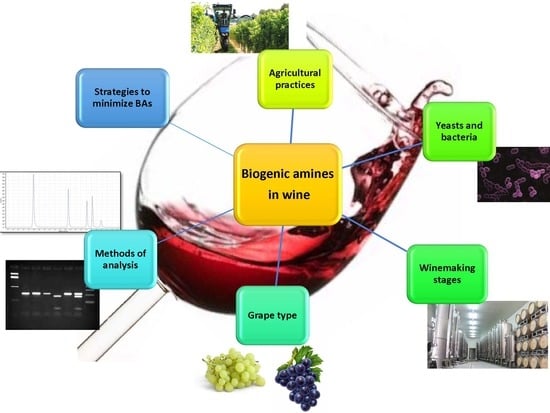
An Overview on Biogenic Amines in Wine
Abstract
1. Introduction
2. Biogenic Amines and Microorganisms
3. Biogenic Amines and the Type of Grape/Wine and Winemaking Stages
4. Agricultural and Oenological Practices
5. Detection and Quantification of BAs
5.1. Enzymatic Methods
5.2. Chromatographic Methods
5.3. Capillary Electrophoresis
5.4. Biosensors
5.5. Molecular Methods
6. BA Degradation
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| Him | Histamine |
| Tyr | Tyramine |
| Put | Putrescine |
| Cad | Cadaverine |
| Met | Methylamine |
| Agm | Agmatine |
| Phe | Phenylethylamine |
| Spm | Spermine |
| Spd | Spermidine |
| Try | Tryptamine |
| Ety | Ethylamine |
| Eth | Ethanolamine |
| Tea | Triethylamine |
| Tma | Trimethylamine |
| Dap | Diamine-propane |
| Ism | Isoamylamine |
References
- Al Bulushi, I.; Poole, S.; Deeth, H.C.; Dykes, G.A. Biogenic amines in fish: Roles in intoxication, spoilage, and nitrosamine formation—A review. Crit. Rev. Food Sci. Nutr. 2009, 49, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Pastore, P.; Favaro, G.; Badocco, D.; Tapparo, A.; Cavalli, S.; Saccani, G. Determination of biogenic amines in chocolate by ion chromatographic separation and pulsed integrated amperometric detection with implemented wave-form at Au disposable electrode. J. Chromatogr. A 2005, 1098, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Innocente, N.; D’Agostin, P. Formation of biogenic amines in a typical semihard Italian cheese. J. Food Prot. 2002, 65, 1498–1501. [Google Scholar] [CrossRef] [PubMed]
- Toro-Funes, N.; Bosch-Fuste, J.; Latorre-Moratalla, M.L.; Veciana-Nogués, M.T.; Vidal-Carou, M.C. Biologically active amines in fermented and non-fermented commercial soybean products from the Spanish market. Food Chem. 2015, 173, 1119–1124. [Google Scholar] [CrossRef] [PubMed]
- Suzzi, G.; Gardini, F. Biogenic amines in dry fermented sausages: A review. Int. J. Food Microbiol. 2003, 88, 41–54. [Google Scholar] [CrossRef]
- Favaro, G.; Pastore, P.; Saccani, G.; Cavalli, S. Determination of biogenic amines in fresh and processed meat by ion chromatography and integrated pulsed amperometric detection on Au electrode. Food Chem. 2007, 105, 1652–1658. [Google Scholar] [CrossRef]
- Maintz, L.; Novak, N. Histamine and histamine intolerance. Am. J. Clin. Nutr. 2007, 85, 1185–1196. [Google Scholar] [CrossRef] [PubMed]
- Smit, A.Y.Y.; Du Toit, W.J.J.; Du Toit, M. Biogenic amines in wine: Understanding the headache. S. Afr. J. Enol. Vitic. 2008, 29, 109–127. [Google Scholar] [CrossRef]
- Bauza, T.; Blaise, A.; Teissedre, P.; Cabanis, J.; Kanny, G.; Moneret-Vautrin, D.A.; Daumas, F. Les amines biogènes du vin. Métabolisme et toxicité. Bulletin de l’O.I.V. 1995, 68, 42–67. [Google Scholar]
- EFSA Panel on Biological Hazards. Scientific opinion on risk based control of biogenic amine formation in fermented foods. EFSA J. 2011, 9, 2393. [Google Scholar] [CrossRef]
- Kanny, G.; Gerbaux, V.; Olszewski, A.; Frémont, S.; Empereur, F.; Nabet, F.; Cabanis, J.-C.C.; Moneret-Vautrin, D.-A.A. No correlation between wine intolerance and histamine content of wine. J. Allergy Clin. Immunol. 2001, 107, 375–378. [Google Scholar] [CrossRef] [PubMed]
- Ramos, R.M.; Valente, I.M.; Rodrigues, J.A. Analysis of biogenic amines in wines by salting-out assisted liquid-liquid extraction and high-performance liquid chromatography with fluorimetric detection. Talanta 2014, 124, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Martuscelli, M.; Arfelli, G.; Manetta, A.C.; Suzzi, G. Biogenic amines content as a measure of the quality of wines of Abruzzo (Italy). Food Chem. 2013, 140, 590–597. [Google Scholar] [CrossRef] [PubMed]
- Bach, B.; Le Quere, S.; Vuchot, P.; Grinbaum, M.; Barnavon, L. Validation of a method for the analysis of biogenic amines: Histamine instability during wine sample storage. Anal. Chim. Acta 2012, 732, 114–119. [Google Scholar] [CrossRef] [PubMed]
- Comuzzo, P.; Rauhut, D.; Werner, M.; Lagazio, C.; Zironi, R. A survey on wines from organic viticulture from different European countries. Food Control 2013, 34, 274–282. [Google Scholar] [CrossRef]
- Preti, R.; Antonelli, M.L.; Bernacchia, R.; Vinci, G. Fast determination of biogenic amines in beverages by a core-shell particle column. Food Chem. 2015, 187, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Glória, M.B.A.; Watson, B.T.; Simon-Sarkadi, L.; Daeschel, M.A. A survey of biogenic amines in Oregon Pinot Noir and Cabernet Sauvignon wines. Am. J. Enol. Vitic. 1998, 49, 279–282. [Google Scholar]
- Vazquez-Lasa, M.B.; Iniguez-Crespo, M.; González-Larraina, M.; González-Guerrero, A. Biogenic amines in Rioja wines. Am. J. Enol. Vitic. 1998, 49, 229. [Google Scholar]
- Busto, O.; Mestres, M.; Guasch, J.; Borrull, F. Determination of biogenic amines in wine after clean-up by solid-phase extraction. Chromatographia 1995, 40, 404–410. [Google Scholar] [CrossRef]
- Ortega-Heras, M.; Pérez-Magariño, S.; Del-Villar-Garrachón, V.; González-Huerta, C.; Moro Gonzalez, L.C.; Guadarrama Rodríguez, A.; Villanueva Sanchez, S.; Gallo González, R.; Martín de la Helguera, S. Study of the effect of vintage, maturity degree, and irrigation on the amino acid and biogenic amine content of a white wine from the Verdejo variety. J. Sci. Food Agric. 2014, 94, 2073–2082. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Arribas, M.V.; Smit, A.Y.; du Toit, M. Biogenic amines and the winemaking process. In Understanding and Managing Wine Quality and Safety; Reynolds, A.G., Ed.; Woodhead Publishing Limited: Cambridge, UK, 2010. [Google Scholar]
- Preti, R.; Vinci, G. Biogenic amine content in red wines from different protected designations of origin of Southern Italy: Chemometric characterization and classification. Food Anal. Methods 2016, 9, 2280–2287. [Google Scholar] [CrossRef]
- Torrea, D.; Ancín, C. Content of biogenic amines in a Chardonnay wine obtained through spontaneous and inoculated fermentations. J. Agric. Food Chem. 2002, 50, 4895–4899. [Google Scholar] [CrossRef] [PubMed]
- Caruso, M.; Fiore, C.; Contursi, M.; Salzano, G.; Paparella, A.; Romano, P. Formation of biogenic amines as criteria for the selection of wine yeasts. World J. Microbiol. Biotechnol. 2002, 18, 159–163. [Google Scholar] [CrossRef]
- Granchi, L.; Romano, P.; Mangani, S.; Guerrini, S.; Vincenzini, M. Production of biogenic amines by wine microorganisms. Bulletin de l’O.I.V. 2005, 78, 595–609. [Google Scholar]
- Torrea, D.; Azpilicueta, C.A. Influence of yeast strain on biogenic amines content in wines: Relationship with the utilization of amino acids during fermentation. Am. J. Enol. Vitic. 2001, 52, 185–190. [Google Scholar]
- Tristezza, M.; Vetrano, C.; Bleve, G.; Spano, G.; Capozzi, V.; Logrieco, A.; Mita, G.; Grieco, F. Biodiversity and safety aspects of yeast strains characterized from vineyards and spontaneous fermentations in the Apulia Region, Italy. Food Microbiol. 2013, 36, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Landete, J.M.; de las Rivas, B.; Marcobal, A.; Muñoz, R. Molecular methods for the detection of biogenic amine-producing bacteria on foods. Int. J. Food Microbiol. 2007, 117, 258–269. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Martin-Alvarez, P.J.; Polo, M.C.; Munoz, R.; Moreno-Arribas, M.V. Formation of Biogenic Amines throughout the Industrial Manufacture of Red Wine. J. Food Prot. 2006, 69, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Arribas, M.V.; Polo, M.C.; Jorganes, F.; Muñoz, R. Screening of biogenic amine production by lactic acid bacteria isolated from grape must and wine. Int. J. Food Microbiol. 2003, 84, 117–123. [Google Scholar] [CrossRef]
- Henríquez-Aedo, K.; Durán, D.; Garcia, A.; Hengst, M.B.; Aranda, M. Identification of biogenic amines-producing lactic acid bacteria isolated from spontaneous malolactic fermentation of chilean red wines. LWT Food Sci. Technol. 2016, 68, 183–189. [Google Scholar] [CrossRef]
- Lucas, P.; Landete, J.; Coton, M.; Coton, E.; Lonvaud-Funel, A. The tyrosine decarboxylase operon of Lactobacillus brevis IOEB 9809: Characterization and conservation in tyramine-producing bacteria. FEMS Microbiol. Lett. 2003, 229, 65–71. [Google Scholar] [CrossRef]
- Romano, A.; Trip, H.; Lonvaud-Funel, A.; Lolkema, J.S.; Lucas, P.M. Evidence of two functionally distinct ornithine decarboxylation systems in lactic acid bacteria. Appl. Environ. Microbiol. 2012, 78, 1953–1961. [Google Scholar] [CrossRef] [PubMed]
- Molenaar, D.; Bosscher, J.S.; Ten Brink, B.; Driessen, A.J.M.; Konings, W.N. Generation of a proton motive force by histidine decarboxylation and electrogenic histidine/histamine antiport in Lactobacillus buchneri. J. Bacteriol. 1993, 175, 2864–2870. [Google Scholar] [CrossRef] [PubMed]
- Cruz Martín, M.; Fernández, M.; Linares, D.M.; Alvarez, M.A. Sequencing, characterization and transcriptional analysis of the histidine decarboxylase operon of Lactobacillus buchneri. Microbiology 2005, 151, 1219–1228. [Google Scholar] [CrossRef] [PubMed]
- Spano, G.; Russo, P.; Lonvaud-Funel, A.; Lucas, P.; Alexandre, H.; Grandvalet, C.; Coton, E.; Coton, M.; Barnavon, L.; Bach, B.; et al. Biogenic amines in fermented foods. Eur. J. Clin. Nutr. 2010, 64, S95–S100. [Google Scholar] [CrossRef] [PubMed]
- Pessione, A.; Lamberti, C.; Pessione, E. Proteomics as a tool for studying energy metabolism in lactic acid bacteria. Mol. Biosyst. 2010, 6, 1419–1430. [Google Scholar] [CrossRef] [PubMed]
- Pessione, E.; Mazzoli, R.; Giuffrida, M.G.; Lamberti, C.; Garcia-Moruno, E.; Barello, C.; Conti, A.; Giunta, C. A proteomic approach to studying biogenic amine producing lactic acid bacteria. Proteomics 2005, 5, 687–698. [Google Scholar] [CrossRef] [PubMed]
- Lonvaud-Funel, A. Lactic acid bacteria in the quality improvement and depreciation of wine. Antonie Van Leeuwenhoek 1999, 76, 317–331. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Arribas, V.; Lonvaud-Funel, A. Purification and characterization of tyrosine decarboxylase of Lactobacillus brevis IOEB 9809 isolated from wine. FEMS Microbiol. Lett. 2001, 195, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Guirard, B.; Snell, E. Purification and Properties of Ornithine Decarboxylase from Lactobacillus sp. 30a. J. Biol. Chem. 1980, 255, 5960–5964. [Google Scholar] [PubMed]
- Marcobal, Á.; Martín-Álvarez, P.J.; Moreno-Arribas, M.V.; Muñoz, R. A multifactorial design for studying factors influencing growth and tyramine production of the lactic acid bacteria Lactobacillus brevis CECT 4669 and Enterococcus faecium BIFI-58. Res. Microbiol. 2006, 157, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Gardini, F.; Özogul, Y.; Suzzi, G.; Tabanelli, G.; Özogul, F. Technological factors affecting biogenic amine content in foods: A review. Front. Microbiol. 2016, 7, 1218. [Google Scholar] [CrossRef] [PubMed]
- Landete, J.M.; Pardo, I.; Ferrer, S. Regulation of hdc expression and HDC activity by enological factors in lactic acid bacteria. J. Appl. Microbiol. 2008, 105, 1544–1551. [Google Scholar] [CrossRef] [PubMed]
- Wibowo, D.; Eschenbruch, R.; Davis, C.R.; Fleet, G.H.; Lee, T.H. Occurrence and growth of lactic acid bacteria in wine: A review. Am. J. Enol. Vitic. 1985, 36, 302–313. [Google Scholar]
- Coton, E.; Rollan, G.; Bertrand, A.; Lonvaud-Funel, A. Histamine-producing lactic acid bacteria in wines: Early detection, frequency and distribution. Am. J. Enol. Vitic. 1998, 49, 199–204. [Google Scholar]
- Guerrini, S.; Mangani, S.; Granchi, L.; Vincenzini, M. Biogenic Amine Production by Oenococcus oeni. Curr. Microbiol. 2002, 44, 374–378. [Google Scholar] [CrossRef] [PubMed]
- Costantini, A.; Cersosimo, M.; Del Prete, V.; Garcia-Moruno, E. Production of biogenic amines by lactic acid bacteria: Screening by PCR, thin-layer chromatography, and high-performance liquid chromatography of strains isolated from wine and must. J. Food Prot. 2006, 69, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Moruno, E.; Muñoz, R. Does Oenococcus oeni produce histamine? Int. J. Food Microbiol. 2012, 157, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; de Las Rivas, B.; Moreno-Arribas, M.V.; Muñoz, R. Identification of the ornithine decarboxylase gene in the putrescine-producer Oenococcus oeni BIFI-83. FEMS Microbiol. Lett. 2004, 239, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Coton, E.; Coton, M. Multiplex PCR for colony direct detection of Gram-positive histamine- and tyramine-producing bacteria. J. Microbiol. Methods 2005, 63, 296–304. [Google Scholar] [CrossRef] [PubMed]
- Lucas, P.M.; Claisse, O.; Lonvaud-Funel, A. High Frequency of Histamine-Producing Bacteria in the Enological Environment and Instability of the Histidine Decarboxylase Production Phenotype. Appl. Environ. Microbiol. 2008, 74, 811–817. [Google Scholar] [CrossRef] [PubMed]
- Costantini, A.; García-Moruno, E.; Moreno-Arribas, M.V. Biochemical Transformations Produced by Malolactic Fermentation; Springer: New York, NY, USA, 2009; ISBN 9780387741161. [Google Scholar]
- Moreno-Arribas, V.; Torlois, S.; Joyeux, A.; Bertrand, A.; Lonvaud-Funel, A. Isolation, properties and behaviour of tyramine-producing lactic acid bacteria from wine. J. Appl. Microbiol. 2000, 88, 584–593. [Google Scholar] [CrossRef] [PubMed]
- Sebastian, P.; Herr, P.; Fischer, U.; König, H. Molecular identification of lactic acid bacteria occurring in must and wine. S. Afr. J. Enol. Vitic. 2011, 32, 300–309. [Google Scholar] [CrossRef]
- Arena, M.E.; Fiocco, D.; Manca de Nadra, M.C.; Pardo, I.; Spano, G. Characterization of a Lactobacillus plantarum Strain Able to Produce Tyramine and Partial Cloning of a Putative Tyrosine Decarboxylase Gene. Curr. Microbiol. 2007, 55, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, N.; Hansen, W.; Engesser, D.; Hammes, W.P.; Holzapfel, W.H. Formation of histamine and tyramine by lactic acid bacteria in decarboxylase assay medium. Lett. Appl. Microbiol. 1990, 11, 278–281. [Google Scholar] [CrossRef]
- Del Prete, V.; Costantini, A.; Cecchini, F.; Morassut, M.; Garcia-Moruno, E. Occurrence of biogenic amines in wine: The role of grapes. Food Chem. 2009, 112, 474–481. [Google Scholar] [CrossRef]
- Zhijun, L.; Yongning, W.; Gong, Z.; Yunfeng, Z.; Changhu, X. A survey of biogenic amines in chinese red wines. Food Chem. 2007, 105, 1530–1535. [Google Scholar] [CrossRef]
- Landete, J.M.; Ferrer, S.; Polo, L.; Pardo, I. Biogenic amines in wines from three Spanish regions. J. Agric. Food Chem. 2005, 53, 1119–1124. [Google Scholar] [CrossRef] [PubMed]
- Soufleros, E.H.; Bouloumpasi, E.; Zotou, A.; Loukou, Z. Determination of biogenic amines in Greek wines by HPLC and ultraviolet detection after dansylation and examination of factors affecting their presence and concentration. Food Chem. 2007, 101, 704–716. [Google Scholar] [CrossRef]
- Pineda, A.; Carrasco, J.; Peña-Farfal, C.; Henríquez-Aedo, K.; Aranda, M. Preliminary evaluation of biogenic amines content in Chilean young varietal wines by HPLC. Food Control 2012, 23, 251–257. [Google Scholar] [CrossRef]
- Cecchini, F.; Morassut, M. Effect of grape storage time on biogenic amines content in must. Food Chem. 2010, 123, 263–268. [Google Scholar] [CrossRef]
- Leitão, M.C.; Marques, A.P.; San Romão, M.V. A survey of biogenic amines in commercial Portuguese wines. Food Control 2005, 16, 199–204. [Google Scholar] [CrossRef]
- Ough, C.S.; Daudt, C.E.; Crowell, E.A. Identification of new volatile amines in grapes and wines. J. Agric. Food Chem. 1981, 29, 938–941. [Google Scholar] [CrossRef] [PubMed]
- Broquedis, M.; Dumery, B.; Boucard, J. Mise en evidence de polyamines (putrescine, cadaverine, norspermidine, spermidine et spermine) dans les feuilles et les grappes de Vitis vinifera L. Connaiss. la Vigne du Vin 1989, 23, 1–6. [Google Scholar]
- Vidal-Carou, M.C.; Codony-Salcedo, R.; Mariné-Font, A. Histamine and tyramine in spanish wines: Relationships with total sulfur dioxide level, volatile acidity and malo-lactic fermentation intensity. Food Chem. 1990, 35, 217–227. [Google Scholar] [CrossRef]
- Herbert, P.; Cabrita, M.J.; Ratola, N.; Laureano, O.; Alves, A. Free amino acids and biogenic amines in wines and musts from the Alentejo region. Evolution of amines during alcoholic fermentation and relationship with variety, sub-region and vintage. J. Food Eng. 2005, 66, 315–322. [Google Scholar] [CrossRef]
- Bauza, T.; Kelly, M.T.; Blaise, A. Study of polyamines and their precursor amino acids in Grenache noir and Syrah grapes and wine of the Rhone Valley. Food Chem. 2007, 105, 405–413. [Google Scholar] [CrossRef]
- Soufleros, E.; Barrios, M.-L.; Bertrand, A. Correlation between the content of biogenic amines and other wine compounds. Am. J. Enol. Vitic. 1998, 49, 266–278. [Google Scholar]
- García-Marino, M.; Trigueros, Á.; Escribano-Bailón, T. Influence of oenological practices on the formation of biogenic amines in quality red wines. J. Food Compos. Anal. 2010, 23, 455–462. [Google Scholar] [CrossRef]
- Gerbaux, V.; Money, C. Biogenic amines in Burgundy wines. Contents and origin in wines. Rev. Fr. Oenol. 2000, 183, 25–28. [Google Scholar]
- Henríquez-Aedo, K.; Galarce-Bustos, O.; Aqueveque, P.; García, A.; Aranda, M. Dynamic of biogenic amines and precursor amino acids during cabernet sauvignon vinification. LWT 2018, 97, 238–244. [Google Scholar] [CrossRef]
- Ordóñez, J.L.; Callejón, R.M.; Troncoso, A.M.; García–Parrilla, M.C. Evaluation of biogenic amines profile in opened wine bottles: Effect of storage conditions. J. Food Compos. Anal. 2017, 63, 139–147. [Google Scholar] [CrossRef]
- Smit, A.Y.; du Toit, W.J.; Stander, M.; du Toit, M. Evaluating the influence of maceration practices on biogenic amine formation in wine. LWT Food Sci. Technol. 2013, 53, 297–307. [Google Scholar] [CrossRef]
- Martínez-Pinilla, O.; Guadalupe, Z.; Hernández, Z.; Ayestarán, B. Amino acids and biogenic amines in red varietal wines: The role of grape variety, malolactic fermentation and vintage. Eur. Food Res. Technol. 2013, 237, 887–895. [Google Scholar] [CrossRef]
- Marques, A.P.; Leitão, M.C.; San Romão, M.V. Biogenic amines in wines: Influence of oenological factors. Food Chem. 2008, 107, 853–860. [Google Scholar] [CrossRef]
- Hernandez-Orte, P.; Guitart, A.; Cacho, J. Changes in the Concentration of Amino Acids During the Ripening of Vitis vinifera Tempranillo Variety from the Denomination d’Origine Somontano (Spain). Am. J. Enol. Vitic. 1999, 50, 144–154. [Google Scholar]
- Soleas, G.J.; Carey, M.; Goldberg, D.M. Method development and cultivar-related differences of nine biogenic amines in Ontario wines. Food Chem. 1999, 64, 49–58. [Google Scholar] [CrossRef]
- Bauza, T.; Blaise, A.; Daumas, F.; Cabanis, J.C. Determination of biogenic amines and their precursor amino acids in wines of the Vallée du Rhône by high-performance liquid chromatography with precolumn derivatization and fluorimetric detection. J. Chromatogr. A 1995, 707, 373–379. [Google Scholar] [CrossRef]
- Martín-Álvarez, P.J.; Marcobal, A.; Polo, C.; Moreno-Arribas, M.V. Influence of technological practices on biogenic amine contents in red wines. Eur. Food Res. Technol. 2006, 222, 420–424. [Google Scholar] [CrossRef]
- Pogorzelski, E. Studies on the formation of histamine in must and wines from elderberry fruit. J. Sci. Food Agric. 1992, 60, 239–244. [Google Scholar] [CrossRef]
- Ancín-Azpilicueta, C.; González-Marco, A.; Jiménez-Moreno, N. Comparative study of the amine concentration in wines obtained from the traditional fermentation and from a more anaerobic fermentation method. LWT Food Sci. Technol. 2010, 43, 771–776. [Google Scholar] [CrossRef]
- Galgano, F.; Caruso, M.; Condelli, N.; Favati, F. Focused review: Agmatine in fermented foods. Front. Microbiol. 2012, 3, 199. [Google Scholar] [CrossRef] [PubMed]
- Tassoni, A.; Tango, N.; Ferri, M. Polyphenol and Biogenic Amine Profiles of Albana and Lambrusco Grape Berries and Wines Obtained Following Different Agricultural and Oenological Practices. Food Nutr. Sci. 2014, 5, 8–16. [Google Scholar] [CrossRef]
- Tassoni, A.; Tango, N.; Ferri, M. Comparison of biogenic amine and polyphenol profiles of grape berries and wines obtained following conventional, organic and biodynamic agricultural and oenological practices. Food Chem. 2013, 139, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Polo, M.C.; Martín-Álvarez, P.J.; Moreno-Arribas, M.V. Biogenic amine content of red Spanish wines: Comparison of a direct ELISA and an HPLC method for the determination of histamine in wines. Food Res. Int. 2005, 38, 387–394. [Google Scholar] [CrossRef]
- Henríquez-Aedo, K.; Vega, M.; Prieto-Rodríguez, S.; Aranda, M. Evaluation of biogenic amines content in chilean reserve varietal wines. Food Chem. Toxicol. 2012, 50, 2742–2750. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Alonso, S.; Hermosín-Gutiérrez, I.; García-Romero, E. Simultaneous HPLC analysis of biogenic amines, amino acids, and ammonium ion as aminoenone derivatives in wine and beer samples. J. Agric. Food Chem. 2007, 55, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Pereira, V.; Pontes, M.; Câmara, J.S.; Marques, J.C. Simultaneous analysis of free amino acids and biogenic amines in honey and wine samples using in loop orthophthalaldeyde derivatization procedure. J Chromatogr A 2008, 1189, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Önal, A.; Tekkeli, S.E.K.; Önal, C. A review of the liquid chromatographic methods for the determination of biogenic amines in foods. Food Chem. 2013, 138, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Bedia Erim, F. Recent analytical approaches to the analysis of biogenic amines in food samples. TrAC Trends Anal. Chem. 2013, 52, 239–247. [Google Scholar] [CrossRef]
- Jiang, H.-L.; Ying, L.-Y.; Zhou, S.-C.; Ying, M.; Shen, W.; Qiu, D.-H. Chromatographic determination of biogenic amines in wines after treatment with ionic liquids as novel media. J. Sep. Sci. 2011, 34, 1055–1062. [Google Scholar] [CrossRef] [PubMed]
- Arrieta, M.P.; Prats-Moya, M.S. Free amino acids and biogenic amines in Alicante Monastrell wines. Food Chem. 2012, 135, 1511–1519. [Google Scholar] [CrossRef] [PubMed]
- Kelly, M.T.; Blaise, A.; Larroque, M. Rapid automated high performance liquid chromatography method for simultaneous determination of amino acids and biogenic amines in wine, fruit and honey. J. Chromatogr. A 2010, 1217, 7385–7392. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-Q.; Ye, D.-Q.; Zhu, B.-Q.; Wu, G.-F.; Duan, C.-Q. Rapid HPLC analysis of amino acids and biogenic amines in wines during fermentation and evaluation of matrix effect. Food Chem. 2014, 163, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Saurina, J.; Hernández-Cassou, S. Chromatographic determination of amino acids by pre-column derivatization using 1,2-naphthoquinone-4-sulfonate as reagent. J. Chromatogr. A 1996, 740, 21–30. [Google Scholar] [CrossRef]
- Latorre, R.M.; Hernández-Cassou, S.; Saurina, J. Strategies for in-capillary derivatization of amino acids in capillary electrophoresis using 1,2-naphthoquinone-4-sulfonate as a labeling reagent. J. Chromatogr. A 2001, 934, 105–112. [Google Scholar] [CrossRef]
- Hlabangana, L.; Hernández-Cassou, S.; Saurina, J. Determination of biogenic amines in wines by ion-pair liquid chromatography and post-column derivatization with 1,2-naphthoquinone-4-sulphonate. J. Chromatogr. A 2006, 1130, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Jastrzębska, A.; Piasta, A.; Kowalska, S.; Krzemiński, M.; Szłyk, E. A new derivatization reagent for determination of biogenic amines in wines. J. Food Compos. Anal. 2016, 48, 111–119. [Google Scholar] [CrossRef]
- García-Moruno, E.; Carrascosa, A.V.; Muñoz, R. A rapid and inexpensive method for the determination of biogenic amines from bacterial cultures by thin-layer chromatography. J. Food Prot. 2005, 68, 625–629. [Google Scholar] [CrossRef] [PubMed]
- Romano, A.; Klebanowski, H.; La Guerche, S.; Beneduce, L.; Spano, G.; Murat, M.-L.; Lucas, P. Determination of biogenic amines in wine by thin-layer chromatography/densitometry. Food Chem. 2012, 135, 1392–1396. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, M.; Lambropoulou, D.; Morrison, C.; Kłodzińska, E.; Namieśnik, J.; Płotka-Wasylka, J. Literature update of analytical methods for biogenic amines determination in food and beverages. TrAC Trends Anal. Chem. 2018, 98, 128–142. [Google Scholar] [CrossRef]
- Ginterová, P.; Marák, J.; Staňová, A.; Maier, V.; Ševčík, J.; Kaniansky, D. Determination of selected biogenic amines in red wines by automated on-line combination of capillary isotachophoresis–capillary zone electrophoresis. J. Chromatogr. B 2012, 904, 135–139. [Google Scholar] [CrossRef] [PubMed]
- Daniel, D.; dos Santos, V.B.; Vidal, D.T.R.; do Lago, C.L. Determination of biogenic amines in beer and wine by capillary electrophoresis–tandem mass spectrometry. J. Chromatogr. A 2015, 1416, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Uzaşçı, S.; Başkan, S.; Erim, F.B. Biogenic Amines in Wines and Pomegranate Molasses—A Non-Ionic Micellar Electrokinetic Chromatography Assay with Laser-Induced Fluorescence Detection. Food Anal. Methods 2012, 5, 104–108. [Google Scholar] [CrossRef]
- Pereira, V.; Andreia, M.; Leca, J.M.; Marques, J.C. Analytical methodologies for the determination of biogenic amines in wines: An overview of the recent trends. J. Anal. Bioanal. Sep. Tech. 2017, 2, 52–57. [Google Scholar]
- Alonso-Lomillo, M.A.; Domínguez-Renedo, O.; Matos, P.; Arcos-Martínez, M.J. Disposable biosensors for determination of biogenic amines. Anal. Chim. Acta 2010, 665, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Di Fusco, M.; Federico, R.; Boffi, A.; Macone, A.; Favero, G.; Mazzei, F.; Di Fusco, M.; Federico, R.; Boffi, A.; Macone, A.; et al. Characterization and application of a diamine oxidase from Lathyrus sativus as component of an electrochemical biosensor for the determination of biogenic amines in wine and beer. Anal. Bioanal. Chem. 2011, 401, 707–716. [Google Scholar] [CrossRef] [PubMed]
- Henao-Escobar, W.; del Torno-de Román, L.; Domínguez-Renedo, O.; Alonso-Lomillo, M.A.; Arcos-Martínez, M.J. Dual enzymatic biosensor for simultaneous amperometric determination of histamine and putrescine. Food Chem. 2016, 190, 818–823. [Google Scholar] [CrossRef] [PubMed]
- Le Jeune, C.; Lonvaud-Funel, A.; ten Brink, B.; Hofstra, H.; van der Vossen, J.M. Development of a detection system for histidine decarboxylating lactic acid bacteria based on DNA probes, PCR and activity test. J. Appl. Bacteriol. 1995, 78, 316–326. [Google Scholar] [CrossRef] [PubMed]
- Lucas, P.; Lonvaud-Funel, A. Purification and partial gene sequence of the tyrosine decarboxylase of Lactobacillus brevis IOEB 9809. FEMS Microbiol. Lett. 2002, 211, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; de las Rivas, B.; Moreno-Arribas, M.V.; Munoz, R. Multiplex PCR Method for the Simultaneous Detection of Histamine-, Tyramine-, and Putrescine-Producing Lactic Acid Bacteria in Foods. J. Food Prot. 2005, 68, 874–878. [Google Scholar] [CrossRef] [PubMed]
- De las Rivas, B.; Marcobal, A.; Carrascosa, A.V.; Munoz, R. PCR Detection of Foodborne Bacteria Producing the Biogenic Amines Histamine, Tyramine, Putrescine, and Cadaverine. J. Food Prot. 2006, 69, 2509–2514. [Google Scholar] [CrossRef] [PubMed]
- Costantini, A.; Doria, F.; Vaudano, E.; Garcia-Moruno, E. Chemical and molecular methods for the control of biogenic amine production by microorganisms. Ann. Microbiol. 2011, 61, 173–178. [Google Scholar] [CrossRef]
- Nannelli, F.; Claisse, O.; Gindreau, E.; De Revel, G.; Lonvaud-Funel, A.; Lucas, P.M. Determination of lactic acid bacteria producing biogenic amines in wine by quantitative PCR methods. Lett. Appl. Microbiol. 2008, 47, 594–599. [Google Scholar] [CrossRef] [PubMed]
- Fernández, M.; del Río, B.; Linares, D.M.; Martín, M.C.; Alvarez, M.A. Real-Time Polymerase Chain Reaction for Quantitative Detection of Histamine-Producing Bacteria: Use in Cheese Production. J. Dairy Sci. 2006, 89, 3763–3769. [Google Scholar] [CrossRef]
- Coton, M.; Coton, E.; Lucas, P.; Lonvaud, A. Identification of the gene encoding a putative tyrosine decarboxylase of Carnobacterium divergens 508. Development of molecular tools for the detection of tyramine-producing bacteria. Food Microbiol. 2004, 21, 125–130. [Google Scholar] [CrossRef]
- Marcobal, Á.; de las Rivas, B.; García-Moruno, E.; Muñoz, R. The Tyrosine Decarboxylation Test Does Not Differentiate Enterococcus faecalis from Enterococcus faecium. Syst. Appl. Microbiol. 2004, 27, 423–426. [Google Scholar] [CrossRef] [PubMed]
- Fernández, M.; Linares, D.M.; Alvarez, M.A. Sequencing of the Tyrosine Decarboxylase Cluster of Lactococcus lactis IPLA 655 and the Development of a PCR Method for Detecting Tyrosine Decarboxylating Lactic Acid Bacteria. J. Food Prot. 2004, 67, 2521–2529. [Google Scholar] [CrossRef] [PubMed]
- Mohedano, M.L.; Spano, G.; López, P.; Russo, P. Controlling the formation of biogenic amines in fermented foods. In Advances in Fermented Foods and Beverages. Improving Quality, Technologies and Health Benefits; Holzapfel, W., Ed.; Woodhead Publishing: Cambridge, UK, 2014; pp. 273–310. [Google Scholar]
- García-Ruiz, A.; González-Rompinelli, E.M.; Bartolomé, B.; Moreno-Arribas, M.V. Potential of wine-associated lactic acid bacteria to degrade biogenic amines. Int. J. Food Microbiol. 2011, 148, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Capozzi, V.; Russo, P.; Ladero, V.; Fernández, M.; Fiocco, D.; Alvarez, M.A.; Grieco, F.; Spano, G. Biogenic Amines Degradation by Lactobacillus plantarum: Toward a Potential Application in Wine. Front. Microbiol. 2012, 3, 122. [Google Scholar] [CrossRef] [PubMed]
- Callejón, S.; Sendra, R.; Ferrer, S.; Pardo, I. Identification of a novel enzymatic activity from lactic acid bacteria able to degrade biogenic amines in wine. Appl. Microbiol. Biotechnol. 2014, 98, 185–198. [Google Scholar] [CrossRef] [PubMed]
- Beneduce, L.; Romano, A.; Capozzi, V.; Lucas, P.; Barnavon, L.; Bach, B.; Vuchot, P.; Grieco, F.; Spano, G. Biogenic amine in wines. Ann. Microbiol. 2010, 60, 573–578. [Google Scholar] [CrossRef]
- Bäumlisberger, M.; Moellecken, U.; König, H.; Claus, H.; Bäumlisberger, M.; Moellecken, U.; König, H.; Claus, H. The Potential of the Yeast Debaryomyces hansenii H525 to Degrade Biogenic Amines in Food. Microorganisms 2015, 3, 839–850. [Google Scholar] [CrossRef] [PubMed]
| Wine | Type of Wine | Biogenic Amine | Reference |
|---|---|---|---|
| Cabernet Franc | red | Eth,Ety, Put, Agm | [58] |
| Carmenere | red | Eth,Ety, Put, Agm | |
| Cesanese d’Affile | red | Eth,Ety, Put, Agm | |
| Merlot | red | Eth,Ety, Put, Agm | |
| Montepulciano | red | Eth,Ety, Put, Agm | |
| Sangiovese | red | Eth,Ety, Put, Agm | |
| Syrah | red | Eth,Ety, Put, Agm | |
| Aglianico (13) | red | Met, Eth, Agm, Pea, Put, Cad, Him, Spd, Tym | [22] |
| Primitivo di Manduria (15) | red | Eth, Agm, Pea, Put, Cad, Him, Spd, Tym | |
| Syrah (15) | red | Met, Eth, Agm, Pea, Put, Cad, Him, Spd, Tym | |
| Etna Rosso (13) | red | Met, Eth, Agm, Pea, Put, Cad, Him, Spd, Tym | |
| Chinese red wines from Shandong | red | Phe, Put, Cad, Him, Tyr, Spd and Spm | [59] |
| Chinese red wines from Beijing | red | Phe, Put, Cad, Him, Tyr, Spd and Spm | |
| Chinese red wines from Tianjin | red | Phe, Put, Cad, Him, Tyr, Spd and Spm | |
| Chinese red wines from Hebei | red | Phe, Put, Cad, Him, Tyr, Spd and Spm | |
| Chinese red wines from Xinjiang | red | Phe, Put, Cad, Him, Tyr | |
| Tempranillo from La Rioja, | red | Him, Tyr, Put, Phe | [60] |
| Tempranillo from Utiel-Requena | red | Him, Tyr, Put, Phe | |
| Tempranillo from Tarragona | red | Him, Tyr, Put, Phe | |
| Bobal | red | Him, Tyr, Put, Phe | |
| Garnacha | red | Him, Tyr, Put, Phe | |
| Xinomavro | red | [61] Note: amines >1 mg/L are indicated | |
| Roditis | white | Phe | |
| Agiorgitiko | red | Phe, Put | |
| Cabernet Sauvignon | red | ||
| Mantilaria | red | Ism, Him, Tym, Spd, Put | |
| Syrah | red | Put | |
| Merlot | red | Ism | |
| Debina | white | Cad | |
| Moshofilero | white | Put | |
| Malagouzia | white | ||
| Asyrtiko | red | Put, Cad | |
| Grenache rouge | red | Put | |
| Chardonnay | white | ||
| Muscat white | white | Ism, Put | |
| Muscat Hamburg | red | ||
| Muscat d’Alexandrie | white | Put, Cad, Him | |
| Limnio | red | Cad | |
| Chilean young wines: | Put, Him, Tyr, Spd | [62] | |
| Cabernet Sauvignon (9) | red | ||
| Merlot (8) | red | ||
| Carménère (10) | red |
| Biogenic Amines (mg/L) | Must | Must 72h | Wine/AF | Wine/MLF |
|---|---|---|---|---|
| Ethanolamine | 7.91 a | 8.73 a | 11.54 b | 11.65 b |
| Agmatime | nd a | nd a | 0.51 a | 6.4 b |
| Ethylamine | 9.83 b | 10.07 b | 1.87 a | 1.96 a |
| Tyramine | nd a | nd a | 0.36 b | 0.069 a |
| Putrescine | 11.23 d | 3.76 c | 1.76 a | 2.26 ab |
| Winery | Sample | Phe | Put | Cad | Him | Tyr | Spd | Spm | Total |
|---|---|---|---|---|---|---|---|---|---|
| A | grapes | nd | nd | nd | nd | nd | tr | tr | tr |
| AF | nd | 29.2 | tr | nd | nd | tr | tr | 29.2 | |
| MLF | tr | 11 | tr | nd | nd | tr | tr | 11 | |
| Wine | nd | 36.7 | tr | nd | nd | 1.6 | tr | 38.3 | |
| B | grapes | nd | nd | nd | nd | tr | tr | tr | tr |
| AF | nd | 4 | tr | nd | tr | tr | tr | 5 | |
| MLF | tr | 9.2 | tr | nd | tr | tr | tr | 9.2 | |
| Wine | nd | 5.5 | tr | nd | tr | tr | tr | 5.5 | |
| C | grapes | tr | 1.2 | tr | 1.9 | nd | 2.2 | 2.5 | 7.8 |
| AF | nd | 9.3 | tr | nd | tr | 1.3 | tr | 10.7 | |
| MLF | nd | 14.1 | tr | nd | nd | tr | tr | 14.2 | |
| Wine | 10 | 142.1 | tr | 8.4 | 8.1 | tr | tr | 168.6 | |
| D | grapes | nd | 8.9 | nd | nd | tr | 2.9 | tr | 11.8 |
| AF | nd | 38.6 | tr | tr | tr | 1.6 | tr | 40.2 | |
| MLF | nd | 11.7 | 1.3 | 1.3 | tr | 2.2 | tr | 13.9 | |
| Wine | tr | 8.5 | 3.5 | 3.5 | 1.5 | 2.7 | tr | 16.2 | |
| E | grapes | nd | 9.8 | tr | nd | tr | 4.1 | 1.2 | 15.1 |
| AF | nd | 14.7 | tr | nd | tr | 1.1 | tr | 15.8 | |
| MLF | nd | 12.9 | tr | nd | tr | 1.3 | tr | 14.2 | |
| Wine | nd | 14.3 | tr | nd | tr | 1.5 | tr | 15.8 |
| Primer | 5′ → 3′Sequence | Coding for | Reference |
|---|---|---|---|
| CL1 | CCWGGWAAWATWGGWAATGGWTA | hdc | [111] |
| CL2 | GAWGCWGTWGTCATATTWATTTGWCC | hdc | [111] |
| JV16HC | AGATGGTATTGTTTCTTATG | hdc | [111] |
| JV17HC | AGACCATACACCATAACCTT | hdc | [111] |
| JV17 | AGACCATACACCATAACCTTG | hdc | [46] |
| CL1mod | CCAGGWAACATTGGTAATGGATA | hdc | [60] |
| HDC3 | GATGGTATTGTTTCKTATGA | hdc | [51] |
| HDC4 | CAAACACCAGCATCTTC | hdc | [51] |
| PHDC1 | CCGTGCGGAAACAAAGAAT | hdc | [48] |
| PHDC2 | CCAAACACCAGCATCTTCA | hdc | [48] |
| HIS1-F | GGNATNGTNWSNTAYGAYMGNGCNGA | hdc | [114] |
| HIS1-R | ATNGCDATNGCNSWCCANACNCCRTA | hdc | [114] |
| Hdc1 | TTGACCGTATCTCAGTGAGTCCAT | hdc | [117] |
| Hdc2 | ACGGTCATACGAAACAATACCATC | hdc | [117] |
| 3 | GTNTTYAAYGCNGAYAARACNTAYTTYGT | odc | [50] |
| 16 | TACRCARAATACTCCNGGNGGRTANGG | odc | [50] |
| 4 | ATNGARTTNAGTTCRCAYTTYTCNGG | odc | [113] |
| 15 | GGTAYTGTTYGAYCGGAAWAAWCAYAA | odc | [113] |
| AODC1 | GMTCGTGAAATYAARCKG | odc | [48] |
| AODC2 | KGRGTTCMGCYGGRGTAT | odc | [48] |
| Put1-F | TWYMAYGCNGAYAARACNTAYYYTGT | odc | [114] |
| Put1-R | ACRCANAGNACNCCNGNGGRTANGG | odc | [114] |
| Put2-F | ATHWGNTWYGGNAAYACNATHAARAA | odc | [114] |
| Put2-R | GCNARNCCNCCRAAYTTNCCDARTC | odc | [114] |
| P2-for | GAYATIATIGGIATIGGIYTIGAYCARG | tdc | [112] |
| P1-rev | CCRTARTCIGGIATIGCRAARTCIGTRTG | tdc | [112] |
| 41 | CAYGTNGAYGCNGCNTAYGGNGG | tdc | [113] |
| 42 | AYRTANCCCATYTTRTGNGGRTC | tdc | [113] |
| Pt3 | TACACGTAGATGCTGCATATG | tdc | [48] |
| Pt4 | ATGGTTGACTATGTTTTAAAAGAA | tdc | [48] |
| p0303 | CCACTGCTGCATCTGTTTG | tdc | [118] |
| TD5 | CAAATGGAAGAAGAAGTAGG | tdc | [118] |
| TD2 | ACATAGTCAACCATRTTGAA | tdc | [118] |
| 57 | ATGAGTGAATCATTGTCG | tdc | [119] |
| 58 | TTATTTTGCTTCGCTTGCC | tdc | [119] |
| TDC1 | AACTATCGTATGGATATCAACG | tdc | [120] |
| TDC2 | TAGTCAACCATATTGAAATCTGG | tdc | [120] |
| TDC-F | TGGYTNGTNCCNCARACNAARCAYTA | tdc | [114] |
| TDC-R | ACRTARTCNACCATRTTRAARTCNGG | tdc | [114] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costantini, A.; Vaudano, E.; Pulcini, L.; Carafa, T.; Garcia-Moruno, E. An Overview on Biogenic Amines in Wine. Beverages 2019, 5, 19. https://doi.org/10.3390/beverages5010019
Costantini A, Vaudano E, Pulcini L, Carafa T, Garcia-Moruno E. An Overview on Biogenic Amines in Wine. Beverages. 2019; 5(1):19. https://doi.org/10.3390/beverages5010019
Chicago/Turabian StyleCostantini, Antonella, Enrico Vaudano, Laura Pulcini, Tommaso Carafa, and Emilia Garcia-Moruno. 2019. "An Overview on Biogenic Amines in Wine" Beverages 5, no. 1: 19. https://doi.org/10.3390/beverages5010019
APA StyleCostantini, A., Vaudano, E., Pulcini, L., Carafa, T., & Garcia-Moruno, E. (2019). An Overview on Biogenic Amines in Wine. Beverages, 5(1), 19. https://doi.org/10.3390/beverages5010019