Lean-NET-Based Local Brain Connectome Analysis for Autism Spectrum Disorder Classification
Abstract
1. Introduction
- A graph learning-based approach (Lean-NET) is employed to construct subject-specific functional brain networks.
- Analysis of localized graph-theoretical features to characterize region-specific connectivity alterations
- Statistical feature selection using Welch’s t-test with FDR correction
- Robust classification using linear SVM with leave-one-out cross-validation
2. Methodology
2.1. Datasets Description
2.2. Data Preprocessing
2.3. Graph Construction and Features Extraction on BOLD Signals Using Lean-NET
2.3.1. Connectome (Graph) Construction
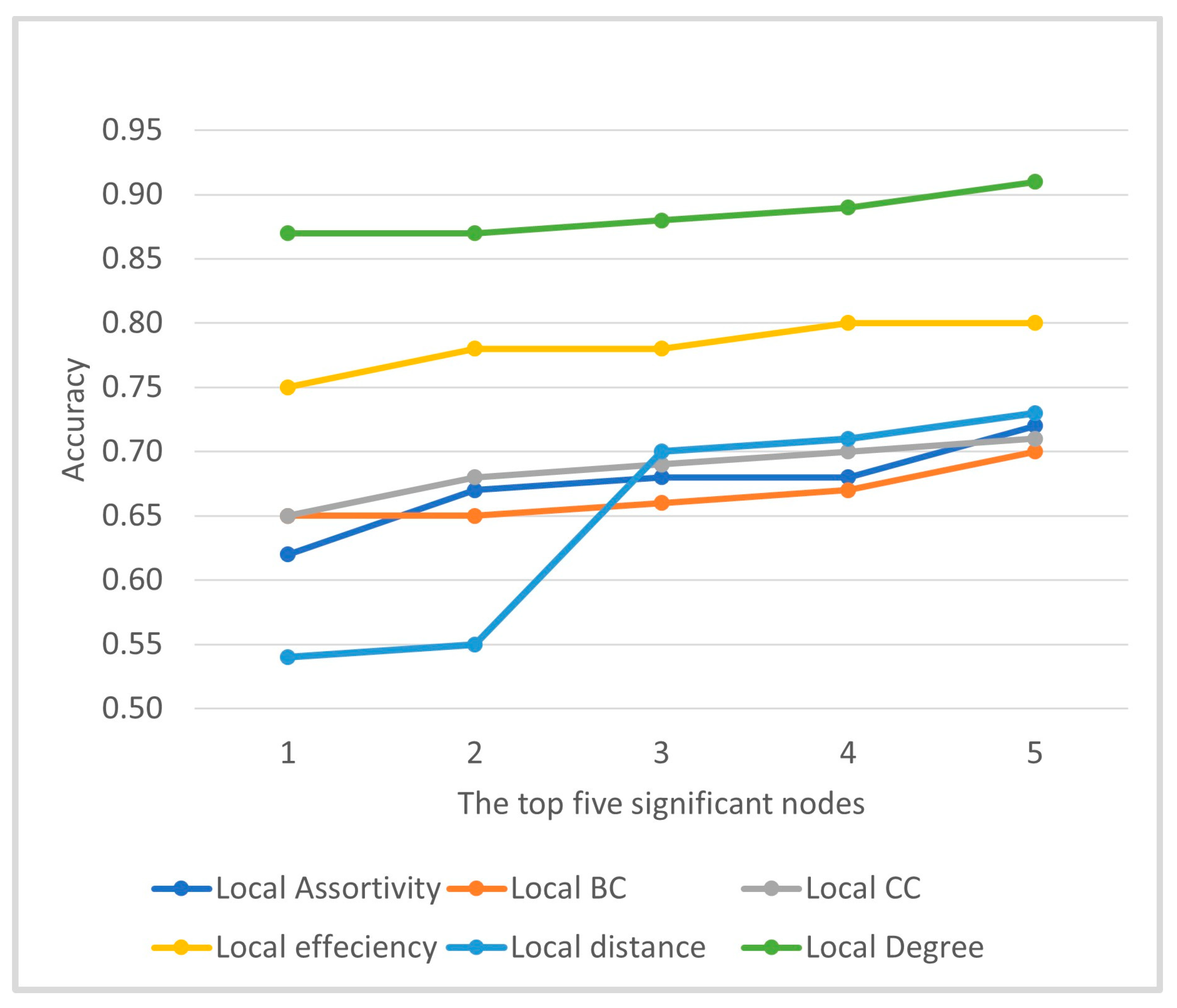
2.3.2. Node Level Graph Metrics
- 1.
- Local (Node) Assortativity (): evaluates the tendency of nodes to connect with other nodes that have similar connectivity degrees, indicating the network’s local structural organization, the local assortativity of node i is defined as [25]:where j is the node’s remaining degree, represent the average remaining degree of the neighbors, and . The is the global average excess degree.
- 2.
- Betweenness centrality (): is a measure of a node’s importance within the network, defined as the fraction of all shortest paths between any two other nodes in the network that pass-through node i. The shortest path lengths were calculated using the inverse of the weighted adjacency matrix W, as the edge lengths, and Bi is computed using an algorithm tailored for weighted graphs as shown in Equation (3) [26].where is the total number of shortest paths between node j and node k and is the number of those paths pass through i.
- 3.
- Clustering coefficient (): The local clustering coefficient measures how tightly connected a node’s neighbors are to each other [26]. A commonly used method for calculating local clustering coefficients is in Equation (4):where the represents then number of edges for node i, and is the degree of node i.
- 4.
- 5.
- Node Degree (): is the most fundamental measure of nodal connectivity and is defined as the number of connections incident to node , reflecting its level of topological integration within the network [26,27]. It is calculating using the binarized version of the adjacency matrix , where if an edge between nodes and is present and otherwise. For a graph G = (V, E) with N nodes, the degree of node i is given by
- 6.
- Local Efficiency (): quantifies the efficiency of information exchange among the immediate neighbors of node i, representing the local fault tolerance of the network [16,26]. The Efficiency is defined as the inverse of the average shortest path length between all pairs of nodes in the subgraph:where is the set of neighbors on node i, represents the shortest path length between the node j and h within the subgraph and is given in Equation (6). So the local is a key measure of functional segregation or modularity.
2.3.3. Statistical Testing and Leakage-Free Feature Selection
2.3.4. Classification
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- IHME. Global Burden of Disease (GBD) 2021; Global Burden of Disease Results Tool (Online Database); Institute for Health Metrics and Evaluation (IHME): Seattle, WA, USA, 2024; Available online: https://ghdx.healthdata.org/gbd-2021 (accessed on 16 July 2025).
- Di Martino, A.; Milham, M. Autism Spectrum Disorder Research and Clinical Program at the Child Study Center; New York University Langone Medical Center: New York, NY, USA, 2017. [Google Scholar]
- Craddock, R.C.; James, G.A.; Holtzheimer, P.E., III; Hu, X.P.; Mayberg, H.S. A whole-brain fMRI atlas generated via spatially constrained spectral clustering. Hum. Brain Mapp. 2012, 33, 1914–1928. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, A.; O’Connor, D.; Chen, B.; Alaerts, K.; Anderson, J.S.; Assaf, M.; Balsters, J.H.; Baxter, L.; Beggiato, A.; Bernaerts, S.; et al. Enhancing studies of the connectome in autism using the Autism Brain Imaging Data Exchange II. Sci. Data 2017, 4, 170010. [Google Scholar] [CrossRef]
- Liu, M.; Li, B.; Hu, D. Autism spectrum disorder studies using fMRI data and machine learning: A review. Front. Neurosci. 2021, 15, 697870. [Google Scholar] [CrossRef]
- Thomas, R.M.; Gallo, S.; Cerliani, L.; Zhutovsky, P.; El-Gazzar, A.; van Wingen, G. Classifying autism spectrum disorder using the temporal statistics of resting-state functional MRI data with 3D convolutional neural networks. Front. Psychiatry 2020, 11, 440. [Google Scholar] [CrossRef]
- Chu, Y.; Wang, G.; Cao, L.; Qiao, L.; Liu, M. Multi-scale graph representation learning for autism identification with functional MRI. Front. Neuroinform. 2021, 15, 802305. [Google Scholar] [CrossRef]
- Tikaram, T.; Raj, U.; Ratnaik, R.; Agastinose Ronickom, J.F. Advancing ASD diagnostic classification using time–frequency spectrograms of fMRI BOLD signals and machine learning. 2024; preprint. [Google Scholar] [CrossRef]
- Abraham, A.; Milham, M.P.; Di Martino, A.; Craddock, R.C.; Samaras, D.; Thirion, B.; Varoquaux, G. Deriving reproducible biomarkers from multi-site resting-state data: An autism-based example. NeuroImage 2017, 147, 736–745. [Google Scholar] [CrossRef] [PubMed]
- Sherkatghanad, Z.; Akhondzadeh, M.; Salari, S.; Zomorodi-Moghadam, M.; Abdar, M.; Acharya, U.R.; Khosrowabadi, R.; Salari, V. Automated detection of autism spectrum disorder using a convolutional neural network. Front. Neurosci. 2020, 13, 1325. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Schrader, P.T.; Zhang, N. A Deep Neural Network Study of the ABIDE Repository on Autism Spectrum Classification. Int. J. Adv. Comput. Sci. Appl. 2020, 11, 401–408. [Google Scholar] [CrossRef]
- Chib, A.S.; Malhotra, D.; Mengi, M. A machine learning approach for autism spectrum disorder detection using BOLD-fMRI signals and the ABIDE-II dataset. In Proceedings of the 2023 International Conference on Computer, Electronics & Electrical Engineering and Their Applications (IC2E3), Srinagar Garhwal, India, 8–9 June 2023; pp. 1–4. [Google Scholar] [CrossRef]
- Bazay, F.E.; Drissi El Maliani, A. Assessing the impact of preprocessing pipelines on fMRI-based autism spectrum disorder classification: ABIDE II results. In Engineering Applications of Neural Networks; Iliadis, L., Maglogiannis, I., Papaleonidas, A., Pimenidis, E., Jayne, C., Eds.; EANN 2024; Communications in Computer and Information Science; Springer: Cham, Switzerland, 2024; Volume 2141. [Google Scholar] [CrossRef]
- Mainas, F.; Golosio, B.; Retico, A.; Oliva, P. Exploring Autism Spectrum Disorder: A Comparative Study of Traditional Classifiers and Deep Learning Classifiers to Analyze Functional Connectivity Measures from a Multicenter Dataset. Appl. Sci. 2024, 14, 7632. [Google Scholar] [CrossRef]
- Dong, Y.; Batalle, D.; Deprez, M. A framework for comparison and interpretation of machine learning classifiers to predict autism on the ABIDE dataset. Hum. Brain Mapp. 2025, 46, e70190. [Google Scholar] [CrossRef]
- Dal, D.Y. Multimodal Statistical Analysis of Brain Connectome with Application to Alzheimer’s Disease; Department of Electrical and Electronics Engineering, Boğaziçi University: Istanbul, Turkey, 2024. [Google Scholar]
- Yang, S.; Yin, Z.; Ma, Y.; Wang, M.; Huang, S.; Zhang, L. M3ASD: Integrating Multi-Atlas and Multi-Center Data via Multi-View Low-Rank Graph Structure Learning for Autism Spectrum Disorder Diagnosis. Brain Sci. 2025, 15, 1136. [Google Scholar] [CrossRef]
- Dong, X.; Thanou, D.; Frossard, P.; Vandergheynst, P. Learning Laplacian Matrix in Smooth Graph Signal Representations. IEEE Trans. Signal Process. 2016, 64, 6160–6173. [Google Scholar] [CrossRef]
- Oztoprak, F.; Nocedal, J.; Rennie, S.; Olsen, P.A. Newton-like methods for sparse inverse covariance estimation. Adv. Neural Inf. Process. Syst. 2012, 7. [Google Scholar]
- Gamgam, G.; Yıldırım, Z.; Kabakçıoğlu, A.; Gürvit, H.; Demiralp, T.; Acar, B. Siamese graph convolutional network quantifies increasing structure–function discrepancy over the cognitive decline continuum. Comput. Methods Programs Biomed. 2024, 254, 108290. [Google Scholar] [CrossRef]
- Smith, S.M.; Miller, K.L.; Salimi-Khorshidi, G.; Webster, M.; Beckmann, C.F.; Nichols, T.E.; Ramsey, J.D.; Woolrich, M.W. Network modelling methods for fMRI. NeuroImage 2011, 54, 875–891. [Google Scholar] [CrossRef]
- Craddock, C.; Sikka, S.; Cheung, B.; Khanuja, R.; Ghosh, S.S.; Yan, C.; Li, Q.; Lurie, D.; Vogelstein, J.; Burns, R.; et al. Towards Automated Analysis of Connectomes: The Configurable Pipeline for the Analysis of Connectomes (C-PAC). Front Neuroinform 2013, 42. [Google Scholar] [CrossRef]
- Abraham, A.; Pedregosa, F.; Eickenberg, M.; Gervais, P.; Mueller, A.; Kossaifi, J.; Gramfort, A.; Thirion, B.; Varoquaux, G. Machine learningmfor neuroimaging with scikit-learn. Front. Neuroinform. 2014, 8, 14. [Google Scholar] [CrossRef] [PubMed]
- Kalofolias, V. How to learn a graph from smooth signals. In Proceedings of the 19th International Conference on Artificial Intelligence and Statistics (AISTATS), Cadiz, Spain, 9–11 May 2016; Volume 51, pp. 920–929. [Google Scholar]
- Thedchanamoorthy, G.; Piraveenan, M.; Kasthuriratna, D.; Senanayake, U. Node assortativity in complex networks: An alternative approach. Procedia Comput. Sci. 2014, 29, 2449–2461. [Google Scholar] [CrossRef]
- Rubinov, M.; Sporns, O. Complex network measures of brain connectivity: Uses and interpretations. NeuroImage 2010, 52, 1059–1069. [Google Scholar] [CrossRef]
- Wu, S.; Zhan, P.; Wang, G.; Yu, X.; Liu, H.; Wang, W. Changes of brain functional network in Alzheimer’s disease and frontotemporal dementia: A graph-theoretic analysis. BMC Neurosci. 2024, 25, 30. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, N.; Schrader, P. A study of brain networks for autism spectrum disorder classification using resting-state functional connectivity. Mach. Learn. Appl. 2022, 8, 100290. [Google Scholar] [CrossRef]
- Kolla, S.; Falakshahi, H.; Abrol, A.; Fu, Z.; Calhoun, V.D. Intra-Atlas Node Size Effects on Graph Metrics in fMRI Data: Implications for Alzheimer’s Disease and Cognitive Impairment. Sensors 2024, 24, 814. [Google Scholar] [CrossRef]
- Deng, Z.; Wang, S. Sex differentiation of brain structures in autism: Findings from a gray matter asymmetry study. Autism Res. 2021, 14, 1115–1126. [Google Scholar] [CrossRef]
- Curtis, D. Welch’s t test is more sensitive to real world violations of distributional assumptions than student’s t test but logistic regression is more robust than either. Stat. Pap. 2024, 65, 3981–3989. [Google Scholar] [CrossRef]
- Li, A.; Barber, R.F. Multiple Testing with the Structure-Adaptive Benjamini–Hochberg Algorithm. J. R. Stat. Soc. Ser. B Stat. Methodol. 2019, 81, 45–74. [Google Scholar] [CrossRef]
- Theng, D.; Bhoyar, K.K. Feature selection techniques for machine learning: A survey of more than two decades of research. Knowl. Inf. Syst. 2024, 66, 1575–1637. [Google Scholar] [CrossRef]
- Lu, C.; Shi, L.; Chen, Z.; Wu, C.; Wierman, A. Overcoming the curse of dimensionality in reinforcement learning through approximate factorization. arXiv 2024, arXiv:2411.07591. [Google Scholar] [CrossRef]
- Patle, A.; Chouhan, D.S. SVM kernel functions for classification. In Proceedings of the 2013 International Conference on Advances in Technology and Engineering (ICATE), Mumbai, India, 23–25 January 2013; pp. 1–9. [Google Scholar] [CrossRef]
- Yüksel Dal, D.; Yıldırım, Z.; Gürvit, H.; Kabakçıoglu, A.; Acar, B. Reorganization of brain connectivity across the spectrum of clinical cognitive decline. Neurol. Sci. 2024, 45, 5719–5730. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Chen, Y.; Zheng, H.; Zhang, B.; Wang, F.; Fang, J.; Li, Y.; Chen, Q.; Zhang, S. Changes in the topological organization of the default mode network in autism spectrum disorder. Brain Imaging Behav. 2021, 15, 1058–1067. [Google Scholar] [CrossRef] [PubMed]
- Gamgam, G.; Kabakcioglu, A.; Yüksel Dal, D.; Acar, B. Disentangled attention graph neural network for Alzheimer’s disease diagnosis. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), Marrakesh, Morocco, 6–10 October 2024; Springer: Cham, Switzerland, 2024; pp. 219–228. [Google Scholar]
- Ayub, R.; Sun, K.L.; Flores, R.E.; Lam, V.T.; Jo, B.; Saggar, M.; Fung, L.K. Thalamocortical connectivity is associated with autism symptoms in high-functioning adults with autism and typically developing adults. Transl. Psychiatry 2021, 11, 93. [Google Scholar] [CrossRef]
- Woodward, N.D.; Giraldo-Chica, M.; Rogers, B.; Cascio, C.J. Thalamocortical dysconnectivity in autism spectrum disorder: An analysis of the autism brain imaging data exchange. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 2017, 2, 76–84. [Google Scholar] [CrossRef]
- Bhamidimarri, P.M.; Alhosani, K.; Cai, H.; Al-Ali, H.; Abukhaled, Y.M.; Tawamie, H.; Abdelaziz, S.; Fawaz, M.; Kashir, J.; Sajjad, Y.; et al. Review on the role of hippocampus in autism spectrum disorder: Recent insights into neuropathology, genetics, and emerging therapeutic strategies. Neurobiol. Dis. 2026, 218, 107227. [Google Scholar] [CrossRef]
- Ong, L.T.; Fan, S.W.D. Morphological and functional changes of the cerebral cortex in autism spectrum disorder. Innov. Clin. Neurosci. 2023, 20, 40–47. [Google Scholar] [PubMed]
- Barreto, C.; Curtin, A.; Topoglu, Y.; Day-Watkins, J.; Garvin, B.; Foster, G.; Ormanoglu, Z.; Sheridan, E.; Connell, J.; Bennett, D.; et al. Prefrontal Cortex Responses to Social Video Stimuli in Young Children with and without autism spectrum disorder. Brain Sci. 2024, 14, 503. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Chen, H.; Wang, H.; Wang, Z. Neural correlates of facial recognition deficits in autism spectrum disorder: A comprehensive review. Front. Psychiatry 2024, 15, 1464142. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, R.-I.; Okada, R.; Aoki, R.; Nakamura, M.; Ohta, H.; Itahashi, T. Functional alterations of lateral temporal cortex for processing voice prosody in adults with autism spectrum disorder. Cereb. Cortex 2024, 34, bhae363. [Google Scholar] [CrossRef]


| ASD (n = 74) | TD (n = 98) | p_Value | |
| Age | 14.7 ± 7.0 | 15.1 ± 6.0 | 0.678 |
| Gender (Male/Female) | 64/10 | 72/26 | 0.572 |
| Full scale IQ | 107.9 ± 16.6 | 113.2 ± 13.1 | 0.045 |
| Metric | Accuracy | Sensitivity | Specificity | Precision | F1-Score |
|---|---|---|---|---|---|
| Assortivity | 0.72 | 0.61 | 0.79 | 0.65 | 0.62 |
| Betweenness centrality | 0.70 | 0.68 | 0.80 | 0.70 | 0.64 |
| Degree | 0.91 | 0.91 | 0.91 | 0.90 | 0.89 |
| Clustering Coefficient | 0.71 | 0.67 | 0.74 | 0.66 | 0.67 |
| Distance | 0.73 | 0.74 | 0.72 | 0.67 | 0.70 |
| Efficiency | 0.80 | 0.68 | 0.80 | 0.83 | 0.75 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chelef, A.; Yuksel Dal, D.; Ozturk, M.; Yousif, M.A.A.; Koc, G. Lean-NET-Based Local Brain Connectome Analysis for Autism Spectrum Disorder Classification. Bioengineering 2026, 13, 99. https://doi.org/10.3390/bioengineering13010099
Chelef A, Yuksel Dal D, Ozturk M, Yousif MAA, Koc G. Lean-NET-Based Local Brain Connectome Analysis for Autism Spectrum Disorder Classification. Bioengineering. 2026; 13(1):99. https://doi.org/10.3390/bioengineering13010099
Chicago/Turabian StyleChelef, Aoumria, Demet Yuksel Dal, Mahmut Ozturk, Mosab A. A. Yousif, and Gokce Koc. 2026. "Lean-NET-Based Local Brain Connectome Analysis for Autism Spectrum Disorder Classification" Bioengineering 13, no. 1: 99. https://doi.org/10.3390/bioengineering13010099
APA StyleChelef, A., Yuksel Dal, D., Ozturk, M., Yousif, M. A. A., & Koc, G. (2026). Lean-NET-Based Local Brain Connectome Analysis for Autism Spectrum Disorder Classification. Bioengineering, 13(1), 99. https://doi.org/10.3390/bioengineering13010099






