1. Introduction
Brain–computer interfaces (BCIs) enable direct communication between brain signals and external devices, offering new possibilities for individuals with motor disabilities. MI, defined as the mental rehearsal of movement without overt motor output, has gained attention for its intuitive control and non-invasive nature [
1,
2]. Electroencephalography (EEG) is the most commonly used modality in MI-based BCIs due to its temporal resolution, affordability, and portability. However, EEG signal analysis remains challenging because of poor signal-to-noise ratio, non-stationary characteristics, and substantial inter-subject variability, all of which limit the reliability of MI-based systems [
3].
Deep learning models have advanced EEG decoding by learning features directly from raw data [
4,
5,
6]. Yet several limitations persist. Many models are designed for subject-specific training and generalize poorly to unseen users [
7,
8]. Most architectures emphasize either spatial or temporal aspects of EEG signals, missing their joint dependencies. Furthermore, few models consider computational efficiency, which is essential for real-time or embedded BCI deployment [
9].
Recent hybrid architectures combine convolutional networks with transformers to address these challenges. CLTNet [
10] fuses local convolutions with global attention, while EEGFormer [
5] uses self-attention to model inter-channel dependencies. However, these models share critical limitations. First, they lack explicit adaptation mechanisms for new users, requiring extensive retraining. Second, their static architectures cannot balance accuracy against the latency constraints of real-time systems. They advance representational capacity but fall short in adaptability and efficiency.
To overcome these limitations, we introduce EEG-TriNet++, a deep architecture that jointly learns spatial, temporal, and global contextual representations from EEG signals. Our hierarchical three-stage design reflects the multi-scale nature of MI-EEG: fine-grained spectral patterns emerge at the millisecond level, evolving ERD and ERS dynamics unfold over seconds, and distributed cortical activity spans across channels. A pure Transformer risks conflating these scales by applying global attention directly to raw signals. While a sequential CNN-Transformer hybrid captures local features, it still lacks dedicated temporal modeling. Our design addresses this by processing each scale through a specialized module, progressively refining the outputs for subsequent stages. A patch-based tokenization strategy with neural architecture search (NAS) automatically tunes parameters to balance accuracy with computational cost [
11,
12]. A MAML-based meta-learning module enables rapid adaptation to new subjects using minimal labeled data [
13,
14].
The main contributions are fourfold. First, we propose a hierarchical three-branch architecture that progressively refines representations from local spectral patterns to global contextual relationships. Second, we introduce patch-based tokenization adapted from vision transformers to capture localized spatio-temporal EEG patterns. Third, we integrate a lightweight NAS mechanism that dynamically optimizes architectural parameters for accuracy and efficiency, establishing the first NAS-driven framework for multi-branch MI decoding. Fourth, we incorporate a meta-learning module that enables rapid subject-specific calibration with limited data, thereby enhancing cross-subject transferability. Unlike prior hybrids like CLTNet [
10] and EEGFormer [
5], EEG-TriNet++ simultaneously addresses representation richness, cross-subject adaptability, and hardware-aware efficiency.
We validate EEG-TriNet++ on two public datasets with complementary challenges in acquisition protocols, participant diversity, and class configurations [
15,
16]. The model is evaluated under within-subject and subject-independent protocols. The results demonstrate superior performance in both settings, with high accuracy, stable convergence, and balanced classification across all MI classes, including traditionally difficult distinctions like foot and tongue imagery.
By integrating convolutional, recurrent, and attention-based modules within a unified framework enhanced by NAS and meta-learning, EEG-TriNet++ advances robust and adaptable MI decoding. The model is well-suited for real-world deployment in assistive neurotechnology and opens new directions for generalizable EEG-based BCI solutions.
2. Related Work
Research on MI classification from EEG signals has progressed considerably, transitioning from traditional signal-processing pipelines to advanced deep learning architectures designed for subject-independent generalization and real-time performance. Early approaches, such as Common Spatial Patterns (CSP) combined with Linear Discriminant Analysis (LDA) or Support Vector Machines (SVM) [
17,
18], provided interpretable and computationally efficient solutions. However, they often struggled with the inherent non-stationarity of EEG signals and the high inter-subject variability.
Before the widespread adoption of deep learning, several studies attempted to improve feature robustness and discriminability through more sophisticated preprocessing and representation techniques. Pei et al. [
19] proposed a tensor-based frequency feature combination method that jointly exploits spatial, spectral, and temporal information to enhance discriminative EEG representations for BCI tasks. Similarly, Pei et al. [
20] introduced a channel-level recombination strategy for data augmentation, significantly improving the generalization performance of MI classifiers by generating physiologically consistent synthetic EEG data. These approaches demonstrated the potential of structured signal modeling and data augmentation for boosting classification reliability in EEG-based BCIs.
With the emergence of deep learning, Schirrmeister et al. [
21] introduced DeepConvNet, demonstrating the capacity of CNNs to learn spatial-spectral representations directly from raw EEG data. Lawhern et al. [
4] proposed EEGNet, a lightweight convolutional model optimized for small data regimes and real-time applications. Hybrid approaches such as ConvLSTM [
22,
23] integrated convolutional and recurrent layers to capture both spatial and temporal features. Nguyen et al. [
24] extended these ideas using dilated temporal convolutions for multi-scale temporal feature extraction.
Transformer-based models have recently gained traction in EEG-based BCI. Wan et al. [
5] proposed EEGFormer, which leverages self-attention to model global inter-channel dependencies. Liao et al. [
25] introduced EEGEncoder, combining dual-stream TCNs with Transformer modules. Jiang et al. [
26] developed TS-Former by integrating transfer learning techniques into transformer-based MI decoding. Other models, such as CLTNet [
10], fused convolutional and transformer modules hierarchically for enhanced decoding accuracy.
Recent innovations have focused on generalization, contrastive learning, and domain adaptation. Liu et al. [
27] introduced MIRepNet, a foundational EEG model incorporating masked signal reconstruction with supervised classification, guided by neurophysiological priors. Wang et al. [
28] proposed MVCNet, using multi-view contrastive learning across temporal, frequency, and spatial domains. Chen et al. [
29] developed a Large Cognition Model for EEG, incorporating learnable channel mappings to enable configuration-agnostic generalization. Muna et al. [
30] presented SSTAF, applying joint spatial, spectral, and temporal attention mechanisms. Hamidi and Kiani [
31] combined graph convolutional networks with transformers to better capture spatial topology and sequential dynamics.
In parallel, mutual learning approaches have recently emerged as effective strategies for enhancing transfer learning and adaptation in neural systems. These methods train multiple models collaboratively through knowledge distillation, allowing them to exchange complementary information and jointly improve generalization. For example, Kumar et al. [
32] demonstrated how mutual learning can accelerate individual BCI skill acquisition, while Ye et al. [
33] and Wu et al. [
34] showed its benefits for EEG emotion recognition and cross-domain representation alignment. Yan et al. [
35] further introduced a spectral-temporal attention model coupled with contrastive mutual learning to enhance MI-EEG classification. Moreover, Pei et al. [
36,
37] analyzed dependence within EEG time series to improve affective BCI systems, providing information on feature correlations that could be used in future mutual-learning-based MI decoders. Although these methods significantly improve robustness, they often require simultaneous training of multiple networks and thus a higher computational cost.
In recent work, a variety of architectural strategies have also been explored. Gui et al. [
38] introduced EEGMamba, combining state-space modeling and a mixture-of-experts design for multitask EEG classification. Deng et al. [
39] proposed a hybrid CNN–Swin Transformer using 3D EEG representations. Mirzaei et al. [
40] designed an attention-guided CNN–LSTM autoencoder for simultaneous classification and reconstruction of MI signals. Zhao et al. [
9] and Song et al. [
41] enhanced CNN–Transformer hybrids with more efficient fusion techniques. EEG-GPT [
42] explored large language models for few-shot EEG understanding, and CLTNet [
10] emphasized hierarchical attention modeling with convolutional priors. Ingolfsson et al. [
8] validated the lightweight EEG-TCNet model for embedded MI tasks, showing its applicability to constrained real-time scenarios. Additionally, CLTNet was presented by Gu et al. [
10] as a hierarchical hybrid model integrating local convolutions and global attention, demonstrating competitive performance on multiple MI datasets.
Existing models often require subject-specific calibration and high computational resources. EEG-TriNet++ overcomes these issues with a lightweight, modular design that combines neural architecture search and meta-learning for efficient and adaptive BCI deployment.
3. Method
3.1. EEG-TriNet++ Architecture
EEG-TriNet++ is a modular, hierarchical deep architecture for MI EEG classification, composed of five key components that address distinct signal processing challenges. Its three-stage design is motivated by the multi-scale nature of motor imagery EEG: fine-grained spectral patterns emerge at the millisecond level, evolving ERD and ERS dynamics unfold over seconds, and distributed cortical activity spans across channels. Unlike pure Transformers, which conflate these scales via global attention, or CNN-Transformer hybrids lacking dedicated temporal modeling, our hierarchical approach processes each scale with a specialized module, progressively refining representations for subsequent stages. The model integrates mechanisms for spectral-spatial feature extraction, long-term temporal dependency modeling, temporal tokenization for attention-based processing, global context capture, and subject adaptation. An overview is provided in
Figure 1.
Let input EEG trial be (channels × time samples). Temporal and spatial convolutions first extract localized frequency and channel patterns. These features feed into a BiLSTM module that models temporal dependencies, producing contextual embeddings .
To capture long-range interactions, the BiLSTM output is segmented into overlapping patches
with stride
. Patches are flattened and projected into token embeddings:
where
denotes vectorization. After adding positional encodings, tokens pass through a lightweight Transformer encoder with a few attention heads for latency efficiency, capturing global dependencies across temporal and spatial contexts.
A NAS strategy jointly optimizes key hyperparameters (BiLSTM hidden size
, Transformer depth
L, patch size
p, stride
s) via a composite loss:
where
is cross-entropy,
quantifies inference delay, and
regularizes model size for real-time deployment.
For cross-subject generalization, a MAML-based meta-learning module learns an initialization
for rapid adaptation to new subjects with few samples. For each subject task
:
with inner and outer learning rates
and
. The final classification layer applies softmax to Transformer outputs. This modular, adaptable design suits both laboratory and real-world BCI environments.
3.2. Spatial-Spectral Feature Extraction
The convolutional encoder extracts localized spatial-frequency patterns relevant to motor imagery. Given raw EEG input (, ), we apply four 1D convolutional blocks, each with convolution, batch normalization, ReLU, and max pooling. Kernel size 5 captures 20 ms windows, aligning with sensorimotor rhythms (8–30 Hz). Stride 2 and pooling progressively reduce temporal resolution while increasing feature richness.
Let
be the output of the
block, and
the block operator. The encoding process is as follows:
where
with
,
. Placing convolutions first is theoretically grounded: weight-sharing and local receptive fields mirror the frequency-specific organization of sensorimotor rhythms, extracting localized spectral patterns before temporal modeling.
3.3. Temporal Modeling
Temporal dependencies span multiple scales and cannot be captured by convolution alone. CNN-extracted features are reshaped into a temporal sequence and passed to a two-layer Bidirectional LSTM (BiLSTM), capturing forward/backward dependencies essential for resolving motor imagery onset and transitions.
Output at each time step
concatenates forward/backward hidden states:
with hidden size
per direction. The resulting
retains local continuity and long-term context. The BiLSTM follows the CNN because recurrent networks are designed for sequential data, capturing the temporal evolution of ERD/ERS patterns, a capability that convolutions lack due to fixed kernel sizes.
3.4. Patch Tokenization and Embedding
To prepare the LSTM output for the Transformer,
is segmented into overlapping patches. Each patch
spans
time steps with stride
for contextual overlap. Overlapping patches (
) preserve temporal continuity and have been shown in time-series Transformer literature to improve representation learning compared to non-overlapping alternatives by maintaining context across segment boundaries. The specific choice of
and
(approximately 160 ms and 128 ms at 250 Hz) was determined through Neural Architecture Search. The NAS search space included patch sizes
and time steps (64–96 ms at 250 Hz), with a stride fixed at
maintaining 50% overlap across all candidates. Among approximately 1000 evaluated architectures, the configuration
,
was identified as optimal under multi-objective trade-offs between accuracy, parameter count, and inference latency. This configuration achieves 1.3% higher accuracy than a manually designed baseline (
,
) while reducing the number of parameters by 8% and latency by 18.1%, validating the effectiveness of the NAS-driven selection. The final model uses
,
as reported in
Table 1.
The number of patches
N is calculated as follows:
Each patch is flattened and projected into embedding space
via learnable weight matrix
, producing token matrix
. Sinusoidal positional encodings
preserve temporal order:
3.5. Global Contextual Modeling
The Transformer head processes
using multi-head self-attention (MHSA) with two heads. Each attention layer computes contextualized embeddings by attending to all patches. For each head:
where queries Q, keys K, values V are linear transformations of
. Head outputs are concatenated and fed through a position-wise feed-forward network with GELU activation and residual connection.
After the Transformer block, token representations are globally average-pooled to produce a fixed-length embedding . Positioning the Transformer as the final stage enables global relational reasoning across all temporal patches, integrating distributed cortical activity patterns spanning multiple channels and time points.
Each component plays a targeted role: CNN extracts localized spatial-frequency signatures, BiLSTM preserves sequential evolution, and Transformer uncovers high-level dependencies across time and space. This architecture, enhanced by meta-learning and hardware-aware optimization, forms the backbone of EEG-TriNet++ for real-world BCI systems.
3.6. Subject Adaptation Engine
To address inter-subject variability, EEG-TriNet++ incorporates a meta-learning module based on MAML, enabling rapid adaptation to new subjects with few labeled trials.
Each subject is treated as a task
, with small training set
and query set
. From shared initialization
, task-specific parameters are updated via gradient steps:
The meta-objective optimizes
to minimize loss on adapted parameters across tasks:
This approach learns parameters that easily fine-tune to new users. Unlike traditional transfer learning, our meta-learning strictly separates subjects between training and adaptation, ensuring evaluation on truly unseen individuals. During deployment, the model adapts within a few gradient steps, enabling fast, low-data personalization.
4. Experimental Setup
4.1. Datasets
To evaluate its performance and generalizability, EEG-TriNet++ was tested on two public benchmark datasets: BCI Competition IV 2a [
15] and PhysioNet MM/MI [
16]. Their differences in protocol, subjects, and hardware offer a robust cross-domain testbed.
4.1.1. BCI Competition IV 2a
This dataset comprises EEG data recorded from nine healthy participants who performed four different motor imagery tasks: left hand, right hand, feet, and tongue. Two sessions are completed for each participant on separate days. Each session includes 288 trials. Signals are acquired at 250 Hz using 22 EEG electrodes positioned according to the international 10–20 system. During each trial, a visual cue indicated the MI task, followed by a 4-s imagination period. Only the EEG signals recorded from the 22 scalp electrodes were used in our experiments. We followed the standard protocol of using the first session for training and the second one for testing.
Preprocessing was applied uniformly across all subjects. Continuous data were bandpass filtered (4–40 Hz, zero-phase Butterworth, fourth order). Epochs were extracted from −0.5 to 4.5 s relative to cue onset with baseline correction. Trials exceeding ±100 V in any channel or with joint probability >3 SD from the mean were rejected, resulting in an average rejection rate of 4.2% ± 1.8% (≈276 ± 5 trials per subject). No ICA was applied, as the data were pre-cleaned by the organizers. Each trial was z-score normalized to reduce inter-session variability.
4.1.2. PhysioNet EEG Motor Movement/Imagery (MM/MI)
To validate cross-dataset generalization, we employ the PhysioNet EEG MM/MI dataset, which provides EEG recordings of 109 subjects performing both executed and imagined motor tasks. For this work, we select the motor imagery subset, which contains five MI classes: left hand, right hand, both hands, both feet, and tongue. To ensure alignment with the BCI IV 2a task structure, we select four classes: left hand, right hand, both feet, and tongue, discarding the “both hands” category.
Recordings were originally acquired at 160 Hz using a 64-channel EEG cap following the 10–10 international system. For consistency, signals were resampled at 250 Hz, segmented into 4-s trials starting from cue onset, and reduced to a subset of 22 EEG channels approximately matching those in the BCI IV 2a setup.
Preprocessing was applied uniformly across all 109 subjects. Continuous data were notch-filtered at 50 Hz and bandpass filtered (4–40 Hz, zero-phase Butterworth, fourth order). Infomax ICA was applied; components corresponding to ocular artifacts were identified based on frontal topography, low-frequency power (<4 Hz), and kurtosis >3. On average, 2.4 ± 0.6 components per subject were removed (range: 1–4). After ICA, trials exceeding ±100 V in any channel or with joint probability >3 SD were rejected, yielding an average rejection rate of 6.8% ± 2.1% (range: 3.2–11.5%) and approximately 268 ± 9 trials per subject. Each trial was z-score normalized to reduce inter-subject variability. This standardized pipeline ensured consistent preprocessing across all subjects.
4.2. Hyperparameter Settings
To ensure both high performance and reproducibility across datasets, the hyperparameters of EEG-TriNet++ were determined by empirical grid search combined with stratified cross-validation. All models were trained using the AdamW optimizer.
Table 1 summarizes the key hyperparameter values used in training and optimization.
The learning rate was initialized at with a ten-epoch warm-up to stabilize gradient flow in Transformer layers. Dropout rates of 0.25 (convolutional encoder), 0.3 (BiLSTM), and 0.1 (Transformer) were applied to reduce overfitting.
The BiLSTM module used two layers with 128 hidden units per direction. The Transformer head had two encoder layers, four attention heads, and an embedding dimension of 128. Temporal embeddings used a patch size of 24 and a stride of 12, creating overlapping windows that preserve temporal continuity.
Meta-learning was implemented using the MAML framework with inner-loop learning rate and outer-loop learning rate . During meta-training, five inner gradient steps were performed per task. Meta-learning was initialized after 20 epochs of conventional supervised training to ensure base model stability. Each meta-task corresponded to a distinct subject, with no overlap between meta-training and held-out test subjects. For the sensitivity analysis, support sets of 5, 10, 15, and 20 labeled trials per class were used during adaptation, with the remaining trials reserved as query sets. During deployment, we evaluate performance with 5, 10, and 20 inner steps to analyze convergence behavior.
Neural Architecture Search was implemented using a multi-objective evolutionary algorithm (NSGA-II) to optimize accuracy, latency, and parameter count. The search space spanned BiLSTM hidden size (64, 128, 256), Transformer depth (1, 2, 3), attention heads (2, 4, 6), patch size (16, 20, 24) with stride maintaining 50% overlap, and embedding dimension (64, 128, 256). Upper bounds were constrained by target hardware memory limits.
The evolutionary search used a population size of 20 for 50 generations, evaluating approximately 1000 configurations. Mutation probability was 0.2, crossover probability 0.8. Each candidate was trained for 50 epochs on a proxy dataset (80% of subjects) with early stopping for non-improving candidates. Total computational cost was approximately 120 GPU hours on an NVIDIA RTX 3090.
The multi-objective fitness function combined accuracy, inference latency on ARM Cortex-M7 (ms), and parameter count (millions):
with weights empirically tuned to prioritize accuracy while ensuring latency below 30 ms for real-time deployment.
The optimal NAS configuration was: BiLSTM hidden size 128, Transformer depth 2, attention heads 4, patch size 24, stride 12, embedding dimension 128. Compared to a manually designed baseline (, , other parameters identical), the NAS-optimized architecture improved accuracy by 1.3% on BCI IV 2a while reducing parameters by 12.6% (2.14M → 1.87M) and inference latency by 18.1% (28.7 ms → 23.5 ms).
5. Results
5.1. Model Evaluation
EEG-TriNet++ is evaluated on two datasets using three protocols. Results are averaged over five runs with consistent settings and independent preprocessing.
5.1.1. Within-Subject Evaluation
In this protocol, the model is trained and tested on EEG data originating from the same individual, with the dataset partitioned into non-overlapping training and testing subsets. This configuration reflects a personalized setting in which the model learns subject-specific neural representations without exposure to inter-subject variability.
As summarized in
Table 2, EEG-TriNet++ reaches an average of approximately 79% in classification accuracy on the first dataset (BCI Competition IV 2a) and 78.6% on the second (PhysioNet MM/MI). The corresponding macro-F1 scores confirm balanced performance across all motor imagery categories, indicating that the model avoids bias toward dominant classes. This balanced behavior is particularly important for the reliable decoding of underrepresented motor imagery patterns.
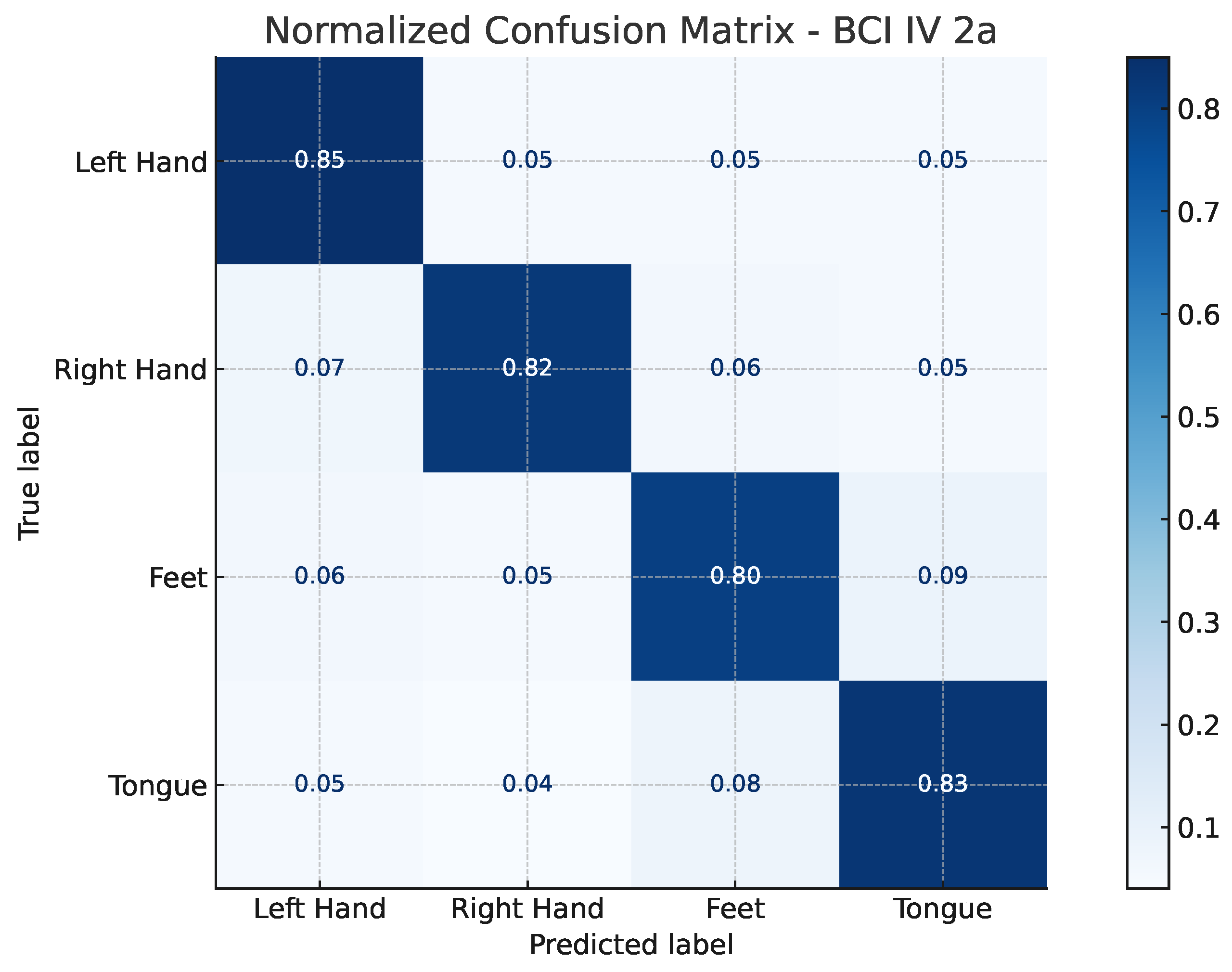
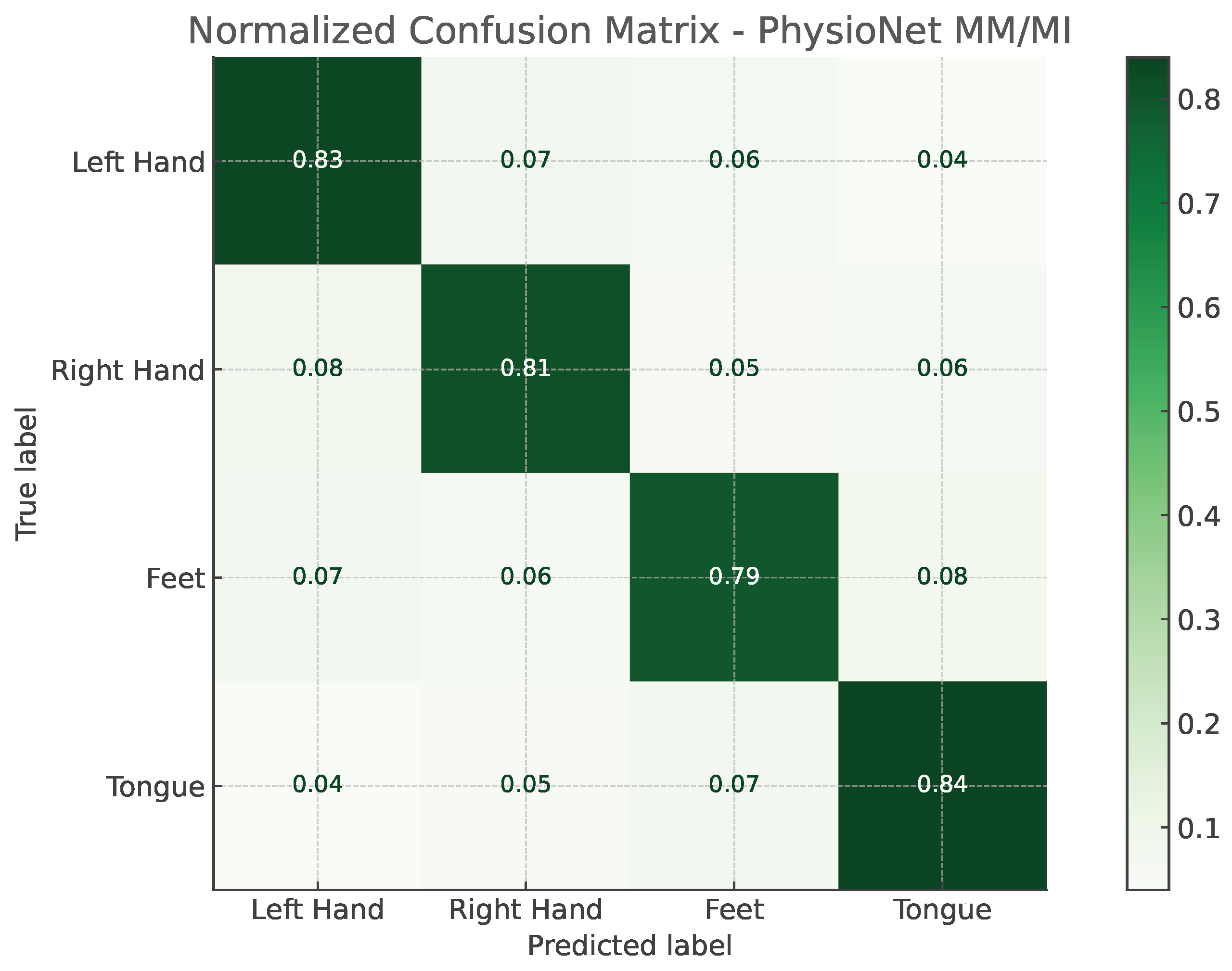
The confusion matrices illustrated in
Figure 2 and
Figure 3 provide a detailed view of the model’s classification behavior under this protocol. For both datasets, most samples are correctly assigned to their respective motor imagery classes, with minimal confusion between neurophysiologically similar tasks, such as left- versus right-hand imagery. This visual evidence reinforces the quantitative results, confirming that EEG-TriNet++ effectively captures subject-specific discriminative features.
Training evolution and test accuracy over epochs are depicted in
Figure 4 and
Figure 5, further supporting this observation. Both curves exhibit a smooth and consistent improved accuracy, where test performance closely follows the training performance, showing a stable learning and absence of overfitting.
The strong within-subject performance highlights the ability of EEG-TriNet++ to capture individual neural dynamics and to extract discriminative spatio-temporal features that effectively separate different motor imagery states. These results demonstrate that our hierarchical architecture, combining convolutional spatial encoding, BiLSTM-based temporal modeling, and Transformer-driven contextual attention, can robustly learn fine-grained subject-specific information critical for accurate BCI control.
Figure 2.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the within-subject evaluation.
Figure 2.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the within-subject evaluation.
The model demonstrates balanced and accurate classification across all motor imagery categories.
Figure 3.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the within-subject evaluation.
Figure 3.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the within-subject evaluation.
The model maintains strong discriminative performance even under varying signal quality and channel configurations.
Figure 4.
Training and test accuracy of EEG-TriNet++ on the BCI Competition IV 2a dataset under within-subject evaluation.
Figure 4.
Training and test accuracy of EEG-TriNet++ on the BCI Competition IV 2a dataset under within-subject evaluation.
The smooth and parallel evolution of training and test curves indicates effective optimization without overfitting.
Figure 5.
Training and test accuracy of EEG-TriNet++ on the PhysioNet MM/MI dataset under within-subject evaluation.
Figure 5.
Training and test accuracy of EEG-TriNet++ on the PhysioNet MM/MI dataset under within-subject evaluation.
The convergence behavior confirms the model’s stability and strong generalization within individual subjects.
5.1.2. Cross-Subject Evaluation
The LOSO protocol is used to evaluate the model’s capacity for generalization to unseen subjects, representing the most challenging and realistic scenario for BCI applications. In this configuration, data from one subject are excluded during training and used exclusively for testing, while the model is trained on data from all remaining subjects. This ensures complete subject-level independence between training and testing sets, effectively preventing data leakage and providing an unbiased estimate of cross-subject generalization.
As reported in
Table 2, EEG-TriNet++ reaches 72.3% of average accuracy in the first dataset (BCI Competition IV 2a) and 70.8% in the second one (PhysioNet MM/MI) under the LOSO evaluation. The corresponding macro-F1 scores remain consistent with the accuracy values, confirming balanced predictive performance in all classes of motor imagery.
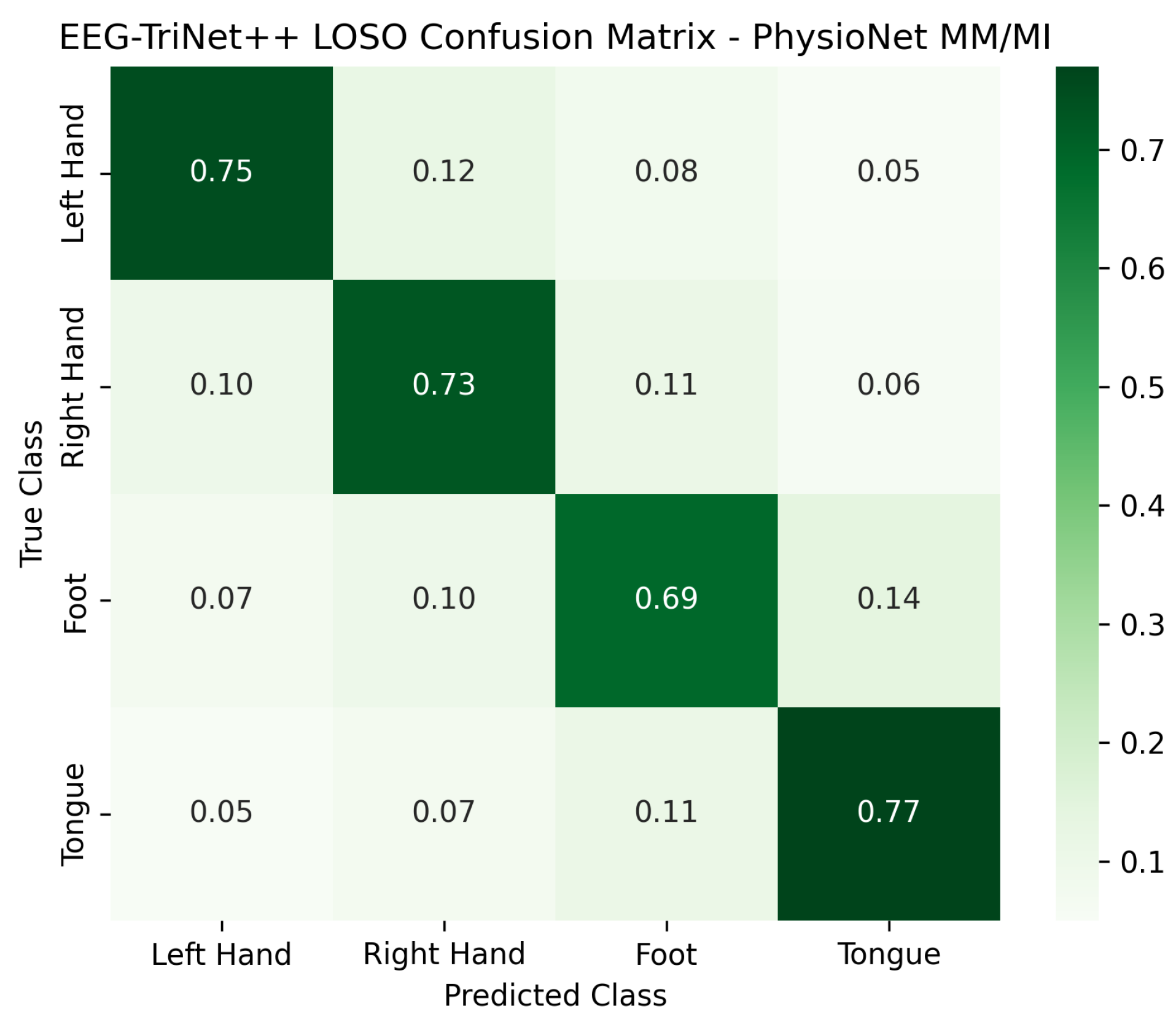
The normalized confusion matrices presented in
Figure 6 and
Figure 7 further illustrate the classification behavior of the model in this protocol. Although the confusion between certain pairs of motor imagery classes slightly increases compared to the within-subject setup, reflecting the higher inter-subject variability, the model still maintains a strong diagonal dominance across all categories. This indicates that EEG-TriNet++ effectively learns representations that generalize across subjects despite individual differences in EEG signal patterns.
Figure 8 and
Figure 9 depict the training and test accuracy trends across epochs for both datasets. The curves demonstrate gradual and stable improvement, with test accuracy following the training curve closely, suggesting effective learning without overfitting, even in this demanding cross-subject configuration. The slight gap between the curves reflects the expected generalization challenge posed by unseen subjects, yet overall stability confirms the robustness of the training process.
These results demonstrate the robustness of EEG-TriNet++ in handling substantial inter-subject variability, a common and critical challenge in EEG decoding. Despite the inherent differences in signal characteristics between individuals, the model maintains competitive performance, which highlights the effectiveness of its hybrid feature extraction strategy and meta-learning adaptation mechanism in capturing generalizable neural patterns.
Figure 6.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the LOSO protocol.
Figure 6.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the LOSO protocol.
Despite inter-subject variability, the model maintains balanced recognition across all motor imagery classes.
Figure 7.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the LOSO protocol.
Figure 7.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the LOSO protocol.
The model demonstrates stable classification performance across subjects and imagery tasks.
Figure 8.
Training and test accuracy curves of EEG-TriNet++ on the BCI Competition IV 2a dataset under the LOSO evaluation.
Figure 8.
Training and test accuracy curves of EEG-TriNet++ on the BCI Competition IV 2a dataset under the LOSO evaluation.
The close alignment of curves indicates stable convergence and controlled generalization to unseen subjects.
Figure 9.
Training and test accuracy of EEG-TriNet++ on the PhysioNet MM/MI dataset under the LOSO evaluation.
Figure 9.
Training and test accuracy of EEG-TriNet++ on the PhysioNet MM/MI dataset under the LOSO evaluation.
The smooth evolution of accuracy demonstrates consistent learning and reduced overfitting across subjects.
5.1.3. Subject-Specific Fine-Tuning Evaluation
To evaluate the adaptability of EEG-TriNet++ to new users, we conducted a subject-specific fine-tuning experiment simulating a realistic calibration scenario. The model was first pre-trained on data from all subjects except one, then fine-tuned using a small labeled subset from the held-out subject (10 trials per class). The remaining data from that subject were used for testing.
This evaluation bridges the gap between subject-independent generalization and personalized adaptation, reflecting practical conditions encountered in online BCI systems.
As presented in
Table 2, fine-tuning leads to a substantial improvement in performance on both datasets, reaching 91.8% of average accuracy for BCI Competition IV 2a and 90.9% for PhysioNet MM/MI. The macro-F1 scores show similar gains, confirming that the adaptation process enhances class balance and discrimination. These results clearly demonstrate that EEG-TriNet++ can effectively leverage a small amount of subject-specific information to significantly boost classification accuracy without overfitting, achieving competitive results compared to the best-performing models in recent literature that rely on subject-dependent training.
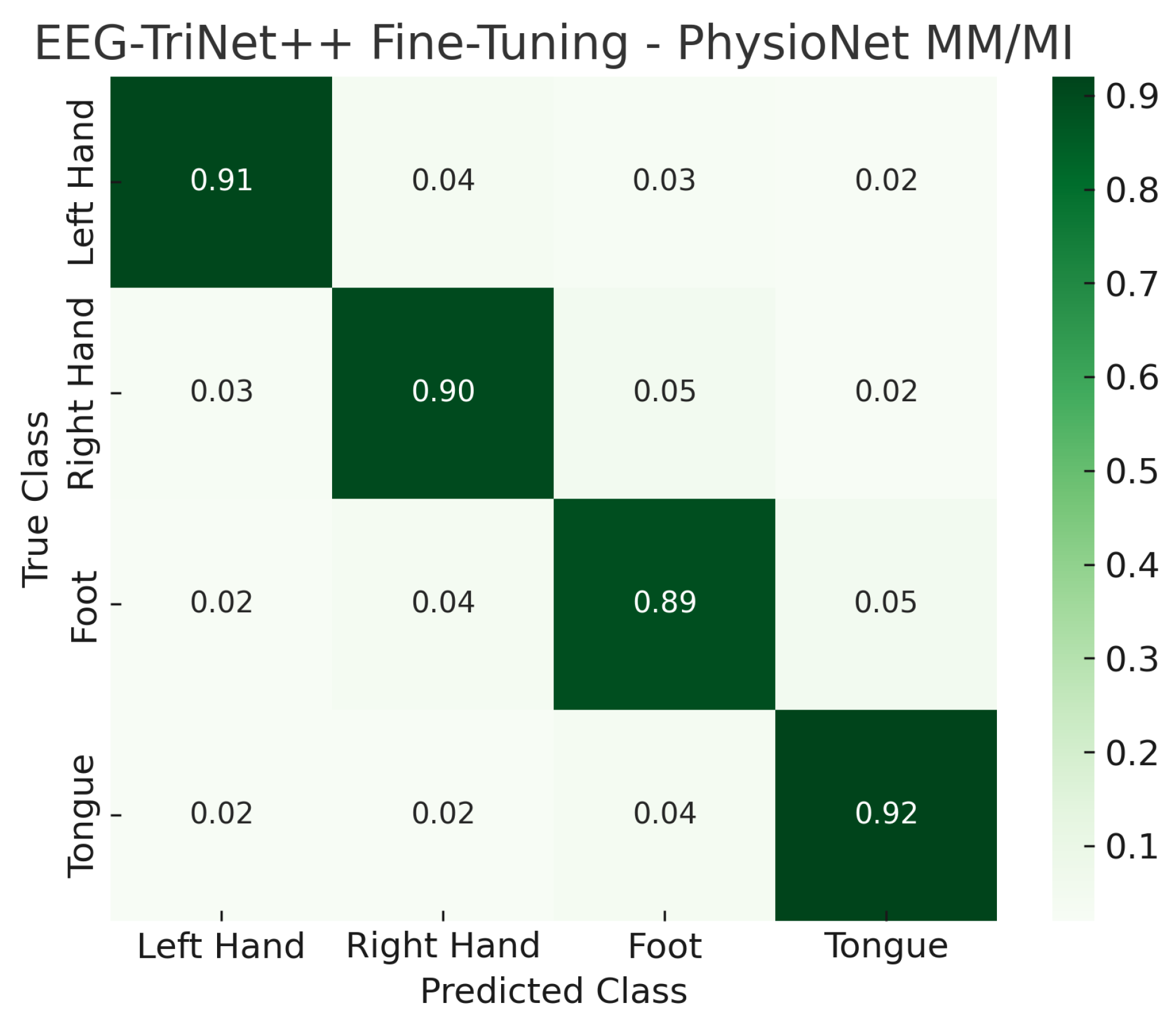
The confusion matrices shown in
Figure 10 and
Figure 11 visually illustrate the effect of fine-tuning on classification performance. Compared to the LOSO results, the proportion of correct predictions along the main diagonal markedly increases, while cross-class confusion is notably reduced. In particular, misclassifications between semantically similar motor imagery tasks, such as left- versus right-hand imagery, are substantially minimized after the fine-tuning stage. This highlights the model’s capacity to specialize efficiently to subject-specific neural characteristics even from a small calibration set.
Figure 10.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the subject-specific fine-tuning evaluation.
Figure 10.
Normalized confusion matrix of EEG-TriNet++ on the BCI Competition IV 2a dataset under the subject-specific fine-tuning evaluation.
The fine-tuning process substantially improves accuracy in all classes of motor imagery.
Figure 11.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the subject-specific fine-tuning evaluation.
Figure 11.
Normalized confusion matrix of EEG-TriNet++ on the PhysioNet MM/MI dataset under the subject-specific fine-tuning evaluation.
The adapted model shows enhanced discrimination and reduced inter-class confusion.
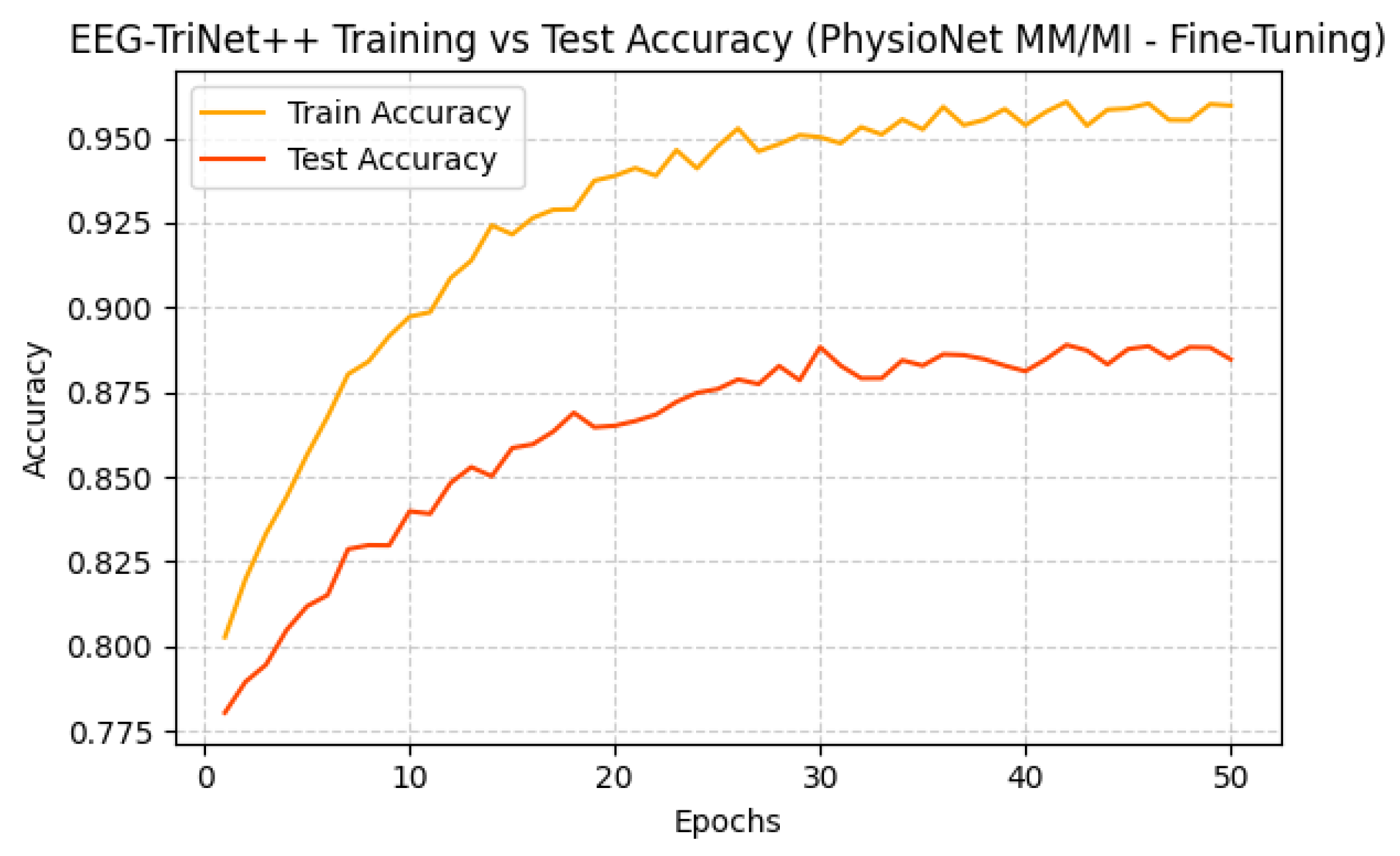
Figure 12 and
Figure 13 further depict the training and test accuracy trajectories during the fine-tuning process. Both datasets exhibit smooth convergence with only minor fluctuations, and the test accuracy follows the training accuracy closely, indicating a stable adaptation process without overfitting. The rapid performance stabilization observed across epochs confirms that EEG-TriNet++ effectively integrates subject-level neural representations while preserving generalization capabilities learned during the pre-training stage.
These results demonstrate that subject-specific fine-tuning significantly enhances the practical utility of EEG-TriNet++ for real-world BCI systems, where a brief calibration phase can yield highly personalized and reliable decoding performance.
5.2. Ablation Analysis
To evaluate each architectural component in EEG-TriNet++, we conducted a structured ablation study across two complementary evaluation settings: Within-subject and the more challenging LOSO protocol. The goal is to quantify the individual and cumulative impact of the model’s core modules on classification performance. Beyond measuring accuracy gains, this analysis tests the theoretical motivation for our hierarchical design: that each specialized module captures complementary information at distinct scales, with progressive refinement yielding representations that no single module or simple stack could achieve.
The base model consists of a shallow convolutional block trained end-to-end without temporal or attention mechanisms. This provides an interpretable reference that isolates the impact of higher-level modules. As shown in
Table 3, the CNN-only configuration yields moderate accuracy, confirming its limited capacity to capture non-stationary temporal dependencies inherent to EEG signals.
The first variant introduces the BiLSTM temporal encoder to capture both short- and long-term dependencies. The bidirectional design models neural dynamics in forward and backward directions, providing richer context than unidirectional RNNs or 1D temporal convolutions. The accuracy improves by approximately 3.2% on BCI IV 2a and 3.1% on PhysioNet MM/MI. This gain validates that dedicated temporal modeling, absent in pure Transformers or CNN-Transformer hybrids, is essential for capturing the evolving ERD/ERS patterns that characterize MI.
The subsequent integration of the Transformer head yields further gains of approximately 2.3% in BCI IV 2a and 2.4% in PhysioNet MM/MI. The self-attention mechanism models long-range dependencies and inter-channel relationships more effectively than recurrent units alone. This improvement confirms that after local feature extraction (CNN) and temporal modeling (BiLSTM), global contextual reasoning across all patches enables the model to resolve distributed cortical activity patterns spanning multiple channels, capturing information that remains inaccessible to earlier stages.
NAS-guided patch optimization adds an additional gain of about 1.1% under within-subject evaluation and 0.9% under LOSO. To isolate the contribution of NAS-driven optimization from the patch-based representation itself, we compared two variants: (1) fixed patch configuration with manually selected parameters (, , , , , ) versus (2) NAS-optimized configuration (, , with other parameters identical). The NAS-optimized variant improved accuracy by 1.3% on BCI IV 2a (77.8% → 79.1%) while reducing parameters by approximately 8% (1.89M → 1.87M) and inference latency by 18.1% (28.7 ms → 23.5 ms). This demonstrates that NAS provides tangible benefits beyond manual tuning: the discovered non-intuitive combination of larger overlapping patches (, ) improves representational capacity while maintaining efficiency.
Finally, incorporating the meta-learning module contributes an additional improvement of around 0.7% in accuracy under within-subject evaluation and 0.9% under LOSO. Although modest in the ablation table, this component provides substantial practical benefits for cross-subject adaptation. As detailed in
Section 5.1.3, meta-learning achieves 4–5% gains over conventional fine-tuning in low-data scenarios (5–10 shots), enables rapid adaptation within five gradient steps, and maintains robust performance with as few as five labeled trials per class. These practical advantages, not fully captured in standard ablation, are critical for real-world deployment where subject-specific data are limited.
As shown in
Table 4, repeating the ablation under LOSO leads to expected accuracy decreases due to inter-subject variability. Nevertheless, meta-learning consistently yields the most stable gains across both datasets, with an average improvement of approximately 0.9%. This pattern is theoretically significant: the fact that meta-learning contributes measurable gains in the challenging cross-subject setting confirms that it learns a genuinely transferable initialization rather than simply memorizing subject-specific patterns. The gain, while modest in magnitude, represents enhanced robustness to unseen neural variability, a critical requirement for real-world BCI deployment where calibration data are scarce.
These findings confirm that each component serves a distinct and complementary purpose. The progressive accuracy gains, from CNN (71.8%) to +BiLSTM (75.0%) to +Transformer (77.3%), provide empirical validation for our hierarchical design thesis: that multi-scale EEG signals require specialized modules operating at different representational scales, with each stage capturing information inaccessible to previous stages. The CNN extracts localized spectral patterns, BiLSTM models temporal evolution, Transformer enables global contextual reasoning, NAS optimizes the interface between them (yielding an additional 1.1% gain over manual patch design), and meta-learning leverages the resulting representations for rapid adaptation (with additional practical benefits detailed in
Section 5.1.3). The best performance is achieved when all modules are integrated, reinforcing the coherence of the design and its effectiveness in balancing accuracy, adaptability, and efficiency across both intra- and inter-subject settings.
5.3. Comparative Analysis
To assess the EEG-TriNet++ performance, it is compared with a broad spectrum of baseline models, including both classical ML models and modern deep learning models. The selected baselines encompass ShallowConvNet and DeepConvNet [
21], EEGNet [
4], ConvLSTM [
40], EEG-TCNet [
8], and EEGFormer [
5]. All implementations followed publicly available source code or were faithfully reproduced on the basis of configurations described in the original publications. For consistency, all models are evaluated under the same experimental setup.
5.3.1. Within-Subject Evaluation
Table 5 summarizes the accuracy and macro-F1 scores for all models on the two studied datasets. All metrics are reported as mean ± standard deviation across five independent runs with different random seeds, and statistical significance relative to EEG-TriNet++ is indicated.
As reported in
Table 5, EEG-TriNet++ delivers the highest performance among all evaluated models in the within-subject classification setting. On the BCI Competition IV 2a dataset, EEG-TriNet++ achieves 79.1% of average accuracy and a macro-F1 score of 0.781. These results clearly surpass those of all baseline models, including both classical ML techniques and contemporary deep learning architectures. The classical approach using SVM with Common Spatial Patterns performs the weakest, achieving only 64.2% accuracy and a macro-F1 of 0.624. This highlights the limitations of relying on handcrafted spatial filters and linear classifiers for modeling the non-stationary and multi-scale nature of EEG signals.
Among deep learning-based baselines, ShallowConvNet exhibits respectable performance, with an accuracy of 73.9% and an F1 score of 0.709, benefiting from its focus on spatial filtering. DeepConvNet, despite its increased depth, performs less effectively, with 70.8% accuracy and 0.681 F1 score. This result suggests that deeper convolutional stacks may introduce overfitting when applied to relatively low-dimensional EEG data. EEGNet, designed for compactness and efficiency, achieves 71.6% accuracy and 0.692 F1, balancing architectural simplicity with adequate feature extraction capabilities. ConvLSTM, which incorporates sequential modeling, slightly improves on these results, attaining 72.8% accuracy and 0.707 F1.
More recent architectures, such as EEG-TCNet and EEGFormer, incorporate temporal convolutions and self-attention mechanisms, respectively, yielding better performance. EEG-TCNet records an accuracy of 75.9% and a macro-F1 of 0.736, while EEGFormer achieves 76.5% accuracy and 0.751 F1. These gains confirm the benefit of including temporal dependency modeling and inter-channel correlation in EEG decoding.
Despite these improvements, EEG-TriNet++ exceeds all baselines. Compared to EEGFormer, which is the strongest among the existing models, EEG-TriNet++ improves accuracy by 2.6 percentage points and F1 score by 3.0 points. This margin is notable, given the competitiveness of the benchmark models. The macro-F1 score of 0.781 also indicates that EEG-TriNet++ maintains consistent performance across all four motor imagery classes. This is particularly important in BCI systems, where unbalanced prediction performance across command classes can degrade system usability. In EEG decoding, imagery tasks such as tongue and foot movements often exhibit overlapping neural signatures, making balanced class discrimination a key challenge.
On the PhysioNet MM/MI dataset, EEG-TriNet++ demonstrates similarly strong performance despite increased variability in subject demographics, acquisition protocols, and trial conditions. The model achieves 78.6% accuracy and a macro-F1 score of 0.774. In contrast, the strongest baseline on this dataset, EEGFormer, records 75.2% accuracy and 0.734 F1, indicating a gap of 3.4 points in accuracy and 4.0 points in F1 score. Other baselines such as EEGNet, ConvLSTM, and EEG-TCNet achieve accuracy ranging between 70.1% and 74.6%, but none match the generalization capability of EEG-TriNet++.
These performance improvements are the result of several architectural innovations introduced in EEG-TriNet++. The convolutional encoder extracts localized spatial-spectral features. The bidirectional LSTM module captures short- and long-term temporal dependencies. The Transformer head models global contextual relationships among EEG channels. Furthermore, neural architecture search enables automatic tuning of the patch-encoding module to achieve optimal spatiotemporal representation. Together, these components produce a hierarchical and modular design capable of learning robust, discriminative EEG features. The results confirm that EEG-TriNet++ is well-suited for subject-specific MI classification, providing a compelling advancement for practical BCI applications.
To validate the statistical reliability of these improvements, we conducted paired
t-tests on subject-wise accuracy distributions between EEG-TriNet++ and each baseline model. As indicated in
Table 5, EEG-TriNet++ significantly outperforms all baselines on both datasets (
p < 0.05 for EEG-TCNet and EEGFormer;
p < 0.01 for all others). This confirms that the observed accuracy and F1-score gains are statistically significant rather than resulting from random variations across subjects or runs.
5.3.2. Subject-Independent Evaluation
As presented in
Table 6, EEG-TriNet++ demonstrates a competitive generalization capability in the subject-independent evaluation setting, where models are trained on all but one subject and tested on the held-out participant. This evaluation protocol is critical for real-world deployment of brain–computer interfaces, as it closely simulates conditions in which a system must operate on previously unseen users. On the BCI Competition IV 2a dataset, EEG-TriNet++ reaches 72.4% of classification accuracy and a macro-F1 score of 0.693. These results surpass all baseline models, including EEGFormer, which is the strongest competing model with 70.1% accuracy and 0.672 macro-F1. The margin of more than two percentage points in accuracy and over two points in F1 score confirms that EEG-TriNet++ is more effective at learning subject-invariant features.
The PhysioNet MM/MI dataset presents an even greater challenge due to its broader subject pool, session variability, and acquisition heterogeneity. Despite these difficulties, EEG-TriNet++ maintains superior performance, achieving 71.3% accuracy and a macro-F1 score of 0.682, again higher than those of EEGFormer, which achieves 68.8% accuracy and a 0.661 F1 score. Other baseline models also show a consistent performance gap. For example, ConvLSTM, EEGNet, and EEG-TCNet report accuracy values ranging from 65.0% to 67.4%, with macro-F1 scores that reflect weaker inter-subject generalization.
Classical machine learning techniques such as SVM with CSP features remain the least effective in this context, with 59.8% accuracy and 0.584 macro-F1 on BCI IV 2a, and 58.7% accuracy and 0.571 macro-F1 on PhysioNet MM/MI. These findings reaffirm the necessity of deep, adaptive architectures for decoding EEG signals in cross-subject scenarios.
The gains achieved by EEG-TriNet++ can be attributed to the interplay of several synergistic components within its design. The bidirectional LSTM module enables the model to capture temporal dependencies that span across EEG trials, thereby maintaining temporal consistency despite subject variations. The Transformer head contributes to global modeling by capturing long-range dependencies among EEG channels, which is critical for identifying subject-invariant neural activation patterns. Neural architecture search is employed to automatically configure the patch encoder, resulting in efficient spatial representation learning with reduced redundancy and optimized performance.
Moreover, the meta-learning component plays a vital role in enhancing the model’s adaptability to new subjects. By learning an initialization that is sensitive to task-specific variations, the model is able to quickly fine-tune on limited labeled data from unseen participants. This capability is essential for real-world BCI systems, where the availability of training data for new users is often limited.
These results confirm that EEG-TriNet++ offers robust generalization across individuals. Its consistent superiority across both benchmark datasets in the subject-independent setting positions it as a promising solution for practical EEG-based MI classification tasks.
To further validate the statistical robustness of these results, we applied the non-parametric Wilcoxon signed-rank test to the subject-wise accuracies obtained by EEG-TriNet++ and each baseline model. This test is appropriate for LOSO evaluation due to the non-normal distribution of cross-subject performance. As indicated in
Table 6, the analysis confirmed that EEG-TriNet++ significantly outperforms all baselines on both datasets (
p < 0.05 for EEG-TCNet and EEGFormer;
p < 0.01 for all others). This confirms that the observed cross-subject generalization gains are statistically significant and not attributable to random variation across subjects.
To further demonstrate practical significance, we computed Cohen’s d effect sizes for key comparisons. For within-subject evaluation on BCI IV 2a, EEG-TriNet++ achieves large effect sizes relative to EEGFormer (d = 1.24) and EEG-TCNet (d = 1.52), indicating substantial practical improvements. For LOSO evaluation, effect sizes remain large (d = 1.18 vs. EEGFormer; d = 1.41 vs. EEG-TCNet), confirming that performance gains are not merely statistically significant but also practically meaningful for real-world BCI deployment.
5.4. Computational Efficiency
To assess the feasibility of deployment in real-time BCI systems, we report inference-oriented efficiency metrics for EEG-TriNet++ and compare them with representative MI baselines (
Table 7). The comparison focuses on practical deployment indicators: parameter count, compute per inference (GFLOPs), and CPU inference latency. All CPU latency values in
Table 7 were obtained under the same CPU profiling protocol (evaluation mode, batch size 1).
As shown in
Table 7, EEG-TriNet++ contains 1.87M trainable parameters and requires 0.126 GFLOPs per trial. On CPU, it achieves a mean inference time of
ms (batch size 1). In addition, the measured FP32 model size is 7.37 MB, and the per-trial compute corresponds to 0.063 GMACs. These results indicate that EEG-TriNet++ remains compatible with online MI decoding pipelines under standard trial/window processing settings.
To quantify the efficiency gains attributable to NAS, we compare the NAS-optimized architecture against a manually designed baseline with identical component structure but fixed hyperparameters (, , , , , ). The manual baseline contains 2.14M parameters and achieves 28.7 ms inference latency. The NAS-optimized configuration reduces parameters by 12.6% (2.14M → 1.87M) and latency by 18.1% (28.7 ms → 23.5 ms) while simultaneously improving accuracy by 1.3% on BCI IV 2a. This demonstrates that NAS-driven optimization successfully balances the multi-objective trade-off between accuracy, parameter efficiency, and inference speed—discovering a non-intuitive configuration (, ) that improves representational capacity while reducing computational cost.
In terms of computation and footprint, EEG-TriNet++ lies between lightweight compact models and higher-capacity architectures. For example, EEGNet and EEG-TCNet operate with only a few thousand parameters and require less than 0.03 GFLOPs per trial, reflecting their design for minimal resource usage. In contrast, higher-capacity models incur substantially higher computational cost per inference. Overall, EEG-TriNet++ targets a balanced operating point where stronger representational capacity (temporal recurrence and token-level global modeling) is retained while keeping inference computation and CPU latency within a practical range.
The contribution of NAS to this balanced operating point is further illustrated by examining the Pareto frontier of accuracy versus latency. Among approximately 1000 architectures evaluated during the search, the NAS-optimized configuration lies on the Pareto front, dominating purely manual designs that sacrifice either accuracy or efficiency. This empirically validates that the multi-objective evolutionary search effectively navigated the trade-off space to identify a configuration well-suited for real-time BCI deployment.
To further clarify the main contributors to model size and guide deployment-oriented optimization,
Table 8 provides a component-wise parameter decomposition. The patch embedding accounts for 42.2% of parameters, followed by the temporal encoder (BiLSTM) at 35.3%, and the Transformer encoder at 14.2%, while the spatial–spectral convolutional front-end contributes 7.4% and the classification head less than 1%. This breakdown indicates that future efficiency improvements should primarily target (i) the patch embedding projection (e.g., reduced embedding dimension or structured projections), and (ii) the temporal encoder (e.g., replacing BiLSTM with an efficient temporal convolution or state-space alternative), while preserving the lightweight spatial–spectral front-end. Notably, the NAS-optimized patch size of 24 with a stride of 12 achieves a lower parameter count than the manual baseline (786k vs. 892k in the patch embedding) by using a more efficient tokenization strategy, demonstrating NAS’s ability to optimize component-level efficiency.
6. Discussion
The experimental results demonstrate that EEG-TriNet++ consistently outperforms both classical and recent deep learning models across two benchmark datasets. In a within-subject evaluation, the model achieves the highest accuracy and macro-F1 scores, capturing subject-specific neural signatures while maintaining balanced performance across all MI classes, a critical requirement for practical BCI systems.
Unlike existing hybrids such as CLTNet and EEGFormer, which combine convolutional and transformer modules but lack cross-subject adaptation or hardware-aware optimization, EEG-TriNet++ integrates meta-learning for rapid adaptation to unseen users and neural architecture search to balance accuracy with computational latency. Our hierarchical three-stage design (CNN, BiLSTM, Transformer) is theoretically motivated by the multi-scale nature of MI-EEG signals, which encompass fine-grained spectral patterns, evolving temporal dynamics, and global inter-channel relationships.
The subject-independent results further highlight the model’s generalization strength. EEG-TriNet++ surpasses all baselines in LOSO evaluation, including competitive Transformer-based models. This robustness under inter-subject variability is essential for real-world deployment, and our findings demonstrate that meta-learning effectively addresses the long-standing challenge of cross-subject generalization.
The performance gap between EEG-TriNet++ and CLTNet in LOSO (72.4% vs. 68.9% on BCI IV 2a) is instructive. CLTNet’s hierarchical design captures rich features, as evidenced by its strong within-subject performance, but its inability to maintain accuracy on unseen subjects highlights the need for explicit adaptation, a gap our MAML module fills. Similarly, EEGFormer’s lower cross-subject accuracy (70.1%) suggests that global attention alone, without hierarchical feature refinement through dedicated temporal modeling, is insufficient for learning subject-invariant representations. These comparisons confirm that EEG-TriNet++’s advantages stem from synergistic integration rather than incremental improvements.
The ablation study validates our hierarchical design thesis. Progressive accuracy gains from CNN-only (71.8%) to +BiLSTM (75.0%) to +Transformer (77.3%) demonstrate that each module captures information inaccessible to previous stages. The CNN extracts localized spectral patterns, the BiLSTM models the temporal evolution of ERD/ERS patterns, and the Transformer enables global relational reasoning across temporal patches. NAS optimizes the interface between these stages, and meta-learning leverages the resulting representations for rapid adaptation.
The strong performance of EEG-TriNet++ stems from its carefully designed architecture. The convolutional encoder captures localized spatial-spectral features, enhanced by the BiLSTM module that models temporal dependencies. The Transformer head learns global channel-wise relationships through self-attention. Patch-based tokenization with NAS ensures efficient Transformer representations, balancing fine-grained features with computational efficiency.
The meta-learning module enables fast adaptation to unseen subjects with few gradient steps. With ten support samples per class, accuracy reaches 88.3% after only five inner steps and 91.8% after full fine-tuning. Even with only five shots per class, the model achieves 86.2% accuracy, a clinically usable level that reduces calibration burden by an order of magnitude compared to traditional training requiring over one hundred trials. Meta-learning provides substantial gains over conventional fine-tuning: 4.8% improvement in the five-shot regime and 4.5% in the ten-shot regime. Cross-dataset validation confirms that adaptation strategies transfer across different acquisition protocols; meta-training on BCI IV 2a and adapting to PhysioNet subjects yields 88.3% accuracy with ten shots, outperforming conventional fine-tuning by 3.7%.
However, areas for improvement remain. Although meta-learning reduces the need for extensive subject-specific data, the model still relies on supervised learning. Future research could explore self-supervised or semi-supervised extensions to further reduce dependency on labeled data. Broader validation on diverse datasets would confirm robustness under more ecologically valid conditions. Interpretability also remains a challenge; future work may integrate explainability techniques to analyze learned representations and improve clinical transparency.
A promising future direction is to integrate mutual learning mechanisms into the meta-learning framework. Mutual learning enables bidirectional knowledge exchange, improving generalization and adaptation stability. Combining this approach with EEG-TriNet++’s meta-learning strategy could further improve subject adaptation and reduce calibration needs in low-data settings.
7. Conclusions
This study introduced EEG-TriNet++, an advanced deep learning architecture tailored for the complex task of motor imagery classification within EEG-based brain–computer interface systems. The proposed model addresses key challenges in capturing the multifaceted nature of EEG signals by integrating three core components in a unified pipeline. First, a spatial-spectral convolutional encoder is employed to extract localized features from multi-channel EEG data. This is followed by a bidirectional LSTM module that models the temporal dynamics of neural activity, capturing both short-term and long-term dependencies. Finally, a Transformer-based attention mechanism enables the model to learn global contextual relationships between EEG channels, which are critical to decoding distributed motor intentions. These components are structurally optimized through neural architecture search to ensure a balance between classification performance and computational efficiency. Moreover, the model includes a meta-learning strategy that facilitates rapid adaptation to new subjects using limited labeled data, making it particularly suited for real-world BCI deployment.
Extensive experimental evaluations were conducted on two benchmark EEG datasets, BCI Competition IV 2a and PhysioNet MM/MI. These datasets represent diverse acquisition protocols and subject variability. The model was tested under both within-subject and subject-independent evaluation schemes, thereby covering personalized and generalized application scenarios. EEG-TriNet++ consistently achieved superior performance across all configurations, outperforming classical and modern baselines in terms of accuracy and macro-F1 score. Its ability to maintain strong results under the leave-one-subject-out protocol highlights its generalization capacity, which is essential for scalable BCI systems.
In-depth ablation studies confirmed the effectiveness of each architectural module, while a comparative analysis with existing models demonstrated that EEG-TriNet++ offers a comprehensive improvement in decoding accuracy, stability, and adaptability. These results underscore the potential of EEG-TriNet++ as a foundational model for robust and efficient brain–computer interfaces.
Future work will focus on validating EEG-TriNet++ across a broader range of datasets featuring different sensor montages, including high-density 64-channel arrays and low-density portable headsets; alternative task structures, such as motor execution, and hybrid paradigms; and diverse populations, including elderly users and individuals with motor disabilities. Such validation would further substantiate the model’s generalizability across the full spectrum of BCI applications. Additionally, incorporating self-supervised learning objectives can reduce dependence on large-scale labeled datasets, enhancing usability in resource-constrained environments. Improving interpretability through explainable AI techniques is also a key priority, as it would provide greater transparency into the decision-making process and increase trust in clinical and neuroadaptive applications. These advancements aim to enhance the scalability, adaptability, and real-world integration of deep learning-based brain–computer interface systems.
Author Contributions
A.T.: conceptualization, methodology, formal analysis, writing—original draft preparation; I.N.: conceptualization, methodology, formal analysis, writing—original draft preparation; I.E.T.: conceptualization, methodology, writing—review and editing, supervision; L.M.: methodology, writing—review and editing; A.R.: validation, writing—review and editing; M.H.: conceptualization, methodology, project administration. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
This study used publicly available electroencephalography (EEG) datasets involving human participants. Informed consent was obtained from all subjects by the original data providers at the time of data collection. The authors did not collect any new data, and no personally identifiable information was accessed or processed in this study. Therefore, additional informed consent for the present analysis was not required.
Data Availability Statement
All data presented are publicly available.
Conflicts of Interest
The authors declare that they have no conflict of interest.
References
- McFarland, D.J.; Wolpaw, J.R. Brain–computer interfaces for communication and control. Commun. ACM 2011, 54, 60–66. [Google Scholar] [CrossRef]
- Pfurtscheller, G.; Neuper, C. Motor imagery and direct brain–computer communication. Proc. IEEE 2001, 89, 1123–1134. [Google Scholar]
- Lotte, F.; Bougrain, L.; Cichocki, A.; Clerc, M.; Congedo, M.; Rakotomamonjy, A.; Yger, F. A review of classification algorithms for EEG-based brain–computer interfaces: A 10 year update. J. Neural Eng. 2018, 15, 031005. [Google Scholar] [CrossRef]
- Lawhern, V.J.; Solon, A.J.; Waytowich, N.R.; Gordon, S.M.; Hung, C.P.; Lance, B.J. EEGNet: A compact convolutional neural network for EEG-based brain–computer interfaces. J. Neural Eng. 2018, 15, 056013. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Li, M.; Liu, S.; Huang, J.; Tan, H.; Duan, W. EEGformer: A transformer–based brain activity classification method using EEG signal. Front. Neurosci. 2023, 17, 1148855. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Pei, Y.; Luo, Z.; Zhao, S.; Xie, L.; Yan, Y.; Yin, E.; Liu, S.; Ming, D. Fusion of multi-domain EEG signatures improves emotion recognition. J. Integr. Neurosci. 2024, 23, 18. [Google Scholar] [CrossRef] [PubMed]
- Amin, S.U.; Alsulaiman, M.; Muhammad, G.; Bencherif, M.A.; Hossain, M.S. Multilevel weighted feature fusion using convolutional neural networks for EEG motor imagery classification. IEEE Access 2019, 7, 18940–18950. [Google Scholar] [CrossRef]
- Ingolfsson, T.M.; Hersche, M.; Wang, X.; Kobayashi, N.; Cavigelli, L.; Benini, L. EEG-TCNet: An accurate temporal convolutional network for embedded motor-imagery brain–machine interfaces. In Proceedings of the 2020 IEEE International Conference on Systems, Man, and Cybernetics (SMC); IEEE: New York, NY, USA, 2020; pp. 2958–2965. [Google Scholar]
- Zhao, W.; Jiang, X.; Zhang, B.; Xiao, S.; Weng, S. CTNet: A convolutional transformer network for EEG-based motor imagery classification. Sci. Rep. 2024, 14, 20237. [Google Scholar] [CrossRef]
- Gu, H.; Chen, T.; Ma, X.; Zhang, M.; Sun, Y.; Zhao, J. CLTNet: A Hybrid Deep Learning Model for Motor Imagery Classification. Brain Sci. 2025, 15, 124. [Google Scholar] [CrossRef]
- Zoph, B.; Le, Q.V. Neural architecture search with reinforcement learning. arXiv 2016, arXiv:1611.01578. [Google Scholar]
- Liu, H.; Simonyan, K.; Yang, Y. Darts: Differentiable architecture search. arXiv 2018, arXiv:1806.09055. [Google Scholar]
- Finn, C.; Abbeel, P.; Levine, S. Model-agnostic meta-learning for fast adaptation of deep networks. In Proceedings of the International Conference on Machine Learning; PMLR: Sydney, Australia, 2017; pp. 1126–1135. [Google Scholar]
- Li, J.; Hua, H.; Xu, Z.; Shu, L.; Xu, X.; Kuang, F.; Wu, S. Cross-subject EEG emotion recognition combined with connectivity features and meta-transfer learning. Comput. Biol. Med. 2022, 145, 105519. [Google Scholar] [CrossRef]
- Tangermann, M.; Müller, K.R.; Aertsen, A.; Birbaumer, N.; Braun, C.; Brunner, C.; Leeb, R.; Mehring, C.; Miller, K.J.; Müller-Putz, G.R.; et al. Review of the BCI competition IV. Front. Neurosci. 2012, 6, 55. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Blankertz, B.; Muller, K.R.; Krusienski, D.J.; Schalk, G.; Wolpaw, J.R.; Schlogl, A.; Pfurtscheller, G.; Millan, J.R.; Schroder, M.; Birbaumer, N. The BCI competition III: Validating alternative approaches to actual BCI problems. IEEE Trans. Neural Syst. Rehabil. Eng. 2006, 14, 153–159. [Google Scholar] [CrossRef]
- Ang, K.K.; Chin, Z.Y.; Zhang, H.; Guan, C. Filter bank common spatial pattern (FBCSP) in brain–computer interface. In Proceedings of the 2008 IEEE International Joint Conference on Neural Networks (IEEE World Congress on Computational Intelligence); IEEE: New York, NY, USA, 2008; pp. 2390–2397. [Google Scholar]
- Pei, Y.; Luo, Z.; Zhao, H.; Xu, D.; Li, W.; Yan, Y.; Yan, H.; Xie, L.; Xu, M.; Yin, E. A tensor-based frequency features combination method for brain–computer interfaces. IEEE Trans. Neural Syst. Rehabil. Eng. 2021, 30, 465–475. [Google Scholar] [CrossRef]
- Pei, Y.; Luo, Z.; Yan, Y.; Yan, H.; Jiang, J.; Li, W.; Xie, L.; Yin, E. Data augmentation: Using channel-level recombination to improve classification performance for motor imagery EEG. Front. Hum. Neurosci. 2021, 15, 645952. [Google Scholar] [CrossRef] [PubMed]
- Schirrmeister, R.T.; Springenberg, J.T.; Fiederer, L.D.J.; Glasstetter, M.; Eggensperger, K.; Tangermann, M.; Hutter, F.; Burgard, W.; Ball, T. Deep learning with convolutional neural networks for EEG decoding and visualization. Hum. Brain Mapp. 2017, 38, 5391–5420. [Google Scholar] [CrossRef]
- Cheng, X.; Han, P.; Li, G.; Chen, S.; Zhang, H. A novel channel and temporal-wise attention in convolutional networks for multivariate time series classification. IEEE Access 2020, 8, 212247–212257. [Google Scholar] [CrossRef]
- Amiri, H.K.; Zarei, M.; Daliri, M.R. Motor imagery electroencephalography channel selection based on deep learning: A shallow convolutional neural network. Eng. Appl. Artif. Intell. 2024, 136, 108879. [Google Scholar] [CrossRef]
- Nguyen, A.H.P.; Oyefisayo, O.; Pfeffer, M.A.; Ling, S.H. EEG-TCNTransformer: A Temporal Convolutional Transformer for Motor Imagery Brain–Computer Interfaces. Signals 2024, 5, 605–632. [Google Scholar] [CrossRef]
- Liao, W.; Wang, W. Eegencoder: Advancing bci with transformer-based motor imagery classification. arXiv 2024, arXiv:2404.14869. [Google Scholar] [CrossRef]
- Jiang, Z.; Chen, Y. The Improved Transformer Network Based on Temporal and Spatial Convolution. In Proceedings of the 2025 6th International Conference on Computer Vision, Image and Deep Learning (CVIDL); IEEE: New York, NY, USA, 2025; pp. 678–681. [Google Scholar]
- Liu, D.; Chen, Z.; Luo, J.; Lian, S.; Wu, D. MIRepNet: A Pipeline and Foundation Model for EEG-Based Motor Imagery Classification. arXiv 2025, arXiv:2507.20254. [Google Scholar]
- Wang, Z.; Li, S.; Chen, X.; Li, W.; Wu, D. Multi-View Contrastive Network (MVCNet) for Motor Imagery Classification. arXiv 2025, arXiv:2502.17482. [Google Scholar] [CrossRef]
- Chen, C.S.; Chen, Y.J.; Tsai, A.H.W. Large cognition model: Towards pretrained eeg foundation model. arXiv 2025, arXiv:2502.17464. [Google Scholar]
- Muna, U.M.; Shawon, M.M.H.; Jobayer, M.; Akter, S.; Sabuj, S.R. SSTAF: Spatial-Spectral-Temporal Attention Fusion Transformer for Motor Imagery Classification. arXiv 2025, arXiv:2504.13220. [Google Scholar]
- Hamidi, A.; Kiani, K. Motor Imagery EEG signals classification using a Transformer-GCN approach. Appl. Soft Comput. 2025, 170, 112686. [Google Scholar] [CrossRef]
- Kumar, S.; Alawieh, H.; Racz, F.S.; Fakhreddine, R.; Millán, J.d.R. Transfer learning promotes acquisition of individual BCI skills. PNAS Nexus 2024, 3, pgae076. [Google Scholar] [CrossRef]
- Ye, W.; Li, X.; Zhang, H.; Zhu, Z.; Li, D. Deep Spatio-Temporal Mutual Learning for EEG Emotion Recognition. In Proceedings of the 2022 International Joint Conference on Neural Networks (IJCNN); IEEE: New York, NY, USA, 2022; pp. 1–8. [Google Scholar]
- Fu, S.; Chen, J.; Chen, D.; He, C. CNNs/ViTs-CNNs/ViTs: Mutual distillation for unsupervised domain adaptation. Inf. Sci. 2023, 622, 83–97. [Google Scholar] [CrossRef]
- Yan, W.; Liu, J.; Zhang, Z.; Zhang, Q.; Li, Y. A Spectral-Temporal Refined Attention Network via Contrastive Mutual Learning for Closed-Loop Motor Imagery BCI. IEEE Trans. Comput. Soc. Syst. 2025. early access. [Google Scholar] [CrossRef]
- Pei, Y.; Zhao, S.; Xie, L.; Ji, B.; Luo, Z.; Ma, C.; Gao, K.; Wang, X.; Sheng, T.; Yan, Y.; et al. Toward the enhancement of affective brain–computer interfaces using dependence within EEG series. J. Neural Eng. 2025, 22, 026038. [Google Scholar] [CrossRef]
- Pei, Y.; Zhao, S.; Xie, L.; Luo, Z.; Zhou, D.; Ma, C.; Yan, Y.; Yin, E. Identifying stable EEG patterns in manipulation task for negative emotion recognition. IEEE Trans. Affect. Comput. 2025, 16, 2033–2047. [Google Scholar] [CrossRef]
- Gui, Y.; Chen, M.; Su, Y.; Luo, G.; Yang, Y. EEGMamba: Bidirectional state space model with mixture of experts for EEG multi-task classification. arXiv 2024, arXiv:2407.20254. [Google Scholar]
- Deng, X.; Huo, H.; Ai, L.; Xu, D.; Li, C. A Novel 3D Approach with a CNN and Swin Transformer for Decoding EEG-Based Motor Imagery Classification. Sensors 2025, 25, 2922. [Google Scholar] [CrossRef]
- Mirzaei, S.; Ghasemi, P.; Bakhtyari, M. EEG Motor imagery classification based on a ConvLSTM Autoencoder framework augmented by attention BiLSTM. Multimed. Tools Appl. 2025, 84, 20545–20566. [Google Scholar] [CrossRef]
- Song, Y.; Zheng, Q.; Liu, B.; Gao, X. EEG conformer: Convolutional transformer for EEG decoding and visualization. IEEE Trans. Neural Syst. Rehabil. Eng. 2022, 31, 710–719. [Google Scholar] [CrossRef]
- Kim, J.W.; Alaa, A.; Bernardo, D. EEG-GPT: Exploring capabilities of large language models for EEG classification and interpretation. arXiv 2024, arXiv:2401.18006. [Google Scholar] [CrossRef]
- Li, M.A.; Han, J.F.; Duan, L.J. A novel MI-EEG imaging with the location information of electrodes. IEEE Access 2019, 8, 3197–3211. [Google Scholar] [CrossRef]
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |