Improved AI-Assisted Image Recognition of Cervical Spine Vertebrae Enables Motion Pattern Analysis in Dynamic X-Ray Recordings
Abstract
1. Introduction
2. Materials and Methods
2.1. Population and Manual Annotation
2.2. Dataset
2.3. Development of the Model
2.4. Histogram Equalization
2.5. Pre-Training
2.6. Model Evaluation Metrics
2.7. Mean Shape
2.8. Outcome
2.9. Analyses
2.10. Correlation Analysis
3. Results
3.1. Segmentation Performance
3.2. Intraclass Correlation Coefficient
3.3. Sensitivity Analyses
3.4. Motion Pattern Analysis
3.5. ICC vs. Segmentation Performance Correlation
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACD | Anterior Cervical Discectomy |
| ACDA | Anterior Cervical Discectomy with Arthroplasty |
| AI | Artificial intelligence |
| DSC | Dice similarity coefficient |
| HD95 | Hausdorff Distance |
| ICC | Intraclass Correlation Coefficient |
| IoU | Intersection over Union |
| sROM | Segmental range of motion |
| SSC | Sequence of segmental contribution |
Appendix A


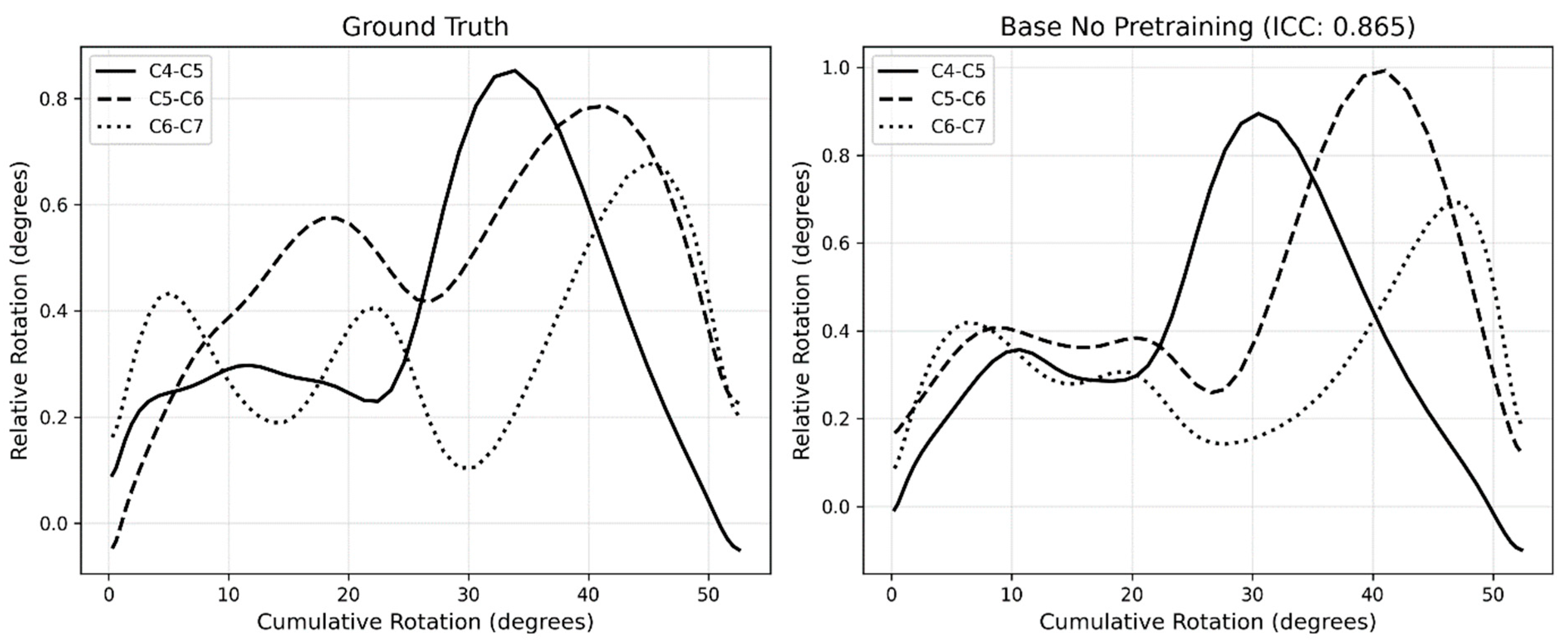



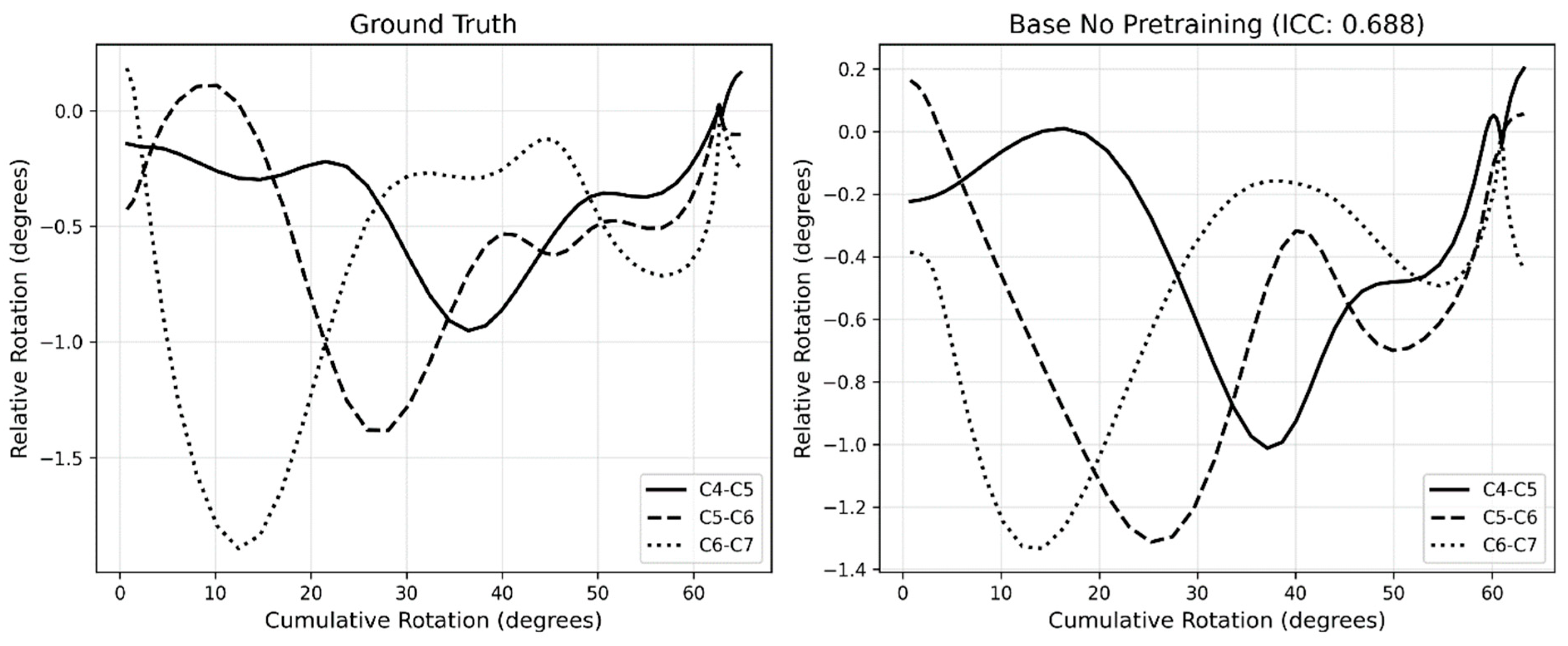





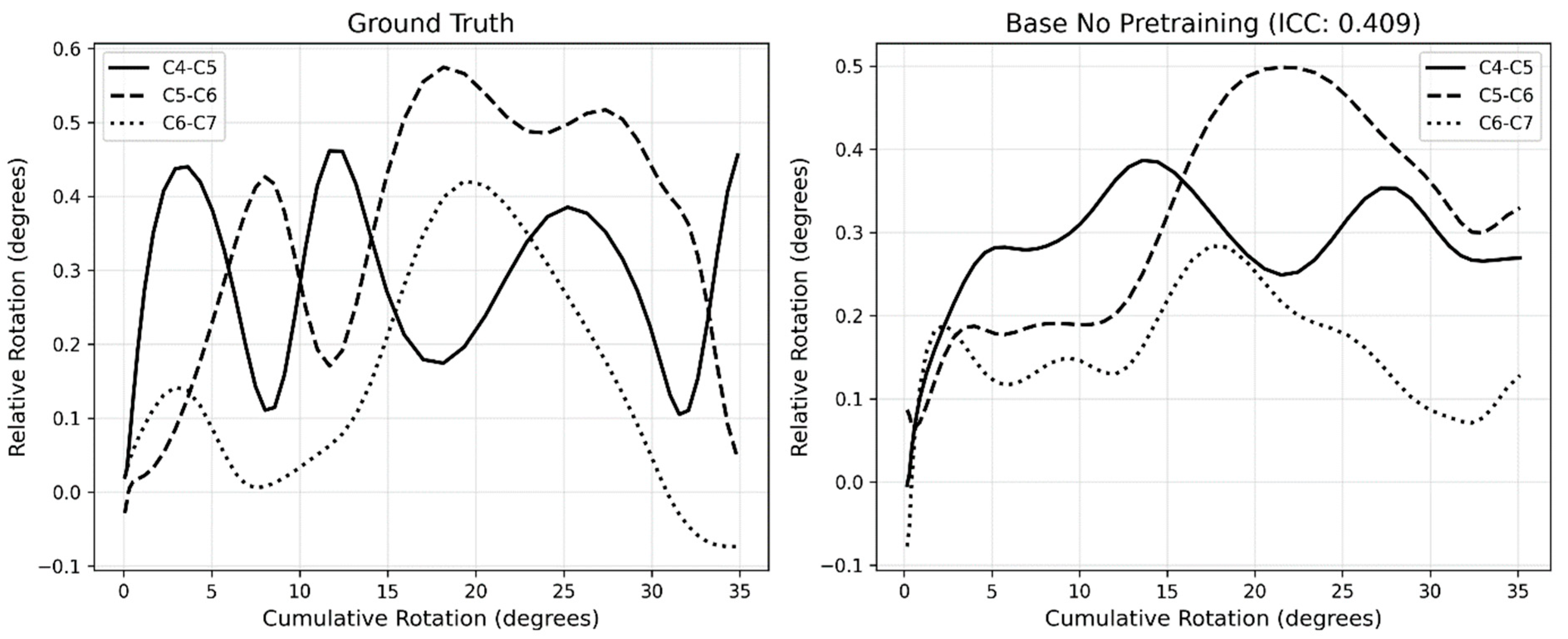









| DSC (SD) | IoU (SD) | HD95 in mm (SD) | |
|---|---|---|---|
| C0 | 0.69 ± 0.25 | 0.57 ± 0.25 | 19.55 ± 16.06 |
| C1 | 0.86 ± 0.08 | 0.76 ± 0.11 | 4.19 ± 2.48 |
| C2 | 0.88 ± 0.06 | 0.79 ± 0.09 | 4.47 ± 2.04 |
| C3 | 0.90 ± 0.05 | 0.82 ± 0.07 | 3.19 ± 2.42 |
| C4 | 0.91 ± 0.05 | 0.84 ± 0.06 | 2.49 ± 1.71 |
| C5 | 0.88 ± 0.07 | 0.78 ± 0.10 | 4.12 ± 3.67 |
| C6 | 0.82 ± 0.15 | 0.72 ± 0.18 | 6.82 ± 5.92 |
| C7 | 0.75 ± 0.19 | 0.63 ± 0.21 | 10.06 ± 8.71 |
| Exclusion | None | n | Low-Quality | n | Low sROM | n | Low-Quality/sROM | n |
|---|---|---|---|---|---|---|---|---|
| Segment | ||||||||
| C1–C2 | 0.589 (−0.074–0.949) | 23 | 0.625 (−0.074–0.949) | 19 | 0.681 (0.165–0.949) | 19 | 0.750 (0.491–0.949) | 15 |
| C2–C3 | 0.423 (−0.444–0.941) | 23 | 0.491 (−0.444–0.941) | 19 | 0.469 (−0.444–0.941) | 20 | 0.539 (−0.444–0.941) | 17 |
| C3–C4 | 0.679 (0.031–0.948) | 23 | 0.728 (0.158–0.948) | 19 | 0.679 (0.031–0.948) | 23 | 0.728 (0.158–0.948) | 19 |
| C4–C5 | 0.681 (0.123–0.963) | 23 | 0.679 (0.123–0.963) | 17 | 0.681 (0.123–0.963) | 23 | 0.679 (0.123–0.963) | 17 |
| C5–C6 | 0.676 (−0.003–0.982) | 23 | 0.695 (−0.003–0.982) | 16 | 0.672 (−0.003–0.982) | 22 | 0.689 (−0.003–0.982) | 15 |
| C6–C7 | 0.385 (−0.431–0.989) | 23 | 0.522 (−0.095–0.989) | 9 | 0.391 (−0.431–0.989) | 21 | 0.556 (−0.095–0.989) | 8 |
References
- Boselie, T.F.M.; van Santbrink, H.; de Bie, R.A.; van Mameren, H. Pilot Study of Sequence of Segmental Contributions in the Lower Cervical Spine During Active Extension and Flexion: Healthy Controls Versus Cervical Degenerative Disc Disease Patients. Spine 2017, 42, E642–E647. [Google Scholar] [CrossRef] [PubMed]
- Van Mameren, H.; Drukker, J.; Sanches, H.; Beursgens, J. Cervical spine motion in the sagittal plane (I) range of motion of actually performed movements, an X-ray cinematographic study. Eur. J. Morphol. 1990, 28, 47–68. [Google Scholar] [PubMed]
- Boselie, T.F.; van Mameren, H.; de Bie, R.A.; van Santbrink, H. Cervical spine kinematics after anterior cervical discectomy with or without implantation of a mobile cervical disc prosthesis; an RCT. BMC Musculoskelet. Disord. 2015, 16, 34. [Google Scholar] [CrossRef] [PubMed]
- Schuermans, V.N.E.; Smeets, A.; Curfs, I.; van Santbrink, H.; Boselie, T.F.M. A randomized controlled trial with extended long-term follow-up: Quality of cervical spine motion after anterior cervical discectomy (ACD) or anterior cervical discectomy with arthroplasty (ACDA). Brain Spine 2024, 4, 102726. [Google Scholar] [CrossRef] [PubMed]
- Schuermans, V.N.E.; Smeets, A.; Breen, A.; Branney, J.; Curfs, I.; van Santbrink, H.; Boselie, T.F.M. An observational study of quality of motion in the aging cervical spine: Sequence of segmental contributions in dynamic fluoroscopy recordings. BMC Musculoskelet. Disord. 2024, 25, 330. [Google Scholar] [CrossRef] [PubMed]
- van Santbrink, E.; Schuermans, V.; Cerfonteijn, E.; Breeuwer, M.; Smeets, A.; van Santbrink, H.; Boselie, T. AI-Assisted Image Recognition of Cervical Spine Vertebrae in Dynamic X-Ray Recordings. Bioengineering 2025, 12, 679. [Google Scholar] [CrossRef] [PubMed]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 2021, 18, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Zhang, Z.; Ding, N.; Gu, Y.; Liu, X.; Huo, Y.; Qiu, J.; Yao, Y.; Zhang, A.; Zhang, L.; et al. Pre-trained models: Past, present and future. AI Open 2021, 2, 225–250. [Google Scholar] [CrossRef]
- Saifullah, S.; Dreżewski, R. Modified Histogram Equalization for Improved CNN Medical Image Segmentation. Procedia Comput. Sci. 2023, 225, 3021–3030. [Google Scholar] [CrossRef]
- Branney, J. An Observational Study of Changes in Cervical Inter-Vertebral Motion and the Relationship with Patient-Reported Outcomes in Patients Undergoing Spinal Manipulative Therapy for Neck Pain. Ph.D. Thesis, Bournemouth University, Poole, UK, 2014. [Google Scholar]
- Branney, J.; Breen, A.C. Does inter-vertebral range of motion increase after spinal manipulation? A prospective cohort study. Chiropr. Man. Ther. 2014, 22, 24. [Google Scholar] [CrossRef] [PubMed]
- Reinartz, R.; Platel, B.; Boselie, T.; van Mameren, H.; van Santbrink, H.; Romeny, B. Cervical vertebrae tracking in video-fluoroscopy using the normalized gradient field. In Medical Image Computing and Computer-Assisted Intervention; Springer: Berlin/Heidelberg, Germany, 2009; pp. 524–531. [Google Scholar] [CrossRef]
- Isensee, F.; Wald, T.; Ulrich, C.; Baumgartner, M.; Roy, S.; Maier-Hein, K.; Jaeger, P.F. nnU-Net Revisited: A Call for Rigorous Validation in 3D Medical Image Segmentation. arXiv 2024, arXiv:2404.09556. [Google Scholar] [CrossRef]
- Ran, Y.; Qin, W.; Qin, C.; Li, X.; Liu, Y.; Xu, L.; Mu, X.; Yan, L.; Wang, B.; Dai, Y.; et al. A high-quality dataset featuring classified and annotated cervical spine X-ray atlas. Sci. Data 2024, 11, 625. [Google Scholar] [CrossRef] [PubMed]
- Boehringer, A.S.; Sanaat, A.; Arabi, H.; Zaidi, H. An active learning approach to train a deep learning algorithm for tumor segmentation from brain MR images. Insights Imaging 2023, 14, 141. [Google Scholar] [CrossRef] [PubMed]
- Gon Park, S.; Park, J.; Rock Choi, H.; Ho Lee, J.; Tae Cho, S.; Goo Lee, Y.; Ahn, H.; Pak, S. Deep Learning Model for Real-time Semantic Segmentation During Intraoperative Robotic Prostatectomy. Eur. Urol. Open Sci. 2024, 62, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Ramieri, A.; Costanzo, G.; Miscusi, M. Functional Anatomy and Biomechanics of the Cervical Spine. In Cervical Spine: Minimally Invasive and Open Surgery; Menchetti, P.P.M., Ed.; Springer International Publishing: Cham, Switzerland, 2022; pp. 11–31. [Google Scholar] [CrossRef]
- Koo, T.K.; Li, M.Y. A Guideline of Selecting and Reporting Intraclass Correlation Coefficients for Reliability Research. J. Chiropr. Med. 2016, 15, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Napravnik, M.; Hržić, F.; Urschler, M.; Miletić, D.; Štajduhar, I. Lessons learned from RadiologyNET foundation models for transfer learning in medical radiology. Sci. Rep. 2025, 15, 21622. [Google Scholar] [CrossRef] [PubMed]
- Barrett, J.F.; Keat, N. Artifacts in CT: Recognition and Avoidance. RadioGraphics 2004, 24, 1679–1691. [Google Scholar] [CrossRef] [PubMed]
- Ravi, N.; Gabeur, V.; Hu, Y.-T.; Hu, R.; Ryali, C.; Ma, T.; Khedr, H.; Rädle, R.; Rolland, C.; Gustafson, L.; et al. SAM 2: Segment Anything in Images and Videos. arXiv 2024, arXiv:2408.00714. [Google Scholar] [CrossRef] [PubMed]
- Machino, M.; Yukawa, Y.; Hida, T.; Ito, K.; Nakashima, H.; Kanbara, S.; Morita, D.; Kato, F. Cervical alignment and range of motion after laminoplasty: Radiographical data from more than 500 cases with cervical spondylotic myelopathy and a review of the literature. Spine 2012, 37, E1243–E1250. [Google Scholar] [CrossRef] [PubMed]
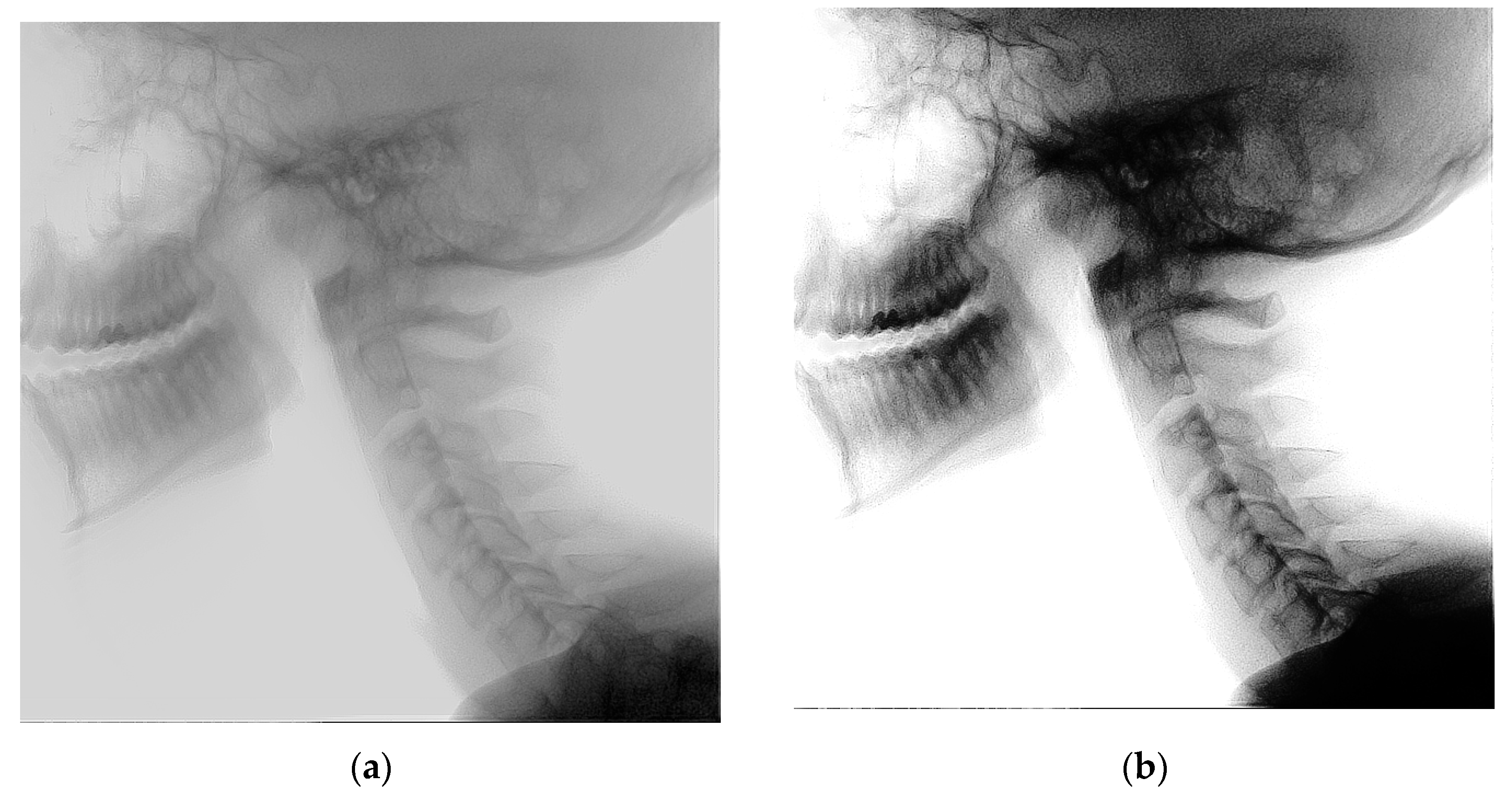




| Data Subset | Individuals (N) | Recordings (N) |
|---|---|---|
| Training (60%) | 36 | 67 |
| Validation (20%) | 12 | 22 |
| Testing (20%) | 14 | 23 |
| DSC (SD) | IoU (SD) | HD95 in mm (SD) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Model | A | B | C | D | A | B | C | D | A | B | C | D |
| C0 | 0.69 ± 0.25 | 0.71 ± 0.22 | 0.69 ± 0.24 | 0.68 ± 0.23 | 0.57 ± 0.25 | 0.58 ± 0.23 | 0.57 ± 0.25 | 0.55 ± 0.23 | 19.55 ± 16.06 | 19.67 ± 18.20 | 19.07 ± 17.94 | 19.25 ± 18.70 |
| C1 | 0.86 ± 0.08 | 0.86 ± 0.08 | 0.85 ± 0.10 | 0.87 ± 0.04 | 0.76 ± 0.11 | 0.76 ± 0.10 | 0.74 ± 0.11 | 0.78 ± 0.06 | 4.19 ± 2.48 | 4.07 ± 1.55 | 4.74 ± 5.56 | 3.93 ± 1.50 |
| C2 | 0.88 ± 0.06 | 0.89 ± 0.04 | 0.87 ± 0.08 | 0.89 ± 0.04 | 0.79 ± 0.09 | 0.80 ± 0.07 | 0.78 ± 0.10 | 0.80 ± 0.06 | 4.47 ± 2.04 | 4.15 ± 1.73 | 6.01 ± 10.52 | 4.31 ± 1.81 |
| C3 | 0.90 ± 0.05 | 0.91 ± 0.03 | 0.90 ± 0.06 | 0.90 ± 0.03 | 0.82 ± 0.07 | 0.83 ± 0.05 | 0.82 ± 0.08 | 0.82 ± 0.05 | 3.19 ± 2.42 | 2.76 ± 1.18 | 3.80 ± 8.35 | 2.99 ± 1.68 |
| C4 | 0.91 ± 0.05 | 0.92 ± 0.03 | 0.90 ± 0.06 | 0.91 ± 0.03 | 0.84 ± 0.06 | 0.85 ± 0.04 | 0.83 ± 0.08 | 0.84 ± 0.05 | 2.49 ± 1.71 | 2.35 ± 1.39 | 3.78 ± 5.85 | 2.58 ± 1.61 |
| C5 | 0.88 ± 0.07 | 0.90 ± 0.04 | 0.87 ± 0.08 | 0.89 ± 0.04 | 0.78 ± 0.10 | 0.82 ± 0.06 | 0.77 ± 0.11 | 0.81 ± 0.06 | 4.12 ± 3.67 | 3.19 ± 1.83 | 5.30 ± 4.78 | 3.29 ± 1.67 |
| C6 | 0.82 ± 0.15 | 0.90 ± 0.04 | 0.80 ± 0.17 | 0.89 ± 0.05 | 0.72 ± 0.18 | 0.82 ± 0.07 | 0.69 ± 0.19 | 0.80 ± 0.07 | 6.82 ± 5.92 | 3.68 ± 3.19 | 7.42 ± 6.35 | 4.42 ± 3.35 |
| C7 | 0.75 ± 0.19 | 0.84 ± 0.10 | 0.73 ± 0.21 | 0.81 ± 0.13 | 0.63 ± 0.21 | 0.73 ± 0.14 | 0.62 ± 0.22 | 0.70 ± 0.16 | 10.06 ± 8.71 | 5.55 ± 4.35 | 11.39 ± 15.79 | 7.81 ± 12.39 |
| Model | A | n | B | n | C | n | D | n |
|---|---|---|---|---|---|---|---|---|
| Segment | ||||||||
| C1–C2 | 0.595 (−0.369–0.981) | 23 | 0.569 (−0.423–0.966) | 22 | 0.476 (−0.271–0.966) | 22 | 0.577 (−0.222–0.967) | 23 |
| C2–C3 | 0.449 (−0.262–0.913) | 23 | 0.546 (−0.275–0.956) | 22 | 0.384 (−0.421–0.858) | 22 | 0.438 (−0.386–0.920) | 23 |
| C3–C4 | 0.697 (−0.212–0.971) | 23 | 0.734 (−0.041–0.967) | 22 | 0.612 (−0.409–0.928) | 22 | 0.724 (−0.055–0.967) | 23 |
| C4–C5 | 0.728 (0.086–0.961) | 23 | 0.715 (0.080–0.965) | 22 | 0.589 (−0.173–0.955) | 22 | 0.742 (0.301–0.952) | 23 |
| C5–C6 | 0.693 (−0.042–0.968) | 23 | 0.733 (−0.198–0.963) | 22 | 0.342 (−0.410–0.971) | 22 | 0.689 (−0.087–0.942) | 23 |
| C6–C7 | 0.449 (−0.480–0.991) | 23 | 0.368 (−0.161–0.990) | 22 | 0.287 (−0.151–0.990) | 22 | 0.382 (−0.435–0.990) | 23 |
| Model | A | n | B | n | C | n | D | n |
|---|---|---|---|---|---|---|---|---|
| Segment | ||||||||
| C1–C2 | 0.711 (−0.228–0.981) | 19 | 0.706 (−0.423–0.966) | 18 | 0.557 (−0.027–0.950) | 18 | 0.720 (−0.208–0.967) | 19 |
| C2–C3 | 0.532 (−0.262–0.913) | 20 | 0.583 (−0.275–0.956) | 19 | 0.455 (−0.421–0.858) | 19 | 0.488 (−0.386–0.920) | 20 |
| C3–C4 | 0.697 (−0.212–0.971) | 23 | 0.734 (−0.041–0.967) | 22 | 0.612 (−0.409–0.928) | 22 | 0.724 (−0.055–0.967) | 23 |
| C4–C5 | 0.728 (0.086–0.961) | 23 | 0.715 (0.080–0.965) | 22 | 0.589 (−0.173–0.955) | 22 | 0.742 (0.301–0.952) | 23 |
| C5–C6 | 0.690 (−0.042–0.968) | 22 | 0.763 (−0.198–0.963) | 21 | 0.330 (−0.410–0.971) | 21 | 0.680 (−0.087–0.942) | 22 |
| C6–C7 | 0.516 (−0.387–0.991) | 21 | 0.399 (−0.161–0.990) | 20 | 0.321 (−0.151–0.968) | 20 | 0.442 (−0.367–0.990) | 21 |
| Model | A | n | B | n | C | n | D | n |
|---|---|---|---|---|---|---|---|---|
| Segment | ||||||||
| C1–C2 | 0.670 (−0.369–0.981) | 19 | 0.605 (−0.375–0.966) | 19 | 0.504 (−0.271–0.950) | 18 | 0.638 (−0.222–0.967) | 19 |
| C2–C3 | 0.549 (−0.148–0.913) | 19 | 0.597 (0.162–0.956) | 19 | 0.498 (−0.082–0.858) | 18 | 0.543 (−0.109–0.920) | 19 |
| C3–C4 | 0.770 (0.302–0.971) | 19 | 0.761 (0.251–0.967) | 19 | 0.689 (0.077–0.928) | 18 | 0.764 (0.304–0.967) | 19 |
| C4–C5 | 0.750 (0.180–0.961) | 17 | 0.727 (0.080–0.965) | 17 | 0.652 (0.091–0.955) | 16 | 0.746 (0.301–0.952) | 17 |
| C5–C6 | 0.715 (−0.042–0.968) | 16 | 0.738 (−0.198–0.963) | 16 | 0.421 (−0.318–0.971) | 16 | 0.694 (−0.087–0.937) | 16 |
| C6–C7 | 0.594 (−0.043–0.991) | 9 | 0.532 (−0.083–0.990) | 9 | 0.427 (−0.112–0.968) | 9 | 0.421 (−0.367–0.990) | 9 |
| Model | A | n | B | n | C | n | D | n |
|---|---|---|---|---|---|---|---|---|
| Segment | ||||||||
| C1–C2 | 0.837 (0.604–0.981) | 15 | 0.780 (0.512–0.966) | 15 | 0.617 (−0.027–0.950) | 14 | 0.835 (0.687–0.967) | 15 |
| C2–C3 | 0.617 (−0.148–0.913) | 17 | 0.636 (0.208–0.956) | 17 | 0.561 (0.050–0.858) | 16 | 0.594 (−0.109–0.920) | 17 |
| C3–C4 | 0.770 (0.302–0.971) | 19 | 0.761 (0.251–0.967) | 19 | 0.689 (0.077–0.928) | 18 | 0.764 (0.304–0.967) | 19 |
| C4–C5 | 0.750 (0.180–0.961) | 17 | 0.727 (0.080–0.965) | 17 | 0.652 (0.091–0.955) | 16 | 0.746 (0.301–0.952) | 17 |
| C5–C6 | 0.711 (−0.042–0.968) | 15 | 0.743 (−0.198–0.963) | 15 | 0.409 (−0.318–0.971) | 15 | 0.680 (−0.087–0.937) | 15 |
| C6–C7 | 0.673 (0.028–0.991) | 8 | 0.609 (−0.066–0.990) | 8 | 0.493 (−0.112–0.968) | 8 | 0.480 (−0.367–0.990) | 8 |
| Model | A | B | C | D | ||||
|---|---|---|---|---|---|---|---|---|
| Metric | n | n | n | n | ||||
| Dice mean | 0.047 | 23 | 0.010 | 22 | 0.654 | 22 | 0.266 | 23 |
| Dice median | 0.031 | 23 | −0.081 | 22 | 0.529 | 22 | 0.028 | 23 |
| Dice std | −0.027 | 23 | −0.041 | 22 | −0.683 | 22 | −0.282 | 23 |
| IoU mean | 0.054 | 23 | −0.008 | 22 | 0.649 | 22 | 0.253 | 23 |
| IoU median | 0.026 | 23 | −0.096 | 22 | 0.528 | 22 | 0.025 | 23 |
| IoU std | −0.001 | 23 | −0.004 | 22 | −0.674 | 22 | −0.290 | 23 |
| HD95 mean | −0.132 | 23 | −0.209 | 22 | −0.710 | 22 | −0.367 | 23 |
| HD95 median | −0.105 | 23 | 0.011 | 22 | −0.600 | 22 | −0.091 | 23 |
| HD95 std | −0.128 | 23 | −0.308 | 22 | −0.510 | 22 | −0.215 | 23 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
van Santbrink, E.; Hijzelaar, T.H.W.; Schuermans, V.N.E.; Smeets, A.Y.J.M.; van Santbrink, H.; de Bie, R.; Veta, M.; Boselie, T.F.M. Improved AI-Assisted Image Recognition of Cervical Spine Vertebrae Enables Motion Pattern Analysis in Dynamic X-Ray Recordings. Bioengineering 2026, 13, 351. https://doi.org/10.3390/bioengineering13030351
van Santbrink E, Hijzelaar THW, Schuermans VNE, Smeets AYJM, van Santbrink H, de Bie R, Veta M, Boselie TFM. Improved AI-Assisted Image Recognition of Cervical Spine Vertebrae Enables Motion Pattern Analysis in Dynamic X-Ray Recordings. Bioengineering. 2026; 13(3):351. https://doi.org/10.3390/bioengineering13030351
Chicago/Turabian Stylevan Santbrink, Esther, Tijmen H. W. Hijzelaar, Valérie N. E. Schuermans, Anouk Y. J. M. Smeets, Henk van Santbrink, Rob de Bie, Mitko Veta, and Toon F. M. Boselie. 2026. "Improved AI-Assisted Image Recognition of Cervical Spine Vertebrae Enables Motion Pattern Analysis in Dynamic X-Ray Recordings" Bioengineering 13, no. 3: 351. https://doi.org/10.3390/bioengineering13030351
APA Stylevan Santbrink, E., Hijzelaar, T. H. W., Schuermans, V. N. E., Smeets, A. Y. J. M., van Santbrink, H., de Bie, R., Veta, M., & Boselie, T. F. M. (2026). Improved AI-Assisted Image Recognition of Cervical Spine Vertebrae Enables Motion Pattern Analysis in Dynamic X-Ray Recordings. Bioengineering, 13(3), 351. https://doi.org/10.3390/bioengineering13030351






