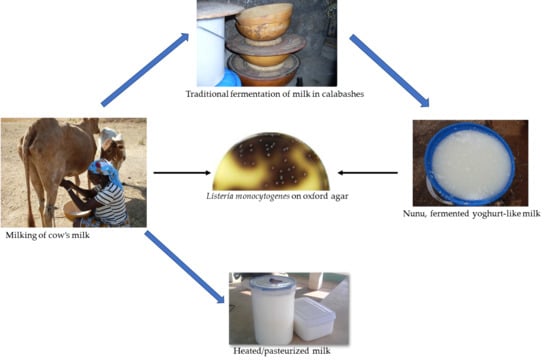
Prevalence and Characteristics of Listeria monocytogenes Isolates in Raw Milk, Heated Milk and Nunu, a Spontaneously Fermented Milk Beverage, in Ghana
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples
2.2. Isolation and Characterization of L. monocytogenes
2.3. PCR-Based Identification of L. monocytogenes
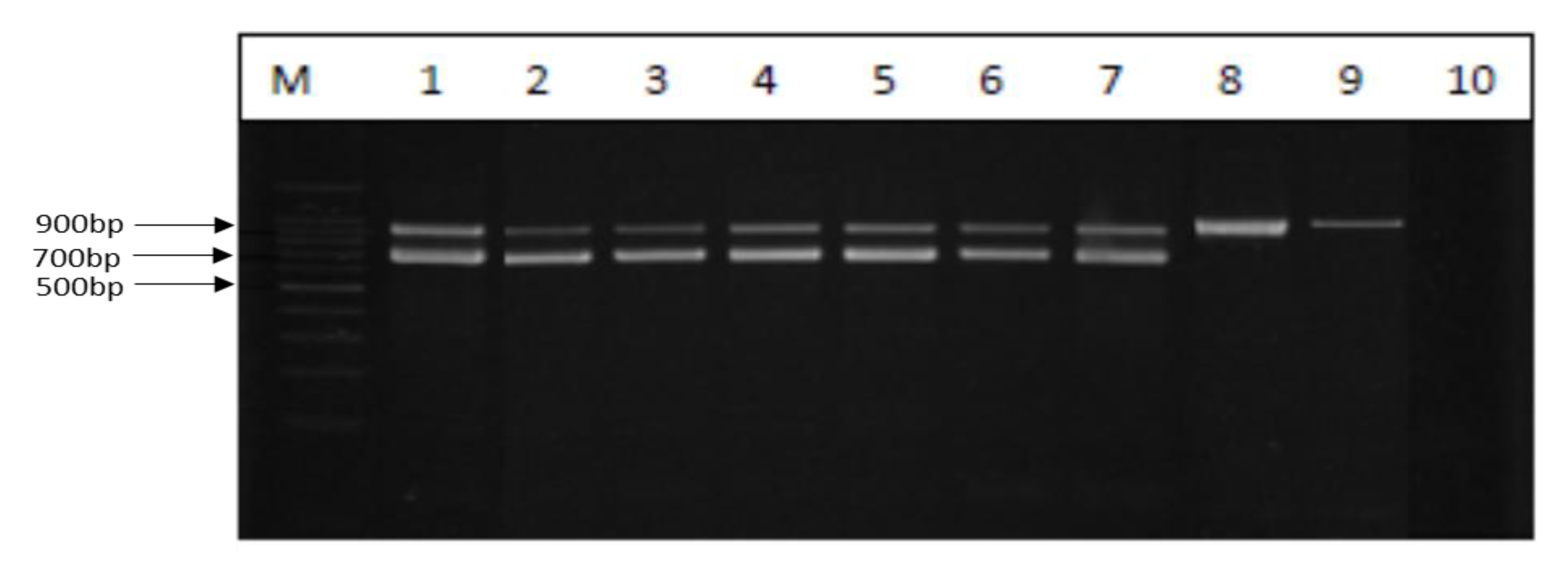
2.4. Molecular Serotyping of L. monocytogenes
2.5. Detection of Virulence Associated Genes in L. monocytogenes
2.6. Determination Antimicrobial Resistance among L. monocytogenes Isolates
3. Results and Discussion
3.1. Prevalence of L. monocytogenes in Raw Milk, Boiled Milk and Nunu
3.2. Molecular Serogroups of L. monocytogenes Isolates
3.3. Prevalence of Virulence-Associated Markers
3.4. Antibiotic Resistance among L. monocytogenes Isolates
4. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Kang, Q.; Guo, Y.; Hong, S.; Jiang, L. Listeria monocytogenes meningitis in infants: Report of two cases. Chin. J. Contemp. Pediatr. 2013, 15, 1150–1152. [Google Scholar]
- Thomas, M.K.; Vriezen, R.; Farber, J.M.; Currie, A.; Schlech, W.; Fazil, A. Economic cost of a Listeria monocytogenes outbreak in Canada, 2008. Foodborne Pathog. Dis. 2015, 12, 966–971. [Google Scholar] [CrossRef] [PubMed]
- Razavilar, V.; Genigeorgis, C. Prediction of Listeria spp. growth as affected by various levels of chemicals, pH, temperature and storage time in a model broth. Int. J. Food Microbiol. 1998, 40, 149–157. [Google Scholar] [CrossRef]
- Jamali, H.; Chai, L.C.; Thong, K.L. Detection and isolation of Listeria spp. and Listeria monocytogenes in ready-to-eat foods with various selective culture media. Food Control 2013, 32, 19–24. [Google Scholar] [CrossRef]
- Rahimi, E.; Ameri, M.; Momtaz, H. Prevalence and antimicrobial resistance of Listeria species isolated from milk and dairy products in Iran. Food Control 2010, 21, 1448–1452. [Google Scholar] [CrossRef]
- Wu, S.; Wu, Q.; Zhang, J.; Chen, M.; Hu, H. Listeria monocytogenes prevalence and characteristics in retail raw foods in China. PLoS ONE 2015, 10, e0136682. [Google Scholar] [CrossRef] [PubMed]
- Scallan, E.; Crim, S.M.; Runkle, A.; Henao, O.L.; Mahon, B.E.; Hoekstra, R.M.; Griffin, P.M. Bacterial enteric infections among older adults in the united states: Foodborne diseases active surveillance network, 1996–2012. Foodborne Pathog. Dis. 2015, 12, 492–499. [Google Scholar] [CrossRef] [PubMed]
- Kathariou, S. Listeria monocytogenes virulence and pathogenicity, a food safety perspective. J. Food Prot. 2002, 65, 1811–1829. [Google Scholar] [CrossRef] [PubMed]
- Salcedo, C.; Arreaza, L.; Alcala, B.; De La Fuente, L.; Vazquez, J. Development of a multilocus sequence typing method for analysis of Listeria monocytogenes clones. J. Clin. Microbiol. 2003, 41, 757–762. [Google Scholar] [CrossRef] [PubMed]
- Jensen, A.; Thomsen, L.E.; Jørgensen, R.L.; Larsen, M.H.; Roldgaard, B.B.; Christensen, B.B.; Vogel, B.F.; Gram, L.; Ingmer, H. Processing plant persistent strains of Listeria monocytogenes appear to have a lower virulence potential than clinical strains in selected virulence models. Int. J. Food Microbiol. 2008, 123, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Rump, L.; Zhang, Y.; Chen, Y.; Wang, X.; Meng, J. Molecular subtyping and virulence gene analysis of Listeria monocytogenes isolates from food. Food Microbiol. 2013, 35, 58–64. [Google Scholar] [CrossRef] [PubMed]
- Soni, D.K.; Singh, M.; Singh, D.V.; Dubey, S.K. Virulence and genotypic characterization of Listeria monocytogenes isolated from vegetable and soil samples. BMC Microbiol. 2014, 14, 241. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wu, Q.; Zhang, J.; Chen, M.; Guo, W. Analysis of multilocus sequence typing and virulence characterization of Listeria monocytogenes isolates from Chineseretail ready-to-eat food. Front. Microbiol. 2016, 7, 168. [Google Scholar] [CrossRef] [PubMed]
- Pesavento, G.; Ducci, B.; Nieri, D.; Comodo, N.; Nostro, A.L. Prevalence and antibiotic susceptibility of Listeria spp. isolated from raw meat and retail foods. Food Control 2010, 21, 708–713. [Google Scholar] [CrossRef]
- Wieczorek, K.; Dmowska, K.; Osek, J. Prevalence, characterization, and antimicrobial resistance of Listeria monocytogenes isolates from bovine hides and carcasses. Appl. Environ. Microbiol. 2012, 78, 2043–2045. [Google Scholar] [CrossRef] [PubMed]
- Bertsch, D.; Muelli, M.; Weller, M.; Uruty, A.; Lacroix, C.; Meile, L. Antimicrobial susceptibility and antibiotic resistance gene transfer analysis of foodborne, clinical, and environmental Listeria spp. isolates including Listeria monocytogenes. MicrobiologyOpen 2014, 3, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wu, Q.; Zhang, J.; Wu, S.; Guo, W. Prevalence, enumeration, and pheno-and genotypic characteristics of Listeria monocytogenes isolated from raw foods in South China. Front. Microbiol. 2015, 6, 1026. [Google Scholar] [CrossRef] [PubMed]
- Soni, D.K.; Singh, R.K.; Singh, D.V.; Dubey, S.K. Characterization of Listeria monocytogenes isolated from Ganges water, human clinical and milk samples at Varanasi, India. Infect. Genet. Evol. 2013, 14, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Akabanda, F.; Owusu-Kwarteng, J.; Tano-Debrah, K.; Glover, R.L.; Nielsen, D.S.; Jespersen, L. Taxonomic and molecular characterization of lactic acid bacteria and yeasts in nunu, a Ghanaian fermented milk product. Food Mcrobiol. 2013, 34, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Appiah, J.; Tano-Debrah, K.; Annor, G.A.; Alpha, M.M.; Makita, K.; Grace, D. Quantitative probabilistic assessment of the risk of Listeriosis from the consumption of milk from informal markets in Ghana. In Proceedings of the International Congress of Pathogens at the Human-Animal Interface (ICOPHAI), Addis Ababa, Ethiopia, 15–17 September 2011; International Livestock Research Institute (ILRI): Nairobi, Kenya, 2011. [Google Scholar]
- Owusu-Kwarteng, J.; Akabanda, F.; Johansen, P.; Jespersen, L.; Nielsen, D.S. Nunu, a West-African fermented yogurt-like milk product. In Yogurt in Health and Disease Prevention; Shah, N.P., Ed.; Elsevier: New York, NY, USA, 2017; pp. 275–283. [Google Scholar]
- Becker, B.; Schuler, S.; Lohneis, M.; Sabrowski, A.; Curtis, G.D.; Holzapfel, W.H. Comparison of two chromogenic media for the detection of Listeria monocytogenes with the plating media recommended by EN/DIN 11290-1. Int. J. Food Microbiol. 2006, 109, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Thong, K.; Lai, M.; Teh, C.; Chua, K. Simultaneous detection of methicillin-resistant Staphylococcus aureus, Acinetobacter baumannii, Escherichia coli, Klebsiella pneumoniae and Pseudomonas aeruginosa by multiplex PCR. Trop. Biomed. 2011, 28, 21–31. [Google Scholar] [PubMed]
- Rossmanith, P.; Krassnig, M.; Wagner, M.; Hein, I. Detection of Listeria monocytogenes in food using a combined enrichment/real-time PCR method targeting the prfA gene. Res. Microbiol. 2006, 157, 763–771. [Google Scholar] [CrossRef] [PubMed]
- Doumith, M.; Buchrieser, C.; Glaser, P.; Jacquet, C.; Martin, P. Differentiation of the major Listeria monocytogenes serovars by multiplex PCR. J. Clin. Microbiol. 2004, 42, 3819–3822. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Lawrence, M.L.; Austin, F.W.; Ainsworth, A.J. A multiplex PCR for species-and virulence-specific determination of Listeria monocytogenes. J. Microbiol. Meth. 2007, 71, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Kalorey, D.; Kurkure, N.; Warke, S.; Rawool, D.; Malik, S.; Barbuddhe, S. Isolation of pathogenic Listeria monocytogenes in faeces of wild animals in captivity. Comp. Immunol. Microbiol. 2006, 29, 295–300. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria; Approved Guideline (M45-A2); Wayne, P.A., Ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2010. [Google Scholar]
- Acar, J.F.; Bergogne-Berezin, E.; Chabbert, Y.; Cluzel, R.; Courtieu, A.; Courvalin, P.; Veron, M. Communiqué 1992 du Comité de L’Antibiogramme de la Societé Francaise de Microbiologie. Pathol. Biol. 1998, 40, 741–748. [Google Scholar]
- Claeys, W.L.; Cardoen, S.; Daube, G.; De Block, J.; Dewettinck, K.; Dierick, K.; De Zutter, L.; Huyghebaert, A.; Imberechts, H.; Thiange, P. Raw or heated cow milk consumption: Review of risks and benefits. Food Control 2013, 31, 251–262. [Google Scholar] [CrossRef]
- Verraes, C.; Vlaemynck, G.; Van Weyenberg, S.; De Zutter, L.; Daube, G.; Sindic, M.; Uyttendaele, M.; Herman, L. A review of the microbiological hazards of dairy products made from raw milk. Int. Dairy J. 2015, 50, 32–44. [Google Scholar] [CrossRef]
- Buchanan, R.L.; Gorris, L.G.; Hayman, M.M.; Jackson, T.C.; Whiting, R.C. A review of Listeria monocytogenes: An update on outbreaks, virulence, dose-response, ecology, and risk assessments. Food Control 2017, 75, 1–13. [Google Scholar] [CrossRef]
- Nayak, D.N.; Savalia, C.; Kalyani, I.; Kumar, R.; Kshirsagar, D. Isolation, identification, and characterization of Listeria spp. from various animal origin foods. Vet. World 2015, 8, 695. [Google Scholar] [CrossRef] [PubMed]
- Stephan, R.; Buhler, K. Prevalence of Campylobacter spp., Salmonella spp. and Listeria monocytogenes in bulk-tank milk samples from north-east Switzerland. Arch. Lebensmittelhyg. 2002, 53, 62–65. [Google Scholar]
- De Reu, K.; Grijspeerdt, K.; Herman, L. A Belgian survey of hygiene indicator bacteria and pathogenic bacteria in raw milk and direct marketing of raw milk farm products. J. Food Safety 2004, 24, 17–36. [Google Scholar] [CrossRef]
- Domínguez, R.L.; Fernández, G.J.; Vazquez, B.J.; Rodriguez, F.E.; Suarez, F.G. Isolation of micro-organisms of the species Listeria from raw milk intended for human consumption. Can. J. Microbiol. 1985, 31, 938–941. [Google Scholar]
- El Marnissi, B.; Bennani, L.; Cohen, N.; Lalami, A.E.O.; Belkhou, R. Presence of Listeria monocytogenes in raw milk and traditional dairy products marketed in the north-central region of Morocco. Afr. J. Food Sci. 2013, 7, 87–91. [Google Scholar] [CrossRef]
- Fenlon, D.; Stewart, T.; Donachie, W. The incidence, numbers and types of Listeria monocytogenes isolated from farm bulk tank milks. Lett. Appl. Microbiol. 1995, 20, 57–60. [Google Scholar] [CrossRef] [PubMed]
- Meyer-Broseta, S.; Diot, A.; Bastian, S.; Rivière, J.; Cerf, O. Estimation of low bacterial concentration: Listeria monocytogenes in raw milk. Int. J. Food Microbiol. 2003, 80, 1–15. [Google Scholar] [CrossRef]
- Waak, E.; Tham, W.; Danielsson-Tham, M.-L. Prevalence and fingerprinting of Listeria monocytogenes strains isolated from raw whole milk in farm bulk tanks and in dairy plant receiving tanks. Appl. Environ. Microbiol. 2002, 68, 3366–3370. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Strawn, L.K.; Fortes, E.D.; Bihn, E.A.; Nightingale, K.K.; Gröhn, Y.T.; Worobo, R.W.; Wiedmann, M.; Bergholz, P.W. Landscape and meteorological factors affecting prevalence of three food-borne pathogens in fruit and vegetable farms. Appl. Environ. Microbiol. 2013, 79, 588–600. [Google Scholar] [CrossRef] [PubMed]
- Carpentier, B.; Cerf, O. Persistence of Listeria monocytogenes in food industry equipment and premises. Int. J. Food Microbiol. 2011, 145, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, V.; Wiedmann, M.; Teixeira, P.; Stasiewicz, M. Listeria monocytogenes persistence in food-associated environments: Epidemiology, strain characteristics, and implications for public health. J. Food Prot. 2014, 77, 150–170. [Google Scholar] [CrossRef] [PubMed]
- WHO. Risk Assessment of Listeria monocytogenes in Ready-to-Eat Foods: Interpretative Summary, Food and Agriculture Organization: Rome, Italy, 2004.
- Althaus, D.; Lehner, A.; Brisse, S.; Maury, M.; Tasara, T.; Stephan, R. Characterization of Listeria monocytogenes strains isolated during 2011–2013 from human infections in Switzerland. Foodborne Pathog. Dis. 2014, 11, 753–758. [Google Scholar] [CrossRef] [PubMed]
- Pontello, M.; Guaita, A.; Sala, G.; Cipolla, M.; Gattuso, A.; Sonnessa, M.; Gianfranceschi, M.V. Listeria monocytogenes serotypes in human infections (Italy, 2000–2010). Ann. Dell’ist. Super. Sanitã 2012, 48, 146–150. [Google Scholar] [CrossRef]
- Yu, T.; Jiang, X. Prevalence and characterization of Listeria monocytogenes isolated from retail food in Henan, China. Food Control 2014, 37, 228–231. [Google Scholar] [CrossRef]
- Korsak, D.; Borek, A.; Daniluk, S.; Grabowska, A.; Pappelbaum, K. Antimicrobial susceptibilities of Listeria monocytogenes strains isolated from food and food processing environment in Poland. Int. J. Food Microbiol. 2012, 158, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Buchrieser, C.; Brosch, R.; Catimel, B.; Rocourt, J. Pulsed-field gel electrophoresis applied for comparing Listeria monocytogenes strains involved in outbreaks. Can. J. Microbiol. 1993, 39, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Camejo, A.; Carvalho, F.; Reis, O.; Leitão, E.; Sousa, S.; Cabanes, D. The arsenal of virulence factors deployed by Listeria monocytogenes to promote its cell infection cycle. Virulence 2011, 2, 379–394. [Google Scholar] [CrossRef] [PubMed]
- Schnupf, P.; Portnoy, D.A. Listeriolysin O: A phagosome-specific lysin. Microbes Infect. 2007, 9, 1176–1187. [Google Scholar] [CrossRef] [PubMed]
- Indrawattana, N.; Nibaddhasobon, T.; Sookrung, N.; Chongsa-nguan, M.; Tungtrongchitr, A.; Makino, S.-I.; Tungyong, W.; Chaicumpa, W. Prevalence of Listeria monocytogenes in raw meats marketed in Bangkok and characterization of the isolates by phenotypic and molecular methods. J. Health Popul. Nutr. 2011, 29, 26. [Google Scholar] [CrossRef] [PubMed]
- Jamali, H.; Paydar, M.; Ismail, S.; Looi, C.Y.; Wong, W.F.; Radmehr, B.; Abedini, A. Prevalence, antimicrobial susceptibility and virulotyping of Listeria species and Listeria monocytogenes isolated from open-air fish markets. BMC Microbiol. 2015, 15, 144. [Google Scholar] [CrossRef] [PubMed]
- Lomonaco, S.; Patti, R.; Knabel, S.J.; Civera, T. Detection of virulence-associated genes and epidemic clone markers in Listeria monocytogenes isolates from PDO Gorgonzola cheese. Int. J. Food Microbiol. 2012, 160, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Sant’Ana, A.S.; Igarashi, M.C.; Landgraf, M.; Destro, M.T.; Franco, B.D. Prevalence, populations and pheno-and genotypic characteristics of Listeria monocytogenes isolated from ready-to-eat vegetables marketed in São Paulo, Brazil. Int. J. Food Microbiol. 2012, 155, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dramsi, S.; Biswas, I.; Maguin, E.; Braun, L.; Mastroeni, P.; Cossart, P. Entry of Listeria monocytogenes into hepatocytes requires expression of inIB, a surface protein of the internalin multigene family. Mol. Microbiol. 1995, 16, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, J.-L.; Berche, P.; Frehel, C.; Gouln, E.; Cossart, P. Entry of L. monocytogenes into cells is mediated by internalin, a repeat protein reminiscent of surface antigens from gram-positive cocci. Cell 1991, 65, 1127–1141. [Google Scholar] [CrossRef]
- Dramsi, S.; Dehoux, P.; Lebrun, M.; Goossens, P.L.; Cossart, P. Identification of four new members of the internalin multigene family of Listeria monocytogenes EGD. Infect. Immun. 1997, 65, 1615–1625. [Google Scholar] [PubMed]
- Raffelsbauer, D.; Bubert, A.; Engelbrecht, F.; Scheinpflug, J.; Simm, A.; Hess, J.; Kaufmann, S.; Goebel, W. The gene cluster inlC2DE of Listeria monocytogenes contains additional new internalin genes and is important for virulence in mice. Mol. Gen. Genet. 1998, 260, 144–158. [Google Scholar] [CrossRef] [PubMed]
- Sabet, C.; Lecuit, M.; Cabanes, D.; Cossart, P.; Bierne, H. LPXTG protein InlJ, a newly identified internalin involved in Listeria monocytogenes virulence. Infect. Immun. 2005, 73, 6912–6922. [Google Scholar] [CrossRef] [PubMed]
- Al-Nabulsi, A.A.; Osaili, T.M.; Shaker, R.R.; Olaimat, A.N.; Jaradat, Z.W.; Elabedeen, N.A.Z.; Holley, R.A. Effects of osmotic pressure, acid, or cold stresses on antibiotic susceptibility of Listeria monocytogenes. Food Microbiol. 2015, 46, 154–160. [Google Scholar] [CrossRef] [PubMed]
- Conter, M.; Paludi, D.; Zanardi, E.; Ghidini, S.; Vergara, A.; Ianieri, A. Characterization of antimicrobial resistance of foodborne Listeria monocytogenes. Int. J. Food Microbiol. 2009, 128, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Lyon, S.A.; Berrang, M.E.; Fedorka-Cray, P.J.; Fletcher, D.L.; Meinersmann, R.J. Antimicrobial resistance of Listeria monocytogenes isolated from a poultry further processing plant. Foodborne Pathog. Dis. 2008, 5, 253–259. [Google Scholar] [CrossRef] [PubMed]
- Troxler, R.; Von Graevenitz, A.; Funke, G.; Wiedemann, B.; Stock, I. Natural antibiotic susceptibility of Listeria species: L. grayi, L. innocua, L. ivanovii, L. monocytogenes, L. seeligeri and L. welshimeri strains. Clin. Microbiol. Infect. 2000, 6, 525–535. [Google Scholar] [CrossRef] [PubMed]
| Product | pH of Samples | Number of Samples | Listeria spp. (%) | Listeria monocytogenes (%) |
|---|---|---|---|---|
| Raw cow milk | 6.7 ± 0.2 | 114 | 20 (17.5) | 10 (8.8) |
| Boiled milk | 6.3 ± 0.4 | 56 | ND | ND |
| Nunu a | 4.1 ± 0.9 | 84 | 11 (13.1) | 4 (4.7) |
| Total | NA | 254 | 44 (12.2) | 14 (5.5) |
| Antimicrobial | a MIC Breakpoints | b Interpretations n (%) | |||
|---|---|---|---|---|---|
| S (≤) | R (≥) | S | I | R | |
| Amoxicillin | 2 | 8 | 62 (100) | 0 | 0 |
| Ampicillin | 4 | 8 | 62 (100) | 0 | 0 |
| Chloramphenicol | 8 | 32 | 60 (96.8) | 2 (3.2) | 0 |
| Ciprofloxacin | 2 | 8 | 55 (88.7) | 7 (11.3) | 0 |
| Clindamycin | 2 | 8 | 51 (82.3) | 11 (17.7) | 0 |
| Doxycycline | 4 | 16 | 58 (93.5) | 4 (6.5) | 0 |
| Erythromycin | 2 | 4 | 62 (100) | N/A | 0 |
| Gentamycin | 2 | 8 | 62 (100) | 0 | 0 |
| Kanamycin | 16 | 64 | 57 (91.9) | 5 (8.1) | 0 |
| Neomycin | 8 | 32 | 21 (33.9) | 3 (4.8) | 38 (61.3) |
| Penicillin | 4 | 8 | 62 (100) | 0 | 0 |
| Rifampicin | 1 | 2 | 62 (100) | N/A | 0 |
| Streptomycin | 8 | 32 | 61 (98.4) | 1 (1.6) | 0 |
| Tetracycline | 2 | 8 | 44 (70.9) | 3 (4.8) | 15 (24.2) |
| Vancomycin | 8 | 32 | 62 (100) | 0 | 0 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Owusu-Kwarteng, J.; Wuni, A.; Akabanda, F.; Jespersen, L. Prevalence and Characteristics of Listeria monocytogenes Isolates in Raw Milk, Heated Milk and Nunu, a Spontaneously Fermented Milk Beverage, in Ghana. Beverages 2018, 4, 40. https://doi.org/10.3390/beverages4020040
Owusu-Kwarteng J, Wuni A, Akabanda F, Jespersen L. Prevalence and Characteristics of Listeria monocytogenes Isolates in Raw Milk, Heated Milk and Nunu, a Spontaneously Fermented Milk Beverage, in Ghana. Beverages. 2018; 4(2):40. https://doi.org/10.3390/beverages4020040
Chicago/Turabian StyleOwusu-Kwarteng, James, Alhassan Wuni, Fortune Akabanda, and Lene Jespersen. 2018. "Prevalence and Characteristics of Listeria monocytogenes Isolates in Raw Milk, Heated Milk and Nunu, a Spontaneously Fermented Milk Beverage, in Ghana" Beverages 4, no. 2: 40. https://doi.org/10.3390/beverages4020040
APA StyleOwusu-Kwarteng, J., Wuni, A., Akabanda, F., & Jespersen, L. (2018). Prevalence and Characteristics of Listeria monocytogenes Isolates in Raw Milk, Heated Milk and Nunu, a Spontaneously Fermented Milk Beverage, in Ghana. Beverages, 4(2), 40. https://doi.org/10.3390/beverages4020040