Characterisation of Milk Microbiota from Subclinical Mastitis and Apparently Healthy Dairy Cattle in Free State Province, South Africa
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Mastitis Screening and Sample Collection
2.2. DNA Extraction and PCR
2.3. 16S rRNA Gene Amplification and Sample Barcoding
2.4. Bioinformatics and Statistical Analyses
3. Results
3.1. Somatic Cell Counts
3.2. Characteristics of the Sequences
3.3. Taxonomic Profile
3.4. Alpha Diversity Determined by 16S Sequencing
3.5. Beta Diversity Analysis Determined by 16S Sequencing
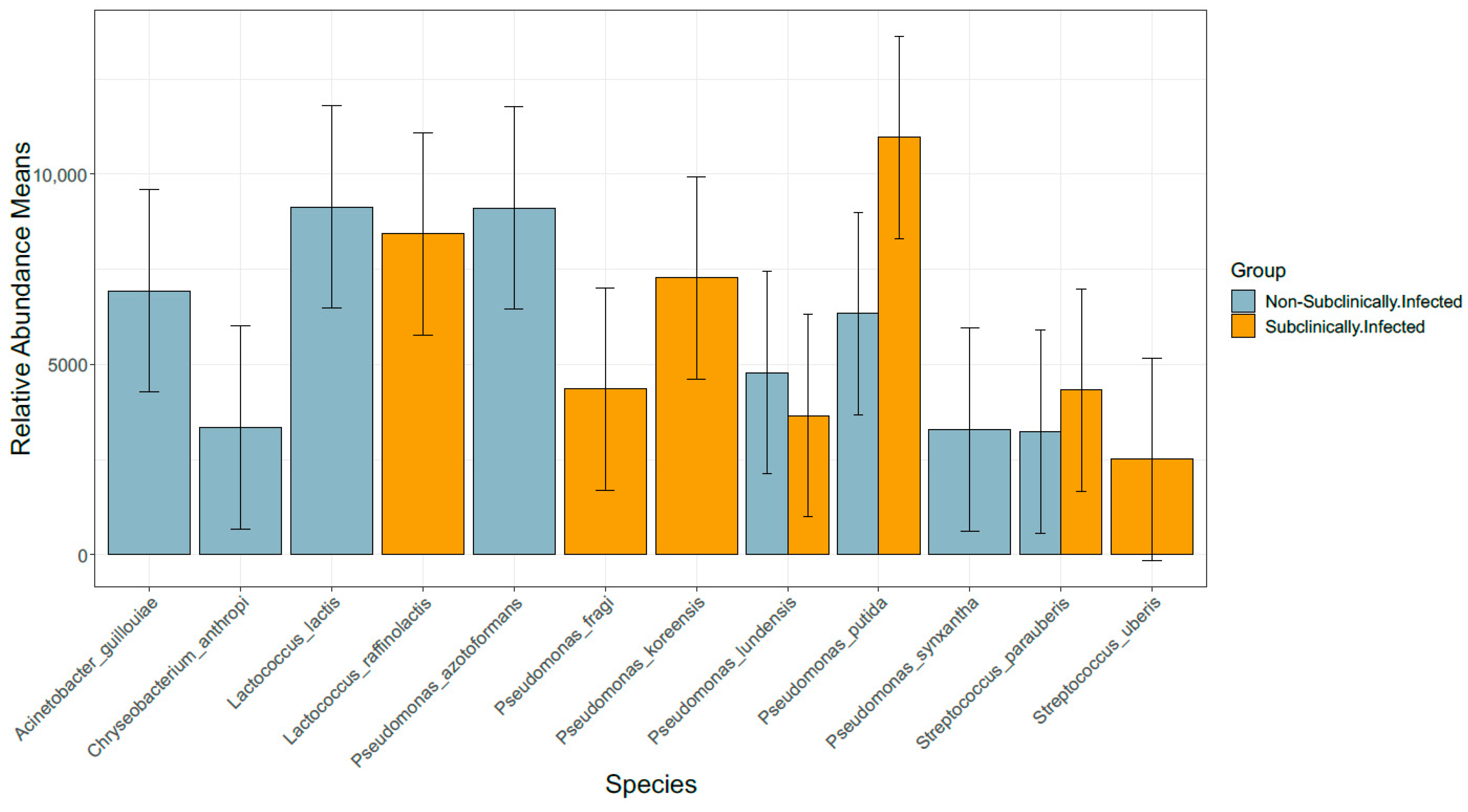
3.6. Comparison of Bacterial Taxonomy Overlaps and Differential Abundance in SCM and Non-SCM
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Khasapane, N.G.; Nkhebenyane, J.S.; Kwenda, S.; Khumalo, Z.T.; Mtshali, P.S.; Taioe, M.O.; Thekisoe, O.M. Application of culture, PCR, and PacBio sequencing for determination of microbial composition of milk from subclinical mastitis dairy cows of smallholder farms. Open Life Sci. 2021, 16, 800–808. [Google Scholar] [CrossRef] [PubMed]
- Mbindyo, C.M.; Gitao, G.C.; Mulei, C.M. Prevalence, etiology, and risk factors of mastitis in dairy cattle in Embu and Kajiado Counties, Kenya. Vet. Med. Int. 2020, 2020, 8831172. [Google Scholar] [CrossRef] [PubMed]
- Hoque, M.N.; Istiaq, A.; Clement, R.A.; Sultana, M.; Crandall, K.A.; Siddiki, A.Z.; Hossain, M.A. Metagenomic deep sequencing reveals association of microbiome signature with functional biases in bovine mastitis. Sci. Rep. 2019, 9, 13536. [Google Scholar] [CrossRef] [PubMed]
- Belay, N.; Mohammed, N.; Seyoum, W. Bovine mastitis: Prevalence, risk factors, and bacterial pathogens isolated in lactating cows in Gamo zone, southern Ethiopia. Vet. Med. Res. Rep. 2022, 9, 19. [Google Scholar] [CrossRef]
- Falentin, H.; Rault, L.; Nicolas, A.; Bouchard, D.S.; Lassalas, J.; Lamberton, P.; Aubry, J.M.; Marnet, P.G.; Le Loir, Y.; Even, S. Bovine teat microbiome analysis revealed reduced alpha diversity and significant changes in taxonomic profiles in quarters with a history of mastitis. Front. Microbiol. 2016, 7, 480. [Google Scholar] [CrossRef]
- Levison, L.J.; Miller-Cushon, E.K.; Tucker, A.L.; Bergeron, R.; Leslie, K.E.; Barkema, H.W.; DeVries, T.J. Incidence rate of pathogen-specific clinical mastitis on conventional and organic Canadian dairy farms. J. Dairy Sci. 2016, 99, 1341–1350. [Google Scholar] [CrossRef]
- Lin, L.; Huang, X.; Yang, H.; He, Y.; He, X.; Huang, J.; Li, S.; Wang, X.; Tang, S.; Liu, G.; et al. Molecular epidemiology, antimicrobial activity, and virulence gene clustering of Streptococcus agalactiae isolated from dairy cattle with mastitis in China. J. Dairy Sci. 2021, 104, 4893–4903. [Google Scholar] [CrossRef]
- Oliver, S.P.; Murinda, S.E. Antimicrobial resistance of mastitis pathogens. Vet. Clin. N. Am. Food Anim. Pract. 2012, 28, 165–185. [Google Scholar] [CrossRef]
- Preethirani, P.L.; Isloor, S.; Sundareshan, S.; Nuthanalakshmi, V.; Deepthikiran, K.; Sinha, A.Y.; Rathnamma, D.; Prabhu, K.N.; Sharada, R.; Mukkur, T.K.; et al. Isolation, biochemical and molecular identification, and in-vitro antimicrobial resistance patterns of bacteria isolated from bubaline subclinical mastitis in South India. PLoS ONE 2015, 10, e0142717. [Google Scholar]
- Rinaldi, M.; Li, R.W.; Capuco, A.V. Mastitis associated transcriptomic disruptions in cattle. Vet. Immunol. Immunopathol. 2010, 138, 267–279. [Google Scholar] [CrossRef]
- Pyörälä, S.; Taponen, S. Coagulase-negative staphylococci—Emerging mastitis pathogens. Vet. Microbiol. 2009, 134, 3–8. [Google Scholar] [CrossRef]
- Lago, A.; Godden, S.M.; Bey, R.; Ruegg, P.L.; Leslie, K. The selective treatment of clinical mastitis based on on-farm culture results: I. Effects on antibiotic use, milk withholding time, and short-term clinical and bacteriological outcomes. J. Dairy Sci. 2011, 94, 4441–4456. [Google Scholar] [CrossRef] [PubMed]
- Oultram, J.W.; Ganda, E.K.; Boulding, S.C.; Bicalho, R.C.; Oikonomou, G. A metataxonomic approach could be considered for cattle clinical mastitis diagnostics. Front. Vet. Sci. 2017, 4, 36. [Google Scholar] [CrossRef] [PubMed]
- D’Argenio, V.; Casaburi, G.; Precone, V.; Salvatore, F. Comparative metagenomic analysis of human gut microbiome composition using two different bioinformatic pipelines. BioMed Res. Int. 2014, 25, 2014. [Google Scholar] [CrossRef] [PubMed]
- Cremonesi, P.; Ceccarani, C.; Curone, G.; Severgnini, M.; Pollera, C.; Bronzo, V.; Riva, F.; Addis, M.F.; Filipe, J.; Amadori, M.; et al. Milk microbiome diversity and bacterial group prevalence in a comparison between healthy Holstein Friesian and Rendena cows. PLoS ONE 2018, 13, e0205054. [Google Scholar] [CrossRef]
- Catozzi, C.; Bonastre, A.S.; Francino, O.; Lecchi, C.; De Carlo, E.; Vecchio, D.; Martucciello, A.; Fraulo, P.; Bronzo, V.; Cuscó, A.; et al. The microbiota of water buffalo milk during mastitis. PLoS ONE 2017, 12, e0184710. [Google Scholar] [CrossRef]
- Hornik, B.; Czarny, J.; Staninska-Pięta, J.; Wolko, Ł.; Cyplik, P.; Piotrowska-Cyplik, A. The Raw Milk Microbiota from SemiSubsistence Farms Characteristics by NGS Analysis Method. Molecules 2021, 26, 5029. [Google Scholar] [CrossRef]
- Oikonomou, G.; Bicalho, M.L.; Meira, E.; Rossi, R.E.; Foditsch, C.; Machado, V.S.; Teixeira, A.G.V.; Santisteban, C.; Schukken, Y.H.; Bicalho, R.C. Microbiota of cow’s milk; distinguishing healthy, sub-clinically and clinically diseased quarters. PLoS ONE 2014, 9, e85904. [Google Scholar] [CrossRef]
- Addis, M.F.; Tanca, A.; Uzzau, S.; Oikonomou, G.; Bicalho, R.C.; Moroni, P. The bovine milk microbiota: Insights and perspectives from-omics studies. Mol. Biosyst. 2016, 12, 2359–2372. [Google Scholar] [CrossRef]
- Derakhshani, H.; Fehr, K.B.; Sepehri, S.; Francoz, D.; De Buck, J.; Barkema, H.W.; Plaizier, J.C.; Khafipour, E. Invited review: Microbiota of the bovine udder: Contributing factors and potential implications for udder health and mastitis susceptibility. J. Dairy Sci. 2018, 101, 10605–10625. [Google Scholar] [CrossRef]
- Angelopoulou, A.; Holohan, R.; Rea, M.C.; Warda, A.K.; Hill, C.; Ross, R.P. Bovine mastitis is a polymicrobial disease requiring a polydiagnostic approach. Int. Dairy J. 2019, 99, 104539. [Google Scholar]
- Gao, J.; Liu, Y.-C.; Wang, Y.; Li, H.; Wang, X.-M.; Wu, Y.; Zhang, D.-R.; Gao, S.; Qi, Z. Impact of yeast and lactic acid bacteria on mastitis and milk microbiota composition of dairy cows. AMB Express 2020, 10, 22. [Google Scholar] [CrossRef]
- Porcellato, D.; Aspholm, M.; Skeie, S.B.; Monshaugen, M.; Brendehaug, J.; Mellegård, H. Microbial diversity of consumption milk during processing and storage. Int. J. Food Microbiol. 2018, 266, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Karzis, J.; Donkin, E.F.; Webb, E.C.; Etter, E.M.; Petzer, I.M. Somatic cell count thresholds in composite and quarter milk samples as indicator of bovine intramammary infection status. Onderstepoort J. Vet. Res. 2017, 84, e1–e10. [Google Scholar]
- Pootakham, W.; Mhuantong, W.; Yoocha, T.; Putchim, L.; Sonthirod, C.; Naktang, C.; Thongtham, N.; Tangphatsornruang, S. High resolution profiling of coral-associated bacterial communities using full-length 16S rRNA sequence data from PacBio SMRT sequencing system. Sci. Rep. 2017, 7, 2774. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PloS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Andersen, K.S.; Kirkegaard, R.H.; Karst, S.M.; Albertsen, M. ampvis2: An R package to analyse and visualise 16S rRNA amplicon data. BioRxiv 2018, 11, 299537. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Conway, J.R.; Lex, A.; Gehlenborg, N. UpSetR: An R package for the visualization of intersecting sets and their properties. Bioinformatics 2017, 33, 2938–2940. [Google Scholar] [CrossRef]
- Sokolov, S.; Fursova, K.; Shulcheva, I.; Nikanova, D.; Artyemieva, O.; Kolodina, E.; Sorokin, A.; Dzhelyadin, T.; Shchannikova, M.; Shepelyakovskaya, A.; et al. Comparative analysis of milk microbiomes and their association with bovine mastitis in two farms in Central Russia. Animals 2021, 11, 1401. [Google Scholar] [CrossRef]
- Birhanu, M.; Leta, S.; Mamo, G.; Tesfaye, S. Prevalence of bovine subclinical mastitis and isolation of its major causes in Bishoftu Town, Ethiopia. BMC Res. Notes 2017, 10, 767. [Google Scholar] [CrossRef]
- Wiegel, J.; Tanner, R.A.; Rainey, F.A. An introduction to the family Clostridiaceae. Prokaryotes 2006, 4, 654–678. [Google Scholar]
- O’Connell, A.; Kelly, A.L.; Tobin, J.; Ruegg, P.L.; Gleeson, D. The effect of storage conditions on the composition and functional properties of blended bulk tank milk. J. Dairy Sci. 2017, 100, 991–1003. [Google Scholar] [CrossRef]
- Kamimura, B.A.; Cabral, L.; Noronha, M.F.; Baptista, R.C.; Nascimento, H.M.; Sant’Ana, A.S. Amplicon sequencing reveals the bacterial diversity in milk, dairy premises and Serra da Canastra artisanal cheeses produced by three different farms. Food Microbiol. 2020, 89, 103453. [Google Scholar] [CrossRef]
- Schukken, Y.; Wilson, D.; Welcome, F.; Garrison-Tikofsky, L.; Gonzalez, R. Monitoring udder health and milk quality using somatic cell counts. Vet. Res. 2003, 34, 579–596. [Google Scholar] [CrossRef] [PubMed]
- Heeschen, W.H. IDF and mastitis–A general review. In Proceedings of the Masfifis Research into Pracfice, 5th IDF Mastitis Conference, Christ Church, New Zealand, 21 March 2010; pp. 21–24. [Google Scholar]
- Bhatt, V.D.; Ahir, V.B.; Koringa, P.G.; Jakhesara, S.J.; Rank, D.N.; Nauriyal, D.S.; Kunjadia, A.P.; Joshi, C.G. Milk microbiome signatures of subclinical mastitis-affected cattle analysed by shotgun sequencing. J. Appl. Microbiol. 2012, 112, 639–650. [Google Scholar] [CrossRef]
- Oikonomou, G.; Addis, M.F.; Chassard, C.; Nader-Macias, M.E.; Grant, I.; Delbès, C.; Bogni, C.I.; Le Loir, Y.; Even, S. Milk microbiota: What are we exactly talking about? Front. Microbiol. 2020, 11, 60. [Google Scholar] [CrossRef]
- Klaas, I.C.; Zadoks, R.N. An update on environmental mastitis: Challenging perceptions. Transbound. Emerg. Dis. 2018, 65, 166–185. [Google Scholar] [CrossRef] [PubMed]
- Lei, X.; Chen, K.; Zhu, L.; Song, E.; Su, F.; Li, S. Treatments for idiopathic granulomatous mastitis: Systematic review and meta-analysis. Breastfeed. Med. 2017, 12, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Kaczorowski, Ł.; Powierska-Czarny, J.; Wolko, Ł.; Piotrowska-Cyplik, A.; Cyplik, P.; Czarny, J. The influence of bacteria causing subclinical mastitis on the structure of the cow’s milk microbiome. Molecules 2022, 27, 1829. [Google Scholar] [CrossRef] [PubMed]
- Sanders, J.W.; Martin, J.W.; Hooke, M.; Hooke, J. Methylobacterium mesophilicum infection: Case report and literature review of an unusual opportunistic pathogen. Clin. Infect. Dis. 2000, 30, 936–938. [Google Scholar] [CrossRef] [PubMed]
- Pang, M.; Xie, X.; Bao, H.; Sun, L.; He, T.; Zhao, H.; Zhou, Y.; Zhang, L.; Zhang, H.; Wei, R.; et al. Insights into the bovine milk microbiota in dairy farms with different incidence rates of subclinical mastitis. Front. Microbiol. 2018, 9, 2379. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, P.; Wang, Y.; Fu, L.; Liu, L.; Xu, D.; Hou, Y.; Li, Y.; Fu, M.; Wang, X.; et al. Capsular serotypes, antimicrobial susceptibility, and the presence of transferable oxazolidinone resistance genes in Streptococcus suis isolated from healthy pigs in China. Vet. Microbiol. 2020, 247, 108750. [Google Scholar] [CrossRef]
- Bi, Y.; Wang, Y.J.; Qin, Y.; Guix Vallverdú, R.; Maldonado García, J.; Sun, W.; Li, S.; Cao, Z. Prevalence of bovine mastitis pathogens in bulk tank milk in China. Plos ONE 2016, 11, e0155621. [Google Scholar] [CrossRef]
- Gonçalves, J.L.; Tomazi, T.; Barreiro, J.R.; Beuron, D.C.; Arcari, M.A.; Lee, S.H.; Martins, C.M.; Junior, J.P.; dos Santos, M.V. Effects of bovine subclinical mastitis caused by Corynebacterium spp. on somatic cell count, milk yield and composition by comparing contralateral quarters. Vet. J. 2016, 209, 87–92. [Google Scholar] [CrossRef]
- Patel, S.H.; Vaidya, Y.H.; Patel, R.J.; Pandit, R.J.; Joshi, C.G.; Kunjadiya, A.P. Culture independent assessment of human milk microbial community in lactational mastitis. Sci. Rep. 2017, 7, 7804. [Google Scholar] [CrossRef]
- Irorere, V.U.; Tripathi, L.; Marchant, R.; McClean, S.; Banat, I.M. Microbial rhamnolipid production: A critical re-evaluation of published data and suggested future publication criteria. Appl. Microbiol. Biotechnol. 2017, 101, 3941–3951. [Google Scholar]
- Hadjisymeou, S.; Loizou, P.; Kothari, P. Lactococcus lactis cremoris infection: Not rare anymore? Case Rep. 2013, 2013, bcr2012008479. [Google Scholar] [CrossRef]
- Inoue, Y.; Kambara, T.; Murata, N.; Komori-Yamaguchi, J.; Matsukura, S.; Takahashi, Y.; Ikezawa, Z.; Aihara, M. Effects of oral administration of Lactobacillus acidophilus L-92 on the symptoms and serum cytokines of atopic dermatitis in Japanese adults: A double-blind, randomized, clinical trial. Int. Arch. Allergy Immunol. 2014, 165, 247–254. [Google Scholar] [CrossRef]
- Kuang, Y.; Tani, K.; Synnott, A.J.; Ohshima, K.; Higuchi, H.; Nagahata, H.; Tanji, Y. Characterization of bacterial population of raw milk from bovine mastitis by culture-independent PCR–DGGE method. Biochem. Eng. J. 2009, 45, 76–81. [Google Scholar] [CrossRef]
- Quigley, L.; O′Sullivan, O.; Stanton, C.; Beresford, T.P.; Ross, R.P.; Fitzgerald, G.F.; Cotter, P.D. The complex microbiota of raw milk. FEMS Microbiol. Rev. 2013, 37, 664–698. [Google Scholar] [CrossRef]
- Alnakip, M.E.; Mohamed, A.S.; Kamal, R.M.; Elbadry, S. Diversity of lactic acid bacteria isolated from raw milk in Elsharkia province, Egypt. Jpn. J. Vet. Res. 2016, 64 (Suppl. S2), S23–S30. [Google Scholar]
- Carvalheira, A.; Ferreira, V.; Silva, J.; Teixeira, P. Enrichment of Acinetobacter spp. from food samples. Food Microbiol. 2016, 55, 123–127. [Google Scholar] [CrossRef]
- Amorim, A.M.; Nascimento, J.D. A highlight for non-Escherichia coli and non-Salmonella sp. Enterobacteriaceae in dairy foods contamination. Front. Microbiol. 2017, 8, 930. [Google Scholar] [CrossRef]
- von Neubeck, M.; Baur, C.; Krewinkel, M.; Stoeckel, M.; Kranz, B.; Stressler, T.; Fischer, L.; Hinrichs, J.; Scherer, S.; Wenning, M. Biodiversity of refrigerated raw milk microbiota and their enzymatic spoilage potential. Int. J. Food Microbiol. 2015, 211, 57–65. [Google Scholar] [CrossRef]
- Cho, G.S.; Li, B.; Rostalsky, A.; Fiedler, G.; Rösch, N.; Igbinosa, E.; Kabisch, J.; Bockelmann, W.; Hammer, P.; Huys, G.; et al. Diversity and antibiotic susceptibility of Acinetobacter strains from milk powder produced in Germany. Front. Microbiol. 2018, 9, 536. [Google Scholar] [CrossRef]
- Straley, B.A.; Donaldson, S.C.; Hedge, N.V.; Sawant, A.A.; Srinivasan, V.; Oliver, S.P.; Jayarao, B.M. Public health significance of antimicrobial-resistant gram-negative bacteria in raw bulk tank milk. Foodborne Pathog. Dis. 2006, 3, 222–233. [Google Scholar] [CrossRef]
- Gomes, F.; Henriques, M. Control of bovine mastitis: Old and recent therapeutic approaches. Curr. Microbiol. 2016, 72, 377–382. [Google Scholar] [CrossRef]
- Lamichhane, J.; Adhikary, S.; Gautam, P.; Maharjan, R.; Dhakal, B. Risk of handling paper currency in circulation chances of potential bacterial transmittance. Nepal J. Sci. Technol. 2009, 10, 161–166. [Google Scholar] [CrossRef]
- EFSA Panel on Biological Hazards (BIOHAZ). Scientific Opinion on the public health risks of bacterial strains producing extended-spectrum β-lactamases and/or AmpC β-lactamases in food and food-producing animals. EFSA J. 2011, 9, 2322. [Google Scholar] [CrossRef]
- Munsch-Alatossava, P.; Alatossava, T. Quality and Safety of Bovine Raw Milk: Present Challenges and Technological Solutions. In Milk Production, Processing and Marketing; IntechOpen: London, UK, 2019. [Google Scholar]
- Poretsky, R.; Rodriguez, L.M.; Luo, C.; Tsementzi, D.; Konstantinidis, K.T. Strengths and limitations of 16S rRNA gene amplicon sequencing in revealing temporal microbial community dynamics. PloS ONE 2014, 9, e93827. [Google Scholar]
- Kumar, V.; Singh, K.; Shah, M.P.; Singh, A.K.; Kumar, A.; Kumar, Y. Application of omics technologies for microbial community structure and function analysis in contaminated environment. In Wastewater Treatment; Elsevier: Amsterdam, The Netherlands, 2021; pp. 1–40. [Google Scholar]
- Rodrigues, M.X. Molecular Characterization of Bacterial Isolates and Microbiome: Study of Mastitic Milk, Bulk Tank Milk, and Cheese Processing Plants. Doctoral Dissertation, Universidade de São Paulo, São Paulo, Brazil, 2016. [Google Scholar]
- Hattingh, A. The Proteolytic Activity in Raw Milk and the Effect of Such Activity on the Stability of Milk Proteins. Doctoral Dissertation, University of the Free State, Bloemfontein, South Africa, 2017. [Google Scholar]
- Tarrah, A.; Callegaro, S.; Pakroo, S.; Finocchiaro, R.; Giacomini, A.; Corich, V.; Cassandro, M. New insights into the raw milk microbiota diversity from animals with a different genetic predisposition for feed efficiency and resilience to mastitis. Sci. Rep. 2022, 12, 13498. [Google Scholar] [CrossRef] [PubMed]
- Tibbetts, L. Mosby’s Comprehensive Review for Veterinary Technicians; Mosby: Maryland Heights, MO, USA, 2019; p. 370. Available online: https://www.amazon.com/Mosbys-Comprehensive-Review-Veterinary-Technicians/dp/0323596150 (accessed on 5 October 2023).








Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khasapane, N.G.; Khumalo, Z.T.H.; Kwenda, S.; Nkhebenyane, S.J.; Thekisoe, O. Characterisation of Milk Microbiota from Subclinical Mastitis and Apparently Healthy Dairy Cattle in Free State Province, South Africa. Vet. Sci. 2023, 10, 616. https://doi.org/10.3390/vetsci10100616
Khasapane NG, Khumalo ZTH, Kwenda S, Nkhebenyane SJ, Thekisoe O. Characterisation of Milk Microbiota from Subclinical Mastitis and Apparently Healthy Dairy Cattle in Free State Province, South Africa. Veterinary Sciences. 2023; 10(10):616. https://doi.org/10.3390/vetsci10100616
Chicago/Turabian StyleKhasapane, N. G., Z. T. H. Khumalo, S. Kwenda, S. J. Nkhebenyane, and O. Thekisoe. 2023. "Characterisation of Milk Microbiota from Subclinical Mastitis and Apparently Healthy Dairy Cattle in Free State Province, South Africa" Veterinary Sciences 10, no. 10: 616. https://doi.org/10.3390/vetsci10100616




