3.4. Results
3.4.1. Results of Main Task: SAD Identification
The average performance metrics derived from five-fold cross-validation were adopted as the final evaluation results, as detailed in
Table 2. The proposed model achieved an overall accuracy of 0.8139 on the current dataset. Notably, the model demonstrated a specificity of 0.8724, indicating a strong capability to exclude non-SAD cases and suggesting potential utility in clinical screening scenarios. Furthermore, the precision score of 0.8186 reflects the reliability of positive predictions, while the sensitivity of 0.7340 and F1 score of 0.7687 indicate a favorable balance between sensitivity and precision.
To validate the effectiveness of the proposed dual-domain complementary deep learning framework, comparative experiments were conducted against classic machine learning algorithms and various mainstream deep learning baselines. These comparative experiments were conducted under identical settings, and the quantitative results are detailed in
Table 3.
Firstly, the XGBoost model, representing traditional machine learning, exhibited limited performance, recording the lowest accuracy of 0.6926 and sensitivity of 0.4834. This limitation may be attributable to the reliance on static and discrete IOS-derived parameters, which may be insufficient to fully capture the dynamic temporal characteristics of airway impedance during respiration.
Secondly, we analyzed the impact of network depth on performance using the ResNet series. The results revealed a non-monotonic trend. The shallow ResNet10 network showed signs of underfitting, which was likely due to constrained feature extraction capabilities. Conversely, although ResNet34 possesses a larger theoretical receptive field, it exhibited performance degradation. This decline appeared to be caused by overfitting given the current dataset size, suggesting that excessively deep architectures may not be optimal for this specific task. Among the series, ResNet18 achieved the best balance between parameter quantity and feature extraction, reaching an accuracy of 0.7291.
Overall, the proposed model outperformed the aforementioned baselines across all evaluated metrics. Notably, our method achieved performance gains even when compared to the Transformer, which was the strongest baseline. This advantage may be attributable to the integration of TFRIM spectrograms with the dual-stream architecture. By preserving the time-domain details of the original signal while effectively leveraging dynamic frequency-domain texture features, our approach realized superior SAD identification capabilities.
We conducted a series of ablation studies to evaluate the specific contributions of different input branches to the SAD identification task and to validate the effectiveness of the multi-branch fusion strategy. These experiments covered various combinations, ranging from single-branch inputs to joint multi-branch inputs. Detailed performance metrics are presented in
Table 4.
First, in the single-branch setting, utilizing only the raw time-series signal or the TFRIM yielded accuracies of 0.7430 and 0.7490, respectively. These results suggest that each individual branch contains certain discriminative patterns. However, single-branch feature extraction appears insufficient to fully capture the complex pathophysiological changes associated with SAD. Furthermore, relying solely on demographic features resulted in performance only slightly above chance level. This implies that demographic features do not directly reflect airway pathological features. Instead, they likely serve as auxiliary calibration information rather than a primary identification basis.
Upon introducing a second input branch, performance improvements were consistently observed. Specifically, combining the raw time-series signal with the TFRIM increased accuracy. This improvement indicates potential complementarity between time-domain respiratory dynamics and time–frequency impedance spectral texture features. Consequently, their combination provides a more comprehensive characterization of airway function. Additionally, incorporating demographic features into either the time-series or TFRIM inputs yielded a performance gain of approximately 2%. This finding supports the efficacy of the proposed DAFM module. It suggests that integrating demographic information reduces the confounding effects of inter-individual variability on SAD identification.
This indicates that the three input branches likely form a synergistic interaction within the feature space. In this synergy, the raw signal provides respiratory dynamics, the TFRIM contributes time–frequency texture information, and demographic features offer global calibration. Such a fusion strategy appears instrumental in enhancing the accuracy of SAD identification.
To evaluate the effectiveness of the proposed DAFM fusion scheme, we compared it with three classical feature fusion strategies. The experimental results are detailed in
Table 5. As a baseline, the concatenation fusion strategy directly joins demographic features with the feature representation
. Although this approach introduces auxiliary information, it relies on simple dimensional stacking. Consequently, it may fail to explicitly capture the specific influence of demographic factors on airway impedance.
The gating fusion strategy performed slightly worse than the concatenation approach. This phenomenon suggests that using only a multiplicative gating mechanism to adjust feature amplitude may be insufficient. Mathematically, it might not effectively characterize the additive shift effect that demographic features exert on the respiratory impedance baseline.
In contrast, the additive bias calibration fusion strategy achieved higher accuracy than both the concatenation and gating methods. This result supports the hypothesis that demographic features primarily serve a baseline correction role in physiological signal processing. It appears that explicit bias calibration effectively guides the model to adapt to distributional differences among individuals.
Finally, the proposed DAFM module demonstrated the most favorable performance among the evaluated methods. This module treats demographic features not as isolated information fragments but as active calibration signals. By mapping these features to adaptive modulation parameters, the DAFM module achieves dynamic, sample-level modulation of airway-impedance features. Functionally, this mechanism can be viewed as analogous to the clinical process where physicians adjust decision thresholds based on patient physique. Therefore, this approach can reduce the confounding effects of inter-individual physiological variability on SAD identification.
3.4.2. Results of Subtasks
To further analyze the performance characteristics of the model, we evaluated its ability to identify abnormalities in three key small airway function indices. These indices include FEF25–75, FEF50, and FEF75.
Table 6 quantifies the model’s performance across individual subtasks. The final decision is obtained by combining subtask predictions via the voting mechanism. In this setting, the subtasks exhibit complementary characteristics across metrics, potentially benefiting overall identification performance. Specifically, the model exhibited high sensitivity (0.8446) in the task of identifying FEF
75 abnormalities. This suggests that the network is capable of capturing feature patterns associated with distal airway flow limitations. Consequently, this capability helps reduce the rate of missed detections for potentially abnormal samples. Conversely, the model demonstrated high specificity (0.9079) when identifying FEF
50 abnormalities. This implies that the model adopts a more conservative approach in assessing mid-airway function. Therefore, it exhibits high reliability in excluding false positive cases.
These distinct, complementary performance characteristics support the rationale of the multi-label voting mechanism introduced in this study. This mechanism effectively integrates the capability of the FEF75 classification head to capture abnormal samples with the ability of the FEF50 classification head to suppress false alarms.
Through this decision-level fusion, the final SAD identification results achieved a favorable balance between sensitivity (0.7340) and specificity (0.8724). This strategy not only surpasses the average performance of individual metrics but also contributes to enhancing the robustness of the identification system in clinical application scenarios.
Figure 4 further presents the performance comparison between the proposed model and existing mainstream baselines on these proxy subtasks using a radar chart. As illustrated, the proposed model encloses a larger area across the evaluation dimensions compared to the baseline models. This observation suggests that our approach yields a more balanced overall performance profile.
Notably, the FEF75 index, which reflects distal airway function, typically presents a greater detection challenge. On this specific metric, both XGBoost and Transformer exhibited certain performance limitations. In contrast, the proposed architecture maintained relatively high recognition accuracy. This robustness is likely attributed to the rich time–frequency texture information provided by the TFRIM, as well as the adaptive calibration capability of the DAFM module. These results imply that the model possesses potential efficacy in capturing subtle airway impedance abnormalities.
3.5. Model Interpretability and Feature Visualization
To provide an intuitive analysis of the high-dimensional feature distribution structure learned by the model, we extracted feature vectors from the penultimate layer. Subsequently, we employed the t-SNE algorithm to project these vectors into a two-dimensional space, as illustrated in
Figure 5. In this visualization, each data point represents an individual subject.
To clearly depict the distributional trend ranging from normal to impaired small airway function, we defined a Small Airway Impairment Index, denoted as
S. This index integrates three soft labels into a single continuous variable. Specifically, drawing upon voting principles commonly used in clinical decision-making, we designed a differentiable fusion function to calculate
S:
This formulation is designed such that the
S value yields a higher response when two or more indicators fall below their respective thresholds. This characteristic maintains consistency with clinical decision-making rules. For visualization purposes, we mapped the color coding to the
S value, achieving a smooth visual transition from normal small airway function (represented by deep blue) to higher degrees of impairment (represented by deep red). An
S value approaching 1 indicates that the three indicators are simultaneously or largely below the threshold, suggesting a higher risk of small airway impairment. Conversely, an
S value approaching 0 implies that the indicators are generally above the threshold, suggesting normal airway function. It is crucial to emphasize that
S is an engineering-based continuous proxy constructed from spirometry thresholds. Its primary purpose is to facilitate visualization and feature structure analysis; therefore, it is not equivalent to a strict clinical severity grading system.
Observing the visualization of the feature space, the samples collectively exhibit a continuous band-like distribution pattern accompanied by a distinct color gradient. The deep blue cluster representing the healthy state and the deep red cluster representing the impaired state occupy opposite ends of the trajectory. Samples in the intermediate region demonstrate a gradual transition from healthy to transitional and then to impaired states, with slight overlapping. This distributional morphology presents a trend consistent with the definition of S. This phenomenon suggests that the feature representation likely encodes information regarding the deviation of small airway-related indicators from their thresholds.
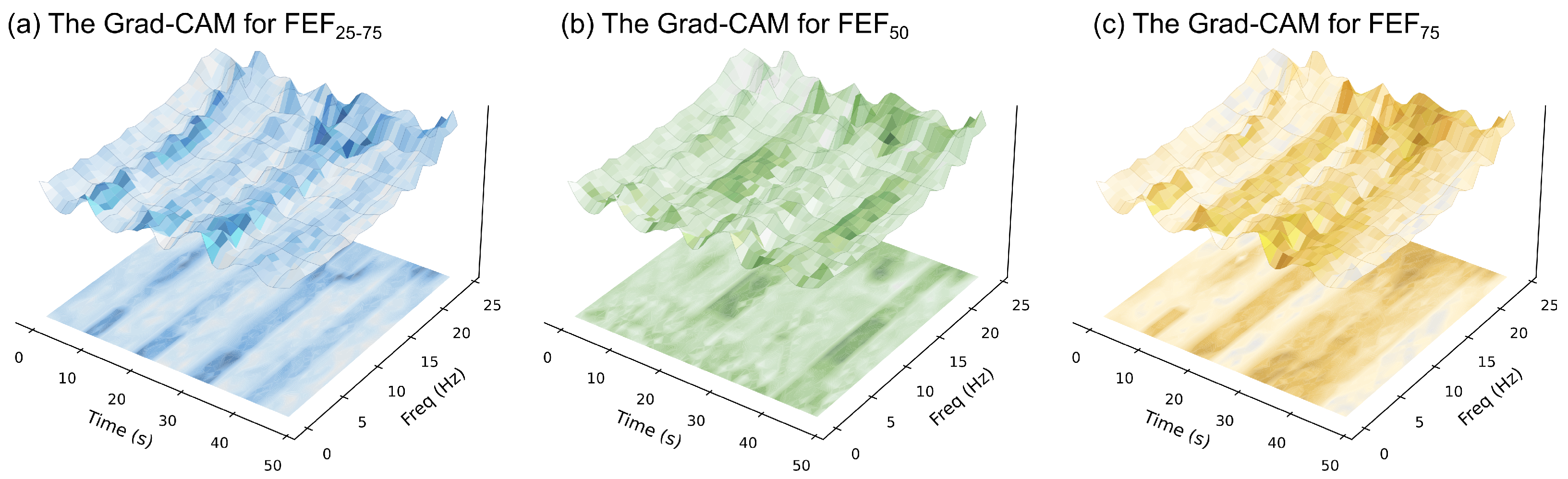
We employed Grad-CAM to analyze the time–frequency texture features from the TFRIM branch. This analysis aimed to interpret the features prioritized by the model.
Figure 6 presents a 3D fusion visualization. It intuitively maps the model’s focus onto the TFRIM. The
z-axis height represents the TFRIM amplitude. Meanwhile, the surface color indicates the distribution of visual attention. Darker colors correlate with a higher degree of model attention.
Although the subtasks target different indices, the model consistently exhibits respiratory phase selectivity. Specifically, the model concentrates on the expiratory phase window. This aligns with the pathophysiological mechanisms of SAD. Abnormal impedance fluctuations occur primarily during exhalation. These manifest as flow limitations [
46,
47]. The model appears to suppress inspiratory background signals and focus on end-expiratory dynamics. This suggests the network may capture transient pathological textures associated with airway collapse.
Furthermore, in the frequency dimension, highlighted regions are concentrated within specific bands. This indicates the network identifies respiratory phases and captures non-linear impedance characteristics. We further observed distinct attention patterns across different subtasks. This pattern suggests that the model may encode subtask-specific information related to airway function. For the FEF75 and FEF25–75 tasks (reflecting small airway function), attention focuses on the low-frequency band. This distribution is consistent with the physics of IOS. According to oscillation mechanics, only low-frequency waves penetrate deep into the lungs. Therefore, variations in this band may reflect the peripheral small airway state. In contrast, for the FEF50 task (reflecting mid-airway function), attention extends to a mixed low-to-mid frequency range.
This spectral differentiation is consistent with the frequency-dependent characteristics of IOS signals. The Grad-CAM results suggest that the discriminatory basis is not uniformly distributed. Instead, it exhibits differentiated attention to low-to-mid frequency information. This alignment with expected signal-sensitive bands supports the interpretability of the model’s performance.
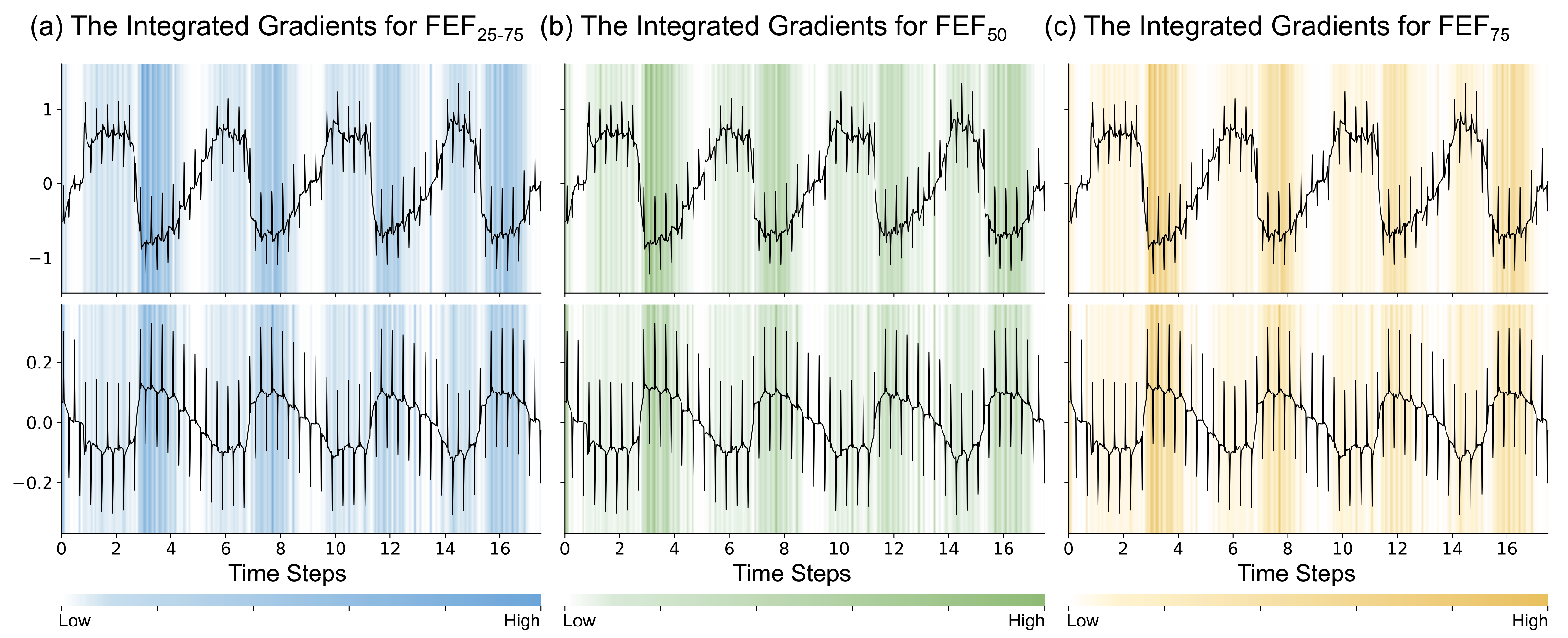
To further investigate the attribution focus of the time-series branch during feature extraction, we employed the Integrated Gradients algorithm to calculate the time-step attribution values of the time-series signals.
Figure 7 visualizes the attribution for raw IOS signals across three subtasks. The background color intensity represents the attribution of each signal point.
Observing the attribution distribution reveals patterns at both macro and micro levels. On a macro scale, high-attribution regions are synchronized with the respiratory cycle. Specifically, attribution values concentrate in the expiratory phase. In contrast, attribution values during the inspiratory phase are lower. This suggests that the model is sensitive to dynamic airway compression. It aligns with the expiratory flow limitation observed in SAD patients.
On a micro scale, the model exhibits selectivity for specific signal components. Input signals contain large-amplitude, low-frequency breathing waves. However, the model assigns low attribution values to these broad contours. Instead, high-frequency oscillatory impulses receive higher attribution values. As shown in
Figure 7, high attribution values cluster at the points where IOS pulses are applied. This implies the network utilizes external oscillatory signals for feature extraction. It does not merely memorize the breathing morphology.
To assess how the DAFM module utilizes demographic information for result calibration, we employed the SHapley Additive exPlanations (SHAP) method to calculate the marginal contribution of each demographic feature to the model’s output.
Figure 8 presents the SHAP summary plots for the three subtasks (identifying abnormalities in FEF
25–75, FEF
75, and FEF
50). In these plots, each row represents a feature, and each dot represents an individual data sample. The color of the dot indicates the magnitude of the feature value, while the horizontal position denotes the direction and intensity of the contribution to the prediction. A positive value indicates an increased probability of SAD identification, whereas a negative value implies the opposite.
The distribution of SHAP values reveals a consistent pattern of feature importance across different subtasks. Among all features, age consistently exhibits the highest importance, with its SHAP values spanning the widest range. Specifically, samples representing older age (indicated in red) are predominantly concentrated in the positive SHAP value region. This suggests that advancing age positively drives the model’s determination of airway dysfunction. This finding aligns with established knowledge in respiratory physiology, which notes that pulmonary function undergoes a physiological decline with age, thereby increasing the baseline probability of pathology.
Weight and height follow in importance, exhibiting substantial contributions to the model output across the subtasks. This reflects the corrective effect of body physique parameters on the feature distribution. In contrast, the marginal contribution of gender is relatively small, with its SHAP values converging near zero. Nevertheless, with males encoded as 1, a slight tendency toward SAD identification for male subjects is observable. This data preference offers interpretability from an epidemiological perspective. It may implicitly reflect the statistically higher risk of small airway dysfunction in the male population, potentially due to factors such as higher smoking rates or occupational exposure.
In summary, the SHAP analysis suggests that the DAFM module has acquired a calibration logic consistent with clinical priors. Specifically, the model appears to dynamically adjust decision boundaries based on the subject’s age and physique characteristics. This feature attribution pattern, which aligns with existing clinical theories, further supports the validity and interpretability of this branch as a personalized calibration unit within the multi-modal network.
The visualization analyses suggest internal consistency in the model’s decision rationale. Although the TFRIM and raw time-series branches operate on distinct representations, their attribution patterns are largely concordant. Specifically, both Grad-CAM and Integrated Gradients consistently assign higher relevance to the expiratory phase and low-frequency oscillatory components. This cross-method agreement, together with the continuous embedding gradient observed in the t-SNE projection and the demographic weighting patterns revealed by SHAP, provides convergent evidence that the model leverages physiologically plausible airway dynamics, thereby reducing the likelihood that the performance is dominated by background noise or spurious correlations.