Abstract
In this research dataset, we summarize for the first time volatile organic compounds (VOCs) emitted in vivo from ripening wine grapes. We studied four cold-hardy cultivars grown in the Midwestern U.S.: St. Croix, Frontenac, Marquette, and La Crescent. These cultivars have gained popularity among local growers and winemakers, but still very little is known about their performance compared with long-established V. vinifera grapes. Volatiles were collected using two novel approaches: biogenic emissions from grape clusters on a vine and single grape berries. A third approach was headspace collection of volatiles from crushed grapes. Solid-phase microextraction (SPME) was used to collect volatiles. Vacuum-assisted SPME was used in the case of single grape berry. Collected VOCs were analyzed using separation and identification on a gas chromatograph mass spectrometer (GC-MS). More than 120 VOCs were identified using mass spectral libraries. The dataset provides evidence that detecting biogenic emissions from growing grapes is feasible. The dataset provides a record of temporal and spatial variability of VOCs, many of which could potentially impart aroma and flavor in the wine. The number of VOCs detected followed the order from single berry (the least) to crushed berry (the most). Thus, more information for potential use in harvesting in order to obtain the desired flavor is found in data from crushed grapes.
Dataset:www.mdpi.com/2306-5729/4/1/22/s1.
Dataset License: CC-BY
1. Summary
Volatile organic compounds (VOCs) emitted in vivo from cold-hardy grape cultivars Frontenac, Marquette, St. Croix, and La Crescent grape berries from veraison to harvest were captured by two novel methods using solid-phase microextraction (SPME) and characterized by a gas chromatograph mass spectrometer (GC-MS). Whole cluster VOC characterization was performed in simple custom-made polyvinyl fluoride (PVF) film bags fitted with a custom SPME sampling port. Single grape berry VOC characterization was performed using a modified glass vial and vacuum-assisted SPME. In addition, berries were collected in the vineyard, crushed, and analyzed using an automated headspace SPME method in the lab. The data is a summary of these biogenic volatiles from all three sampling methods, from two different vineyards, four cold-hardy grape cultivars, and selected sampling timepoints approximately corresponding to a 2-degree rise in Brix (as % sugar content).
This dataset can be used to link biogenic amines from ripening grapes to the aroma profile of the wines made from these grapes. An automated SPME method for aroma analysis in wines has been described elsewhere [1], and used to characterize aromas in “Frontenac’ and “Marquette” wines [2], and “Brianna” and “Frontenac gris” wines [3], and even concentrations of pigments and tannins in “Frontenac” and “Marquette” berries [4] harvested at different stages of berry ripening. It is important to understand berry chemistry (i.e., aroma, pH, titratable acidity, Brix) in order to make quality wines, and this dataset serves as the starting point to help growers farm for flavor.
2. Data Description
The data provided in Supplementary Materials (Table S1) in spreadsheet format is the summary of biogenic volatiles emitted from four selected cold-hardy grape cultivars during ripening. Each row contains information about the Chemical Abstracts Service (CAS) number, volatile compound name, gas chromatography (GC) retention time in minutes, site location (Iowa or South Dakota), sampling method (whole cluster, single berry, crushed berries), sampling date, relative abundance of the volatile compound (in peak area counts), percent match to mass spectral libraries, and published aroma descriptors from two databases [5,6], as read from left to right in the spreadsheet. Each row represents data for a single vine, that is, three or four replicates were samples at the same timepoint. Blanks (i.e., ambient air samples collected at each vineyard) were used to account for potential interfering compounds. Small variations in the GC column retention time in Table S1 are typical with manual injection of SPME-based samples. A key is also provided in the sample table footer (Table 1). Data analysis including multivariate analyses of the entire dataset in Table S1 is provided elsewhere [7].

Table 1.
Example of a summary of biogenic volatiles emitted from four cold-hardy grape cultivars during ripening. Complete summary (~4800 rows) is in Table S1 (Supplementary Materials).
3. Methods
3.1. Overview
Research vineyards are located at NE Hansen research farm at South Dakota State University (SDSU, Brookings, SD, USA) and Iowa State University Horticulture Farm (ISU, Ames, IA, USA). Cultivars were grown in a randomized complete block design in both test vineyards in 2008 as part of the NE-1020 Cold Hardy Wine Grape Cultivar Trial. Grape clusters were randomly selected and tagged, so the same clusters were sampled from veraison to harvest for VOCs emitted in vivo from ripening grapes. Two in vivo sampling methods utilizing SPME were used, namely, extractions from whole air chambers enveloping ripening clusters in the 2012 season and vacuum-assisted extractions from single berries using modified glass vials in the 2013 season, respectively. Volatiles present in the headspace of crushed berries (i.e., skin, pulp, and seeds) were also investigated for the 2012 and 2013 growing seasons for comparisons with in vivo sampling. Berries from each cultivar were collected on the same day as in vivo sampling, collected from untagged clusters, crushed, and analyzed in comparison. These berries were kept in the dark and frozen at −20 °C for batch analysis after harvest.
A 65 µm polydimethylsiloxane/divinylbenzene (PDMS/DVB) fused silica SPME fiber (P/N 57310-U, Sigma-Aldrich, St. Louis, MO, USA) was conditioned according to the manufacturer’s specifications. Cleaned fibers were stored and transported in aluminum foil packets and sealed in mason jars. Foil and mason jars had been thermally cleaned overnight in an oven (at 110 °C) to minimize interferences from the environment. After sampling, described in the following sections, SPME fibers are wrapped in clean aluminum foil, placed and sealed in clean glass mason jars, and transported on ice packs to the lab for analysis on the same day. Samples of the volatiles collected on SPME fibers in Brookings, SD were prepared and used in the same manner and sent to Ames, IA via an overnight carrier for analyses. Volatiles emitted from whole clusters of Frontenac and Marquette cultivars were investigated for the 2012 growing season in both Iowa and South Dakota. Volatiles emitted from single berries of St. Croix and La Crescent cultivars were investigated for the 2013 growing season.
3.2. Non-Destructive Sampling of Biogenic Volatiles
The feasibility of “in vivo” sampling with SPME fibers and the optimization of coatings and extraction times were investigated prior to field sampling using Chilean seeded table grapes from a local grocery store (Figure A1, Figure A2 and Figure A3). Based on the results of the feasibility test, a 65 μm PDMS/DVB SPME fiber coating type and 30-min sampling time were selected as parameters that could enable the collection of the largest number of VOCs, with the least variability in the least amount of time. Thus, SPME fibers were exposed for 30 min for static collection and pre-concentration of VOCs in the “headspace” of both PVF chambers and glass vials. Fibers were cleaned by thermal desorption at 260 °C under a flow of clean helium prior to each sampling.
3.3. Biogenic Volatiles from Single Berry Grape from a Cluster on the Vine
Borosilicate glass vials were modified by removing flat bottoms of standard 2 mL screw top vials fitted with 9 mm PTFE/silicone lined septa (Fisher Scientific, Waltham, MA, USA). The edges of the modified vials were flared and rounded at a chemistry glass shop. A half-hole, cylindrical septa (P/N 20668, Sigma-Aldrich, St. Louis, MO, USA) was added to the outside of the screw top to support the SPME fiber assembly (without a manual holder) during sampling. The SPME fiber was placed through the septa prior to sampling. After assembly and placement of the vial apparatus on the individual berry (Figure 1), 5 mL of air was pulled from the vial using a syringe. Care was taken not to disturb the SPME fiber with the syringe needle. The resultant vacuum held the apparatus in place (i.e., sealed by suction onto the berry surface) while the SPME fiber was exposed for vacuum-assisted VOC sampling for 30 min. The single berry sampling vials were cleaned prior to each sampling by rinsing in deionized water and oven baked overnight at 107 °C. Cleaned vials were transported in an aluminum lined box. PTFE screw tops were replaced after each sampling.
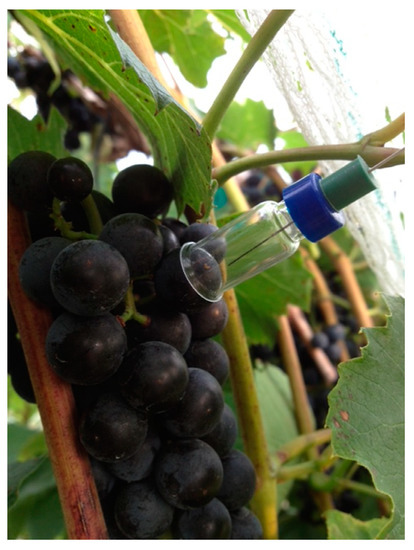

Figure 1.
Biogenic volatiles were collected by solid-phase microextraction (SPME) released from grape skin. A glass vial was modified for vacuum-assisted SPME of volatile organic compounds (VOC)s from a single grape. The vacuum was created with a syringe prior to SPME, so the vial was adhering to the skin and sealing headspace for SPME.
3.4. Biogenic Volatiles from Whole Grape Clusters on the Vine
Sampling chambers for the non-destructive detection of in vivo volatile metabolite were made from a rectangular sheet of PVF film, folded over itself, and heat-sealed on three sides (Figure 2). The PVF chambers were held open with an inner aluminum cage frame, allowing the chambers to be supported during sampling, and providing a consistent ~5 L headspace for sampling capacity. The PVF chambers were fitted with custom SPME sampling ports, consisting of an 11 mm polytetrafluoroethylene (PTFE) port and PTFE-lined silicone septa. These chambers with attached sampling ports for SPME insertion were preconditioned by thermal desorption before first use at 107 °C for 48 h. The PVF chambers were also cleaned prior to each sampling by rinsing in deionized water and oven baked overnight at 107 °C. Cleaned chambers were wrapped in aluminum foil for transport and to minimize the risk of light-induced changes to sample integrity. Aluminum wire support cages were also kept in an aluminum lined box. PTFE-lined silicone septa were replaced after each sampling (lasting 30 min).

Figure 2.
Biogenic volatiles were collected with SPME using an enclosure made from a polyvinyl fluoride (PVF) film to temporarily envelop a whole growing cluster of grapes (left). SPME is inserted into enclosure via a sampling port (right).
3.5. Volatiles Released from Crushed Grape Berries
Berries were collected from each cultivar on the same day and time of in vivo sampling. Five berries were collected from clusters adjacent to the cluster tagged for in vivo sampling (i.e., from the same vine but a different cluster than the in vivo sampled berries). Collected berries were frozen prior to analysis and stored in a −20 °C freezer. Berries collected in South Dakota were also frozen and shipped on blue ice blocks overnight for analysis in Iowa. Frozen berries were hand-crushed in the lab, placed into 20 mL amber screw top vials (P/N: 16-6000, Wheaton, Millville, NJ USA) with PTFE/silicone septa. A CTC CombiPal (LEAP Technologies, Carrboro, NC, USA) was used for automated SPME sampling. Briefly, the vials were agitated and heated to 50 °C for 10 min, followed by 30 min agitated headspace sampling using a 65 μm PDMS/DVB SPME fiber. The fiber was thermally desorbed under a flow of helium prior to each sample exposure. Optimized SPME parameters (sampling time, extraction temperature) were determined, data not shown.
3.6. Data Acquisition and Analysis
A custom multidimensional GC was used (Microanalytics, a part of Volatile Analysis Corporation, Round Rock, TX, USA), built on a standard Agilent 6890 platform (Agilent Technologies, Santa Clara, CA, USA). System automation and data acquisition software were MultiTrax v. 6.00.1 (Microanalytics, Round Rock, TX, USA) and ChemStation E.01.01.335 (Agilent Technologies, Santa Clara, CA, USA). Chromatography was performed on two capillary columns connected in series. The use of conventional retention indexes (e.g., Kovats retention index) is not appropriate for identification in this type of column configuration. The first column was 5% phenyl polysilphenylene-siloxane (30 m × 0.53 mm inner diameter × 0.5 μm thickness, Trajan Scientific, Austin, TX, USA) with a fixed restrictor pre-column. The second polar column was bonded polyethylene glycol in a Sol-Gel matrix (30 m × 0.53 mm inner diameter × 0.5 μm thickness, Trajan Scientific, Austin, TX, USA). The midpoint between the two columns was maintained at a constant pressure of 0.39 atm by a pneumatic switch. In this research, all effluent from the first column was directed into the second analytical column, that is, no heartcutting was performed. True multidimensional analyses were not performed (i.e., the system was used in full heartcut mode), meaning separation was performed on both columns in series. Effluent from the second polar column was simultaneously directed to a single quadrupole mass spectrometer (MS) (Model 5973N, Agilent Technologies, Santa Clara, CA, USA) and an olfactometry (sniff) port via an open spit interface at atmospheric pressure. The sniff port was equipped with a purge flow controller and supplied with humidified air at 0.54 atm. Flow to the MS and sniff port was determined by fixed restrictor columns, one part to the MS and three parts to the sniff port. Olfactometry was not utilized in the research. The GC inlet was operated in splitless mode at 250 °C. GC oven parameters started with an initial temperature of 40 °C, held for 3.0 min, followed by a 7 °C per min ramp to 240 °C, held for 8.43 min. Total run time was 40 min. Carrier gas was ultra-high purity (UHP) helium (99.999%, Airgas, Des Moines, IA, USA). Temperatures of the sniff port and MS transfer lines were 240 °C and 280 °C, respectively. MS full scan range was set from 34 m/z to 350 m/z. Scans were collected in electron ionization (EI) mode with an ionization energy of 70 eV. MS heated zones for quadrupole and source were 150 °C and 230 °C, respectively. Daily tuning of the MS was performed with perfluorotributylamine (PFTBA) before each analysis.
Identification of compounds was performed using the Automated Mass Spectral Deconvolution & Identification System (AMDIS) target library search with at least 80% mass spectral match [8]. Target libraries included (a) the 6 libraries that are included with the AMDIS program, (b) an onsite library created from analysis of pure standards (200+ compounds), and (c) NIST11 mass spectral library. Retention times were also verified with pure standards.
4. User Notes
Table S1 presented in spreadsheet format in Supplemental Materials has the same layout of columns and column headings as in Table 1. The data in Table S1 can be sorted and ordered. Nearly 4800 rows of data are available.
Supplementary Materials
The following are available online at http://www.mdpi.com/2306-5729/4/1/22/s1, Table S1: Summary of biogenic volatiles emitted from four cold-hardy grape cultivars during ripening.
Author Contributions
Conceptualization, J.A.K., S.R., A.F. and M.D.; methodology, S.R.; validation, S.R. and J.A.K.; formal analysis, S.R.; investigation, S.R. and D.L.M.; resources, D.L.M., S.R., A.F., J.A.K., and M.D.; data curation, S.R. and J.A.K.; writing—original draft preparation, S.R.; writing—review and editing, S.R., A.F., J.A.K., and M.D.; visualization, S.R.; supervision, J.A.K. and M.D.; project administration, J.A.K. and M.D.; funding acquisition, M.D. and J.A.K.
Funding
This research was funded by the United States Department of Agriculture’s Specialty Crops Research Initiative Program of the National Institute for Food and Agriculture, grant number 2011-51181-30850, titled “Northern grapes: integrating viticulture, winemaking, and marketing of new cold-hardy cultivars supporting new and growing rural wineries.” In addition, this project was partially supported by the Iowa Agriculture and Home Economics Experiment Station, Ames, Iowa and USDA National Institute of Food and Agriculture, Hatch project SD00H449-12 and SD Agriculture Experiment Station. Project No. IOW05556 (Future Challenges in Animal Production Systems: Seeking Solutions through Focused Facilitation) is sponsored by Hatch Act and State of Iowa funds.
Acknowledgments
The authors are thankful to Jason Vallone for his support with sample collection and laboratory analyses.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Appendix A
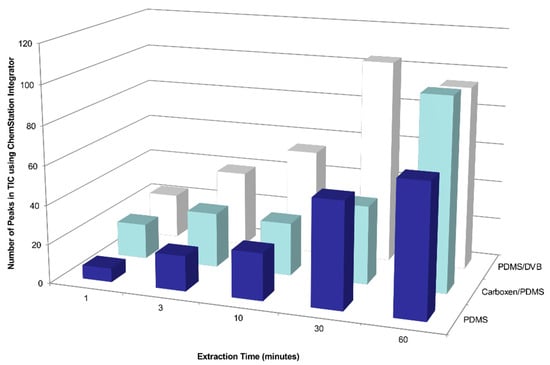
Figure A1.
Effects of SPME sampling time and coating type on the number of detected compounds emitted by a Chilean table grape cluster enclosed in a gas sampling bag made from polyvinyl fluoride film.
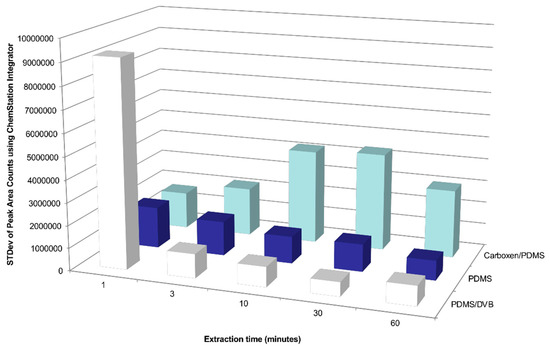
Figure A2.
Effects of SPME sampling time and coating type on standard deviation (STDev) of the total peak area counts of detected compounds emitted by a Chilean table grape cluster enclosed in a gas sampling bag made from polyvinyl fluoride film.
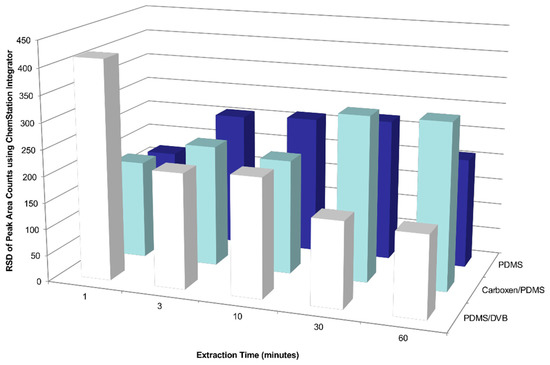
Figure A3.
Effects of SPME sampling time and coating type on the relative standard deviation (RSD) of the total peak area counts of detected compounds emitted by a Chilean table grape cluster enclosed in a gas sampling bag made from polyvinyl fluoride film.
References
- Cai, L.; Rice, S.; Koziel, J.A.; Dharmadhikari, M. Development of an automated method for aroma analysis of red wines from cold-hardy grapes using simultaneous solid-phase microextraction-multidimensional gas chromatography-mass spectrometry–olfactometry. Separations 2017, 4, 24. [Google Scholar] [CrossRef]
- Rice, S.; Lutt, N.; Koziel, J.A.; Dharmadhikari, M.; Fennell, A. Determination of selected aromas in Marquette and Frontenac wine using headspace-SPME coupled with GC-MS and simultaneous olfactometry. Separations 2018, 5, 20. [Google Scholar] [CrossRef]
- Rice, S.; Tursumbayeva, M.; Clark, M.; Greenlee, D.; Dharmadhikari, M.; Fennell, A.; Koziel, J.A. Effects of harvest time on aroma of white wines made from cold-hardy Brianna and Frontenac gris grapes using headspace solid-phase microextraction and gas-chromatography-mass-spectrometry-olfactometry. Foods 2019, 8, 29. [Google Scholar] [CrossRef] [PubMed]
- Rice, S.; Koziel, J.A.; Dharmadhikari, M.; Fennell, A. Evaluation of tannins and anthocyanins in Marquette, Frontenac, and St. Croix cold-hardy grape cultivars. Fermentation 2017, 3, 47. [Google Scholar] [CrossRef]
- Acree, T.; Arn, H. Flavornet and Human Odor Space. Available online: http://www.flavornet.org (accessed on 28 December 2018).
- The Good Scents Company Information System. Available online: http://www.thegoodscentscompany.com/index.html (accessed on 20 August 2018).
- Rice, S.; Maurer, D.L.; Fennell, A.; Dharmadhikari, M.; Koziel, J.A. Evaluation of volatile metabolites emitted in-vivo from cold-hardy grapes during ripening using SPME and GC-MS: A proof-of-concept. Molecules 2019. accepted. [Google Scholar]
- AMDIS. Automated Mass Spectral Decomposition and Identification System. Available online: http://amdis.net/ (accessed on 28 December 2018).
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).