1. Introduction
In recent years, unprecedented advances in medical imaging have significantly transformed cancer diagnosis. Techniques such as magnetic resonance imaging (MRI), computed tomography (CT), and positron emission tomography (PET) provide detailed spatial and functional information, making them crucial for the detection, characterization, and monitoring of tumors, as well as a guide for therapeutic planning [
1]. Advances in diagnostic technologies require that image processing not be limited to isolated technical solutions, rather that it be integrated into them. In this context, multimodal and multiomic integration is increasingly recognized as a core paradigm in modern biomedical research [
2], such as the integration of pathology and radiology images with genomic and clinical data [
3], as it enables a more comprehensive characterization of complex phenotypes by integrating complementary signals across different scales (the molecular and cellular, and at the tissue level). Moreover, this transition to quantitative biomedical imaging enables us to leverage the ability of deep learning models to extract morphological details, textures, and complex patterns that often exceed the visual perception of human experts, enabling more objective clinical decision-making [
4,
5].
Currently, histopathology remains the gold standard for tumor diagnosis. Pathologists can identify and categorize neoplastic lesions with high specificity by looking at cell morphology and tissue architecture [
6,
7]. However, histopathological assessment is still largely manual and time-consuming, as well as being heavily dependent on the training and expertise of each pathologist. In addition, it requires a series of molecular markers that are quite time-consuming and significantly increase the cost of diagnosis and routine pathology services. This complicates efforts to scale or standardize diagnostics, especially in high-volume or resource-limited settings. In response to these challenges, deep learning (DL)-based tools have been applied for assisted image diagnosis, enabling faster and more reproducible analysis of whole-slide image (WSI) captures [
8] while maintaining high diagnostic performance, as shown in recent GBM studies [
9,
10].
Although all these advances in DL-based pathological diagnostic tools are crucial, they are all based on information from standard red, green, and blue (RGB) cameras, which limits their ability to capture biochemical tissue properties beyond what the professional eye can see. In this context, histopathology can benefit substantially from the application of hyperspectral imaging (HSI). This technology preserves one of the fundamental pillars of histopathology, which is the morphological analysis of tissues through spatial information, while simultaneously enabling the investigation of the tissue’s other spectral information. Moreover, optical microscopy environments provide nearly ideal acquisition conditions for hyperspectral imaging due to controlled illumination and minimal external light interference. The data acquired by this type of imagery are called hyperspectral cubes, and consist of spatial images with an additional spectral dimension spanning multiple contiguous wavelength bands. Building upon these technical capabilities, HSI has gained increasing attention. Variations in spectral signatures can reveal subtle biochemical and metabolic alterations reflecting changes in the tissue [
11]. An alteration in tissue that leads to cancer often affects the morphology of tumor cells depending on proliferation, tumor microenvironment resources, and specific metabolic dispositions [
12]. For instance, HSI has been applied to quantitatively measure oxygenation levels in tissues being tested for various pathologies [
13]. These findings highlight the ability of HSI to detect subtle biochemical changes beyond morphological changes at the cellular level, reinforcing its potential as a powerful diagnostic tool.
Indeed, hyperspectral imaging has been increasingly explored in biomedical applications. For instance, Karim et al. [
14] provide a comprehensive review of medical fields in which HSI is currently being applied, while Mangotra et al. [
15] present an extensive compilation of preprocessing techniques, evaluation metrics, and clinical applications of HSI for early disease diagnosis. Within the context of conventional histopathology based on hematoxylin and eosin (H&E) staining, recent studies have demonstrated the feasibility and reliability of HSI for histological analysis. Yan et al. [
16], for example, highlight the applicability of HSI to lung cancer specimens and propose labeling strategies aimed at reducing the annotation burden of pathologists. Jong et al. [
17] investigate hyperspectral analysis of surgical margins in breast cancer resections using H&E-stained samples and spectral unmixing techniques. Alternative hyperspectral modalities have also been explored, such as in the work by Muniz et al. [
18], which analyzes healthy and tumor tissues using Fourier transform infrared hyperspectral imaging (FTIR-HSI). In this context, hyperspectral microscopic imaging (HMI) is one of the most promising tools assisting pathologists in the accurate detection of abnormalities in the tissues analyzed.
The aim of this paper is to establish a reliable, repeatable, and effective methodology that allows for the generation of hyperspectral databases of cell nuclei regardless of the tissue analyzed, in order to enable subsequent analyses based on spectral properties. In histopathology, the cell nucleus plays a central role in disease diagnosis and grading as nuclear morphology reflects fundamental biological processes such as chromatin organization, cell cycle progression, proliferation, and genetic instability. Indeed, many diagnostic criteria routinely used by pathologists, such as nuclear size, shape, density, pleomorphism, and chromatin texture, are primarily derived from nuclear features rather than from cytoplasmic or stromal components. In order to extract spectral information solely from cell nuclei, we have proposed three different segmentation methodologies that were compared by evaluating both pixel-wise segmentation accuracy and instance-level nucleus separation. We have also analyzed the practical implications of each method for downstream spectral analysis.
To the best of the authors’ knowledge, the systematic evaluation of nuclei segmentation strategies for hyperspectral histopathological images, specifically in the context of reliable nuclear spectral database generation, has not yet been thoroughly explored. Existing works have addressed related problems from different perspectives but fall short of this specific goal. For instance, Sultana et al. [
19] propose a U-Net-based pipeline for cytoplasm segmentation, emphasizing hyperspectral-specific data augmentation strategies that simulate instrumental noise from raw CMOS sensor data to improve robustness. Ma et al. [
20] introduce a patch-based classification framework in which hyperspectral cubes are reduced using principal component analysis (PCA) and subsequently classified using neural networks, demonstrating the relevance of hyperspectral information for tissue diagnosis; however, nuclear regions are not explicitly segmented, limiting the correct extraction of nucleus-specific spectral signatures. Chen et al. [
21] present a segmentation approach based on binarization of a specific spectral band exploiting chromatin distribution, followed by cell detection using support vector machines (SVMs), but this method is highly dependent on acquisition conditions and staining consistency, restricting its generalizability. Finally, Yun et al. [
22] propose SpecTr, a U-Net-based segmentation framework that employs transformers to filter redundant or noisy spectral bands; nevertheless, its reported segmentation performance remains limited. Collectively, these studies highlight the growing interest in hyperspectral histopathology while underscoring the lack of a comprehensive and comparative analysis focused on robust nucleus segmentation for hyperspectral signature extraction.
Therefore, the main contributions of this research work are as follows:
The development of a complete methodology for the generation and evaluation of hyperspectral databases of cell nuclei from histopathological samples.
The proposal and evaluation of three nucleus segmentation strategies: a purely spectral approach, a purely spatial approach, and a combined spatial–spectral approach.
A systematic comparison of the proposed methods using pixel-wise segmentation metrics, instance-level cell count analysis against ground truth, and qualitative evaluation of the resulting segmentation maps.
The identification of an optimal pipeline for hyperspectral nucleus database generation based on a 3D U-Net architecture, achieving a Dice similarity coefficient (DSC) of 73.13% for nucleus segmentation and a cell count deviation of approximately 4% relative to ground truth, indicating robust instance-level nucleus separation.
4. Discussion
Hyperspectral imaging is a powerful technology for clinical histology, as it enables spatial analysis comparable to conventional RGB imaging while substantially extending the available information by capturing a certain range of the electromagnetic spectrum. To perform reliable spectral analysis of pathological samples, it is essential to generate databases of spectral signatures specifically associated with cell nuclei, since the background in histological images is typically highly heterogeneous and spectrally noisy. Nuclear morphology is also key in the diagnosis of tumor cells and can enable standardization of diagnosis. It also tends to show small changes that are not observable to the pathologist’s eye, as they are associated with structural loss, changes in DNA quantity, or even chromatin condensation. For all these reasons, this work presents a comprehensive methodology for constructing hyperspectral databases of cell nuclei from histopathological samples.
The primary objective of the proposed methodology is to accurately extract meaningful spectral information from raw hyperspectral data cubes. To this end,
Section 2.1.2 and
Section 2.1.3 describe the hyperspectral acquisition system and its validated preprocessing pipeline. Once the hyperspectral cubes are obtained, the critical step becomes the accurate segmentation of cell nuclei. Accordingly, three segmentation strategies are investigated: a spectral-only approach that relies exclusively on pixel-wise spectral information (
Section 2.2.1), a spatial-only approach based on features extracted from synthetic RGB images derived from the hyperspectral cubes (
Section 2.2.2), and a spatial–spectral approach that jointly exploits spatial and spectral information (
Section 2.2.3). Regardless of the segmentation method chosen, each one produces produces a nuclei segmentation map, from which individual cells are identified by grouping spatially adjacent nuclear pixels. In the final step of the proposed pipeline, the average spectral signature of each segmented nucleus is computed and automatically labeled using the available clinical information, resulting in a structured and clinically meaningful hyperspectral database.
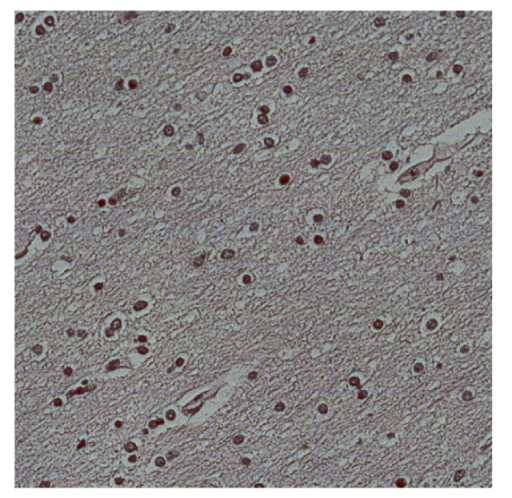
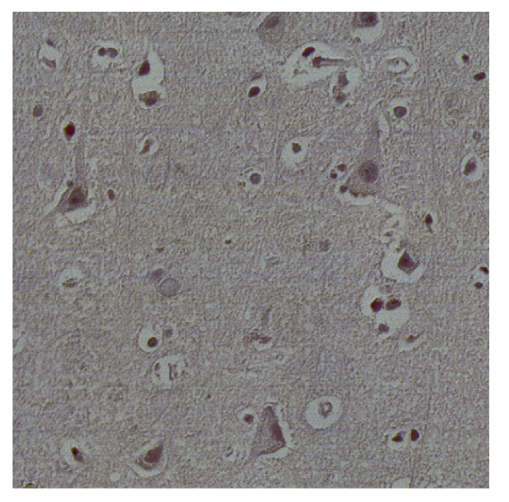
To evaluate the proposed segmentation strategies, a proprietary hyperspectral dataset of brain histopathological samples comprising both tumor and healthy tissue was constructed. The dataset includes 30 hyperspectral image cubes (10 tumor and 20 healthy), which were split into 20 cubes for training, four for validation, and six for testing (see
Table 3). Three expert pathologists independently annotated the dataset, providing the ground truth segmentation masks. Each method was trained by optimizing the DSC for the nuclei class, and performance was evaluated on the test set. Quantitative segmentation metrics and qualitative visual results are reported in
Section 3.1 and
Section 3.3, respectively. In addition, an instance-level analysis was conducted by comparing the number of segmented nuclei with the ground truth cell counts, as presented in
Section 3.2.
Together, these results provide a comprehensive characterization of the strengths and limitations of each segmentation approach. The spectral-only method (Method 1) exhibits highly noisy behavior across both healthy and tumor tissues, with particularly extreme deviations in healthy samples. As shown in
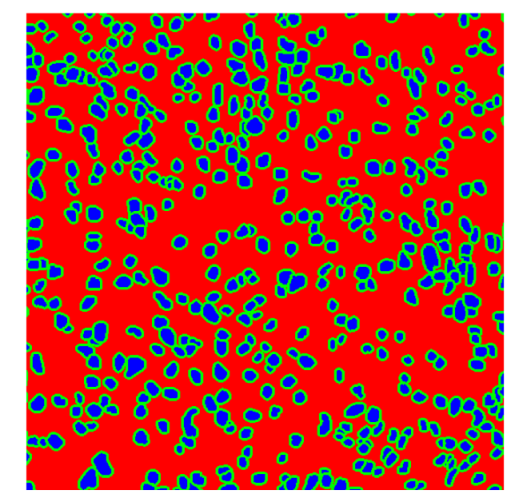
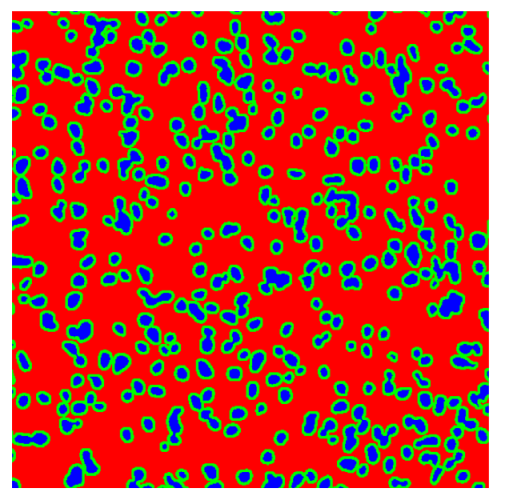
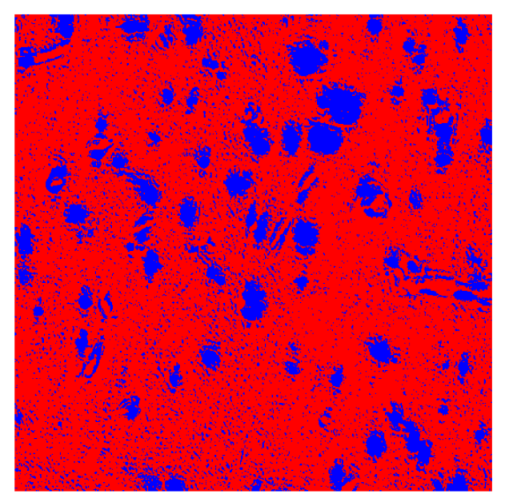
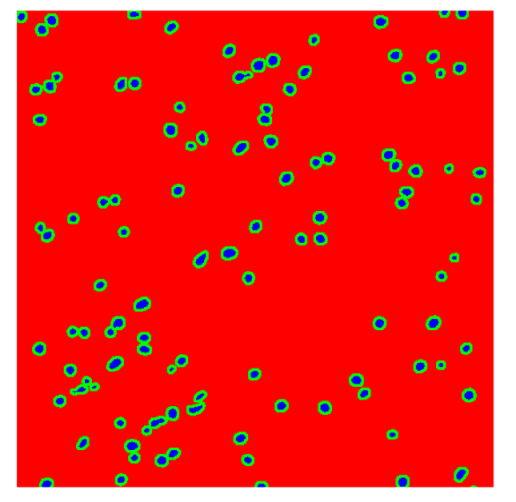
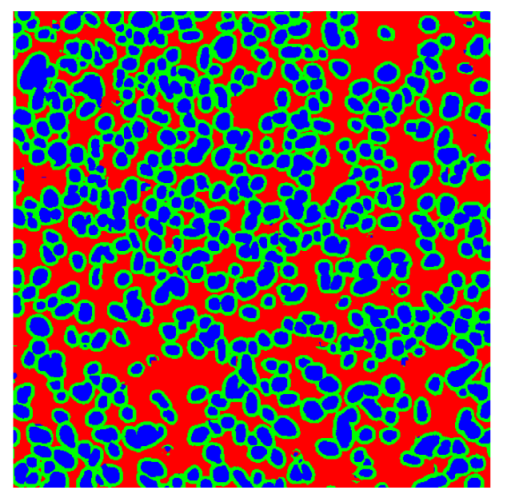
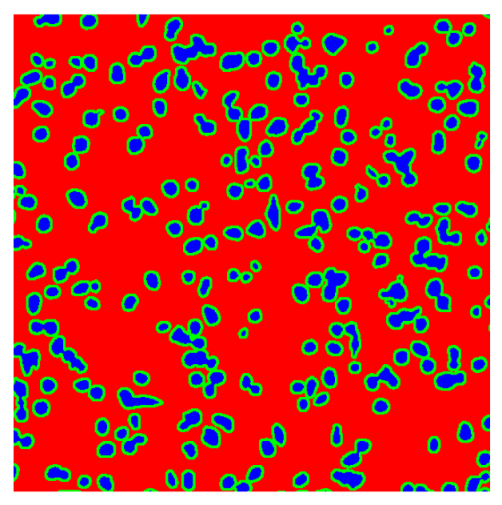
Table 8, the number of detected nuclei in healthy cubes exceeds the ground truth by several orders of magnitude (up to 7702%), indicating severe fragmentation of homogeneous nuclear regions into multiple small disconnected components. This behavior is consistent with the qualitative segmentation maps shown in
Table 9, where it can be observed how there is a very significant number of pixels marked as background that are incorrectly predicted as core by the algorithm, especially in the healthy cubes (4, 5, and 6). Despite being the only method optimized using the Optuna framework to maximize the DSC of the nucleus class in the validation set, this approach yields the poorest overall performance. In particular, it achieves a DSC of only 61.89% for the nucleus class (
Table 7). For this method to be viable in a production setting, a robust post-processing pipeline would be required, primarily focused on aggressive denoising of the predicted segmentation maps. Even with such corrections, the limited ability of the method to accurately separate individual nuclei would likely remain a critical bottleneck.
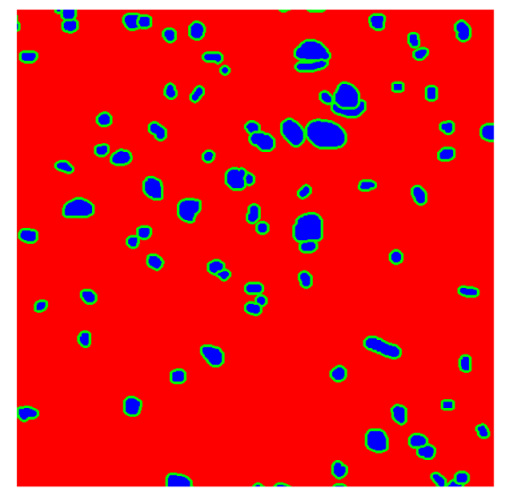
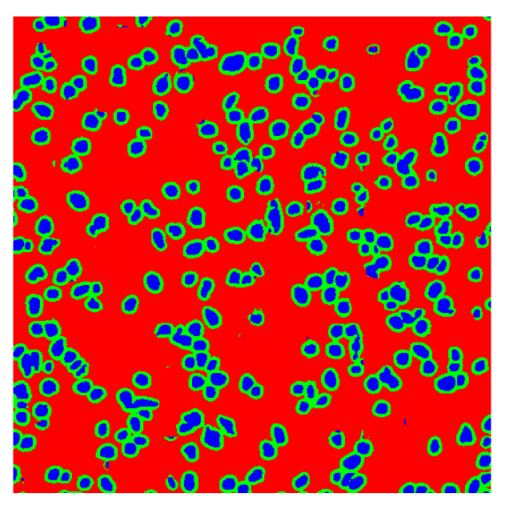
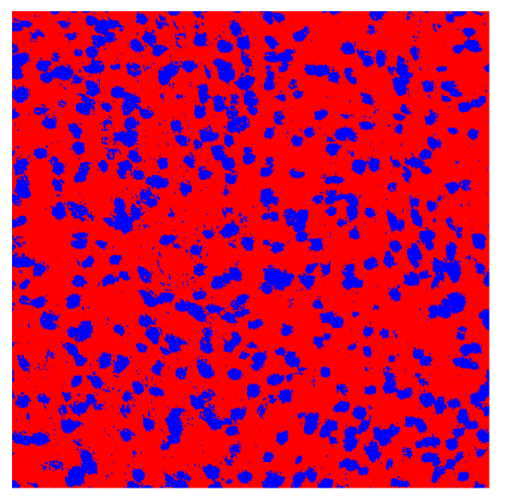
In contrast, the spatial-only (Method 2) and spatial–spectral (Method 3) approaches benefit from explicitly modeling the spatial context. Both methods incorporate an auxiliary nuclei border class, which enables the neural networks to learn the physical contours of nuclei and improves their separation during the spectral signature extraction stage. Method 2 achieves the highest pixel-wise segmentation performance, with DSC values of 78.97% for the nucleus class and 94.32% for the background. However, the cell count analysis reveals a substantial limitation: this method underestimates the number of nuclei by approximately 30% in tumor tissue, which typically exhibits a higher cell density than healthy tissue. Inspection of the corresponding segmentation maps indicates that this behavior arises from the tendency of the model to merge adjacent nuclei, thereby reducing its reliability for instance-level analysis and morphological studies. By comparison, Method 3 achieves slightly lower pixel-wise performance, with DSC values of 73.13% for nuclei and 93.16% for background, but demonstrates significantly greater robustness in correctly separating individual cell nuclei. Its average deviation from the ground truth cell count is approximately 4%, indicating a much closer correspondence with expert annotations. This improved instance-level accuracy is further corroborated by qualitative comparisons, where the segmentation maps produced by Method 3 closely resemble the ground truth delineations and avoid the large nuclear clusters observed with Method 2.
From a broader biomedical imaging perspective, these findings reinforce the principle that performance metrics must be interpreted in the context of their downstream analytical implications. Systematic reviews in quantitative biomedical imaging have emphasized that preprocessing and segmentation stages fundamentally shape the robustness, reproducibility, and translational value of derived biomarkers and structured databases [
39,
40]. Furthermore, Ghai et al. [
41] reinforces the idea that the use of standardized metrics not only improves reproducibility but also has the potential to optimize patient care by facilitating more accurate comparisons between different domains and populations in medical research. Inadequate instance-level delineation may introduce systematic bias in feature extraction, compromise statistical validity, and limit cross-cohort comparability. Therefore, segmentation strategies should be evaluated not only according to conventional pixel-wise metrics but also based on their capacity to preserve biologically and clinically meaningful structures, particularly when the ultimate objective is database construction or biomarker development.