CaosDB—Research Data Management for Complex, Changing, and Automated Research Workflows
Abstract
1. Introduction
- Scientists use specific or customized tools, software and data formats with good reason. A research data management system (RDMS) must be built around their workflows and practices and be open for change. It furthermore should assist in integrating data, even if stored in propriertary formats. Enforcing one of the individual systems cannot work in a heterogeneous scientific environment.
- If the RDMS requires too much extra work for learning and understanding, it is likely that the individual scientist will be unable to use it efficiently or just be unwilling to use it at all. This holds in particular for the construction of queries which should retrieve data according to powerful criteria while being simple and intuitive at the same time. For this reason it is unlikely that a scientist without computer science background will be able to use Structured Query Language (SQL) or SPARQL efficiently [8,9,10].
- The system should strongly encourage to develop and use standards for workflows and data models without being overly restrictive. Users of the database can only profit from the system when data is organized sufficiently structured to enable everyone to search and retrieve data easily and to understand the structure of the data intuitively. At the same time the database has to be prepared for constantly evolving data models and standards [2].
- File systems and other types of storage usually organize data into some kind of hierarchy which can be folders or projects. In scientific environments this can raise issues, especially when data belongs to multiple projects or is part of cooperations.
- Simple and intuitive access to complex and changing data models
- Reuse and integration of existing workflows and technologies
2. Results
2.1. Requirements
- Findable (F1–F4)
- Accessible (A1–A2)
- Interoperable (I1–I3)
- Reusable (R1–R1.3)
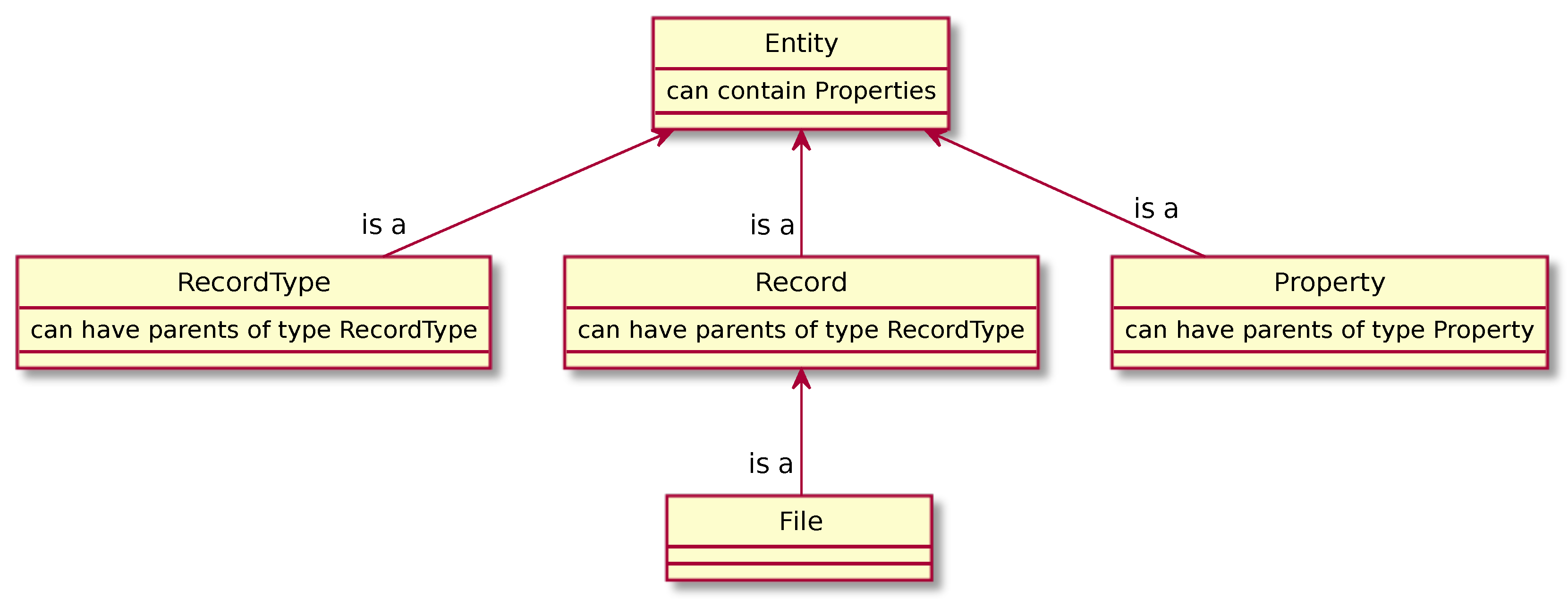
- Entities with subtyping
- User-defined n-ary relationships and properties
- Integration of files and directories as entities
- Native support for primitive data types which include several numeric data types with their physical units and uncertainties, standard compliant date and time values, booleans, strings, and undefined values
- Compound data types for lists, sets, tuples, and dictionaries
2.2. Implementation
SELECT flavour, rating, ingredients FROM Experiment WHICH HAS A room_temperature > 26C AND WHICH IS REFERENCED BY ExperimentSeries WHICH HAS A name LIKE *ice cream testing*
- offers a flexible data model
- can be seamlessly integrated into existing workflows
- allows search for values of specific fields (not just full text search), with automatic unit conversion and search for (back)references of linked objects.
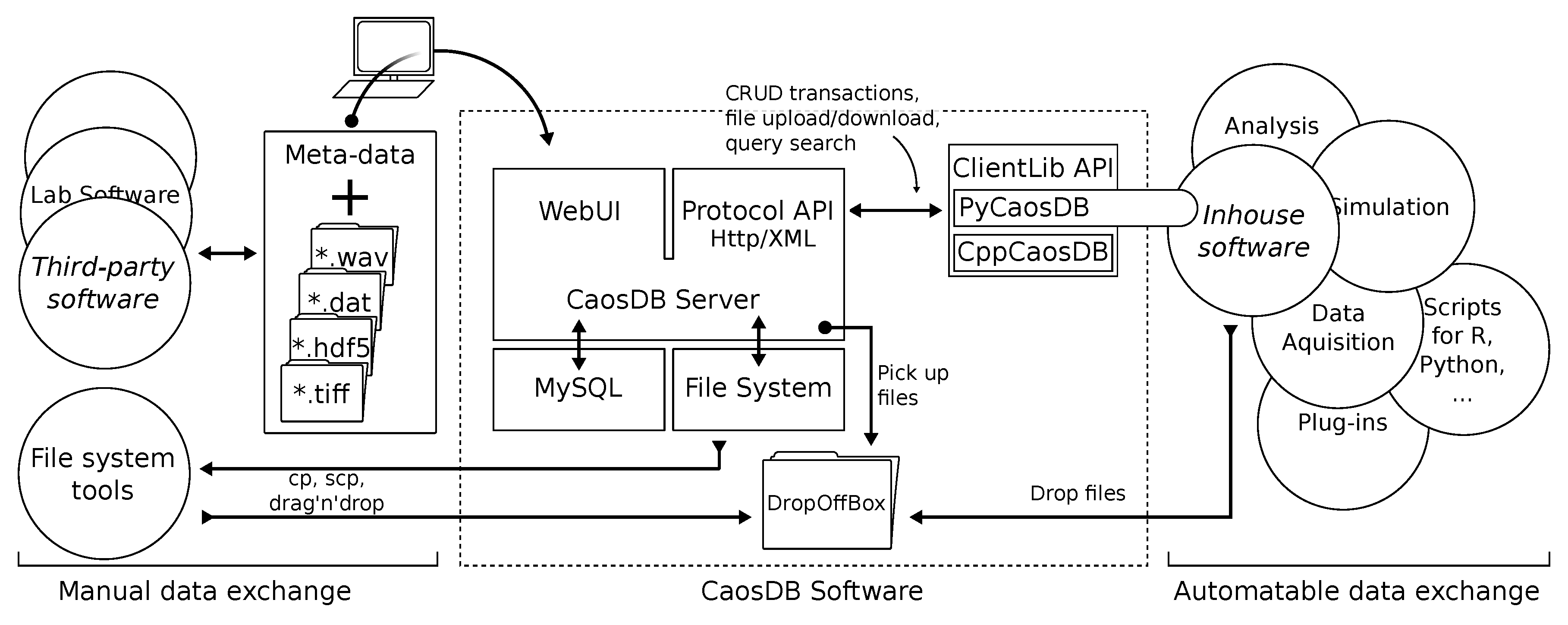
2.3. Architecture
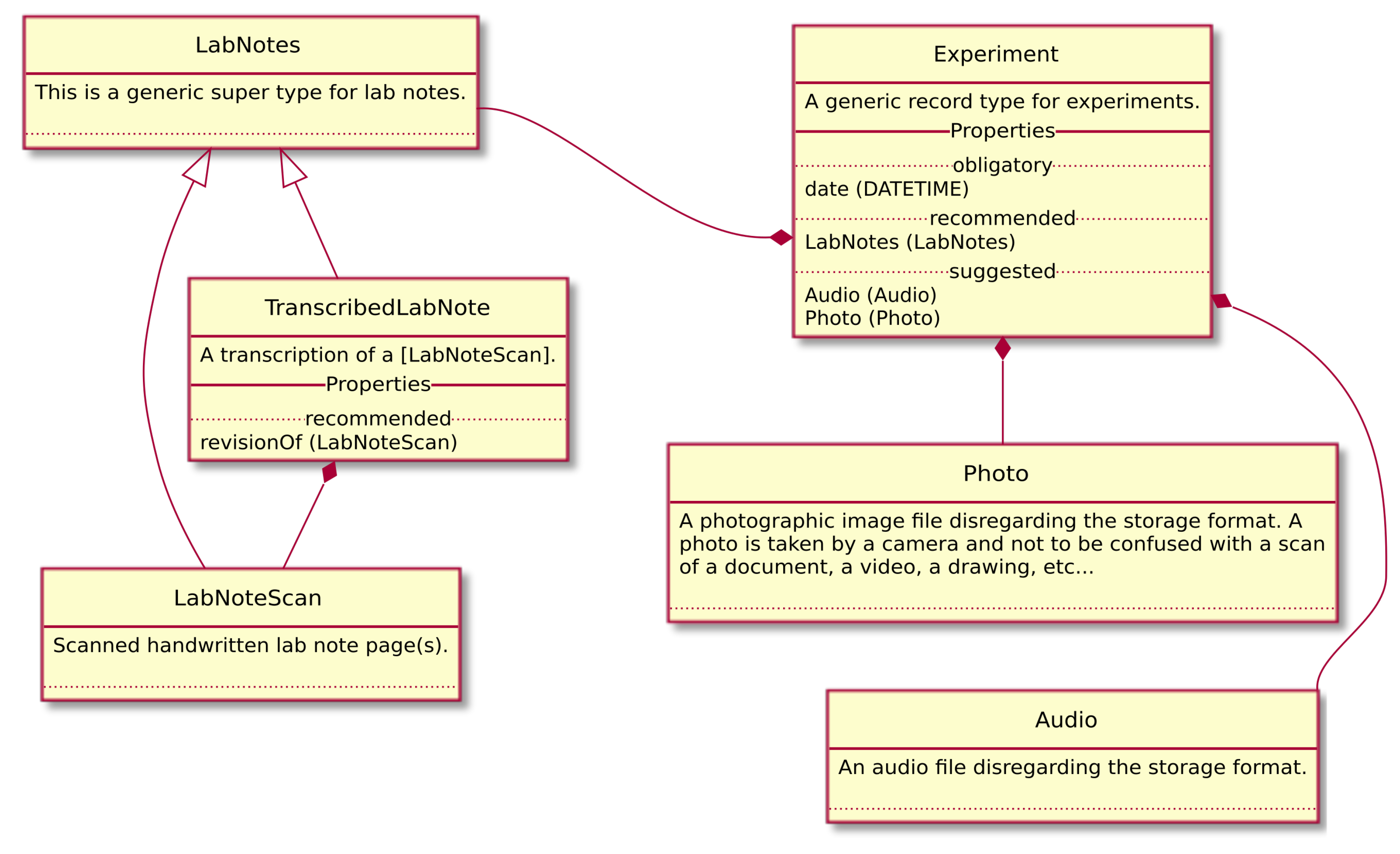
2.4. Data Model
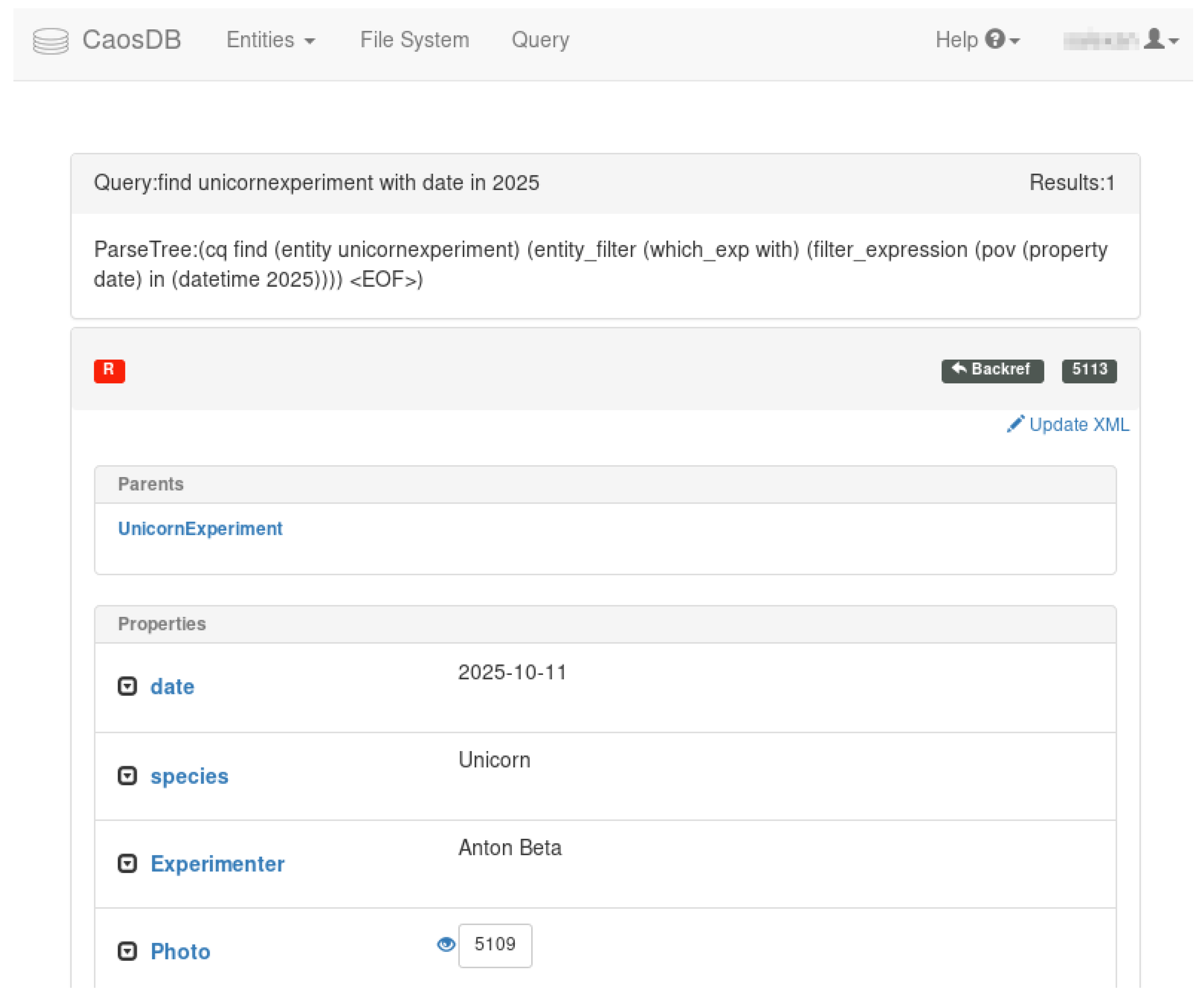
2.5. Query Language
(?date >= xsd:date("2017-01-01") && ?date < xsd:date("2018-01-01"))
The implementation of a temperature filter covering unit conversion would even rely on external unit conversion extensions.Find Experiment with date in 2017 and room temperature=293.15K
- A query starting with Count returns a non-negative integer.
- A query starting with Find returns a list of entities.
- A query starting with Select returns a table containing the values of selected Properties.
COUNT Experiment with date in 2017will return the number of experiments from 2017. In this query, Experiment is typically the name of a Record Type with a possibly large number of subtypes and instances. All Entities which have the name Experiment or have a parent with this name are filtered for those which have a Property with the name date and a date value in the year 2017. CQL filters can also express the equivalence of complex SQL joins in an easily understandable syntax:
FIND Person which is referenced as an Author by an Article which has aTitle like *terminating ventricular fibrillation*
SELECT first name, family name from person with date of birth > 2000will appear as an HTML table in the WebUI (downloadable as a tsv table), with three columns—id, first name, and family name. This feature is intended to provide one of the interfaces between CaosDB and existing scientific workflows.
2.6. User Management and Access Control
3. Discussion
Future Work
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| RDMS | Research Data Management System |
| FAIR | Findable, Accessible, Interoperable and Reusable |
| REST | Representational State Transfer |
| CRUD | Create/Read/Update/Delete |
| ACID | Atomicity, Consistency, Isolation, Durability |
| API | Application Programming Interface |
| XML | Extensible Markup Language |
| HTTP | Hypertext Transfer Protocol |
| XSLT | Extensible Stylesheet Language Transformations |
| HTML | Hypertext Markup Language |
| SQL | Structured Query Language |
| UI | User Interface |
| CQL | CaosDB Query Language |
| GNU | GNU’s not Unix |
| PAM | Pluggable Authentication Modules |
| IP | Internet Protocol |
| EBNF | Extended Backus-Naur Form |
| RDF | Resource Description Framework |
| OWL | Web Ontology Language |
| UML | Unified Modeling Language |
| SPARQL | SPARQL Protocol and RDF Query Language |
References
- Nelson, E.K.; Piehler, B.; Eckels, J.; Rauch, A.; Bellew, M.; Hussey, P.; Ramsay, S.; Nathe, C.; Lum, K.; Krouse, K.; et al. LabKey Server: An open source platform for scientific data integration, analysis and collaboration. BMC Bioinform. 2011, 12, 71. [Google Scholar] [CrossRef] [PubMed]
- Anderson, N.R.; Lee, E.S.; Brockenbrough, J.S.; Minie, M.E.; Fuller, S.; Brinkley, J.; Tarczy-Hornoch, P. Issues in Biomedical Research Data Management and Analysis: Needs and Barriers. J. Am. Med. Inform. Assoc. 2007, 14, 478–488. [Google Scholar] [CrossRef] [PubMed]
- Wruck, W.; Peuker, M.; Regenbrecht, C.R. Data management strategies for multinational large-scale systems biology projects. Brief. Bioinform. 2014, 15, 65–78. [Google Scholar] [CrossRef] [PubMed]
- Rocha da Silva, J.; Aguiar Castro, J.; Ribeiro, C.; Correia Lopes, J. Dendro: Collaborative Research Data. In The Semantic Web: ESWC 2014 Satellite Events; Presutti, V., Blomqvist, E., Troncy, R., Sack, H., Papadakis, I., Tordai, A., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 483–487. [Google Scholar]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.W.; da Silva Santos, L.B.; Bourne, P.E.; et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 2016, 3, 160018. [Google Scholar] [CrossRef] [PubMed]
- Marcus, D.S.; Olsen, T.; Ramaratnam, M.; Buckner, R.L. XNAT: A software framework for managing neuroimaging laboratory data. In Proceedings of the 12th Annual Meeting of the Organization for Human Brain Mapping, Florence, Italy, 11–15 June 2006. [Google Scholar]
- Kanza, S.; Willoughby, C.; Gibbins, N.; Whitby, R.; Frey, J.G.; Erjavec, J.; Zupančič, K.; Hren, M.; Kovač, K. Electronic lab notebooks: Can they replace paper? J. Cheminform. 2017, 9, 31. [Google Scholar] [CrossRef] [PubMed]
- Schweiger, D.; Trajanoski, Z.; Pabinger, S. SPARQLGraph: A web-based platform for graphically querying biological Semantic Web databases. BMC Bioinform. 2014, 15, 279. [Google Scholar] [CrossRef] [PubMed]
- Haag, F.; Lohmann, S.; Siek, S.; Ertl, T. QueryVOWL: A Visual Query Notation for Linked Data. In The Semantic Web: ESWC 2015 Satellite Events; Gandon, F., Guéret, C., Villata, S., Breslin, J., Faron-Zucker, C., Zimmermann, A., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 387–402. [Google Scholar]
- Chochiang, K.; Grabmann, B.; Betbeder, M.L.; Lapayre, J.C.; Sa-ngiamsak, C. OntoQuer: A Tool for Building SPARQL Query Automatically Applying with Our Ontologies. J. Softw. 2017, 12, 145–152. [Google Scholar]
- Fielding, R.T. Architectural Styles and the Design of Network-Based Software Architectures. Ph.D. Thesis, University of California, Irvine, CA, USA, 2000. [Google Scholar]
- Fitschen, T.; Hornung, D.; Schlemmer, A.; tom Wörden, H. CaosDB. 2018. Available online: http://dx.doi.org/10.17617/3.1s (accessed on 12 October 2018).
- Føllesdal, D.; Hilpinen, R. Deontic Logic: An Introduction. In Deontic Logic: Introductory and Systematic Readings; Hilpinen, R., Ed.; Springer: Dordrecht, The Netherlands, 1971; pp. 1–35. [Google Scholar]
- Free Software Foundation. GNU Affero General Public License; Free Software Foundation, Inc.: Boston, MA, USA, 2018. [Google Scholar]
| 1 | Atomicity, Consistency, Isolation, Durability. |
| 2 | Representational State Transfer. |
| 3 | Extended Backus-Naur Form. |

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fitschen, T.; Schlemmer, A.; Hornung, D.; tom Wörden, H.; Parlitz, U.; Luther, S. CaosDB—Research Data Management for Complex, Changing, and Automated Research Workflows. Data 2019, 4, 83. https://doi.org/10.3390/data4020083
Fitschen T, Schlemmer A, Hornung D, tom Wörden H, Parlitz U, Luther S. CaosDB—Research Data Management for Complex, Changing, and Automated Research Workflows. Data. 2019; 4(2):83. https://doi.org/10.3390/data4020083
Chicago/Turabian StyleFitschen, Timm, Alexander Schlemmer, Daniel Hornung, Henrik tom Wörden, Ulrich Parlitz, and Stefan Luther. 2019. "CaosDB—Research Data Management for Complex, Changing, and Automated Research Workflows" Data 4, no. 2: 83. https://doi.org/10.3390/data4020083
APA StyleFitschen, T., Schlemmer, A., Hornung, D., tom Wörden, H., Parlitz, U., & Luther, S. (2019). CaosDB—Research Data Management for Complex, Changing, and Automated Research Workflows. Data, 4(2), 83. https://doi.org/10.3390/data4020083




