G.; Marchand, B.; Das, K.; Himmel, D.M.; Parniak, M.A.; Hughes, S.H.; Arnold, E. Structure and function of HIV-1 reverse transcriptase: Molecular mechanisms of polymerization and inhibition. J. Mol. Biol. 2009, 385, 693–713. [Google Scholar] [CrossRef] [PubMed]
- Luciw, P. Human Immunodeficiency Viruses and Their Replication, 3rd ed.; Raven Press: Philadelphia, PA, USA, 1996; pp. 1881–1952. [Google Scholar]
- Vaishnav, Y.; Wong-Staal, F. The Biochemistry of AIDS. Annu. Rev. Biochem. 1991, 60, 577–630. [Google Scholar] [CrossRef] [PubMed]
- Kalland, K.H.; Szilvay, A.M.; Brokstad, K.A.; Saetrevik, W.; Haukenes, G. The human immunodeficiency virus type 1 Rev protein shuttles between the cytoplasm and nuclear compartments. Mol. Biol. Cell. 1994, 14, 7436–7444. [Google Scholar]
- Meyer, B.E.; Malim, M.H. The HIV-1 Rev trans-activator shuttles between the nucleus and the cytoplasm. Genes Dev. 1994, 8, 1538–1547. [Google Scholar] [CrossRef] [PubMed]
- Daly, T.J.; Cook, K.S.; Gray, G.S.; Maione, T.E.; Rusche, J.R. Specific binding of HIV-1 recombinant Rev protein to the Rev-responsive element in vitro. Nature 1989, 342, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, M.; Asano, S.; Nakamura, T.; Adachi, M.; Yoshida, M.; Yanagida, M.; Nishida, E. CRM1 is responsible for intracellular transport mediated by the nuclear export signal. Nature 1997, 390, 308–311. [Google Scholar] [PubMed]
- Simon, J.H.; Malim, M.H. The human immunodeficiency virus type 1 Vif protein modulates the postpenetration stability of viral nucleoprotein complexes. J. Virol. 1996, 70, 5297–5305. [Google Scholar] [PubMed]
- Fouchier, R.A.; Simon, J.H.; Jaffe, A.B.; Malim, M.H. Human immunodeficiency virus type 1 Vif does not influence expression or virion incorporation of gag-, pol-, and env-encoded proteins. J. Virol. 1996, 70, 8263–8269. [Google Scholar] [PubMed]
- Simon, J.H.; Miller, D.L.; Fouchier, R.A.; Malim, M.H. Virion incorporation of human immunodeficiency virus type-1 Vif is determined by intracellular expression level and may not be necessary for function. Virology 1998, 248, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Goila-Gaur, R.; Strebel, K. HIV-1 Vif, APOBEC, and intrinsic immunity. Retrovirology 2008. [Google Scholar] [CrossRef] [PubMed]
- Sheehy, A.M.; Gaddis, N.C.; Choi, J.D.; Malim, M.H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 2002, 418, 646–650. [Google Scholar] [CrossRef] [PubMed]
- Desimmie, B.A.; Delviks-Frankenberrry, K.A.; Burdick, R.C.; Qi, D.; Izumi, T.; Pathak, V.K. Multiple APOBEC3 restriction factors for HIV-1 and one Vif to rule them all. J. Mol. Biol. 2014, 426, 1220–1245. [Google Scholar] [CrossRef] [PubMed]
- Holmes, R.K.; Malim, M.H.; Bishop, K.N. APOBEC-mediated viral restriction: Not simply editing? Trends Biochem. Sci. 2007, 32, 118–128. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Ikeuchi, K.; Byrn, R.; Groopman, J.; Baltimore, D. Lack of a negative influence on viral growth by the nef gene of human immunodeficiency virus type 1. Proc. Natl. Acad. Sci. USA 1989, 86, 9544–9548. [Google Scholar] [CrossRef] [PubMed]
- Pawlak, E.N.; Dikeakos, J.D. HIV-1 Nef: a master manipulator of the membrane trafficking machinery mediating immune evasion. Biochim. Biophys. Acta 2015, 1850, 733–741. [Google Scholar] [CrossRef] [PubMed]
- Piguet, V.; Wan, L.; Borel, C.; Mangasarian, A.; Demaurex, N.; Thomas, G.; Trono, D. HIV-1 Nef protein binds to the cellular protein PACS-1 to downregulate class I major histocompatibility complexes. Nat. Cell. Biol. 2000, 2, 163–167. [Google Scholar] [PubMed]
- Atkins, K.M.; Thomas, L.; Youker, R.T.; Harriff, M.J.; Pissani, F.; You, H.; Thomas, G. HIV-1 Nef binds PACS-2 to assemble a multikinase cascade that triggers major histocompatibility complex class I (MHC-I) down-regulation: analysis using short interfering RNA and knock-out mice. J. Biol. Chem. 2008, 283, 11772–11784. [Google Scholar] [CrossRef] [PubMed]
- Dikeakos, J.D.; Atkins, K.M.; Thomas, L.; Emert-Sedlak, L.; Byeon, I.J.; Jung, J.; Ahn, J.; Wortman, M.D.; Kukull, B.; Saito, M.; et al. Small molecule inhibition of HIV-1-induced MHC-I down-regulation identifies a temporally regulated switch in Nef action. Mol. Biol. Cell. 2010, 21, 3279–3292. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.H.; Thomas, L.; Ruby, C.E.; Atkins, K.M.; Morris, N.P.; Knight, Z.A.; Scholz, I.; Barklis, E.; Weinberg, A.D.; Shokat, K.M.; et al. HIV-1 Nef assembles a Src family kinase-ZAP-70/Syk-PI3K cascade to downregulate cell-surface MHC-I. Cell Host Microbe 2007, 1, 121–133. [Google Scholar] [CrossRef]
- Schaefer, M.R.; Wonderlich, E.R.; Roeth, J.F.; Leonard, J.A.; Collins, K.L. HIV-1 Nef targets MHC-I and CD4 for degradation via a final common beta-COP-dependent pathway in T cells. PLoS Pathog 2008. [Google Scholar] [CrossRef] [PubMed]
- Swann, S.A.; Williams, M.; Story, C.M.; Bobbitt, K.R.; Fleis, R.; Collins, K.L. HIV-1 Nef blocks transport of MHC class I molecules to the cell surface via a PI 3-kinase-dependent pathway. Virology 2001, 282, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Matthias, G.; Oliver, T.F.; Matija, P. Structure- function relationships in HIV-1 Nef. EMBO Rep. 2001, 2, 580–585. [Google Scholar]
- Roth, W.W.; Khan, M.; Geleziunas, R.; Stringer, H.G.; Zuberi, J.A.; Greene, W.C.; Powell, M.; Bond, V.C. Functionally-impaired HIV-1 Nef alleles from a mother-child transmission pair. Int. J. Mol. Sci. 2002, 3, 1058–1072. [Google Scholar] [CrossRef]
- Jere, A.; Fujita, M.; Adachi, A.; Nomaguchi, M. Role of HIV-1 Nef protein for virus replication in vitro. Microb. Infect. 2010, 12, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Fackler, O.; Lu, X.; Frost, J.; Geyer, M.; Jiang, B.; Luo, W.; Abo, A.; Alberts, A.; Peterlin, M. p21-activated kinase 1 plays a critical role in cellular activation by Nef. Mol. Biol. Cell. 2000, 20, 2619–2627. [Google Scholar] [CrossRef]
- Wolf, D.; Witte, V.; Laffert, B.; Blume, K.; Stromer, E.; Trapp, S.; d'Aloja, P.; Schurmann, A.; Baur, S. HIV-1 Nef associated PAK and PI3-kinases stimulate Akt-independent Bad-phosphorylation to induce anti-apoptotic signals. Nat. Med. 2001, 7, 1217–1224. [Google Scholar] [CrossRef] [PubMed]
- Groschel, B.; Bushman, F. Cell cycle arrest in G2/M promotes early steps of infection by human immunodeficiency virus. J. Virol. 2005, 79, 5695–5704. [Google Scholar] [CrossRef] [PubMed]
- Cohen, E.A. From arrest to escape: HIV-1 Vpr cuts a deal. Cell Host Microbe 2014, 15, 125–127. [Google Scholar] [CrossRef] [PubMed]
- Murakami, T.; Aida, Y. Visualizing Vpr-induced G2 arrest and apoptosis. PloS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
- Philippon, V.; Matsuda, Z.; Essex, M. Transactivation is a conserved function among primate lentivirus Vpr proteins but is not shared by Vpx. J. Hum. Virol. 1999, 2, 167–174. [Google Scholar] [PubMed]
- Mirani, M.; Elenkov, I.; Volpi, S.; Hiroi, N.; Chrousos, G.; Kino, T. HIV-1 protein Vpr suppresses IL-12 production from human monocytes by enhancing glucocorticoid action: Potential implications of Vpr coactivator activity for the innate and cellular immunity deficits observed in HIV-1 infection. J. Immunol. 2002, 169, 6361–6368. [Google Scholar] [CrossRef] [PubMed]
- Hoch, J.; Lang, S.M.; Weeger, M.; Stahl-Hennig, C.; Coulibaly, C.; Dittmer, U.; Hunsmann, G.; Fuchs, D.; Muller, J.; Sopper, S.; et al. vpr deletion mutant of simian immunodeficiency virus induces AIDS in rhesus monkeys. J. Virol. 1995, 69, 4807–4813. [Google Scholar] [PubMed]
- Lang, S.M.; Weeger, M.; Stahl-Hennig, C.; Coulibaly, C.; Hunsmann, G.; Muller, J.; Muller-Hermelink, H.; Fuchs, D.; Wachter, H.; Daniel, M.M.; et al. Importance of vpr for infection of rhesus monkeys with simian immunodeficiency virus. J. Virol. 1993, 67, 902–912. [Google Scholar] [PubMed]
- Bouzar, A.B.; Guiguen, F.; Morin, T.; Villet, S.; Fornazero, C.; Garnier, C.; Gallay, K.; Gounel, F.; Favier, C.; Durand, J.; et al. Specific G2 arrest of caprine cells infected with a caprine arthritis encephalitis virus expressing vpr and vpx genes from simian immunodeficiency virus. Virology 2003, 309, 41–52. [Google Scholar]
- Bouzar, A.B.; Villet, S.; Morin, T.; Rea, A.; Genestier, L.; Guiguen, F.; Garnier, C.; Mornex, J.F.; Narayan, O.; Chebloune, Y. Simian immunodeficiency virus Vpr/Vpx proteins kill bystander noninfected CD4+ T-lymphocytes by induction of apoptosis. Virology 2004, 326, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Arrode-Brusés, G.; Halloway, N.; Narayan, O.; Chebloune, Y. Changes of biological properties and pathogenesis of CAEV chimeras expressing Nef and Vpx/Vpr accessory proteins in infected goats. Retrovirology 2009. [Google Scholar] [CrossRef]
- Cullen, B.R. HIV-1 auxiliary proteins: Making connections ina dying cell. Cell. 1998, 93, 685–692. [Google Scholar] [CrossRef]
- Emerman, M.; Malim, M. HIV-1 regulatory/accessory genes: Key to unraveling viral and host cell biology. Science 1998, 280, 1880–1884. [Google Scholar] [CrossRef] [PubMed]
- Garcia, J.; Harrich, D.; Pearson, L.; Mitsuyasu, R.; Gaynor, R. Functional domains required for Tat-induced transcriptional activation of the HIV-1 long terminal repeats. EMBO J. 1988, 7, 3143–3147. [Google Scholar] [PubMed]
- Hauber, J.; Malim, M.; Cullen, B.R. Mutational analysis of the conserved basic domain of human immunodeficiency virus Tat protein. J. Virol. 1989, 63, 1181–1187. [Google Scholar] [PubMed]
- Tang, H.; Kuhen, K.; Wong-Staal, F. Lentivirus replication and regulation. Annu. Rev. Genet. 1999, 33, 133–170. [Google Scholar] [CrossRef] [PubMed]
- Arya, S.; Guo, C.; Josephs, S.; Wong-Staal, F. Trans-activator gene of human T-lymphotropic virus type III (HTLV-III). Science 1985, 229, 69–73. [Google Scholar] [CrossRef] [PubMed]
- Saltarelli, M.J.; Schoborg, R.; Gdovin, S.L.; Clements, J.E. The CAEV tat gene transactivates the viral LTR and is necessary for efficient viral replication. Virology 1993, 197, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Neuveut, C.; Vigne, R.; Clements, J.E.; Sire, J. The visna transcriptional activator Tat: Effects on the viral LTR and on cellular genes. Virology 1993, 197, 236–244. [Google Scholar] [CrossRef] [PubMed]
- Carruth, L.M.; Hardwick, J.; Morse, B.A.; Clements, J.E. Visna virus Tat protein: a potent transcription factor with both activator and suppressor domains. J. Virol. 1994, 68, 6137–6146. [Google Scholar] [PubMed]
- Villet, S.; Faure, C.; Bouzar, B.A.; Morin, T.; Verdier, G.; Chebloune, Y.; Legras, C. Lack of trans-activation function for Maedi Visna virus and Caprine arthritis encephalitis virus Tat proteins. Virology 2003, 307, 317–327. [Google Scholar] [CrossRef]
- Villet, S.; Bouzar, B.A.; Morin, T.; Verdier, G.; Legras, C.; Chebloune, Y. Maedi-Visna virus and caprine arthritis encephalitis virus genomes encode a Vpr-Like but No Tat protein. J. Virol. 2003, 77, 9632–9638. [Google Scholar] [CrossRef] [PubMed]
- Rea-Boutrois, A.; Villet, S.; Greenland, T.; Mehlen, P.; Chebloune, Y.; Verdier, G.; Legras-Lachuer, C. Small ruminant lentivirus Tat protein induces apoptosis in caprine cells in vitro by the intrinsic pathway. Virology 2009, 383, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Rea-Boutrois, A.; Pontini, G.; Greenland, T.; Mehlen, P.; Chebloune, Y.; Verdier, G.; Legras-Lachuer, C. Caprine arthritis-encephalitis virus induces apoptosis in infected cells in vitro through the intrinsic pathway. Virology 2008, 375, 452–463. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P.M.; Hahn, B.H. Origins of HIV and the AIDS pandemic. Cold Spring Harb. Perspect. Med. 2011. [Google Scholar] [CrossRef] [PubMed]
- Delaugerre, C.; De Oliveira, F.; Lascoux-Combe, C.; Plantier, J.C.; Simon, F. HIV-1 group N: Travelling beyond Cameroon. Lancet 2011, 378, 1894. [Google Scholar] [CrossRef]
- Plantier, J.C.; Leoz, M.; Dickerson, J.E.; De Oliveira, F.; Cordonnier, F.; Lemee, V.; Damond, F.; Robertson, D.L.; Simon, F. A new human immunodeficiency virus derived from gorillas. Nat. Med. 2009, 15, 871–872. [Google Scholar] [CrossRef] [PubMed]
- Vallari, A.; Holzmayer, V.; Harris, B.; Yamaguchi, J.; Ngansop, C.; Makamche, F.; Mbanya, D.; Kaptue, L.; Ndembi, N.; Gurtler, L.; et al. Confirmation of putative HIV-1 group P in Cameroon. J. Virol. 2011, 85, 1403–1407. [Google Scholar] [CrossRef] [PubMed]
- D’Arc, M.; Ayouba, A.; Esteban, A.; Learn, G.H.; Boue, V.; Liegeois, F.; Etienne, L.; Tagg, N.; Leendertz, F.H.; Boesch, C.; et al. Origin of the HIV-1 group O epidemic in western lowland gorillas. Proc. Natl. Acad. Sci. USA 2015, 112, 1343–1352. [Google Scholar] [CrossRef] [PubMed]
- Shaw, G.M.; Hunter, E. HIV transmission. Cold Spring Harb. Perspect. Med. 2012. [Google Scholar] [CrossRef] [PubMed]
- Touloumi, G.; Hatzakis, A. Natural history of HIV-1 infection. Clin. Dermatol. 2000, 18, 389–399. [Google Scholar] [CrossRef]
- Mitchell, D.T. Investigations into Jaagziekte or Chronic Catarrhal Pneumonia of Sheep; Directory Veterinary Education Research: Union of South Africa, UK, 1915; p. 585. [Google Scholar]
- Palsson, P.A. Maedi and visna in sheep. Front. Biol. 1976, 44, 17–43. [Google Scholar] [PubMed]
- Marsh, H. Progressive pneumonia in sheep. J. Am. Vet. Med. Assoc. 1923, 62, 458–473. [Google Scholar]
- Gislason, G. Thaettir urn Indutning Bdfjar og Karak~Ilsjdkdbma; Icelandic Department of Agricultural Publication: Reykjavik, Iceland, 1947; pp. 235–254. (In Icelandic)
- Palsson, P.A. Maedi-visna. J. Clin. Pathol. 1972, 6, 115–120. [Google Scholar] [CrossRef]
- Stunzi, H.; Buchi, H.F.; LeRoy, H.L.; Leeman, W. Endemische arthritis chronic a bei zeiegen. Schw. Arch. Tier. 1959, 106, 778–788. [Google Scholar]
- Stavrou, D.; Deutschlander, N.; Dahme, E. Granulomatous encephalomyelitis in goats. J. Comp. Pathol. 1969, 79, 393–396. [Google Scholar] [CrossRef]
- Cork, L.C.; Hadlow, W.J.; Crawford, T.B.; Gorham, J.R.; Piper, R.C. Infectious leukoencephalomyelitis of young goats. J. Infect. Dis. 1974, 129, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Crawford, T.B.; Adams, D.S.; cheevers, W.; Cork, L.C. Chronic arthritis in goats caused by a retrovirus. Science 1980, 207, 997–999. [Google Scholar] [CrossRef] [PubMed]
- Narayan, O.; Clements, J.E.; Strandberg, J.D.; Cork, L.C.; Griffin, D.E. Biological characterization of the virus causing leukoencephalitis and arthritis in goats. J. Gen. Virol. 1980, 50, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Peluso, R.; Haase, A.; Stowring, L.; Edwards, M.; ventura, P. A trojan hosre mechanism for the spread of visna virus in monocytes. Virology 1985, 147, 231–236. [Google Scholar] [CrossRef]
- Gendelman, H.E.; Narayan, O.; Molineaux, S.; Clements, J.E.; Ghotbi, Z. Slow, persistent replication of lentiviruses: role of tissue macrophages and macrophage precursors in bone marrow. Proc. Natl. Acad. Sci. USA 1985, 82, 7086–7090. [Google Scholar] [CrossRef] [PubMed]
- Lairmore, M.D.; Akita, G.Y.; Russell, H.I.; DeMartini, J.C. Replication and cytopathic effects of ovine lentivirus strains in alveolar macrophages correlate with in vivo pathogenicity. J. Virol. 1987, 61, 4038–4042. [Google Scholar] [PubMed]
- Narayan, O.; Kennedy-Stoskopf, S.; Sheffer, D.; Griffin, D.E.; Clements, J.E. Activation of caprine arthritis-encephalitis virus expression during maturation of monocytes to macrophages. Infect. Immun. 1983, 41, 67–73. [Google Scholar] [PubMed]
- Blacklaws, B.A. Small ruminant lentiviruses: immunopathogenesis of visna-maedi and caprine arthritis and encephalitis virus. Comp. Immunol. Microbiol. Infect. Dis. 2012, 35, 259–269. [Google Scholar] [CrossRef] [PubMed]
- Carrozza, M.L.; Mazzei, M.; Bandecchi, P.; Arispici, M.; Tolari, F. In situ PCR-associated immunohistochemistry identifies cell types harbouring the Maedi-Visna virus genome in tissue sections of sheep infected naturally. J. Virol. Methods 2003, 107, 121–127. [Google Scholar] [CrossRef]
- Sanna, E.; Sanna, M.P.; Vitali, C.G.; Renzoni, G.; Sanna, L.; Spano, S.; Rossi, G.; Leoni, A. Proviral DNA in the brains of goats infected with caprine arthritis-encephalitis virus. J. Comp. Pathol. 1999, 121, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Pautrat, G.; Filippi, P. Evidence for the production of a fusion factor during in vitro infection of sheep choroid plexus cells by visna virus. C. R. Seances Soc. Biol. Fil. 1979, 173, 811–817. [Google Scholar] [PubMed]
- Sihvonen, L.; Veijalainen, P. Kinetics of maedi virus production in sheep choroid plexus cells. Vet. Microbiol. 1981, 6, 1–8. [Google Scholar] [CrossRef]
- Ryan, S.; Tiley, L.; McConnell, I.; Blacklaws, B. Infection of dendritic cells by the Maedi-Visna lentivirus. J. Virol. 2000, 74, 10096–10103. [Google Scholar] [CrossRef] [PubMed]
- Lechat, E.; Milhau, N.; Brun, P.; Bellaton, C.; Greenland, T.; Mornex, J.F.; le Jan, C. Goat endothelial cells may be infected in vitro by transmigration of caprine arthritis-encephalitis virus-infected leucocytes. Vet. Immunol. Immunopathol. 2005, 104, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Keele, B.F.; Giorgi, E.E.; Salazar-Gonzalez, J.F.; Decker, J.M.; Pham, K.T.; Salazar, M.G.; Sun, C.; Grayson, T.; Wang, S.; Li, H.; et al. Identification and characterization of transmitted and early founder virus envelopes in primary HIV-1 infection. Proc. Natl. Acad. Sci. USA. 2008, 105, 7552–7557. [Google Scholar] [CrossRef] [PubMed]
- Schuitemaker, H.; Koot, M.; Kootstra, N.A.; Dercksen, M.W.; de Goede, R.E.; van Steenwijk, R.P.; Lange, J.M.; Schattenkerk, J.K.; Miedema, F.; Tersmette, M. Biological phenotype of human immunodeficiency virus type 1 clones at different stages of infection: progression of disease is associated with a shift from monocytotropic to T-cell-tropic virus population. J. Virol. 1992, 66, 1354–1360. [Google Scholar] [PubMed]
- McCune, J.M. The dynamics of CD4+ T-cell depletion in HIV disease. Nature 2001, 410, 974–979. [Google Scholar] [CrossRef] [PubMed]
- Khan, K.A.; Abbas, W.; Varin, A.; Kumar, A.; Di Martino, V.; Dichamp, I.; Herbein, G. HIV-1 Nef interacts with HCV Core, recruits TRAF2, TRAF5 and TRAF6, and stimulates HIV-1 replication in macrophages. J. Innate Immun 2013, 5, 639–656. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.; Zhang, H.; Lopez, P.; Pardo, C.A.; Gartner, S. In vitro modeling of the HIV-1 macrophage reservoir. J. Leukoc. Biol. 2006, 80, 1127–1135. [Google Scholar] [CrossRef] [PubMed]
- Aquaro, S.; Calio, R.; Balzarini, J.; Bellocchi, M.; Garaci, E.; Perno, C. Macrophages and HIV infection: therapeutical approaches toward this strategic virus reservoir. Antiviral Res. 2002, 55, 209–225. [Google Scholar] [CrossRef]
- Crowe, S.; Zhu, T.; muller, W. The contribution of monocyte infection and trafficking to viral persistence, and maintenance of the viral reservoir in HIV infection. J. Leukoc. Biol. 2003, 74, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Lambotte, O.; Taoufik, Y.; de Goer, M.; Wallon, C.; Goujard, C.; Delfraissy, J. Detection of infectious HIV in circulating monocytes from patients on prolonged highly active antiretroviral therapy. J. Acquir. Immune Defic. Syndr. 2000, 23, 114–119. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Muthui, D.; Holte, S.; Nickle, D.; Feng, F.; Brodie, S.; Hwangbo, Y.; Mullins, J.I.; Corey, L. Evidence for Human Immunodeficiency Virus Type 1 Replication In Vivo in CD14+ Monocytes and Its Potential Role as a Source of Virus in Patients on Highly Active Antiretroviral Therapy. J. Virol. 2002, 76, 707–716. [Google Scholar] [CrossRef] [PubMed]
- Le Douce, V.; Herbein, G.; Rohr, O.; Schwartz, C. Molecular mechanisms of HIV-1 persistence in the monocyte-macrophage lineage. Retrovirology 2010. [Google Scholar] [CrossRef] [PubMed]
- Zink, M.C.; Yager, J.A.; Myers, J.D. Pathogenesis of caprine arthritis encephalitis virus. Cellular localization of viral transcripts in tissues of infected goats. Am. J. Pathol. 1990, 136, 843. [Google Scholar] [PubMed]
- Clements, J.E.; Zink, M.C.; Narayan, O.; Gabuzda, D. Lentiviral infection of macrophages. Immunol. Ser. 1994, 60, 589–600. [Google Scholar]
- Coleman, C.M.; Wu, L. HIV interactions with monocytes and dendritic cells: Viral latency and reservoirs. Retrovirology 2009. [Google Scholar] [CrossRef] [PubMed]
- Lore, K.; Smed-sorensen, A.; Vasudevan, J.; Mascola, J.; Koup, R.A. Myeloid and plasmacytoid dendritic cells transfer HIV-1 preferentially to antigen-specific CD4+ T cells. J. Exp. Med. 2005, 201, 2023–2033. [Google Scholar] [CrossRef] [PubMed]
- Smed-Sörensen, A.; Loré, K.; Vasudevan, J.; Louder, M.K.; Andersson, J.; Mascola, J.R.; Spetz, A.-L.; Koup, R.A. Differential susceptibility to human immunodeficiency virus type 1 infection of myeloid and plasmacytoid dendritic cells. J. Virol. 2005, 79, 8861–8869. [Google Scholar] [CrossRef] [PubMed]
- Burleigh, L.; Lozach, P.-Y.; Schiffer, C.; Staropoli, I.; Pezo, V.; Porrot, F.; Canque, B.; Virelizier, J.-L.; Arenzana-Seisdedos, F.; Amara, A. Infection of dendritic cells (DCs), not DC-SIGN-mediated internalization of human immunodeficiency virus, is required for long-term transfer of virus to T cells. J. Virol. 2006, 80, 2949–2957. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Kwon, D.S.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Middel, J.; Cornelissen, I.L.; Nottet, H.S.; KewalRamani, V.N.; Littman, D.R.; et al. DC-SIGN, a dendritic cell-specific HIV-1-binding protein that enhances trans-infection of T cells. Cell. 2000, 100, 587–597. [Google Scholar] [CrossRef]
- Evans, V.A.; Kumar, N.; Filali, A.; Procopio, F.A.; Yegorov, O.; Goulet, J.P.; Saleh, S.; Haddad, E.K.; da Fonseca Pereira, C.; Ellenberg, P.C.; et al. Myeloid dendritic cells induce HIV-1 latency in non-proliferating CD4+ T cells. PLoS Pathog. 2013, 9, e1003799. [Google Scholar] [CrossRef] [PubMed]
- Banks, K.L.; Adams, D.S.; McGuire, T.C.; Carlson, J. Experimental infection of sheep by caprine arthritis-encephalitis virus and goats by progressive pneumonia virus. Am. J. Vet. Res. 1983, 44, 2307–2311. [Google Scholar] [PubMed]
- Leroux, C.; Cruz, J.C.; Mornex, J.F. SRLVs: A genetic continuum of lentiviral species in sheep and goats with cumulative evidence of cross species transmission. Curr. HIV Res. 2010, 8, 94–100. [Google Scholar] [PubMed]
- Karr, B.M.; Chebloune, Y.; Leung, K.; Narayan, O. Genetic characterization of two phenotypically distinct North American ovine lentiviruses and their possible origin from caprine arthritis-encephalitis virus. Virology 1996, 225, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Shah, C.; Huder, J.B.; Boni, J.; Schonmann, M.; Muhlherr, J.; Lutz, H.; Schupbach, J. Direct evidence for natural transmission of small-ruminant lentiviruses of subtype A4 from goats to sheep and vice versa. J. Virol. 2004, 78, 7518–7522. [Google Scholar] [CrossRef] [PubMed]
- Guiguen, F.; Mselli-Lakhal, L.; Durand, J.; Du, J.; Favier, C.; Fornazero, C.; Grezel, D.; Balleydier, S.; Hausmann, E.; Chebloune, Y. Experimental infection of Mouflon-domestic sheep hybrids with caprine arthritis-encephalitis virus. Am. J. Vet. Res. 2000, 61, 456–461. [Google Scholar] [CrossRef]
- Morin, T.; Guiguen, F.; Bouzar, B.A.; Villet, S.; Greenland, T.; Grezel, D.; Gounel, F.; Gallay, K.; Garnier, C.; Durand, J.; et al. Clearance of a Productive Lentivirus Infection in Calves Experimentally Inoculated with Caprine Arthritis-Encephalitis Virus. J. Virol. 2003, 77, 6430–6437. [Google Scholar] [CrossRef] [PubMed]
- Erhouma, E.; Guiguen, F.; Chebloune, Y.; Gauthier, D.; Lakhal, L.M.; Greenland, T.; Mornex, J.F.; Leroux, C.; Alogninouwa, T. Small ruminant lentivirus proviral sequences from wild ibexes in contact with domestic goats. J. Gen. Virol. 2008, 89, 1478–1484. [Google Scholar] [CrossRef] [PubMed]
- Pisoni, G.; Bertoni, G.; Puricelli, M.; Maccalli, M.; Moroni, P. Demonstration of coinfection with and recombination by caprine arthritis-encephalitis virus and maedi-visna virus in naturally infected goats. J. Virol. 2007, 81, 4948–4955. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.S.; Klevjer-Anderson, P.; Carlson, J.L.; McGuire, T.C.; Gorham, J.R. Transmission and control of caprine arthritis-encephalitis virus. Am. J. Vet. Res. 1983, 44, 1670–1675. [Google Scholar] [PubMed]
- Knight, A.; Jokinen, M. Caprine arthritis-encephalitis. Compend. Contin. Educ. Pract. Vet. 1982, 4, 263–269. [Google Scholar]
- Lloyd, S. Goat medicine and surgery. Br. Vet. J. 1982, 138, 70–85. [Google Scholar] [PubMed]
- Sherman, D.M. CAE: Caprine arthritis encephalitis—a growing concern. DGJ 1983, 61, 93–95. [Google Scholar]
- Preziuso, S.; Renzoni, G.; Allen, T.E.; Taccini, E.; Rossi, G.; DeMartini, J.C.; Braca, G. Colostral transmission of maedi visna virus: sites of viral entry in lambs born from experimentally infected ewes. Vet. Microbiol. 2004, 104, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Cork, L.C.; Narayan, O.; Strandberg, J.; Clements, J.E.; Griffin, D. Viral leukoencephalomyelitis-arthritis of goats: Pathogenesis of the persistent viral infection. J. Neuropath. Exp. Neur. 1980. [Google Scholar] [CrossRef]
- Rimstad, E.; East, N.E.; Torten, M.; Higgins, J.; DeRock, E.; Pedersen, N.C. Delayed seroconversion following naturally acquired caprine arthritis-encephalitis virus infection in goats. Am. J. Vet. Res. 1993, 54, 1858–1862. [Google Scholar] [PubMed]
- Cork, L.C. Pathology and epidemiology of lentiviral infection of goats. In Maedi-Visna and Related Diseases; Pétursson, G., Hoff-Jørgensen, R., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1990; pp. 119–127. [Google Scholar]
- Daas, E.; Moudgil, T.; Meyer, R. Transient high levels of viremia in patients with primary human immunodeficiency virus type 1 infection. N. Engl. J. Med. 1991, 324, 961–964. [Google Scholar]
- Piatak, M.; saag, M.; yang, L. High levels of HIV-1 in plasma during all stages of infection determined by competitive PCR. Science 1993, 259, 1749–1754. [Google Scholar] [PubMed]
- Mehandru, S.; Poles, M.A.; Tenner-Racz, K.; Horowitz, A.; Hurley, A.; Hogan, C.; Boden, D.; Racz, P.; Markowitz, M. Primary HIV-1 infection is associated with preferential depletion of CD4+ T lymphocytes from effector sites in the gastrointestinal tract. J. Exp. Med. 2004, 200, 761–770. [Google Scholar] [CrossRef] [PubMed]
- Saksena, N.; Potter, S. Reservoirs of HIV-1 in vivo: implications for antiretroviral therapy. AIDS Rev. 2003, 5, 3–18. [Google Scholar] [PubMed]
- Busch, M.; El-Amad, Z.; Sheppard, H.; Ascher, M. Primary HIV-1 infection. N. Engl. J. Med. 1991, 325, 733–735. [Google Scholar]
- Clark, S.; Saag, M.; Decker, W. High titers of cytopathic virus in plasma of patients with symptomatic primary infection. N. Engl. J. Med. 1991, 324, 954–960. [Google Scholar] [CrossRef] [PubMed]
- Tindall, B.; Cooper, D. Primary HIV infection: Host responses and intervention strategies. AIDS 1991, 5, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Kingsley, L.; Armstrong, J.; Ding, M.; Cottrill, M.; Rinaldo, C. Enhanced expression of human immunodeficiency virus type 1 correlates with development of AIDS. Virology 1993, 196, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Henrad, D.; Philips, J.; Muenz, L.; Blattner, W.A.; Wiesner, D.; Eyster, M.E. Natural history of HIV-1 cell free viremia. J. Am. Med. Assoc. 1995, 274, 554–558. [Google Scholar] [CrossRef]
- Jurriaans, S.; van German, B.; Weverling, G.J.; van Strijp, D.; Nara, P.; Coutinho, R.; Koot, M.; Schuitemaker, H.; Goudsmit, J. The natural history of HIV-1 infection: Virus load and virus phenotype independent determinants of clinical course? Virology 1994, 204, 223–233. [Google Scholar] [CrossRef] [PubMed]
- Cozzi-Lepri, A.; Sabin, C.; Tezzotti, P. Is there a general tendency for rate of CD41 lymphocyte count decline to speed up during HIV infection? Evidence from the Italian Seroconversion Study. J. Infect. Dis. 1997, 175, 775–780. [Google Scholar] [CrossRef]
- Lang, W.; Perkins, H.; Anderson, R.; Royce, R.; Jewell, N. Patterns of T lymphocyte changes with human immunodeficiency virus infection: From seroconversion to the development of AIDS. J. Acquir. Immune Defic. Syndr. 1989, 2, 63–69. [Google Scholar] [PubMed]
- Phillips, A.N.; Lee, C.A.; Elford, J.; Janossy, G.; Timms, A.; Bofill, M.; Kernoff, P.B. Serial CD4 lymphocyte counts and development of AIDS. Lancet 1991, 337, 389–392. [Google Scholar] [PubMed]
- Touloumi, G.; Hatzakis, A.; Rosenberg, P.S.; O’Brien, T.R.; Goedert, J.J. Effects of age at seroconversion and baseline HIV-RNA level on the loss of CD41 cells among persons with hemophilia. AIDS 1998, 12, 1691–1697. [Google Scholar] [CrossRef] [PubMed]
- Lewthwaite, P.; Wilkins, E. Natural history of HIV/AIDS. Medicine 2005, 33, 10–13. [Google Scholar] [CrossRef]
- Lewthwaite, P.; Wilkins, E. Natural history of HIV/AIDS. Medicine 2009, 37, 333–337. [Google Scholar] [CrossRef]
- Dean, M.; Carrington, M.; Winkler, C.; Huttley, G.A.; Smith, M.W.; Allikmets, R.; Goedert, J.J.; Buchbinder, S.P.; Vittinghoff, E.; Gomperts, E.; et al. Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. hemophilia growth and development study, multicenter AIDS cohort study, multicenter hemophilia cohort study, San Francisco city cohort, ALIVE study. Science 1996, 273, 1856–1862. [Google Scholar] [PubMed]
- Al-Jabri, A.A. Mechanisms of host resistance against HIV infection and progression to AIDS. Sultan Qaboos Univ. Med. J. 2007, 7, 82–96. [Google Scholar] [PubMed]
- Crowe, S.; Carlin, J.; Steward, K.; Lucas, C.R. Predictive value of CD4 lymphocyte numbers for the development of opportunistic infections and malignacies in HIV-infected persons. J. Acquir. Immune Defic. Syndr. 1991, 4, 770–776. [Google Scholar] [PubMed]
- Hoover, D.R.; Rinaldo, C.; He, Y.; Phair, J.; Fahey, J.; Graham, N.M. Long-term survival without clinical AIDS after CD4+ cell fall below 200 × 106/L. AIDS 1995, 9, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Mocroft, A.; Johnson, M.; Sabin, C.; Lipman, M.; Elford, J.; Emery, V.; Morcinek, J.; Youle, M.; Janossy, G.; Lee, C.A. Staging system for clinical AIDS patients. Lancet 1995, 346, 12–17. [Google Scholar] [CrossRef]
- Orenstein, R. Presenting Syndromes of Human Immunodeficiency Virus; Elsevier: Amsterdam, The Netherlands, 2002; pp. 1097–1102. [Google Scholar]
- O’Sullivan, B.M.; Eaves, F.W.; Baxendell, S.A.; Rowan, K.J. Leucoencephalomyelitis of goat kids. Aust. Vet. J. 1978, 54, 479–483. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.S.; Crawford, T.B.; Klevjer-Anderson, P. A pathogenetic study of the early connective tissue lesions of viral caprine arthritis-encephalitis. Amer. J. Pathol. 1980, 99, 257–278. [Google Scholar]
- Sigurdsson, B.; Palsson, P.A. Visna of sheep. A slow demyelinating infection. Brit. J. Exp. Pathol. 1958, 39, 519–528. [Google Scholar]
- Ruprecht, R.M.; Baba, T.W.; Liska, V.; Ray, N.B.; Martin, L.N.; Murphey-Corb, M.; Rizvi, T.A.; Bernacky, B.J.; Bernacky, M.E.; McClure, H.M.; et al. Oral transmission of primate lentiviruses. J. Infect. Dis. 1999, 179, 408–412. [Google Scholar] [CrossRef] [PubMed]
- Parrish, N.F.; Gao, F.; Li, H.; Giorgi, E.E.; Barbian, H.J.; Parrish, E.H.; Zajic, L.; Iyer, S.S.; Decker, J.M.; Kumar, A.; et al. Phenotypic properties of transmitted founder HIV-1. Proc. Natl. Acad. Sci. USA 2013, 110, 6626–6633. [Google Scholar] [CrossRef] [PubMed]
- Pope, M.; Betjes, M.; Romani, N.; Hirmand, H.; Cameron, P.; Hoffman, L.; Gezelter, S.; Schuler, G.; Steinman, R. Conjugates of dendritic cells and memory T lymphocytes from skin facilitate productive infection with HIV-1. Cell. 1994, 78, 389–398. [Google Scholar] [CrossRef]
- Neutra, M.; Frey, A.; Kraehenbuhl, J. Epithelial M cells: Gateways for mucosal infection and immunization. Cell. 1996, 86, 345–348. [Google Scholar] [CrossRef]
- Hladik, F.; Hope, T.J. HIV infection of the genital mucosa in women. Curr. HIV/AIDS Rep. 2009, 6, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Stieh, D.J.; Maric, D.; Kelley, Z.L.; Anderson, M.R.; Hattaway, H.Z.; Beilfuss, B.A.; Rothwangl, K.B.; Veazey, R.S.; Hope, T.J. Vaginal challenge with an SIV-based dual reporter system reveals that infection can occur throughout the upper and lower female reproductive tract. PLoS Pathog. 2014. [Google Scholar] [CrossRef] [PubMed]
- Dandekar, S. Pathogenesis of HIV in the Gastrointestinal tract. Curr HIV/AIDS Rep. 2007, 4, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Veazey, R.S. Getting to the Guts of HIV Pathogenesis. J. Exp. Med. 2004, 200, 697–700. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, T.; Spencer, J. Lymphoid cells and tissues of the gastrointestinal tract; Cambridge University Press: Cambridge, UK, 1994; pp. 1–23. [Google Scholar]
- Schieferdecker, H.; Ullrich, R.; Hirseland, H.; Zeitz, M. T cell differentiation antigens on lymphocytes in the human intestinal lamina propria. J. Immunol. 1992, 149, 2816–2822. [Google Scholar] [PubMed]
- Brenchley, J.; Schacker, T.; Ruff, L.; Price, D.A.; Taylor, J.H.; Beilman, G.J.; Nguyen, P.L.; Khoruts, A.; Larson, M.; Haase, A.T.; et al. CD4+T cell depletion during all stages of HIV disease occurs predominantly in the gastrointestinal tract. J. Exp. Med. 2004, 200, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Veazey, R.S.; Marx, P.; Lackner, A. Importance of the state of activation and/or differentiation of CD4+ T cells in AIDS pathogenesis. Trends Immunol. 2002, 23, 128–129. [Google Scholar] [CrossRef]
- Mattapallil, J.; Douek, D.; Hill, B.; Nishimura, Y.; Martin, M.; Roederer, M. Massive infection and loss of memory CD4+ T cells in multiple tissues during acute SIV infection. Nature 2005, 434, 1093–1097. [Google Scholar] [CrossRef] [PubMed]
- Lackner, A.A.; Mohan, M.; Veazey, R.S. The Gastrointestinal Tract and AIDS Pathogenesis. Gastroenterology 2009, 136, 1966–1978. [Google Scholar] [CrossRef]
- Lackner, A.A.; Lederman, M.M.; Rodriguez, B. HIV pathogenesis: The host. Cold Spring Harb. Perspect. Med. 2012. [Google Scholar] [CrossRef] [PubMed]
- Katsikis, P.D.; Mueller, Y.M.; Villinger, F. The cytokine network of acute HIV Infection: A promising target for vaccines and therapy to reduce viral set-point? PLoS Pathog. 2011. [Google Scholar] [CrossRef] [PubMed]
- Deeks, S.G. HIV Infection, inflammation, immunosenescence, and aging. Annu. Rev. Med. 2011, 62, 141–155. [Google Scholar] [CrossRef] [PubMed]
- Elbirt, D.; Mahlab-Guri, K.; Bezalel-Rosenberg, S.; Gill, H.; Attali, M.; Asher, I. HIV-associated neurocognitive disorders (HAND). Isr. Med. Assoc. J. 2015, 17, 54–59. [Google Scholar]
- Albright, A.V.; Soldan, S.S.; Gonzalez-Scarano, F. Pathogenesis of human immunodeficiency virus-induced neurological disease. J. Neurovirol. 2003, 9, 222–227. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Scarano, F.; Martin-Garcia, J. The neuropathogenesis of AIDS. Nat. Rev. Immunol. 2005, 5, 69–81. [Google Scholar] [CrossRef] [PubMed]
- Kaul, M.; Lipton, S.A. Chemokines and activated macrophages in HIV gp120-induced neuronal apoptosis. Proc. Natl. Acad. Sci. USA 1999, 96, 8212–8216. [Google Scholar] [CrossRef] [PubMed]
- Lindl, K.A.; Marks, D.R.; Kolson, D.L.; Jordan-Sciutto, K.L. HIV-associated neurocognitive disorder: Pathogenesis and therapeutic opportunities. J. Neuroimmune Pharmacol. 2010, 5, 294–309. [Google Scholar] [CrossRef] [PubMed]
- Price, R.W.; Brew, B.; Sidtis, J.; Rosenblum, M.; Scheck, A.C.; Cleary, P. The brain in AIDS: Central nervous system HIV-1 infection and AIDS dementia complex. Science 1988, 239, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Valcour, V.; Sithinamsuwan, P.; Letendre, S.; Ances, B. Pathogenesis of HIV in the Central Nervous System. Curr. HIV/AIDS Rep. 2011, 8, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Fieni, F.; Rowe, J.; Van Hoosear, K.; Burucoa, C.; Oppenheim, S.; Anderson, G.; Murray, J.; BonDurant, R. Presence of caprine arthritis-encephalitis virus (CAEV) proviral DNA in genital tract tissues of superovulated dairy goat does. Theriogenology 2003, 59, 1515–1523. [Google Scholar] [CrossRef]
- Petursson, G.; Nathanson, N.; Georgsson, G.; Panitch, H.; Palsson, P.A. Pathogenesis of visna Sequential virologic, serologic and pathologic studies. Lab. Invest. 1976, 35, 402–412. [Google Scholar] [PubMed]
- Adams, D.S.; Crawford, T.B.; Klevjer-Anderson, P. A pathogenetic study of the early connective tissue lesions of viral caprine arthritis-encephalitis. Am. J. Pathol. 1980, 99, 257–278. [Google Scholar]
- Narayan, O.; Cork, L.C. Lentiviral diseases of sheep and goats: Chronic pneumonia leukoencephalomyelitis and arthritis. Rev. Infect. Dis. 1985, 7, 89–98. [Google Scholar] [CrossRef] [PubMed]
- Cork, L.C.; Davis, W.C. Ultrastructural features of viral leukoencephalomyelitis of goats. Lab. Invest. 1975, 32, 359–365. [Google Scholar] [PubMed]
- Ellis, T.M.; Robinson, W.F.; Wilcox, G.E. The pathology and aetiology of lung lesions in goats infected with caprine arthritis-encephalitis virus. Aust. Vet. J. 1988, 65, 69–73. [Google Scholar] [CrossRef] [PubMed]
- Sigurdsson, B.; Grimsson, H.; Palsson, P.A. Maedi, a chronic, progressive infection of sheep’s lungs. J. Infect. Dis. 1952, 90, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Cutlip, R.C.; Lehmkuhl, H.D.; Brogden, K.A.; Bolin, S.R. Mastitis associated with ovine progressive pneumonia virus infection in sheep. Am. J. Vet. Res. 1985, 46, 326–328. [Google Scholar] [PubMed]
- Clapham, P.R.; McKnight, A. HIV-1 receptors and cell tropism. Brit. Med. Bull. 2001, 58, 43–59. [Google Scholar] [CrossRef] [PubMed]
- Michael, N. Host genetic influences on HIV-1 pathogenesis. Curr. Opin. Immunol. 1999, 11, 466–474. [Google Scholar] [CrossRef]
- Crespo, H.; Jáuregui, P.; Glaria, I.; Sanjose, L.; Polledo, L.; García-Marin, J.; Luján, L.; de Andrés, D.; Amorena, B.; Reina, R. Mannose receptor may be involved in small ruminant lentivirus pathogenesis. Vet. Res. 2012. [Google Scholar] [CrossRef] [PubMed]
- Crespo, H.; Reina, R.; Glaria, I.; Ramírez, H.; de Andrés, X.; Jáuregui, P.; Luján, L.; Martinez- Pomares, L.; Amorena, B.; de Andrés, D. Identification of the ovine mannose receptor and its possible role in Visna/Maedi virus infection. Vet. Res. 2011. [Google Scholar] [CrossRef] [PubMed]
- Narayan, O.; Wolinsky, J.; Clements, J.E.; Strandberg, J.; Griffin, D.; Cork, L.C. Slow virus replication: The role of macrophages in the persistence and expression of visna viruses of sheep and goats. J. Gen. Virol. 1982, 59, 345–356. [Google Scholar] [CrossRef]
- Chen, M.; Westmoreland, S.; Ryzhova, E.; MArtin, G.J.; Soldan, S.; Lackner, A.A.; González-Scarano, F. Simian immunodeficiency virus envelope compartmentalizes in brain regions independent of neuropathology. J. Neurovirol. 2006, 12, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Hudson, L.; Tompkins, M.; Vahlenkamp, T.; Meeker, R. Compartmentalization and evolution of feline immunodeficiency virus between the central nervous system and periphery following intracerebroventricular or systemic inoculation. J. Neurovirol. 2006, 12, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Zárate, S.; Kosakovsky, P.; Shapshak, F. Comparative study of methods for detecting sequence compartmentalization in human immunodeficiency virus type 1. J. Virol. 2007, 81, 6643–6651. [Google Scholar] [CrossRef]
- Ramirez, H.; Reina, R.; Bertolotti, L.; Cenoz, A.; Hernandez, M.; San Roman, B.; Glaria, I.; de Andrés, X.; Crespo, H.; Jauregui, P. Study of compartmentalization in the visna clinical form of small ruminant lentivirus infection in sheep. BMC Vet. Res. 2012. [Google Scholar] [CrossRef] [PubMed]
- Korber, B.; Kunstman, K.; Patterson, B.; Furtado, M.; McEvilly, M.; Levy, R.; Wolinsky, S. Genetic differences between blood- and brain-derived viral sequences from human immunodeficiency virus type-1 infected patients: Evidence of conserved elements in the V3 region of the Envelope protein of brain-derived sequences. J. Virol. 1994, 68, 7467–7481. [Google Scholar]
- Hotzel, I.; Cheevers, W. Sequence similarity between the envelope surface unit (SU) glycoproteins of primate and small ruminant lentiviruses. Virus Res. 2000, 69, 47–54. [Google Scholar] [CrossRef]
- Mwaengo, D.; Grant, R.; DeMartini, J.; Carlson, J. Envelope glycoprotein nucleotide sequence and genetic characterization of North American ovine lentiviruses. Virology 1997, 238, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Blackard, J.T. HIV compartmentalization: a review on a clinically important phenomenon. Curr HIV Res. 2012, 10, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Deeks, S.G.; Autran, B.; Berkhout, B.; Benkirane, M.; Cairns, S.; Chomont, N.; Chun, T.W.; Churchill, M.; di Mascio, M.; Katlama, C.; et al. Towards an HIV cure: A global scientific strategy. Nat. Rev. Immunol. 2012, 12, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Deeks, S.G.; Wrin, T.; Liegler, T.; Hoh, R.; Hayden, M.; Barbour, J.D.; Hellmann, N.S.; Petropoulos, C.J.; McCune, J.M.; Hellerstein, M.K.; et al. Virologic and immunologic consequences of discontinuing combination antiretroviral-drug therapy in HIV-infected patients with detectable viremia. New Engl. J. Med. 2001, 344, 472–480. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, S.K.; Rezk, N.L.; Ince, W.L.; Cheema, M.; Zhang, L.; Su, L.; Swanstrom, R.; Kashuba, A.D.; Margolis, D.M. Suppression of human immunodeficiency virus type 1 (HIV-1) viremia with reverse transcriptase and integrase inhibitors, CD4+ T-cell recovery, and viral rebound upon interruption of therapy in a new model for HIV treatment in the humanized Rag2-/-γc-/- mouse. J. Virol. 2009, 83, 8254–8258. [Google Scholar] [PubMed]
- Chun, T.W.; Engel, D.; Berrey, M.M.; Shea, T.; Corey, L.; Fauci, A.S. Early establishment of a pool of latently infected, resting CD4(+) T cells during primary HIV-1 infection. Proc. Natl. Acad. Sci. USA 1998, 95, 8869–8873. [Google Scholar] [CrossRef] [PubMed]
- Davis, L.E.; Hjelle, B.L.; Miller, V.E.; Palmer, D.L.; Llewellyn, A.L.; Merlin, T.L.; Young, S.A.; Mills, R.G.; Wachsman, W.; Wiley, C.A. Early viral brain invasion in iatrogenic human immunodeficiency virus infection. Neurology 1992, 42, 1736–1739. [Google Scholar] [CrossRef] [PubMed]
- Chun, T.-W.; Fauci, A.S. Latent reservoirs of HIV: obstacles to the eradication of virus. Proc. Natl. Acad. Sci. USA 1999, 96, 10958–10961. [Google Scholar] [CrossRef] [PubMed]
- Chun, T.-W.; Fauci, A.S. HIV reservoirs: pathogenesis and obstacles to viral eradication and cure. AIDS 2012, 26, 1261–1268. [Google Scholar] [CrossRef] [PubMed]
- Samuel, W.; Warner, G. Host factor regulating post integration latency of HIV. Trends Microbiol. 2005, 13, 137–139. [Google Scholar]
- du Chéné, I.; Basyuk, E.; Lin, Y.-L.; Triboulet, R.; Knezevich, A.; Chable-Bessia, C.; Mettling, C.; Baillat, V.; Reynes, J.; Corbeau, P.; et al. Suv39H1 and HP1γ are responsible for chromatin-mediated HIV-1 transcriptional silencing and post-integration latency. EMBO J. 2007, 26, 424–435. [Google Scholar] [CrossRef] [PubMed]
- Van Lint, C. Molecular control of HIV-1 postintegration latency: Implications for therapeutic strategies. Retrovirology 2012. [Google Scholar] [CrossRef]
- Karn, J.; Stoltzfus, C.M. Transcriptional and posttranscriptional regulation of HIV-1 gene expression. Cold Spring Harb. Perspect. Med. 2012. [Google Scholar] [CrossRef] [PubMed]
- Shan, L.; Siliciano, R.F. From reactivation of latent HIV-1 to elimination of the latent reservoir: The presence of multiple barriers to viral eradication. Bioessays 2013, 35, 544–552. [Google Scholar] [CrossRef] [PubMed]
- Siliciano, J.D.; Siliciano, R.F. Recent developments in the search for a cure for HIV-1 infection: Targeting the latent reservoir for HIV-1. J. Allergy Clin. Immun. 2014, 134, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Lassen, K.; Han, Y.; Zhou, Y.; Siliciano, J.; Siliciano, R.F. The multifactorial nature of HIV-1 latency. Trends Mol. Med. 2004, 10, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Sharp, P. Novel mechanisms and factors for regulation by HIV-1 Tat. EMBO J. 1995, 14, 321–328. [Google Scholar] [PubMed]
- Jordan, A.; Defechereux, P.; Verdin, E. The site of HIV-1 integration in the human genome determines basal transcriptional activity and response to Tat transactivation. EMBO J. 2001, 20, 1726–1738. [Google Scholar] [CrossRef] [PubMed]
- Osorio, A.A.; Munoz, A.; Torres-Romero, D.; Bedoya, L.M.; Perestelo, N.R.; Jimenez, I.A.; Alcami, J.; Bazzocchi, I.L. Olean-18-ene triterpenoids from Celastraceae species inhibit HIV replication targeting NF-kB and Sp1 dependent transcription. Eur. J. Med. Chem. 2012, 52, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Dandekar, D.H.; Ganesh, K.N.; Mitra, D. HIV-1 Tat directly binds to NFκB enhancer sequence: Role in viral and cellular gene expression. Nucleic Acids Res. 2004, 32, 1270–1278. [Google Scholar] [CrossRef] [PubMed]
- Margolis, D.M.; Somasundaran, M.; Green, M.R. Human transcription factor YY1 represses human immunodeficiency virus type 1 transcription and virion production. J. Virol. 1994, 68, 905–910. [Google Scholar] [PubMed]
- Tyagi, M.; Karn, J. CBF-1 promotes transcriptional silencing during the establishment of HIV-1 latency. EMBO J. 2007, 26, 4985–4995. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Lin, Y.; An, W.; Xu, J.; yahg, H.; O’Connell, K.; Dordai, D.; Boeke, J.; Siliciano, J.D.; Siliciano, R.F. Orientation-dependent regulation of integrated HIV-1 expression by host gene transcriptional readthrough. Cell Host Microbe 2008, 4, 134–146. [Google Scholar] [CrossRef] [PubMed]
- Powell, D.M.; Amaral, M.C.; Wu, J.Y.; Maniatis, T.; Greene, W.C. HIV Rev-dependent binding of SF2/ASF to the Rev response element: possible role in Rev-mediated inhibition of HIV RNA splicing. Proc. Natl. Acad. Sci. USA 1997, 94, 973–978. [Google Scholar] [CrossRef] [PubMed]
- Brady, J.; Kashanchi, F. Tat gets the “green” light on transcription initiation. Retrovirology 2005. [Google Scholar] [CrossRef] [PubMed]
- Barboric, M.; Peterlin, B.M. A New Paradigm in Eukaryotic Biology: HIV Tat and the Control of Transcriptional Elongation. PloS Biol. 2005. [Google Scholar] [CrossRef] [PubMed]
- Haase, A.; Stowring, L.; Narayan, O.; Griffin, D.; Price, D. Slow persistant infection caused by visna virus role of host restriction. Science 1977, 195, 175–177. [Google Scholar] [CrossRef] [PubMed]
- Thormar, H. Physical, Chemical, Biological Properties of Visna Virus and Its Relationship to Other Animal Viruses. In Slow, Latent and Temprate Virus Infection; The National Institute of Neurological Disorders and Blindness: Washington, DC, USA, 1966; pp. 335–340. [Google Scholar]
- Thormar, H. Maedi-visna virus and its relationship to human immunodeficiency virus. AIDS Rev. 2005, 7, 233–245. [Google Scholar] [PubMed]
- Pereira, L.A.; Bentley, K.; Peeters, A.; Churchill, M.J.; Deacon, N.J. A compilation of cellular transcription factor interactions with the HIV-1 LTR promoter. Nucleic Acids Res. 2000, 28, 663–668. [Google Scholar] [CrossRef] [PubMed]
- Gracia, J.; Harrich, D.; Soultanakis, E.; Wu, F.; Mitsuyasu, R.; Gaynor, R. Human immunodeficiency virus type 1 LTR TATA and TAR region sequences required for transcriptional regulation. EMBO J. 1989, 8, 765–778. [Google Scholar]
- Harrich, D.; Gracia, J.; Mitsuyasu, R.; Gaynor, R. TAR independent activation of the human immunodeficiency virus in phorbol ester stimulated T lymphocytes. EMBO J. 1990, 9, 4417–4423. [Google Scholar] [PubMed]
- Olsen, H.S.; Rosen, C. Contribution of the TATA motif to Tat-mediated transcriptional activation of the human immunodeficiency virus gene expression. J. Virol. 1992, 66, 5594–5597. [Google Scholar] [PubMed]
- Danino, Y.M.; Even, D.; Ideses, D.; Juven-Gershon, T. The core promoter: At the heart of gene expression. Biochim. Biophys. Acta 2015, 1849, 1116–1131. [Google Scholar] [CrossRef] [PubMed]
- Dahabieh, M.S.; Ooms, M.; Malcolm, T.; Simon, V.; Sadowski, I. Identification and functional analysis of a second RBF-2 binding site within the HIV-1 promoter. Virology 2011, 418, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, E.; Doyle, M.C.; Nzaramba, I.; Magdzinski, A.; Dumais, N.; Bell, B. CTGC motifs within the HIV core promoter specify Tat-responsive pre-initiation complexes. Retrovirology 2012. [Google Scholar] [CrossRef] [PubMed]
- Miller-Jensen, K.; Skupsky, R.; Shah, P.S.; Arkin, A.P.; Schaffer, D.V. Genetic selection for context-dependent stochastic phenotypes: Sp1 and TATA mutations increase phenotypic noise in HIV-1 gene expression. PLoS Comput. Biol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Duverger, A.; Wolschendorf, F.; Zhang, M.; Wagner, F.; Hatcher, B.; Jones, J.; Cron, R.Q.; van der Sluis, R.M.; Jeeninga, R.E.; Berkhout, B.; et al. An AP-1 binding site in the enhancer/core element of the HIV-1 promoter controls the ability of HIV-1 to establish latent infection. J. Virol. 2013, 87, 2264–2277. [Google Scholar] [CrossRef] [PubMed]
- Duverger, A.; Jones, J.; May, J.; Bibollet-Ruche, F.; Wagner, F.A.; Cron, R.Q.; Kutsch, O. Determinants of the establishment of human immunodeficiency virus type 1 latency. J. Virol. 2009, 83, 3078–3093. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.A.; Kadonaga, J.T.; Luciw, P.A.; Tjian, R. Activation of the AIDS retrovirus promoter by the cellular transcription factor, Sp1. Science 1986, 232, 755–759. [Google Scholar] [CrossRef] [PubMed]
- Rittner, K. The human immunodeficiency virus long terminal repeat includes a specialised initiator element which is required for Tat responsive transcription. J. Mol. Biol. 1995, 248, 562–580. [Google Scholar] [CrossRef] [PubMed]
- Ross, E.K.; Buckler-White, A.J.; Rabson, A.B.; Englund, G.; Martin, M.A. Contribution of NF-κB and Sp1 binding motifs to the replicative capacity of human immunodeficiency virus type 1: Distinct patterns of viral growth are determined by T-cell types. J. Virol. 1991, 65, 4350–4358. [Google Scholar] [PubMed]
- Dahmus, M. Phosphorylation of C-terminal domain of RNA polymerase II. Biochim. Biophys. Acta 1995, 171–182. [Google Scholar] [CrossRef]
- Kim, Y.K.; Bourgeois, C.F.; Isel, C.; Churcher, M.J.; Karn, J. Phosphorylation of the RNA polymerase II carboxyl-terminal domain by CDK9 is directly responsible for human immunodeficiency virus type 1 Tat-activated transcriptional elongation. Mol. Biol. Cell. 2002, 22, 4622–4637. [Google Scholar] [CrossRef]
- Guo, J.; Price, D.H. RNA polymerase II transcription elongation control. Chem. Rev. 2013, 113, 8583–8603. [Google Scholar] [CrossRef]
- Jadlowsky, J.K.; Wong, J.Y.; Graham, A.C.; Dobrowolski, C.; Devor, R.L.; Adams, M.D.; Fujinaga, K.; Karn, J. Negative elongation factor is required for the maintenance of proviral latency but does not induce promoter-proximal pausing of RNA polymerase II on the HIV long terminal repeat. Mol. Biol. Cell. 2014, 34, 1911–1928. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, M.; Schiralli Lester, G.M.; Lee, C.; Missra, A.; Wasserman, G.A.; Steffen, M.; Gilmour, D.S.; Henderson, A.J. Negative elongation factor (NELF) coordinates RNA polymerase II pausing, premature termination, and chromatin remodeling to regulate HIV transcription. J. Biol. Chem. 2013, 288, 25995–26003. [Google Scholar] [CrossRef] [PubMed]
- Cheng, B.; Price, D.H. Analysis of factor interactions with RNA polymerase II elongation complexes using a new electrophoretic mobility shift assay. Nucleic Acids Res. 2008, 36, e135. [Google Scholar] [CrossRef] [PubMed]
- Missra, A.; Gilmour, D.S. Interactions between DSIF (DRB sensitivity inducing factor), NELF (negative elongation factor), and the Drosophila RNA polymerase II transcription elongation complex. Proc. Natl. Acad. Sci. USA 2010, 107, 11301–11306. [Google Scholar] [CrossRef] [PubMed]
- Pagano, J.M.; Kwak, H.; Waters, C.T.; Sprouse, R.O.; White, B.S.; Ozer, A.; Szeto, K.; Shalloway, D.; Craighead, H.G.; Lis, J.T. Defining NELF-E RNA binding in HIV-1 and promoter-proximal pause regions. PLoS Genet. 2014. [Google Scholar] [CrossRef]
- Feng, S.; Holland, E.C. HIV-1 Tat trans-activation requires the loop sequence within tar. Nature 1988, 334, 165–167. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Babayeva, N.D.; Suwa, Y.; Baranovskiy, A.G.; Price, D.H.; Tahirov, T.H. Crystal structure of HIV-1 Tat complexed with human P-TEFb and AFF4. Cell. Cycle 2014, 13, 1788–1797. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, C.; Rice, A. Lentivirus Tat proteins specifically associate with a cellular protein- kinase, TAK, that hyperphosphorylates the carboxyl-terminal domain of the large subunit of RNA polymerase II: Candidate for a Tat cofactor. J. Virol. 1995, 69, 1612–1620. [Google Scholar] [PubMed]
- Montanuy, I.; Torremocha, R.; Hernandez-Munain, C.; Sune, C. Promoter influences transcription elongation: TATA-box element mediates the assembly of processive transcription complexes responsive to cyclin-dependent kinase 9. J. Biol. Chem. 2008, 283, 7368–7378. [Google Scholar] [CrossRef] [PubMed]
- Wei, P.; Garber, M.E.; Fang, S.M.; Fischer, W.H.; Jones, K.A. A novel CDK9-associated C-type cyclin interacts directly with HIV-1 Tat and mediates its high-affinity, loop-specific binding to TAR RNA. Cell 1998, 92, 451–462. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, T.; Price, D.H. RNA polymerase II elongation control. Annu. Rev. Biochem. 2012, 81, 119–143. [Google Scholar] [CrossRef] [PubMed]
- Itzen, F.; Greifenberg, A.K.; Bosken, C.A.; Geyer, M. Brd4 activates P-TEFb for RNA polymerase II CTD phosphorylation. Nucleic Acids Res. 2014, 42, 7577–7590. [Google Scholar] [CrossRef] [PubMed]
- Marshall, N.F.; Price, D. Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol. Biol. Cell. 1992, 12, 2078–2090. [Google Scholar]
- Fujinaga, K. Dynamics of human immunodeficiency virus transcription: P-TEFb phosphorylates RD and dissociates negative effectors from the transactivation response element. Mol. Biol. Cell. 2004, 24, 787–795. [Google Scholar] [CrossRef]
- Mbonye, U.R.; Gokulrangan, G.; Datt, M.; Dobrowolski, C.; Cooper, M.; Chance, M.R.; Karn, J. Phosphorylation of CDK9 at Ser175 enhances HIV transcription and is a marker of activated P-TEFb in CD4(+) T lymphocytes. PLoS Pathog. 2013. [Google Scholar] [CrossRef] [PubMed]
- Battistini, A.; Sgarbanti, M. HIV-1 Latency: An Update of Molecular Mechanisms and Therapeutic Strategies. Viruses 2014, 6, 1715–1758. [Google Scholar] [CrossRef] [PubMed]
- Mbonye, U.; Karn, J. Transcriptional control of HIV latency: Cellular signaling pathways, epigenetics, happenstance and the hope for a cure. Virology 2014, 454–455, 328–339. [Google Scholar] [CrossRef] [PubMed]
- Karn, J. Tat, A Novel Regulator of HIV Transcription and Latency. In HIV Sequence Compendium 2000; Kuiken, C.M.F., Foley, B., Mellors, J.W., Hahn, B., Mullins, J., Marx, P., Wolinsky, S., Eds.; Theoretical Biology and Biophysics Group Los Alamos National Laboratory: Los Alamos, NM, USA, 2000; pp. 2–18. [Google Scholar]
- Karn, J. Tackling Tat. J. Mol. Biol. 1999, 293, 235–254. [Google Scholar] [CrossRef] [PubMed]
- Siliciano, R.F.; Greene, W.C. HIV Latency. Cold Spring Harb. Perspect. Med. 2011. [Google Scholar] [CrossRef] [PubMed]
- Strebel, K. Virus-host interactions: role of HIV proteins Vif, Tat, and Rev. AIDS 2003, 17, 25–34. [Google Scholar] [CrossRef]
- Taube, R.; Peterlin, M. Lost in Transcription: Molecular Mechanisms that Control HIV Latency. Viruses 2013, 5, 902–927. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, M.; Bukrinsky, M. Human immunodeficiency virus (HIV) latency: The major hurdle in HIV eradication. Mol. Med. 2012, 18, 1096–1108. [Google Scholar] [CrossRef] [PubMed]
- Schiralli Lester, G.M.; Henderson, A.J. Mechanisms of HIV transcriptional regulation and their contribution to latency. Mol. Biol. Int. 2012, 2012, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ruelas, D.S.; Greene, W.C. An integrated overview of HIV-1 latency. Cell 2013, 155, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Gdovin, S.; Clements, J.E. Molecular mechanisms of visna virus Tat: Identification of the targets for transcriptional activation and evidence for a post-transcriptional effect. Virology 1992, 188, 438–450. [Google Scholar] [CrossRef]
- Carruth, L.M.; Morse, B.A.; Clements, J.E. The leucine domain of the visna virus Tat protein mediates targeting to an AP-1 site in the viral long terminal repeat. J. Virol. 1996, 70, 4338–4344. [Google Scholar] [PubMed]
- Morse, B.A.; Carruth, L.M.; Clements, J.E. Targeting of the visna virus Tat protein to AP-1 sites: Interactions with the bZIP domains of fos and jun in vitro and in vivo. J. Virol. 1999, 73, 37–45. [Google Scholar] [PubMed]
- Barber, S.A.; Bruett, L.; Douglass, B.R.; Herbst, D.S.; Zink, M.C.; Clements, J.E. Visna virus-induced activation of MAPK is required for virus replication and correlates with virus-induced neuropathology. J. Virol. 2002, 76, 817–828. [Google Scholar] [CrossRef] [PubMed]
- Hess, J.L.; Small, J.; Clements, J.E. Sequences in the visna virus long terminal repeat that control transcriptional activity and respond to viral trans-activation: involvement of AP-1 sites in basal activity and trans-activation. J. Virol. 1989, 63, 3001–3015. [Google Scholar] [PubMed]
- Harmache, A.; Vitu, C.; Russo, P.; Bouyac, M.; Hieblot, C.; Peveri, P.; Vigne, R.; Suzan, M. The caprine arthritis encephalitis virus tat gene is dispensable for efficient viral replication in vitro and in vivo. J. Virol. 1995, 69, 5445–5454. [Google Scholar] [PubMed]
- Murphy, B.G.; Hotzel, I.; Jasmer, D.P.; Davis, W.C.; Knowles, D. TNFalpha and GM-CSF-induced activation of the CAEV promoter is independent of AP-1. Virology 2006, 352, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Sepp, T.; Tong-Starksen, S.E. STAT1 pathway is involved in activation of caprine arthritis-encephalitis virus long terminal repeat in monocytes. J. Virol. 1997, 71, 771–777. [Google Scholar] [PubMed]
- Guo, D.; Dunbar, J.D.; Yang, C.H.; Pfeffer, L.M.; Donner, D.B. Induction of Jak/STAT signaling by activation of the type 1 TNF receptor. J. Immunol. 1998, 160, 2742–2750. [Google Scholar] [PubMed]
- Jackson, S.H.; Yu, C.R.; Mahdi, R.M.; Ebong, S.; Egwuagu, C.E. Dendritic cell maturation requires STAT1 and is under feedback regulation by suppressors of cytokine signaling. J. Immunol. 2004, 172, 2307–2315. [Google Scholar] [CrossRef] [PubMed]
- Coccia, E.M.; Del Russo, N.; Stellacci, E.; Testa, U.; Marziali, G.; Battistini, A. STAT1 activation during monocyte to macrophage maturation: role of adhesion molecules. Int. Immunol. 1999, 11, 1075–1083. [Google Scholar] [CrossRef] [PubMed]
- Reina, R.; Grego, E.; Bertolotti, L.; De Meneghi, D.; Rosati, S. Genome analysis of small-ruminant lentivirus genotype E: a caprine lentivirus with natural deletions of the dUTPase subunit, vpr-like accessory gene, and 70-base-pair repeat of the U3 region. J. Virol. 2009, 83, 1152–1155. [Google Scholar] [CrossRef] [PubMed]
- Barros, S.C.; Andresdottir, V.; Fevereiro, M. Cellular specificity and replication rate of Maedi Visna virus in vitro can be controlled by LTR sequences. Arch. Virol. 2005, 150, 201–213. [Google Scholar] [CrossRef] [PubMed]
- Oskarsson, T.; Hreggvidsdottir, H.S.; Agnarsdottir, G.; Matthiasdottir, S.; Ogmundsdottir, M.H.; Jonsson, S.R.; Georgsson, G.; Ingvarsson, S.; Andresson, O.S.; Andresdottir, V. Duplicated sequence motif in the long terminal repeat of maedi-visna virus extends cell tropism and is associated with neurovirulence. J. Virol. 2007, 81, 4052–4057. [Google Scholar] [CrossRef] [PubMed]
- Agnarsdottir, G.; Thorsteinsdottir, H.; Oskarsson, T.; Matthiasdottir, S.; Haflidadottir, B.S.; Andresson, O.S.; Andresdottir, V. The long terminal repeat is a determinant of cell tropism of maedi-visna virus. J. Gen. Virol. 2000, 81, 1901–1905. [Google Scholar] [CrossRef] [PubMed]
- Murphy, B.; McElliott, V.; Vapniarsky, N.; Oliver, A.; Rowe, J. Tissue tropism and promoter sequence variation in caprine arthritis encephalitis virus infected goats. Virus Res. 2010, 151, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, A.O.; Barr, B.; Gomez-Lucia, E.; Murphy, B. A polytropic caprine arthritis encephalitis virus promoter isolated from multiple tissues from a sheep with multisystemic lentivirus-associated inflammatory disease. Viruses 2013, 5, 2005–2018. [Google Scholar] [CrossRef] [PubMed]
- Murphy, B.; Hillman, C.; Castillo, D.; Vapniarsky, N.; Rowe, J. The presence or absence of the gamma-activated site determines IFN gamma-mediated transcriptional activation in CAEV promoters cloned from the mammary gland and joint synovium of a single CAEV-infected goat. Virus Res. 2012, 163, 537–545. [Google Scholar] [CrossRef] [PubMed]
- Mbonye, U.; Karn, J. Control of HIV latency by epigenetic and non-epigenetic mechanisms. Curr. HIV Res. 2011, 9, 554–567. [Google Scholar] [PubMed]
- Narlikar, G.J.; Fan, H.-Y.; Kingston, R.E. Cooperation between complexes that regulate chromatin structure and transcription. Cell. 2002, 108, 475–487. [Google Scholar] [CrossRef]
- Wolffe, A. Nucleosome positioning and modification: chromatin structures that potentiate transcription. Trends Biochem. Sci. 1994, 19, 240–244. [Google Scholar] [CrossRef]
- Richman, D.D.; Margolis, D.M.; Delaney, M.; Greene, W.C.; Hazuda, D.; Pomerantz, R.J. The challenge of finding a cure for HIV infection. Science 2009, 323, 1304–1307. [Google Scholar] [PubMed]
- Van Lint, C.; Emiliani, S.; Ott, M.; Verdin, E. Transcriptional activation and chromatin remodeling of the HIV-1 promoter in response to histone acetylation. EMBO J. 1996, 15, 1112–1120. [Google Scholar] [PubMed]
- Verdin, E. DNase I-hypersensitive sites are associated with both long terminal repeats and with the intragenic enhancer of integrated human immunodeficiency virus type 1. J. Virol. 1991, 65, 6790–6799. [Google Scholar] [PubMed]
- Verdin, E.; Paras, P., Jr.; Van Lint, C. Chromatin disruption in the promoter of human immunodeficiency virus type 1 during transcriptional activation. EMBO J. 1993, 12, 3249–3259. [Google Scholar] [PubMed]
- Coull, J.J.; Romerio, F.; Sun, J.-M.; Volker, J.L.; Galvin, K.M.; Davie, J.R.; Shi, Y.; Hansen, U.; Margolis, D.M. The Human Factors YY1 and LSF Repress the Human Immunodeficiency Virus Type 1 Long Terminal Repeat via Recruitment of Histone Deacetylase 1. J. Virol. 2000, 74, 6790–6799. [Google Scholar] [CrossRef] [PubMed]
- Imai, K.; Okamoto, T. Transcriptional Repression of Human Immunodeficiency Virus Type 1 by AP-4. J. Biol. Chem. 2006, 281, 12495–12505. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Espeseth, A.; Hazuda, D.J.; Margolis, D.M. c-Myc and Sp1 contribute to proviral latency by recruiting histone deacetylase 1 to the human immunodeficiency virus type 1 promoter. J. Virol. 2007, 81, 10914–10923. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Margolis, D.M. Counterregulation of chromatin deacetylation and histone deacetylase occupancy at the integrated promoter of human immunodeficiency virus type 1 (HIV-1) by the HIV-1 repressor YY1 and HIV-1 activator Tat. Mol. Biol. Cell 2002, 22, 2965–2973. [Google Scholar] [CrossRef]
- Williams, S.; Chen, L.; Kwon, C.; Ruiz-Jarabo, E.; Greene, W. NF-κB p50 promotes HIV latency through HDAC recruitment and repression of transcriptional initiation. EMBO J. 2006, 25, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Keedy, K.S.; Archin, N.M.; Gates, A.T.; Espeseth, A.; Hazuda, D.J.; Margolis, D.M. A limited group of class I histone deacetylases acts to repress human immunodeficiency virus type 1 expression. J. Virol. 2009, 83, 4749–4756. [Google Scholar] [CrossRef] [PubMed]
- Nabel, G.; Baltimore, D. An inducible transcription factor activates expression of human immunodeficiency virus in T cells. Nature 1987, 326, 711–713. [Google Scholar] [CrossRef] [PubMed]
- Dorr, A.; Kiermer, V.; Pedal, A.; Rackwitz, H.-R.; Henklein, P.; Schubert, U.; Zhou, M.-M.; Verdin, E.; Ott, M. Transcriptional synergy between Tat and PCAF is dependent on the binding of acetylated Tat to the PCAF bromodomain. EMBO J. 2002, 21, 2715–2723. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudi, T.; Parra, M.; Vries, R.G.J.; Kauder, S.E.; Verrijzer, C.P.; Ott, M.; Verdin, E. The SWI/SNF chromatin-remodeling complex is a cofactor for Tat transactivation of the HIV promoter. J. Biol. Chem. 2006, 281, 19960–19968. [Google Scholar] [CrossRef]
- Tréand, C.; du Chéné, I.; Brès, V.; Kiernan, R.; Benarous, R.; Benkirane, M.; Emiliani, S. Requirement for SWI/SNF chromatin-remodeling complex in Tat-mediated activation of the HIV-1 promoter. EMBO J. 2006, 25, 1690–1699. [Google Scholar] [PubMed]
- Agbottah, E.; Deng, L.; Dannenberg, L.O.; Pumfery, A.; Kashanchi, F. Effect of SWI/SNF chromatin remodeling complex on HIV-1 Tat activated transcription. Retrovirology 2006. [Google Scholar] [CrossRef] [PubMed]
- Ott, M.; Geyer, M.; Zhou, Q. The control of HIV transcription: Keeping RNA Polymerase II on track. Cell Host Microbe 2011, 10, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Col, E. The Histone Acetyltransferase, hGCN5, interacts with and acetylates the HIV transactivator, Tat. J. Biol. Chem. 2001, 276, 28179–28184. [Google Scholar] [CrossRef] [PubMed]
- Pagans, S.; Pedal, A.; North, B.J.; Kaehlcke, K.; Marshall, B.L.; Dorr, A.; Hetzer-Egger, C.; Henklein, P.; Frye, R.; McBurney, M.W.; et al. SIRT1 Regulates HIV Transcription via Tat Deacetylation. PloS Biol. 2005. [Google Scholar] [CrossRef] [PubMed]
- Sakane, N.; Kwon, H.-S.; Pagans, S.; Kaehlcke, K.; Mizusawa, Y.; Kamada, M.; Lassen, K.G.; Chan, J.; Greene, W.C.; Schnoelzer, M.; et al. Activation of HIV Transcription by the Viral Tat Protein Requires a Demethylation Step Mediated by Lysine-specific Demethylase 1 (LSD1/KDM1). PLoS Pathog. 2011. [Google Scholar] [CrossRef] [PubMed]
- Ott, M.; Dorr, A.; Hetzer-Egger, C.; Kaehlcke, K.; Schnolzer, M.; Henklein, P.; Cole, P.; Zhou, M.-M.; Verdin, E. Tat acetylation: a regulatory switch between early and late phases in HIV transcription elongation. Novartis Found. Symp. 2004, 259, 182–193. [Google Scholar] [PubMed]
- Pagans, S.; Kauder, S.E.; Kaehlcke, K.; Sakane, N.; Schroeder, S.; Dormeyer, W.; Trievel, R.C.; Verdin, E.; Schnolzer, M.; Ott, M. The Cellular lysine methyltransferase Set7/9-KMT7 binds HIV-1 TAR RNA, monomethylates the viral transactivator Tat, and enhances HIV transcription. Cell. Host Microbe 2010, 7, 234–244. [Google Scholar] [CrossRef] [PubMed]
- Sabo, A.; Lusic, M.; Cereseto, A.; Giacca, M. Acetylation of conserved lysines in the catalytic core of cyclin-dependent kinase 9 inhibits kinase activity and regulates transcription. Mol. Biol. Cell. 2008, 28, 2201–2212. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.; Cho, W.-K.; Chu, C.K.; Keedy, K.S.; Archin, N.M.; Margolis, D.M.; Karn, J. Epigenetic silencing of HIV-1 by the histone H3 lysine 27 methyltransferase enhancer of zeste 2. J. Virol. 2011, 85, 9078–9089. [Google Scholar] [CrossRef] [PubMed]
- Enderle, D.; Beisel, C.; Stadler, M.B.; Gerstung, M.; Athri, P.; Paro, R. Polycomb preferentially targets stalled promoters of coding and noncoding transcripts. Genome Res. 2011, 21, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Imai, K.; Togami, H.; Okamoto, T. Involvement of Histone H3 Lysine 9 (H3K9) Methyltransferase G9a in the maintenance of HIV-1 latency and its reactivation by BIX01294. J. Biol. Chem. 2010, 285, 16538–16545. [Google Scholar] [CrossRef] [PubMed]
- Abbas, W.; Herbein, G. Molecular Understanding of HIV-1 Latency. Adv. Virol. 2012, 2012, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Groen, J.; Morris, K. Chromatin, Non-Coding RNAs, and the Expression of HIV. Viruses 2013, 5, 1633–1645. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Abbas, W.; Herbein, G. HIV-1 latency in monocytes/macrophages. Viruses 2014, 6, 1837–1860. [Google Scholar] [CrossRef]
- Eisele, E.; Siliciano, R.F. Redefining the viral reservoirs that prevent HIV-1 eradication. Immunity 2012, 37, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.A.; Tolstrup, M.; Winckelmann, A.; Ostergaard, L.; Sogaard, O.S. Eliminating the latent HIV reservoir by reactivation strategies: Advancing to clinical trials. Hum. Vaccin. Immunother. 2013, 9, 790–799. [Google Scholar] [CrossRef] [PubMed]
- Barton, K.M.; Burch, B.D.; Soriano-Sarabia, N.; Margolis, D.M. Prospects for treatment of latent HIV. Clin. Pharmacol. Ther. 2013, 93, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Durand, C.M.; Blankson, J.N.; Siliciano, R.F. Developing strategies for HIV-1 eradication. Trends Immuno.l 2012, 33, 554–562. [Google Scholar] [CrossRef] [PubMed]
- Archin, N.M.; Liberty, A.L.; Kashuba, A.D.; Choudhary, S.K.; Kuruc, J.D.; Crooks, A.M.; Parker, D.C.; Anderson, E.M.; Kearney, M.F.; Strain, M.C.; et al. Administration of vorinostat disrupts HIV-1 latency in patients on antiretroviral therapy. Nature 2012, 487, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Levy, Y.; Sereti, I.; Tambussi, G.; Routy, J.P.; Lelievre, J.D.; Delfraissy, J.F.; Molina, J.M.; Fischl, M.; Goujard, C.; Rodriguez, B.; et al. Effects of recombinant human interleukin 7 on T-cell recovery and thymic output in HIV-infected patients receiving antiretroviral therapy: Results of a phase I/IIa randomized, placebo-controlled, multicenter study. Clin. Infect. Dis. 2012, 55, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Strong, C.L.; Guerra, H.P.; Mathew, K.R.; Roy, N.; Simpson, L.R.; Schiller, M.R. Damaging the integrated HIV proviral DNA with TALENs. PloS ONE 2015. [Google Scholar] [CrossRef] [PubMed]
- Mariyanna, L.; Priyadarshini, P.; Hofmann-Sieber, H.; Krepstakies, M.; Walz, N.; Grundhoff, A.; Buchholz, F.; Hildt, E.; Hauber, J. Excision of HIV-1 proviral DNA by recombinant cell permeable tre-recombinase. PloS ONE 2012. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, I.; Hauber, I.; Hauber, J.; Buchholz, F. HIV-1 proviral DNA excision using an evolved recombinase. Science 2007, 316, 1912–1915. [Google Scholar] [CrossRef] [PubMed]
- June, C. Introduction of Acquired CCR5 Deficiency with Zinc Finger Nuclease-Modified Autologous CD4 T Cells (SB-728-T) Correlates with Increases in CD4 Count and Effects on Viral Load in HIV-Infected Subjects. In Proceedings of 19th Conference on Retroviruses and Opportunistic Infections, Seattle, USA, 5–8 March 2012.
- Lalezari, J. A Single Infusion of Zinc Finger Nuclease CCR5 Modified Autologous CD4 T Cells (SB-728-T) Increases CD4 Counts and Leads to Decrease in HIV Proviral Load in an Aviremic HIV-Infected Subject. In Proceedings of 19th Conference on Retroviruses and Opportunistic Infections, Seattle, USA, 5–8 March 2012.
- Allers, K.; Hutter, G.; Hofmann, J.; Loddenkemper, C.; Rieger, K.; Thiel, E.; Schneider, T. Evidence for the cure of HIV infection by CCR5Delta32/Delta32 stem cell transplantation. Blood 2011, 117, 2791–2799. [Google Scholar] [CrossRef] [PubMed]
- Hutter, G.; Ganepola, S. Eradication of HIV by transplantation of CCR5-deficient hematopoietic stem cells. ScientificWorldJournal 2011, 11, 1068–1076. [Google Scholar] [CrossRef]
- Hutter, G.; Nowak, D.; Mossner, M.; Ganepola, S.; Mussig, A.; Allers, K.; Schneider, T.; Hofmann, J.; Kucherer, C.; Blau, O.; et al. Long-term control of HIV by CCR5 Delta32/Delta32 stem-cell transplantation. N. Engl. J. Med. 2009, 360, 692–698. [Google Scholar] [CrossRef] [PubMed]
- Mens, H.; Kearney, M.; Wiegand, A.; Shao, W.; Schonning, K.; Gerstoft, J.; Obel, N.; Maldarelli, F.; Mellors, J.W.; Benfield, T.; et al. HIV-1 continues to replicate and evolve in patients with natural control of HIV infection. J. Virol. 2010, 84, 12971–12981. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, K.A.; Brennan, T.P.; Bailey, J.R.; Ray, S.C.; Siliciano, R.F.; Blankson, J.N. Control of HIV-1 in elite suppressors despite ongoing replication and evolution in plasma virus. J. Virol. 2010, 84, 7018–7028. [Google Scholar] [CrossRef] [PubMed]
- Rosenblatt, J.; Glotzbecker, B.; Mills, H.; Vasir, B.; Tzachanis, D.; Levine, J.D.; Joyce, R.M.; Wellenstein, K.; Keefe, W.; Schickler, M.; et al. PD-1 blockade by CT-011, anti-PD-1 antibody, enhances ex vivo T-cell responses to autologous dendritic cell/myeloma fusion vaccine. J. Immunother. 2011, 34, 409–418. [Google Scholar] [CrossRef] [PubMed]
- Yukl, S.A.; Boritz, E.; Busch, M.; Bentsen, C.; Chun, T.W.; Douek, D.; Eisele, E.; Haase, A.; Ho, Y.C.; Hutter, G.; et al. Challenges in detecting HIV persistence during potentially curative interventions: a study of the Berlin patient. PLoS Pathog. 2013. [Google Scholar] [CrossRef] [PubMed]
- Henrich, T.J.; Hu, Z.; Li, J.Z.; Sciaranghella, G.; Busch, M.P.; Keating, S.M.; Gallien, S.; Lin, N.H.; Giguel, F.F.; Lavoie, L.; et al. Long-term reduction in peripheral blood HIV type 1 reservoirs following reduced-intensity conditioning allogeneic stem cell transplantation. J. Infect. Dis. 2013, 207, 1694–1702. [Google Scholar] [CrossRef] [PubMed]
- Henrich, T.J.; Hanhauser, E.; Sirignano, M.N.; Li, J.Z.; Lichterfeld, M.; Marty, F.M.; Armand, P.; Soiffer, R.J.; Altfeld, M.; Kuritzkes, D.R. HIV-1 Rebound Following Allogeneic Stem Cell Transplantation and Treatment Interruption. In Proceeding of 21st Conference on Retroviruses and Opportunistic Infections, Boston, MA, USA, 3–6 March 2014.
- Frange, P.; Avettand-Fenoel, A.F.V.; Bellaton, E.; Deschamps, D.; Angin, M.; Caillat-Zucman, S.G.; Peytavin, J.; Chenadec, L.; Warszawski, J.; Rouzioux, C.; Saez-Cirion, A. HIV-1 Virological Remission for More Than 11 Years after Interruption of Early Initiated Antiretroviral Therapy in A Perinatally-Infected Child. In Proceedings of IAS 2015—8th IAS Conference on HIV Pathogenesis, Treatment and Prevention, Vancouver, Canada, 22 July 2015.
- Persaud, D.; Gay, H.; Ziemniak, C.; Chen, Y.H.; Piatak, M., Jr.; Chun, T.W.; Strain, M.; Richman, D.; Luzuriaga, K. Absence of detectable HIV-1 viremia after treatment cessation in an infant. N. Engl. J. Med. 2013, 369, 1828–1835. [Google Scholar] [CrossRef] [PubMed]
- Saez-Cirion, A.; Bacchus, C.; Hocqueloux, L.; Avettand-Fenoel, V.; Girault, I.; Lecuroux, C.; Potard, V.; Versmisse, P.; Melard, A.; Prazuck, T.; et al. Post-treatment HIV-1 controllers with a long-term virological remission after the interruption of early initiated antiretroviral therapy ANRS VISCONTI Study. PLoS Pathog. 2013. [Google Scholar] [CrossRef] [PubMed]
- Klatzmann, D.; Barre-Sinoussi, F.; Nugeyre, M. Selective tropism of lymphadenopathy associated virus (LAV) for helper-inducer T lymphocytes. Science 1984, 225, 59–63. [Google Scholar] [CrossRef] [PubMed]
- Klatzmann, D.; Champagne, E.; Chamaret, S.; Gruest, J.; Guetard, D.; Hercend, T.; Gluckman, J.C.; Montagnier, L. T-lymphocyte T4 molecule behaves as the receptor for human retrovirus LAV. Nature 1984, 312, 767–768. [Google Scholar] [CrossRef] [PubMed]
- Ho, D.D.; Rota, T.; Hirsch, M. Infection of monocyte/macrophages by human T lymphotropic virus type III. J. Clin. Invest. 1986, 77, 1712–1715. [Google Scholar] [CrossRef] [PubMed]
- Chomont, N.; El-Far, M.; Ancuta, P.; Trautmann, L.; Procopio, F.A.; Yassine-Diab, B.; Boucher, G.; Boulassel, M.R.; Ghattas, G.; Brenchley, J.M.; et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat. Med. 2009, 15, 893–900. [Google Scholar] [CrossRef] [PubMed]
- Swiggard, W.J.; Baytop, C.; Yu, J.J.; Dai, J.; Li, C.; Schretzenmair, R.; Theodosopoulos, T.; O’Doherty, U. Human immunodeficiency virus type 1 can establish latent infection in resting CD4+ T cells in the absence of activating stimuli. J. Virol. 2005, 79, 14179–14188. [Google Scholar] [CrossRef] [PubMed]
- Chanel, C.; Staropoli, I.; Baleux, F.; Amara, A.; Valenzuela-Fernandez, A.; Virelizier, J.L.; Arenzana-Seisdedos, F.; Altmeyer, R. Low levels of co-receptor CCR5 are sufficient to permit HIV envelope-mediated fusion with resting CD4 T cells. AIDS 2002, 16, 2337–2340. [Google Scholar] [CrossRef] [PubMed]
- Eckstein, D.A.; Penn, M.L.; Korin, Y.D.; Scripture-Adams, D.D.; Zack, J.A.; Kreisberg, J.F.; Roederer, M.; Sherman, M.P.; Chin, P.S.; Goldsmith, M.A. HIV-1 actively replicates in naive CD4(+) T cells residing within human lymphoid tissues. Immunity 2001, 15, 671–682. [Google Scholar] [CrossRef]
- Chu, T.; Carruth, L.M.; Finzi, D. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature 1997, 387, 183–188. [Google Scholar]
- Siliciano, J.D.; Kajdas, J.; Finzi, D.; Quinn, T.C.; Chadwick, K.; Margolick, J.B.; Kovacs, C.; Gange, S.J.; Siliciano, R.F. Long-term follow-up studies confirm the stability of the latent reservoir for HIV-1 in resting CD4+ T cells. Nat. Med. 2003, 9, 727–728. [Google Scholar] [CrossRef] [PubMed]
- Finzi, D.; Balnkson, J.; Siliciano, J. Latent infection of CD4+T cells provides a mechanism for life long persistence of HIV-1, even in patients on effective combination therapy. Nat. Med. 1999, 5, 512–517. [Google Scholar] [PubMed]
- Siliciano, J.; Siliciano, R.F. Latency and viral persistence in HIV-1 infection. J. Clin. Invest. 2000, 106, 823–825. [Google Scholar] [CrossRef] [PubMed]
- Blankson, J.; Persaud, D.; Siliciano, R.F. The challenge of viral reservoirs in HIV-1 infection. Annu. Rev. Med. 2002, 53, 557–593. [Google Scholar] [CrossRef]
- Williams, K.C.; Hickey, W.F. Central nervous system damage, monocytes and macrophages, and neurological disorders in AIDS. Annu. Rev. Neurosci. 2002, 25, 537–562. [Google Scholar] [CrossRef] [PubMed]
- Garden, G.A. Microglia in human immunodeficiency virus-associated neurodegeneration. Glia 2002, 40, 240–251. [Google Scholar] [CrossRef]
- Smith, P.D.; Meng, G.; Salazar-Gonzalez, J.F.; Shaw, G.M. Macrophage HIV-1 infection and the gastrointestinal tract reservoir. J. Leukoc. Biol. 2003, 74, 642–649. [Google Scholar] [CrossRef] [PubMed]
- Shen, R.; Meng, G.; Ochsenbauer, C.; Clapham, P.R.; Grams, J.; Novak, L.; Kappes, J.C.; Smythies, L.E.; Smith, P.D. Stromal down-regulation of macrophage CD4/CCR5 expression and NF-κB activation mediates HIV-1 non-permissiveness in intestinal macrophages. PLoS Pathog. 2011. [Google Scholar] [CrossRef] [PubMed]
- Abbas, W.; Tariq, M.; Iqbal, M.; Kumar, A.; Herbein, G. Eradication of HIV-1 from the Macrophage Reservoir: An Uncertain Goal? Viruses 2015, 7, 1578–1598. [Google Scholar] [CrossRef] [PubMed]
- Alexaki, A.; Liu, Y.; Wigdahl, B. Cellular reservoirs of HIV-1 and their role in viral persistence. Curr. HIV Res. 2008, 6, 388–400. [Google Scholar] [CrossRef] [PubMed]
- Saksena, N.K.; Wang, B.; Zhou, L.; Soedjono, M.; Ho, Y.S.; Conceicao, V. HIV reservoirs in vivo and new strategies for possible eradication of HIV from the reservoir sites. HIV/AIDS 2010, 2, 103–122. [Google Scholar] [CrossRef] [PubMed]
- Haase, A.T.; Henry, K.; Zupancic, M. Quantitative image analysis of HIV-1 infection in lymphoid tissue. Science 1996, 274, 985–989. [Google Scholar] [CrossRef] [PubMed]
- Crespo, H.; Bertolotti, L.; Juganaru, M.; Glaria, I.; de Andres, D.; Amorena, B.; Rosati, S.; Reina, R. Small ruminant macrophage polarization may play a pivotal role on lentiviral infection. Vet. Res. 2013. [Google Scholar] [CrossRef] [PubMed]
- Gray, L.R.; Roche, M.; Flynn, J.K.; Wesselingh, S.L.; Gorry, P.R.; Churchill, M.J. Is the central nervous system a reservoir of HIV-1? Curr. Opin. HIV AIDS 2014, 9, 552–558. [Google Scholar] [CrossRef]
- Churchill, M.; Nath, A. Where does HIV hide? A focus on the central nervous system. Curr. Opin. HIV AIDS 2013, 8, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.W.; Veenstra, M.; Gaskill, P.J.; Morgello, S.; Calderon, T.M.; Berman, J.W. Monocytes mediate HIV neuropathogenesis: mechanisms that contribute to HIV associated neurocognitive disorders. Curr. HIV Res. 2014, 12, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, D.; Major, E. HIV-1 and the brain: connections between HIV-1-associated dementia, neuropathology and neuroimmunology. Microb. Infect. 2002, 4, 301–308. [Google Scholar] [CrossRef]
- Hong, S.; Banks, W.A. Role of the immune system in HIV-associated neuroinflammation and neurocognitive implications. Brain. Behav. Immun. 2015, 45, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Benavides, J.; Garcia-Pariente, C.; Fuertes, M.; Ferreras, M.C.; Garcia-Marin, J.F.; Juste, R.A.; Perez, V. Maedi-visna: the meningoencephalitis in naturally occurring cases. J. Comp. Pathol. 2009, 140, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Benavides, J.; Fuertes, M.; Garcia-Pariente, C.; Ferreras, M.C.; Garcia Marin, J.F.; Perez, V. Natural cases of visna in sheep with myelitis as the sole lesion in the central nervous system. J. Comp. Pathol. 2006, 134, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Yukl, S.A.; Gianella, S.; Sinclair, E.; Epling, L.; Li, Q.; Duan, L.; Choi, A.L.; Girling, V.; Ho, T.; Li, P.; et al. Differences in HIV burden and immune activation within the gut of HIV-positive patients receiving suppressive antiretroviral therapy. J. Infect. Dis. 2010, 202, 1553–1561. [Google Scholar] [CrossRef] [PubMed]
- Cerf-Bensussan, N.; Guy-Grand, D. Intestinal intraepithelial lymphocytes. Gastroenterol. Clin. North. Am. 1991, 20, 549–576. [Google Scholar] [PubMed]
- Schneider, T.; Jahn, H.U.; Schmidt, W.; Riecken, E.O.; Zeitz, M.; Ullrich, R. Loss of CD4 T lymphocytes in patients infected with human immunodeficiency virus type 1 is more pronounced in the duodenal mucosa than in the peripheral blood. Berlin diarrhea/wasting syndrome study group. Gut 1995, 37, 524–529. [Google Scholar] [CrossRef] [PubMed]
- Olech, M.; Kubis, P.; Lipecka, C.; Junkuszew, A.; Gruszecki, T.M.; Kuźmak, J. Presence of specific antibodies and proviral DNA of small ruminant lentiviruses in lambs in their first weeks of life. Bull. Vet. Inst. Pulawy 2014, 58, 507–511. [Google Scholar] [CrossRef]
- Grossi, P.; Giudice, C.; Bertoletti, I.; Cioccarelli, G.; Brocchi, E.; Cammarata, G.; Gelmetti, D. Immunohistochemical detection of the p27 capsid protein of caprine arthritis-encephalitis virus (CAEV) in bone-marrow cells of seropositive goats. J. Comp. Pathol. 2005, 133, 197–200. [Google Scholar] [CrossRef] [PubMed]
- Ravazzolo, A.P.; Nenci, C.; Vogt, H.-R.; Waldvogel, A.; Obexer-Ruff, G.; Peterhans, E.; Bertoni, G. Viral load, organ distribution, histopathological lesions, and cytokine mRNA expression in goats infected with a molecular clone of the caprine arthritis encephalitis virus. Virology 2006, 350, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Lambert-Niclot, S.; Tubiana, R.; Beaudoux, C.; Lefebvre, G.; Caby, F.; Bonmarchand, M.; Naouri, M.; Schubert, B.; Dommergues, M.; Calvez, V.; et al. Detection of HIV-1 RNA in seminal plasma samples from treated patients with undetectable HIV-1 RNA in blood plasma on a 2002–2011 survey. AIDS 2012, 26, 971–975. [Google Scholar] [CrossRef] [PubMed]
- Marcelin, A.G.; Tubiana, R.; Lambert-Niclot, S.; Lefebvre, G.; Dominguez, S.; Bonmarchand, M.; Vauthier-Brouzes, D.; Marguet, F.; Mousset-Simeon, N.; Peytavin, G.; et al. Detection of HIV-1 RNA in seminal plasma samples from treated patients with undetectable HIV-1 RNA in blood plasma. AIDS 2008, 22, 1677–1679. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.; Van Heeswijk, R.; Hoetelmans, R. Concentration of nevirapine, lamivudine and stavudine in semen of HIV-1 infected men. AIDS 2000, 1979–1984. [Google Scholar] [CrossRef]
- Sheth, P.M.; Kovacs, C.; Kemal, K.S.; Jones, R.B.; Raboud, J.M.; Pilon, R.; la Porte, C.; Ostrowski, M.; Loutfy, M.; Burger, H.; et al. Persistent HIV RNA shedding in semen despite effective antiretroviral therapy. AIDS 2009, 23, 2050–2054. [Google Scholar] [CrossRef] [PubMed]
- Le Tortorec, A.; Dejucq-Rainsford, N. HIV infection of the male genital tract—consequences for sexual transmission and reproduction. Int. J. Androl. 2010, 33, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Dejucq-Rainsford, N.; Jegou, B. Viruses in semen and male genital tissues--consequences for the reproductive system and therapeutic perspectives. Curr. Pharm. Des. 2004, 10, 557–575. [Google Scholar] [CrossRef] [PubMed]
- Lasheeb, A.S.; King, J.; Ball, J.K.; Curran, R.; Barratt, C.L.; Afnan, M.; Pillay, D. Semen characteristics in HIV-1 positive men and the effect of semen washing. Genitourin. Med. 1997, 73, 303–305. [Google Scholar] [CrossRef] [PubMed]
- Pudney, J.; Anderson, D. Orchitis and human immunodeficiency virus type 1 infected cells in reproductive tissues from men with the acquired immune deficiency syndrome. Am. J. Pathol. 1991, 139, 149–160. [Google Scholar] [PubMed]
- Muciaccia, B.; Filippini, A.; Ziparo, E.; Colelli, F.; Baroni, C.D.; Stefanini, M. Testicular germ cells of HIV-seropositive asymptomatic men are infected by the virus. J. Reprod. Immunol. 1998, 41, 81–93. [Google Scholar] [CrossRef]
- Muciaccia, B.; Corallini, S.; Vicini, E.; Padula, F.; Gandini, L.; Liuzzi, G.; Lenzi, A.; Stefanini, M. HIV-1 viral DNA is present in ejaculated abnormal spermatozoa of seropositive subjects. Hum. Reprod. 2007, 22, 2868–2878. [Google Scholar] [CrossRef] [PubMed]
- Galvin, S.R.; Cohen, M.S. Genital tract reservoirs. Curr. Opin. HIV AIDS 2006, 1, 162–166. [Google Scholar] [PubMed]
- Bull, M.E.; Learn, G.H.; McElhone, S.; Hitti, J.; Lockhart, D.; Holte, S.; Dragavon, J.; Coombs, R.W.; Mullins, J.I.; Frenkel, L.M. Monotypic human immunodeficiency virus type 1 genotypes across the uterine cervix and in blood suggest proliferation of cells with provirus. J. Virol. 2009, 83, 6020–6028. [Google Scholar] [CrossRef]
- Si-Mohamed, A.; Kazatchkine, M.D.; Heard, I.; Goujon, C.; Prazuck, T.; Aymard, G.; Cessot, G.; Kuo, Y.H.; Bernard, M.C.; Diquet, B.; et al. Selection of drug-resistant variants in the female genital tract of human immunodeficiency virus type 1-infected women receiving antiretroviral therapy. J. Infect. Dis. 2000, 182, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Ali Al Ahmad, M.Z.; Dubreil, L.; Chatagnon, G.; Khayli, Z.; Theret, M.; Martignat, L.; Chebloune, Y.; Fieni, F. Goat uterine epithelial cells are susceptible to infection with Caprine Arthritis Encephalitis Virus (CAEV) in vivo. Vet. Res. 2012. [Google Scholar] [CrossRef] [PubMed]
- Lamara, A.; Fieni, F.; Mselli-Lakhal, L.; Tainturier, D.; Chebloune, Y. Efficient replication of caprine arthritis-encephalitis virus in goat granulosa cells. Virus Res. 2001, 79, 165–172. [Google Scholar] [CrossRef]
- Lamara, A.; Fieni, F.; Mselli-Lakhal, L.; Tainturier, D.; Chebloune, Y. Epithelial cells from goat oviduct are highly permissive for productive infection with caprine arthritis-encephalitis virus (CAEV). Virus Res. 2002, 87, 69–77. [Google Scholar] [CrossRef]
- Ali Al Ahmad, M.Z.; Fieni, F.; Guiguen, F.; Larrat, M.; Pellerin, J.L.; Roux, C.; Chebloune, Y. Cultured early goat embryos and cells are susceptible to infection with caprine encephalitis virus. Virology 2006, 353, 307–315. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lamara, A.; Fieni, F.; Chatagnon, G.; Larrat, M.; Dubreil, L.; Chebloune, Y. Caprine arthritis encephalitis virus (CAEV) replicates productively in cultured epididymal cells from goats. Comp. Immunol. Microbiol. Infect. Dis. 2013, 36, 397–404. [Google Scholar] [CrossRef]
- Ali Al Ahmad, M.Z.; Fieni, F.; Pellerin, J.L.; Guiguen, F.; Cherel, Y.; Chatagnon, G.; Bouzar, A.B.; Chebloune, Y. Detection of viral genomes of caprine arthritis-encephalitis virus (CAEV) in semen and in genital tract tissues of male goat. Theriogenology 2008, 69, 473–480. [Google Scholar] [CrossRef] [PubMed]
- Turchetti, A.P.; Paniago, J.J.; da Costa, L.F.; da Cruz, J.C.; Braz, G.F.; Gouveia, A.M.; Paixao, T.A.; Santos, R.L.; Heinemann, M.B. Distribution of caprine arthritis encephalitis virus provirus, RNA, and antigen in the reproductive tract of one naturally and seven experimentally infected bucks. Theriogenology 2013, 80, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Fieni, F.; Pellerin, J.L.; Roux, C.; Poulin, N.; Baril, G.; Fatet, A.; Valas, S.; Chatagnon, G.; Mermillod, P.; Guignot, F. Can caprine arthritis encephalitis virus (CAEV) be transmitted by in vitro fertilization with experimentally infected sperm? Theriogenology 2012, 77, 644–651. [Google Scholar] [CrossRef] [PubMed]
- Alimohammadi, A.; Coker, R.; Miller, R.; Mitchell, D.; Williamson, J.; Clarke, J. Genotypic variants of HIV-1 from peripheral blood and lungs of AIDS patients. AIDS 1997, 11, 831–832. [Google Scholar] [PubMed]
- White, N.; Israel-Biet, D.; coker, R.; Mitchell, D.; Weber, J.; Clarke, J. Different resistance mutations can be detected simultaneously in the blood and the lung of HIV-1 infected individuals on antiretroviral therapy. J. Med. Virol. 2004, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Cribbs, S.K.; Lennox, J.; Caliendo, A.M.; Brown, L.A.; Guidot, D.M. Healthy HIV-1-infected individuals on highly active antiretroviral therapy harbor HIV-1 in their alveolar macrophages. AIDS Res. Hum. Retroviruses 2015, 31, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Almodovar, S. The complexity of HIV persistence and pathogenesis in the lung under antiretroviral therapy: challenges beyond AIDS. Viral Immunol. 2014, 27, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Costiniuk, C.T.; Jenabian, M.A. The lungs as anatomical reservoirs of HIV infection. Rev. Med. Virol. 2014, 24, 35–54. [Google Scholar] [CrossRef] [PubMed]
- Bruggeman, L.; Ross, M.; Tanji, N. Renal epithelium is a previously unrecognized site of HIV-1 infection. J. Am. Soc. Nephrol. 2000, 2079–2087. [Google Scholar]
- Marras, D.; Bruggeman, L.; Gao, F. Replication and compartmentalization of HIV-1 in kidney epithelium of patients with HIV-associated nephropathy. Nat. Med. 2002, 522–526. [Google Scholar] [CrossRef] [PubMed]
- Blasi, M.; Balakumaran, B.; Chen, P.; Negri, D.R.; Cara, A.; Chen, B.K.; Klotman, M.E. Renal epithelial cells produce and spread HIV-1 via T-cell contact. AIDS 2014, 28, 2345–2353. [Google Scholar] [CrossRef] [PubMed]
- Canaud, G.; Dejucq-Rainsford, N.; Avettand-Fenoel, V.; Viard, J.P.; Anglicheau, D.; Bienaime, F.; Muorah, M.; Galmiche, L.; Gribouval, O.; Noel, L.H.; et al. The kidney as a reservoir for HIV-1 after renal transplantation. J. Am. Soc. Nephrol. 2014, 25, 407–419. [Google Scholar] [CrossRef] [PubMed]
- Stock, P.G.; Barin, B.; Hatano, H.; Rogers, R.L.; Roland, M.E.; Lee, T.H.; Busch, M.; Deeks, S.G. Reduction of HIV persistence following transplantation in HIV-infected kidney transplant recipients. Am. J. Transplant. 2014, 14, 1136–1141. [Google Scholar] [CrossRef] [PubMed]
- McNeilly, T.N.; Baker, A.; Brown, J.K.; Collie, D.; Maclachlan, G.; Rhind, S.M.; Harkiss, G.D. Role of alveolar macrophages in respiratory transmission of visna/maedi virus. J. Virol. 2008, 82, 1526–1536. [Google Scholar] [CrossRef] [PubMed]
- McNeilly, T.N.; Tennant, P.; Lujan, L.; Perez, M.; Harkiss, G.D. Differential infection efficiencies of peripheral lung and tracheal tissues in sheep infected with Visna/maedi virus via the respiratory tract. J. Gen. Virol. 2007, 88, 670–679. [Google Scholar] [CrossRef] [PubMed]
- Zang, Z.; Watt, N.; Hopkins, J.; Harkiss, G.; Woodall, C. Quantitative analysis of Maedi Visna virus DNA load in peripheral blood monocytes and alveolar macrophages. J. Virol. Methods 2000, 13–20. [Google Scholar] [CrossRef]
- Watt, N.J.; MacIntyre, N.; Collie, D.; Sargan, D.; McConnell, I. Phenotypic analysis of lymphocyte populations in the lungs and regional lymphoid tissue of sheep naturally infected with maedi visna virus. Clin. Exp. Immunol. 1992, 90, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Patton, K.M.; Bildfell, R.J.; Anderson, M.L.; Cebra, C.K.; Valentine, B.A. Fatal Caprine arthritis encephalitis virus-like infection in 4 Rocky Mountain goats (Oreamnos americanus). J. Vet. Diagn. Invest. 2012, 24, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Arrode-Brusés, G.; Hegde, R.; Jin, Y.; Liu, Z.; Narayan, O.; Chebloune, Y. Immunogenicity of a lentiviral-based DNA vaccine driven by the 5′LTR of the naturally attenuated caprine arthritis encephalitis virus (CAEV) in mice and macaques. Vaccine 2012, 30, 2956–2962. [Google Scholar] [CrossRef] [PubMed]
- Chebloune, Y.; Moussa, M.; Arrode-Bruses, G.; Gagnon, J. Cowpox helped against smallpox; will the goat lentivirus (Caprine Arthritis Encephalitis Virus) help against HIV-1? AIDS Res. Hum. Retroviruses 2015. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
