Synthetic Inflammation Imaging with PatchGAN Deep Learning Networks
Abstract
1. Introduction
- To our knowledge, this proof-of-concept study is the first application of DL techniques for generating synthetic post-contrast images for MSK inflammatory imaging.
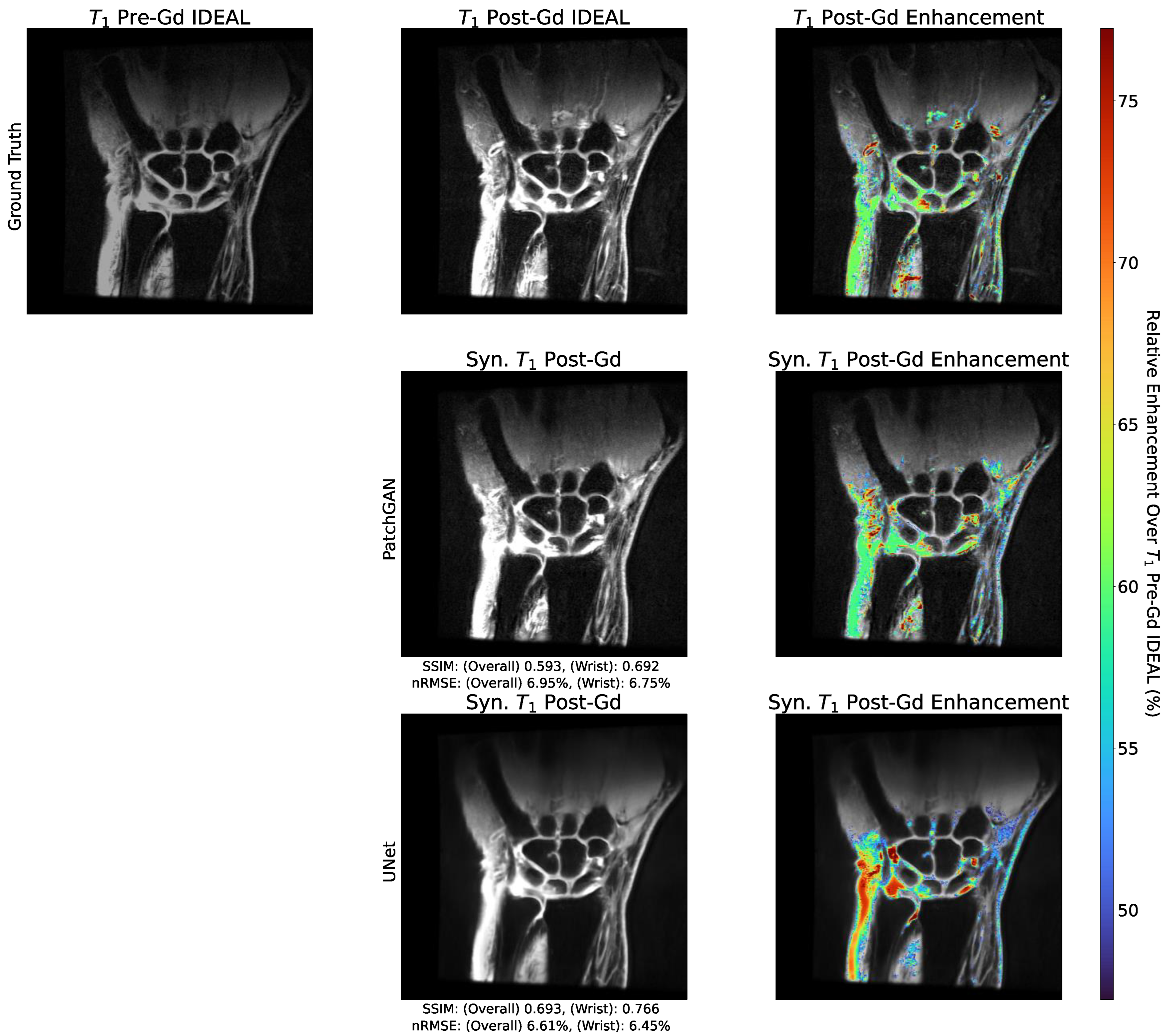
- We show that our trained pipelines perform strongly with regards to predicting post-contrast image appearance, particularly in regions afflicted with synovitis, where these models would see the most clinical utility.
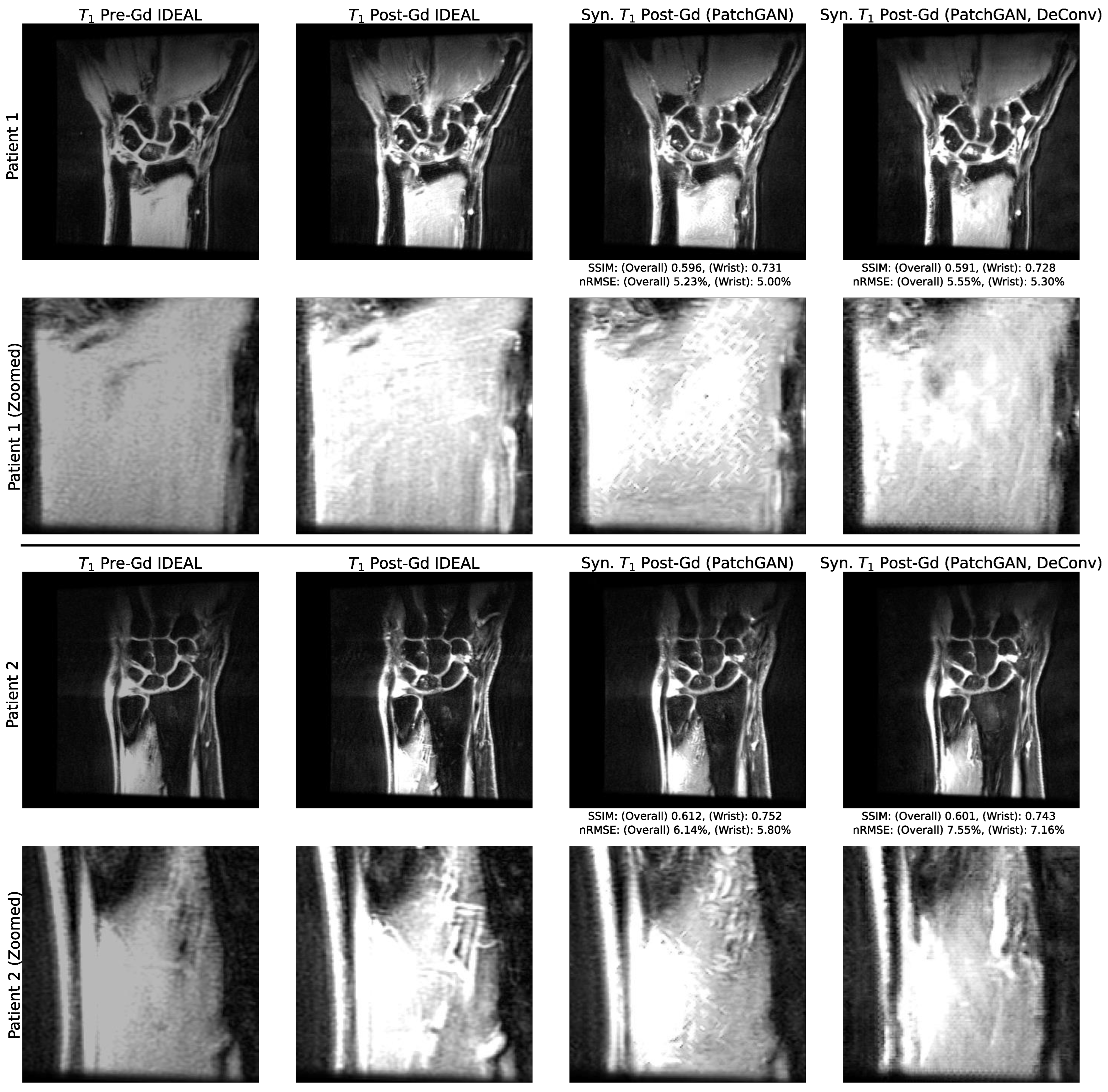
- We investigate the deconvolution operator, checkerboarding artifacts that can be intrinsic to architectures that use it, and how they surface in conventional and adversarial network training schemes.
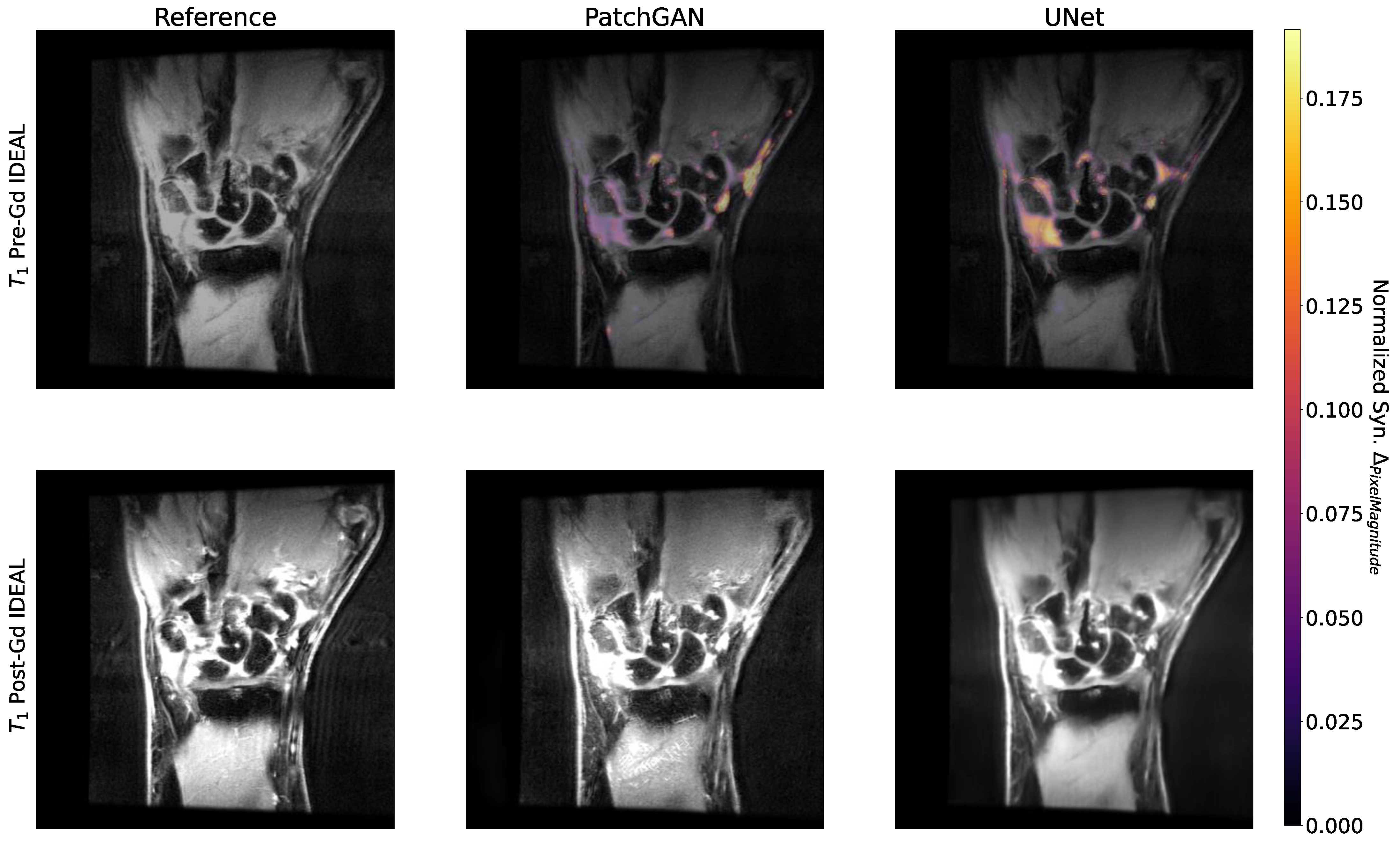
- We conduct a rigorous analysis of model predictions, identifying regions in pre-contrast image inputs that were most important to predicted post-contrast images, and regions in which predictions were most uncertain. This provides a straightforward framework that can be used to understand predictions made by image synthesis architectures in biomedical imaging applications.
2. Materials and Methods
2.1. Study Group
2.2. MR Acquisition
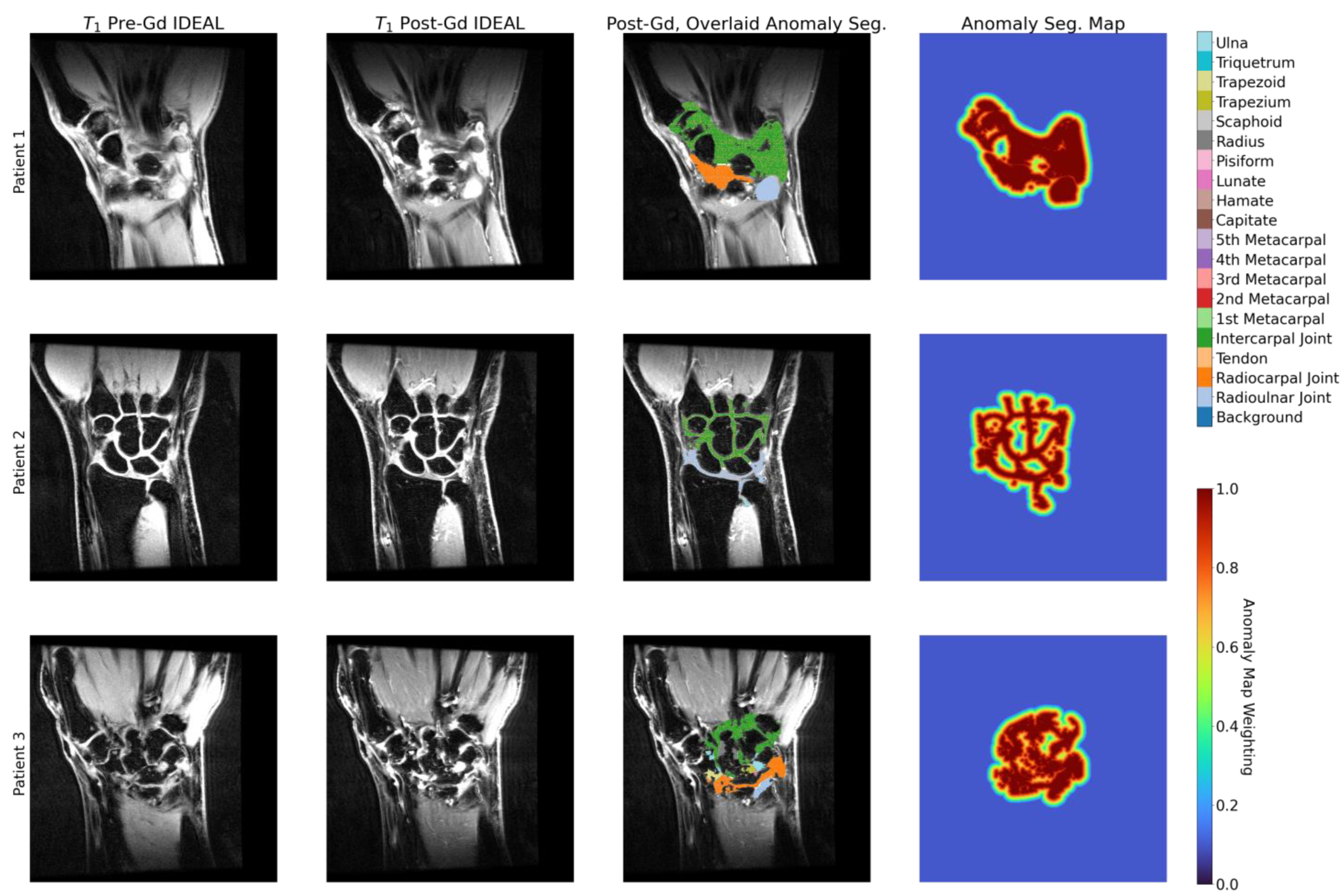
2.3. Anomaly Segmentations and Evaluations
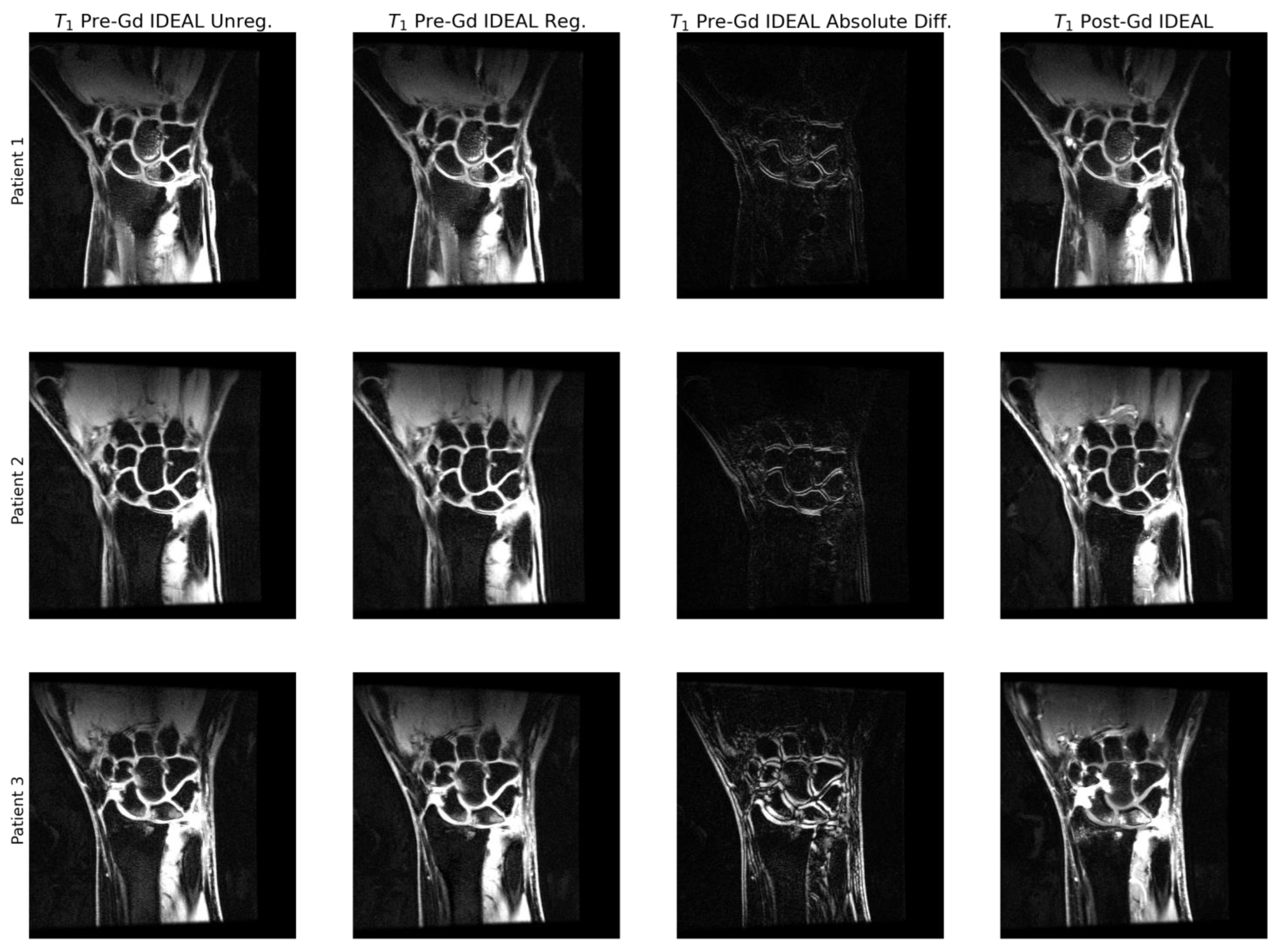
2.4. Image Preprocessing
2.5. Data Partitioning
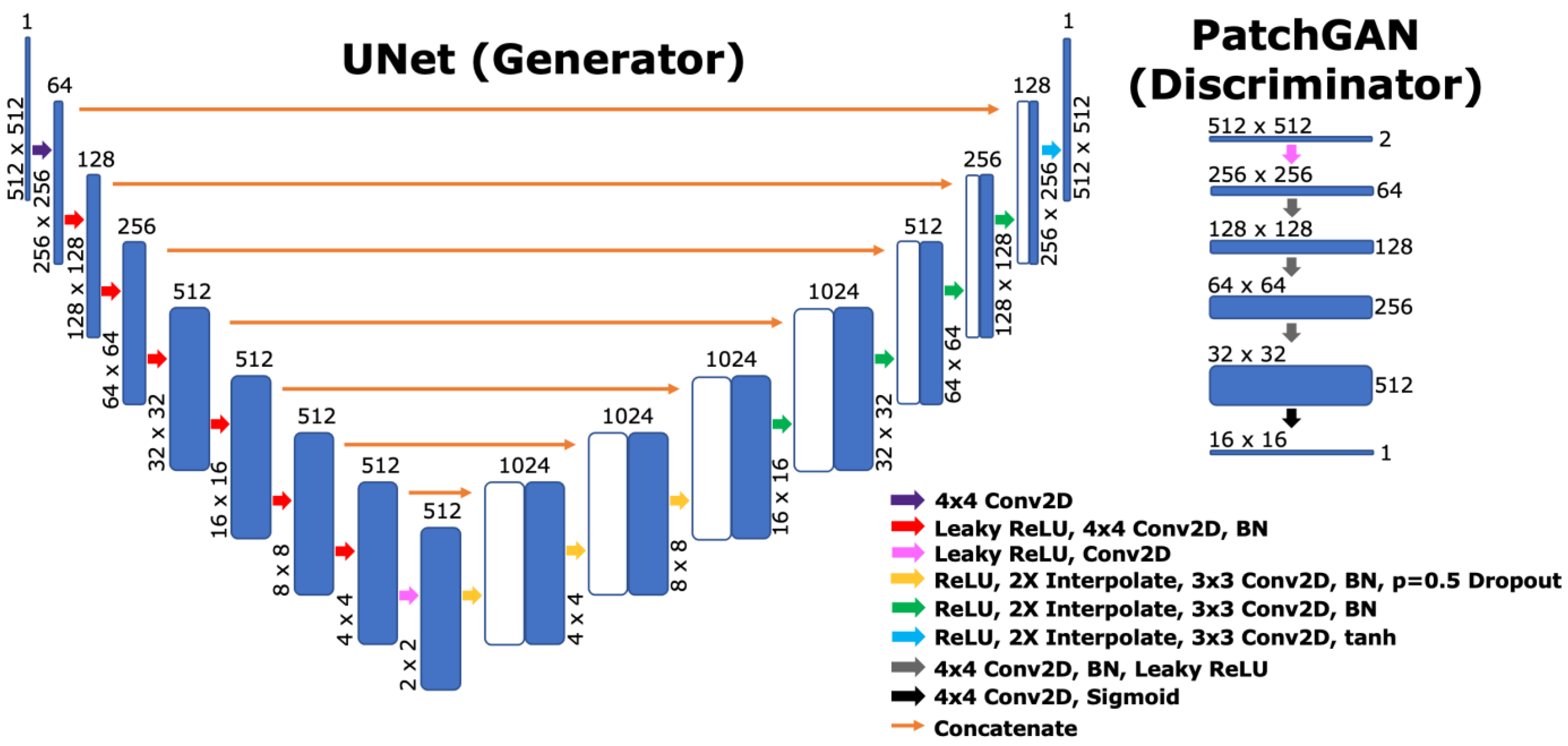
2.6. Network Architecture
2.7. Training Details
2.8. Hyperparameter Search and Model Selection
2.9. Model Performance Evaluation
2.10. Enhancement Maps
2.11. Occlusion Maps
2.12. Uncertainty Maps
2.13. Statistical Analysis
3. Results
3.1. Model Parameter Selection
- PatchGAN, no deconvolutions: = 0.05, = 0.01;
- PatchGAN, with deconvolutions: = 0.15, = 0.001;
- UNet, no deconvolutions: = 0.05;
- UNet, with deconvolutions: = 0.15.
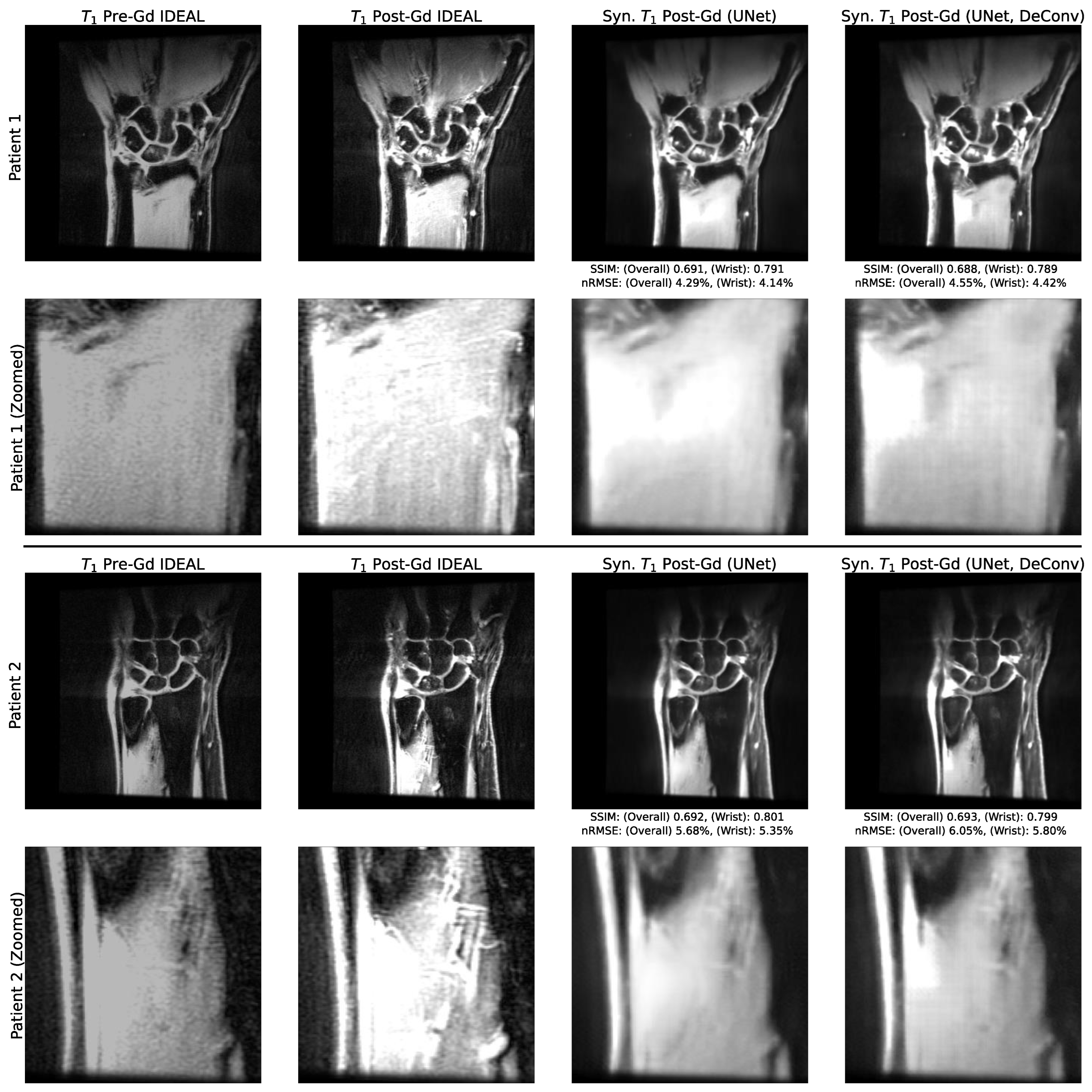
3.2. Utility of Deconvolution Operators in Baseline UNet and PatchGAN Generator Decoders
3.3. Standard Reconstruction Metrics Performance
3.4. Comparison of Reconstruction Performance across Synovitis Severity
3.5. Enhancement Maps Analysis
3.6. Occlusion and Uncertainty Maps Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A

| Scanner | GE Signa Discovery MR750w |
| Coil | 8-channel HD Wrist Array |
| Field Strength | 3T |
| Slice Thickness | 2 mm |
| Spacing between Slices | 2 mm |
| TR | 457–793 ms |
| TE | 10.06–12.48 ms |
| Frequency | 127.8 Hz |
| Bandwidth | 195.3 Hz (384 × 256, n = 58), 390.6 Hz (256 × 224, n = 6) |
| Acquisition Matrix | 384 × 256 (n = 58), 256 × 224 (n = 6) |
| Flip Angle | 90 (n = 2) or 111 (n = 62) |
| SAR | 1.578–3.259 |
| Pixel Spacing | 0.234 × 0.234 mm (n = 58), 0.469 × 0.469 mm (n = 6) |
| λB | ||||||||
| λGAN | 0 | 0.025 | 0.050 | 0.075 | 0.100 | 0.150 | 0.200 | |
| 0.001 | Full | 0.596 | 0.590 | 0.593 | 0.560 | 0.584 | 0.573 | 0.572 |
| Wrist | 0.713 | 0.713 | 0.718 | 0.708 | 0.711 | 0.712 | 0.705 | |
| 0.002 | Full | 0.555 | 0.553 | 0.548 | 0.537 | 0.567 | 0.557 | 0.556 |
| Wrist | 0.697 | 0.709 | 0.673 | 0.662 | 0.715 | 0.694 | 0.690 | |
| 0.003 | Full | 0.538 | 0.577 | 0.541 | 0.590 | 0.538 | 0.634 | 0.592 |
| Wrist | 0.681 | 0.703 | 0.689 | 0.712 | 0.665 | 0.747 | 0.725 | |
| 0.004 | Full | 0.496 | 0.550 | 0.582 | 0.594 | 0.548 | 0.555 | 0.581 |
| Wrist | 0.713 | 0.680 | 0.705 | 0.711 | 0.709 | 0.707 | 0.709 | |
| 0.005 | Full | 0.591 | 0.554 | 0.585 | 0.575 | 0.584 | 0.622 | 0.578 |
| Wrist | 0.721 | 0.698 | 0.708 | 0.692 | 0.721 | 0.738 | 0.716 | |
| 0.006 | Full | 0.519 | 0.527 | 0.596 | 0.583 | 0.565 | 0.505 | 0.577 |
| Wrist | 0.695 | 0.687 | 0.705 | 0.703 | 0.698 | 0.687 | 0.703 | |
| 0.007 | Full | 0.557 | 0.587 | 0.607 | 0.491 | 0.579 | 0.555 | 0.56 |
| Wrist | 0.712 | 0.714 | 0.722 | 0.682 | 0.703 | 0.695 | 0.685 | |
| 0.008 | Full | 0.561 | 0.567 | 0.585 | 0.553 | 0.565 | 0.526 | 0.567 |
| Wrist | 0.694 | 0.732 | 0.715 | 0.688 | 0.717 | 0.675 | 0.681 | |
| 0.009 | Full | 0.560 | 0.510 | 0.624 | 0.550 | 0.571 | 0.580 | 0.507 |
| Wrist | 0.684 | 0.674 | 0.748 | 0.705 | 0.693 | 0.704 | 0.703 | |
| 0.010 | Full | 0.518 | 0.496 | 0.574 | 0.582 | 0.502 | 0.567 | 0.632 |
| Wrist | 0.689 | 0.720 | 0.733 | 0.701 | 0.646 | 0.694 | 0.737 | |
| λB | ||||||||
| λGAN | 0 | 0.025 | 0.050 | 0.075 | 0.100 | 0.150 | 0.200 | |
| 0.001 | Full | 11.5 | 6.1 | 11.4 | 7.8 | 14.8 | 8.7 | 24.1 |
| Wrist | 11.1 | 5.8 | 10.8 | 7.4 | 14.5 | 8.2 | 23.1 | |
| 0.002 | Full | 12.6 | 9.7 | 6.2 | 10 | 23.9 | 20.3 | 20.3 |
| Wrist | 12 | 9.1 | 5.9 | 9.5 | 23.7 | 20 | 20.1 | |
| 0.003 | Full | 10.6 | 12 | 9.9 | 10.6 | 38.4 | 10.1 | 12.6 |
| Wrist | 10.2 | 11.2 | 9.5 | 10 | 38.1 | 9.6 | 11.7 | |
| 0.004 | Full | 9.2 | 18.5 | 22.5 | 11.6 | 10 | 33.8 | 10 |
| Wrist | 8.8 | 18 | 21.8 | 11.1 | 9.7 | 33.4 | 9.5 | |
| 0.005 | Full | 12.8 | 16.7 | 11.8 | 11.3 | 15.3 | 24.5 | 18.9 |
| Wrist | 12.4 | 15.9 | 11.3 | 10.6 | 14.9 | 24.2 | 18.3 | |
| 0.006 | Full | 21.2 | 6.2 | 17.9 | 13.1 | 5.8 | 14.5 | 12.3 |
| Wrist | 20.9 | 5.9 | 17 | 12.6 | 5.5 | 13.9 | 11.1 | |
| 0.007 | Full | 10.1 | 6.4 | 22 | 11.4 | 10.4 | 8.9 | 19.1 |
| Wrist | 9.5 | 6.1 | 21.4 | 11 | 9.9 | 8.4 | 18.1 | |
| 0.008 | Full | 9.9 | 11.4 | 9.4 | 8.9 | 9.6 | 9.8 | 7.9 |
| Wrist | 9.3 | 10.9 | 9 | 8.4 | 9.2 | 8.9 | 7.5 | |
| 0.009 | Full | 15.3 | 47 | 10.5 | 12.7 | 9.9 | 10.7 | 13.2 |
| Wrist | 14.7 | 7.4 | 10 | 12.4 | 9.5 | 10.3 | 12.6 | |
| 0.010 | Full | 10.9 | 12.9 | 11 | 10.6 | 15.4 | 6.8 | 8.7 |
| Wrist | 10.5 | 12.6 | 10.4 | 10.1 | 14.7 | 6.4 | 8.4 | |
| RAMRIS = 0 | RAMRIS > 0 | ||||||
|---|---|---|---|---|---|---|---|
| Full | Wrist Only | Synovial Joints | Full | Wrist Only | Synovial Joints | ||
| Pre-Gd | nRMSE | 23.95 ± 4.71 | 23.72 ± 4.83 | 133.52 ± 43.31 | 27.24 ± 10.27 | 26.94 ± 10.43 | 310.93 ± 159.54 |
| PSNR | 17.76 ± 0.38 | 17.89 ± 0.38 | 9.38 ± 0.40 | 17.77 ± 1.10 | 17.95 ± 1.23 | 8.77 ± 1.90 | |
| SSIM | 0.62 ± 0.03 | 0.75 ± 0.01 | 0.59 ± 0.03 | 0.73 ± 0.01 | |||
| PatchGAN Reg. | nRMSE | 7.13 ± 0.80 | 6.85 ± 0.77 | 28.20 ± 11.85 | 6.55 ± 0.75 | 6.26 ± 0.73 | 21.12 ± 2.37 |
| PSNR | 20.44 ± 0.52 | 20.68 ± 0.51 | 11.53 ± 2.19 | 20.90 ± 0.64 | 21.16 ± 0.65 | 12.33 ± 0.64 | |
| SSIM | 0.58 ± 0.02 | 0.72 ± 0.00 | 0.58 ± 0.02 | 0.73 ± 0.01 | |||
| PatchGAN Unreg. | nRMSE | 8.93 ± 0.69 | 8.65 ± 0.68 | 36.44 ± 17.23 | 8.28 ± 1.09 | 7.96 ± 1.06 | 25.98 ± 2.54 |
| PSNR | 19.54 ± 0.34 | 19.73 ± 0.33 | 10.60 ± 2.45 | 19.98 ± 0.75 | 20.21 ± 0.75 | 11.49 ± 0.76 | |
| SSIM | 0.56 ± 0.02 | 0.70 ± 0.00 | 0.56 ± 0.02 | 0.71 ± 0.01 | |||
| UNet Reg. | nRMSE | 7.26 ± 0.20 | 7.00 ± 0.17 | 28.74 ± 10.61 | 5.90 ± 0.73 | 5.67 ± 0.76 | 25.15 ± 5.39 |
| PSNR | 21.36 ± 0.07 | 21.57 ± 0.04 | 11.26 ± 1.58 | 22.30 ± 0.49 | 22.54 ± 0.51 | 11.71 ± 0.40 | |
| SSIM | 0.68 ± 0.02 | 0.78 ± 0.01 | 0.69 ± 0.02 | 0.79 ± 0.01 | |||
| UNet Unreg. | nRMSE | 8.35 ± 0.45 | 8.11 ± 0.45 | 31.51 ± 9.96 | 7.48 ± 1.09 | 7.24 ± 1.09 | 28.96 ± 6.28 |
| PSNR | 20.82 ± 0.15 | 21.00 ± 0.14 | 10.67 ± 1.36 | 21.36 ± 0.67 | 21.57 ± 0.67 | 11.10 ± 0.51 | |
| SSIM | 0.67 ± 0.02 | 0.77 ± 0.01 | 0.68 ± 0.02 | 0.78 ± 0.01 | |||
References
- Silman, A.J.; Pearson, J.E. Epidemiology and Genetics of Rheumatoid Arthritis. Arthritis Res. 2002, 4 (Suppl. S3), S265–S272. [Google Scholar] [CrossRef]
- Aletaha, D.; Smolen, J.S. Diagnosis and Management of Rheumatoid Arthritis: A Review. JAMA 2018, 320, 1360–1372. [Google Scholar] [CrossRef]
- Taylor, P.C. Update on the Diagnosis and Management of Early Rheumatoid Arthritis. Clin. Med. 2020, 20, 561. [Google Scholar] [CrossRef]
- Goekoop-Ruiterman, Y.P.M.; De Vries-Bouwstra, J.K.; Allaart, C.F.; Van Zeben, D.; Kerstens, P.J.S.M.; Hazes, J.M.W.; Zwinderman, A.H.; Ronday, H.K.; Han, K.H.; Westedt, M.L.; et al. Clinical and Radiographic Outcomes of Four Different Treatment Strategies in Patients with Early Rheumatoid Arthritis (the BeSt Study): A Randomized, Controlled Trial. Arthritis Rheumatol. 2005, 52, 3381–3390. [Google Scholar] [CrossRef]
- Kgoebane, K.; Ally, M.M.T.M.; Duim-Beytell, M.C.; Suleman, F.E. The Role of Imaging in Rheumatoid Arthritis. S. Afr. J. Radiol. 2018, 22, 6. [Google Scholar] [CrossRef]
- De Schepper, A.M.; De Beuckeleer, L.; Vandevenne, J.; Somville, J. Magnetic Resonance Imaging of Soft Tissue Tumors. Eur. Radiol. 2000, 10, 213–222. [Google Scholar] [CrossRef]
- Rubin, D.A. MRI and Ultrasound of the Hands and Wrists in Rheumatoid Arthritis. I. Imaging Findings. Skelet. Radiol. 2019, 48, 677–695. [Google Scholar] [CrossRef]
- Zhou, Z.; Lu, Z.R. Gadolinium-Based Contrast Agents for MR Cancer Imaging. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2012, 5, 1–18. [Google Scholar] [CrossRef]
- Eshed, I.; Feist, E.; Althoff, C.E.; Hamm, B.; Konen, E.; Burmester, G.R.; Backhaus, M.; Hermann, K.G.A. Tenosynovitis of the Flexor Tendons of the Hand Detected by MRI: An Early Indicator of Rheumatoid Arthritis. Rheumatology (Oxford) 2009, 48, 887–891. [Google Scholar] [CrossRef]
- Tamai, M.; Kawakami, A.; Uetani, M.; Fukushima, A.; Arima, K.; Fujikawa, K.; Iwamoto, N.; Aramaki, T.; Kamachi, M.; Nakamura, H.; et al. Magnetic Resonance Imaging (MRI) Detection of Synovitis and Bone Lesions of the Wrists and Finger Joints in Early-Stage Rheumatoid Arthritis: Comparison of the Accuracy of Plain MRI-Based Findings and Gadolinium-Diethylenetriamine Pentaacetic Acid-Enhanced MRI-Based Findings. Mod. Rheumatol. 2012, 22, 654–658. [Google Scholar] [CrossRef]
- Boyd, A.S.; Zic, J.A.; Abraham, J.L. Gadolinium Deposition in Nephrogenic Fibrosing Dermopathy. J. Am. Acad. Dermatol. 2007, 56, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Murata, N.; Gonzalez-Cuyar, L.F.; Murata, K.; Fligner, C.; Dills, R.; Hippe, D.; Maravilla, K.R. Macrocyclic and Other Non-Group 1 Gadolinium Contrast Agents Deposit Low Levels of Gadolinium in Brain and Bone Tissue: Preliminary Results from 9 Patients with Normal Renal Function. Investig. Radiol. 2016, 51, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Gulani, V.; Calamante, F.; Shellock, F.G.; Kanal, E.; Reeder, S.B. Gadolinium Deposition in the Brain: Summary of Evidence and Recommendations. Lancet Neurol. 2017, 16, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Tseng, H.Y.; Lee, H.Y.; Jiang, L.; Yang, M.H.; Yang, W. RetrieveGAN: Image Synthesis via Differentiable Patch Retrieval. Lect. Notes Comput. Sci. (Incl. Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinform.) 2020, 12353, 242–257. [Google Scholar] [CrossRef]
- Ramesh, A.; Pavlov, M.; Goh, G.; Gray, S.; Voss, C.; Radford, A.; Chen, M.; Sutskever, I. Zero-Shot Text-to-Image Generation. Int. Conf. Mach. Learn. 2021, 139, 8821–8831. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef]
- Zeiler, M.D.; Fergus, R. Visualizing and Understanding Convolutional Networks. Lect. Notes Comput. Sci. (Incl. Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinform.) 2013, 8689 LNCS, 818–833. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. LNCS 2015, 9351, 234–241. [Google Scholar]
- Huang, H.; Yu, P.S.; Wang, C. An Introduction to Image Synthesis with Generative Adversarial Nets. arXiv 2018, arXiv:1803.04469. [Google Scholar] [CrossRef]
- Fulgeri, F.; Fabbri, M.; Alletto, S.; Calderara, S.; Cucchiara, R. Can Adversarial Networks Hallucinate Occluded People with a Plausible Aspect? Comput. Vis. Image Underst. 2019, 182, 71–80. [Google Scholar] [CrossRef]
- Kingma, D.P.; Welling, M. Auto-Encoding Variational Bayes. In Proceedings of the 2nd International Conference on Learning Representations, ICLR 2014, Banff, AB, Canada, 14–16 April 2014. [Google Scholar]
- Esser, P.; Rombach, R.; Ommer, B. Taming Transformers for High-Resolution Image Synthesis. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 13–19 June 2020; pp. 12868–12878. [Google Scholar] [CrossRef]
- Calabrese, E.; Rudie, J.D.; Rauschecker, A.M.; Villanueva-Meyer, J.E.; Cha, S. Feasibility of Simulated Postcontrast Mri of Glioblastomas and Lower-Grade Gliomas by Using Three-Dimensional Fully Convolutional Neural Networks. Radiol. Artif. Intell. 2021, 3, e200276. [Google Scholar] [CrossRef]
- Gong, E.; Pauly, J.M.; Wintermark, M.; Zaharchuk, G. Deep Learning Enables Reduced Gadolinium Dose for Contrast-Enhanced Brain MRI. J. Magn. Reson. Imaging 2018, 48, 330–340. [Google Scholar] [CrossRef]
- Pasumarthi, S.; Tamir, J.I.; Christensen, S.; Zaharchuk, G.; Zhang, T.; Gong, E. A Generic Deep Learning Model for Reduced Gadolinium Dose in Contrast-Enhanced Brain MRI. Magn. Reason. Med. 2021, 86, 1687–1700. [Google Scholar] [CrossRef]
- Xie, H.; Lei, Y.; Wang, T.; Roper, J.; Axente, M.; Bradley, J.D.; Liu, T.; Yang, X. Magnetic Resonance Imaging Contrast Enhancement Synthesis Using Cascade Networks with Local Supervision. Med. Phys. 2022, 49, 3278–3287. [Google Scholar] [CrossRef]
- Xu, C.; Xu, L.; Gao, Z.; Zhao, S.; Zhang, H.; Zhang, Y.; Du, X.; Zhao, S.; Ghista, D.; Liu, H.; et al. Direct Delineation of Myocardial Infarction without Contrast Agents Using a Joint Motion Feature Learning Architecture. Med. Image Anal. 2018, 50, 82–94. [Google Scholar] [CrossRef]
- Zhang, N.; Yang, G.; Gao, Z.; Xu, C.; Zhang, Y.; Shi, R.; Keegan, J.; Xu, L.; Zhang, H.; Fan, Z.; et al. Deep Learning for Diagnosis of Chronic Myocardial Infarction on Nonenhanced Cardiac Cine MRI. Radiology 2019, 291, 606–607. [Google Scholar] [CrossRef]
- Wang, P.; Hu, S.; Wang, X.; Ge, Y.; Zhao, J.; Qiao, H.; Chang, J.; Dou, W.; Zhang, H. Synthetic MRI in Differentiating Benign from Metastatic Retropharyngeal Lymph Node: Combination with Diffusion-Weighted Imaging. Eur. Radiol. 2023, 33, 152–161. [Google Scholar] [CrossRef]
- Kausar, T.; Kausar, A.; Ashraf, M.A.; Siddique, M.F.; Wang, M.; Sajid, M.; Siddique, M.Z.; Haq, A.U.; Riaz, I. SA-GAN: Stain Acclimation Generative Adversarial Network for Histopathology Image Analysis. Appl. Sci. 2021, 12, 288. [Google Scholar] [CrossRef]
- Cross-Zamirski, J.O.; Mouchet, E.; Williams, G.; Schönlieb, C.B.; Turkki, R.; Wang, Y. Label-Free Prediction of Cell Painting from Brightfield Images. Sci. Rep. 2022, 12, 10001. [Google Scholar] [CrossRef]
- Vassa, R.; Garg, A.; Omar, I.M. Magnetic Resonance Imaging of the Wrist and Hand. Pol. J. Radiol. 2020, 85, e461. [Google Scholar] [CrossRef]
- Reeder, S.B.; McKenzie, C.A.; Pineda, A.R.; Yu, H.; Shimakawa, A.; Brau, A.C.; Hargreaves, B.A.; Gold, G.E.; Brittain, J.H. Water–Fat Separation with IDEAL Gradient-Echo Imaging. J. Magn. Reson. Imaging 2007, 25, 644–652. [Google Scholar] [CrossRef]
- Østergaard, M.; Peterfy, C.G.; Bird, P.; Gandjbakhch, F.; Glinatsi, D.; Eshed, I.; Haavardsholm, E.A.; Lillegraven, S.; Bøyesen, P.; Ejbjerg, B.; et al. The OMERACT Rheumatoid Arthritis Magnetic Resonance Imaging (MRI) Scoring System: Updated Recommendations by the OMERACT MRI in Arthritis Working Group. J. Rheumatol. 2017, 44, 1706–1712. [Google Scholar] [CrossRef]
- Mattes, D.; Haynor, D.; Vesselle, H.; Lewellyn, T.; Eubank David Mattes, W.; Haynor, D.R.; Lewellyn, T.K.; Eubank, W. Nonrigid Multimodality Image Registration. Med. Imaging 2001 Image Process. 2001, 4322, 1609–1620. [Google Scholar] [CrossRef]
- Dosselmann, R.; Yang, X.D. A Comprehensive Assessment of the Structural Similarity Index. Signal Image Video Process 2009, 5, 81–91. [Google Scholar] [CrossRef]
- Beare, R.; Lowekamp, B.; Yaniv, Z. Image Segmentation, Registration and Characterization in R with SimpleITK. J. Stat. Softw. 2018, 86, 8. [Google Scholar] [CrossRef]
- Yaniv, Z.; Lowekamp, B.C.; Johnson, H.J.; Beare, R. SimpleITK Image-Analysis Notebooks: A Collaborative Environment for Education and Reproducible Research. J. Digit. Imaging 2018, 31, 290. [Google Scholar] [CrossRef]
- Lowekamp, B.C.; Chen, D.T.; Ibáñez, L.; Blezek, D. The Design of SimpleITK. Front. Neuroinform. 2013, 7, 45. [Google Scholar] [CrossRef]
- Isola, P.; Zhu, J.Y.; Zhou, T.; Efros, A.A. Image-to-Image Translation with Conditional Adversarial Networks. In Proceedings of the 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017, Honolulu, HI, USA, 21–26 July 2016; pp. 5967–5976. [Google Scholar] [CrossRef]
- Dong, C.; Loy, C.C.; He, K.; Tang, X. Image Super-Resolution Using Deep Convolutional Networks. IEEE Trans. Pattern Anal. Mach. Intell. 2016, 38, 295–307. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J.L. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference on Learning Representations, ICLR 2015, San Diego, CA, USA, 7–9 May 2015. [Google Scholar] [CrossRef]
- Horé, A.; Ziou, D. Is There a Relationship between Peak-Signal-to-Noise Ratio and Structural Similarity Index Measure? IET Image Process. 2013, 7, 12–24. [Google Scholar] [CrossRef]
- Knoll, F.; Murrell, T.; Sriram, A.; Yakubova, N.; Zbontar, J.; Rabbat, M.; Defazio, A.; Muckley, M.J.; Sodickson, D.K.; Zitnick, C.L.; et al. Advancing Machine Learning for MR Image Reconstruction with an Open Competition: Overview of the 2019 FastMRI Challenge. Magn. Reason. Med. 2020, 84, 3054–3070. [Google Scholar] [CrossRef]
- Mason, A.; Rioux, J.; Clarke, S.E.; Costa, A.; Schmidt, M.; Keough, V.; Huynh, T.; Beyea, S. Comparison of Objective Image Quality Metrics to Expert Radiologists’ Scoring of Diagnostic Quality of MR Images. IEEE Trans. Med. Imaging 2020, 39, 1064–1072. [Google Scholar] [CrossRef]
- Tomczak, A.; Gupta, A.; Ilic, S.; Navab, N.; Albarqouni, S. What Can We Learn About a Generated Image Corrupting Its Latent Representation? Lect. Notes Comput. Sci. (Incl. Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinform.) 2022, 13436 LNCS, 505–515. [Google Scholar]
- De Winter, J.C.F. Using the Student’s t-Test with Extremely Small Sample Sizes. Pract. Assess. Res. Eval. Pract. Assess. 2013, 18, 10. [Google Scholar] [CrossRef]
- Nahler, G. Bonferroni Correction. Dict. Pharm. Med. 2009, 18. [Google Scholar] [CrossRef]
- Adamson, P.M.; Gunel, B.; Dominic, J.; Desai, A.D.; Spielman, D.; Vasanawala, S.; Pauly, J.M.; Chaudhari, A. SSFD: Self-Supervised Feature Distance as an MR Image Reconstruction Quality Metric. In Proceedings of the NeurIPS 2021 Workshop on Deep Learning and Inverse Problems, Virtual, 6–14 December 2021. [Google Scholar]
- Shi, W.; Caballero, J.; Huszar, F.; Totz, J.; Aitken, A.P.; Bishop, R.; Rueckert, D.; Wang, Z. Real-Time Single Image and Video Super-Resolution Using an Efficient Sub-Pixel Convolutional Neural Network. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition 2016, Las Vegas, NV, USA, 27–30 June 2016; pp. 1874–1883. [Google Scholar] [CrossRef]
- Sugawara, Y.; Shiota, S.; Kiya, H. Checkerboard Artifacts Free Convolutional Neural Networks. APSIPA Trans. Signal Inf. Process. 2019, 8, e9. [Google Scholar] [CrossRef]
- Kamrul Hasan, S.M.; Linte, C.A. U-NetPlus: A Modified Encoder-Decoder U-Net Architecture for Semantic and Instance Segmentation of Surgical Instruments from Laparoscopic Images. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS 2019, Berlin, Germany, 23–27 July 2019; pp. 7205–7211. [Google Scholar] [CrossRef]
- Guermazi, A.; Roemer, F.W.; Hayashi, D.; Crema, M.D.; Niu, J.; Zhang, Y.; Marra, M.D.; Katur, A.; Lynch, J.A.; El-Khoury, G.Y.; et al. Assessment of Synovitis with Contrast-Enhanced MRI Using a Whole-Joint Semiquantitative Scoring System in People with, or at High Risk of, Knee Osteoarthritis: The MOST Study. Ann. Rheum. Dis. 2011, 70, 805–811. [Google Scholar] [CrossRef]
- Amini, B.; Murphy, W.A.; Haygood, T.M.; Kumar, R.; McEnery, K.W.; Madewell, J.E.; Mujtaba, B.M.; Wei, W.; Costelloe, C.M. Gadolinium-Based Contrast Agents Improve Detection of Recurrent Soft-Tissue Sarcoma at Mri. Radiol. Imaging Cancer 2020, 2, e190046. [Google Scholar] [CrossRef]
- Crema, M.D.; Felson, D.T.; Roemer, F.W.; Niu, J.; Marra, M.D.; Zhang, Y.; Lynch, J.A.; El-Khoury, G.Y.; Lewis, C.E.; Guermazi, A. Peripatellar Synovitis: Comparison between Non-Contrast-Enhanced and Contrast-Enhanced MRI and Association with Pain. The MOST Study. Osteoarthr. Cartil. 2013, 21, 413–418. [Google Scholar] [CrossRef]
- Lee, J.H.; Dyke, J.P.; Ballon, D.; Ciombor, D.M.K.; Tung, G.; Aaron, R.K. Assessment of Bone Perfusion with Contrast-Enhanced Magnetic Resonance Imaging: Imaging of Bone Marrow Edema Associated with Osteoarthritis and Avascular Necrosis. Orthop. Clin. N. Am. 2009, 40, 249. [Google Scholar] [CrossRef]
- Salaffi, F.; Ceccarelli, L.; Carotti, M.; Di Carlo, M.; Polonara, G.; Facchini, G.; Golfieri, R.; Giovagnoni, A. Differentiation between Infectious Spondylodiscitis versus Inflammatory or Degenerative Spinal Changes: How Can Magnetic Resonance Imaging Help the Clinician? Radiol. Med. 2021, 126, 843. [Google Scholar] [CrossRef]
- Tolpadi, A.A.; Bharadwaj, U.; Gao, K.T.; Bhattacharjee, R.; Gassert, F.G.; Luitjens, J.; Giesler, P.; Morshuis, J.N.; Fischer, P.; Hein, M.; et al. K2S Challenge: From Undersampled K-Space to Automatic Segmentation. Bioengineering 2023, 10, 267. [Google Scholar] [CrossRef]
- Zhao, R.; Yaman, B.; Zhang, Y.; Stewart, R.; Dixon, A.; Knoll, F.; Huang, Z.; Lui, Y.W.; Hansen, M.S.; Lungren, M.P. FastMRI+: Clinical Pathology Annotations for Knee and Brain Fully Sampled Multi-Coil MRI Data. Sci. Data. 2022, 9, 152. [Google Scholar] [CrossRef]
- Peterfy, C.G.; Schneider, E.; Nevitt, M. The Osteoarthritis Initiative: Report on the Design Rationale for the Magnetic Resonance Imaging Protocol for the Knee. Osteoarthr. Cartil. 2008, 16, 1433–1441. [Google Scholar] [CrossRef]


| Train | Validation | Test | Full | |
|---|---|---|---|---|
| Age | 53.38 ± 13.50 | 45.94 ± 16.16 | 52.12 ± 18.60 | 52.41 ± 14.65 |
| BMI | 29.35 ± 8.90 | 25.32 ± 3.06 | 28.33 ± 1.26 | 28.79 ± 8.03 |
| ESR [mm/h] | 29.06 ± 26.07 | 32.00 ± 24.00 | 27.00 ± 20.12 | 29.05 ± 25.32 |
| RAMRIS Synovitis | 4.57 ± 2.13 | 2.33 ± 2.62 | 1.67 ± 1.25 | 4.00 ± 2.37 |
| Slices | 783 | 87 | 105 | 975 |
| Volumes | 51 | 6 | 7 | 64 |
| Full | Wrist Only | Synovial Joints | ||
|---|---|---|---|---|
| Pre-Gd | nRMSE | 26.30 ± 9.16 | 17.82 ± 6.31 | 260.24 ± 158.56 |
| PSNR | 17.77 ± 0.95 | 22.99 ± 0.91 | 8.94 ± 1.64 | |
| SSIM | 0.60 ± 0.03 | 0.94 ± 0.00 | ||
| PatchGAN Registered | nRMSE | 6.72 ± 0.81 *** | 6.07 ± 1.22 *** | 23.14 ± 7.37 ** |
| PSNR | 20.77 ± 0.65 *** | 25.40 ± 1.24 ** | 12.10 ± 1.34 ** | |
| SSIM | 0.58 ± 0.02 | 0.94 ± 0.01 | ||
| PatchGAN Unregistered | nRMSE | 8.46 ± 1.03 *** | 7.68 ± 1.41 ** | 28.96 ± 10.57 ** |
| PSNR | 19.85 ± 0.69 *** | 24.38 ± 1.21 * | 11.23 ± 1.52 * | |
| SSIM | 0.56 ± 0.02 * | 0.94 ± 0.01 | ||
| UNet Registered | nRMSE | 6.29 ± 0.88 *** | 4.36 ± 0.60 *** | 26.18 ± 7.45 ** |
| PSNR | 22.03 ± 0.60 *** | 27.13 ± 0.69 *** | 11.58 ± 0.93 ** | |
| SSIM | 0.69 ± 0.02 *** | 0.95 ± 0.00 ** | ||
| UNet Unregistered | nRMSE | 7.73 ± 1.03 *** | 5.38 ± 0.73 *** | 29.69 ± 7.60 ** |
| PSNR | 21.20 ± 0.62 *** | 26.20 ± 0.77 *** | 10.98 ± 0.87 * | |
| SSIM | 0.68 ± 0.02 *** | 0.95 ± 0.01 * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tolpadi, A.A.; Luitjens, J.; Gassert, F.G.; Li, X.; Link, T.M.; Majumdar, S.; Pedoia, V. Synthetic Inflammation Imaging with PatchGAN Deep Learning Networks. Bioengineering 2023, 10, 516. https://doi.org/10.3390/bioengineering10050516
Tolpadi AA, Luitjens J, Gassert FG, Li X, Link TM, Majumdar S, Pedoia V. Synthetic Inflammation Imaging with PatchGAN Deep Learning Networks. Bioengineering. 2023; 10(5):516. https://doi.org/10.3390/bioengineering10050516
Chicago/Turabian StyleTolpadi, Aniket A., Johanna Luitjens, Felix G. Gassert, Xiaojuan Li, Thomas M. Link, Sharmila Majumdar, and Valentina Pedoia. 2023. "Synthetic Inflammation Imaging with PatchGAN Deep Learning Networks" Bioengineering 10, no. 5: 516. https://doi.org/10.3390/bioengineering10050516
APA StyleTolpadi, A. A., Luitjens, J., Gassert, F. G., Li, X., Link, T. M., Majumdar, S., & Pedoia, V. (2023). Synthetic Inflammation Imaging with PatchGAN Deep Learning Networks. Bioengineering, 10(5), 516. https://doi.org/10.3390/bioengineering10050516







