Structure-Based Screening and Molecular Dynamics of Rifampicin Analogues Targeting InhA of Mycobacterium tuberculosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Receptor Structure Preparation
2.2. Ligand Structure Preparation
2.3. Evaluation of the Drug-Likeness Properties of Selected Compounds
2.4. ADMET Profiling
2.5. Molecular Docking
2.6. Molecular Dynamics Simulation
2.7. Free Energy Landscapes
2.8. Molecular Mechanics Poisson–Boltzmann Surface Area (MM/PBSA) Analysis
3. Results
3.1. Docking Validation and Benchmarking
3.2. Molecular Docking Interaction
3.3. Pharmacokinetic and Physicochemical Analysis
3.4. Stability Analysis
3.4.1. Root Mean Squared Deviation
3.4.2. Solvent Accessible Surface Area (SASA)
3.4.3. Radius of Gyration (Rg)
3.4.4. Root Mean Squared Fluctuations (RMSF)
3.5. Hydrogen Bond Analysis
3.6. Free Energy Landscape Analysis
3.7. MM/PBSA Binding–Free Energy Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mohammadnabi, N.; Shamseddin, J.; Emadi, M.; Bodaghi, A.B.; Varseh, M.; Shariati, A.; Rezaei, M.; Dastranj, M.; Farahani, A. Mycobacterium tuberculosis: The mechanism of pathogenicity, immune responses, and diagnostic challenges. J. Clin. Lab. Anal. 2024, 38, e25122. [Google Scholar] [CrossRef] [PubMed]
- Black, R.E.; Perin, J.; Yeung, D.; Rajeev, T.; Miller, J.; Elwood, S.E.; Platts-Mills, J.A. Estimated global and regional causes of deaths from diarrhoea in children younger than 5 years during 2000–21: A systematic review and Bayesian multinomial analysis. Lancet Glob. Health 2024, 12, e919–e928. [Google Scholar] [CrossRef] [PubMed]
- Çelik, M.; Döker, M.F.; Kırlangıçoğlu, C.; Ünsal, Ö.; Gökçeoğlu, S.; Ceylan, M.R.; Karabay, O. Comprehensive spatial investigation of tuberculosis dynamics and affecting factors in Şanlıurfa, Türkiye (2016–2023). GeoHealth 2025, 9, e2024GH001235. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Wang, T.; Du, J.; Sun, L.; Wang, G.; Ni, R.; An, Y.; Fan, X.; Li, Y.; Guo, R.; et al. Decoding the WHO Global Tuberculosis Report 2024: A critical analysis of global and Chinese key data. Zoonoses 2025, 5, 999. [Google Scholar] [CrossRef]
- Katende-Kyenda, L.N. Determinants of non-adherence to anti-tuberculosis treatment in a public primary healthcare clinic in South Africa: Improving the quality of long-term care. Int. J. Environ. Res. Public Health 2025, 22, 1209. [Google Scholar] [CrossRef]
- Olivier, C.; Luies, L. WHO goals and beyond: Managing HIV/TB co-infection in South Africa. SN Compr. Clin. Med. 2023, 5, 251. [Google Scholar] [CrossRef]
- Brode, S.K.; Dwilow, R.; Kunimoto, D.; Menzies, D.; Khan, F.A. Drug-resistant tuberculosis. Can. J. Respir. Crit. Care Sleep Med. 2022, 6, 109–128. [Google Scholar] [CrossRef]
- Chopra, H.; Mohanta, Y.K.; Rauta, P.R.; Ahmed, R.; Mahanta, S.; Mishra, P.K.; Panda, P.; Rabaan, A.A.; Alshehri, A.A.; Othman, B.; et al. Advances in nanotechnology-based therapeutics, drug delivery, diagnostics and vaccines: Multidimensional applications in tuberculosis disease management. Pharmaceuticals 2023, 16, 581. [Google Scholar] [CrossRef]
- Islam, M.N.; Mili, M.A.; Jahan, I.; Chakma, C.; Munalisa, R. Immunological and neurological signatures of the co-infection of HIV and HTLV: Current insights and future perspectives. Viruses 2025, 17, 545. [Google Scholar] [CrossRef]
- Abdul-Ghani, R.; Al-Awadi, A.; Al-Aghbari, N.; Al-Mikhlafy, A.A.; Abdulmoghni, S.S.; Al-Dobai, S.S.; Nauman, N.F. Latent tuberculosis infection and diagnostic performance of the tuberculin skin test among type 2 diabetics in Sana’a city, Yemen. BMC Infect. Dis. 2024, 24, 1005. [Google Scholar] [CrossRef]
- Chen, Q.; Ren, N.; Liu, S.; Qian, Z.; Li, M.; Mustapha, A.; Luo, W.; Li, J.; Wang, W.; Hao, C. Prevalence of tuberculosis among migrants under national screening programs: A systematic review and meta-analysis. Glob. Health Res. Policy 2025, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Lange, C.; Dheda, K.; Chesov, D.; Mandalakas, A.M.; Udwadia, Z.; Horsburgh, C.R. Management of drug-resistant tuberculosis. Lancet 2019, 394, 953–966. [Google Scholar] [CrossRef]
- Zein-Eddine, R.; Ramuz, M.; Refrégier, G.; Lutzeyer, J.F.; Aleksandrov, A.; Myllykallio, H. Key challenges in tuberculosis drug discovery: What does the future hold? Expert Opin. Drug Discov. 2025, 20, 1115–1130. [Google Scholar] [CrossRef]
- Tiberi, S.; Utjesanovic, N.; Galvin, J.; Centis, R.; D’AMbrosio, L.; Boom, M.v.D.; Zumla, A.; Migliori, G.B. Drug-resistant TB: Latest developments in epidemiology, diagnostics and management. Int. J. Infect. Dis. 2022, 124, S20–S25. [Google Scholar] [CrossRef] [PubMed]
- Patra, J.; Irving, H.; Maini, P.; Liang, J.; Patra, A.; Paradkar, M.; Rehm, J. Treatment outcomes among children and adolescents with XDR and pre-XDR tuberculosis: A systematic review and meta-analysis. PLoS Glob. Public Health 2025, 5, e0003754. [Google Scholar] [CrossRef]
- Wallis, R.S.; Kim, P.; Cole, S.; Hanna, D.; Andrade, B.B.; Maeurer, M.; Schito, M.; Zumla, A. Tuberculosis biomarker discovery: Developments, needs, and challenges. Lancet Infect. Dis. 2013, 13, 362–372. [Google Scholar] [CrossRef]
- Abdallah, E.M.; Alhatlani, B.Y.; Menezes, R.d.P.; Martins, C.H.G. Back to nature: Medicinal plants as promising sources for antibacterial drugs in the post-antibiotic era. Plants 2023, 12, 3077. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 2007, 70, 461–477. [Google Scholar] [CrossRef]
- Gangwal, A.; Lavecchia, A. Artificial intelligence in natural product drug discovery: Current applications and future perspectives. J. Med. Chem. 2025, 68, 3948–3969. [Google Scholar] [CrossRef]
- Neumann, A.; Marrison, L.; Klein, R. Relevance of the trillion-sized chemical space “eXplore” as a source for drug discovery. ACS Med. Chem. Lett. 2023, 14, 466–472. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Gelpi, J.; Hospital, A.; Goñi, R.; Orozco, M. Molecular dynamics simulations: Advances and applications. Adv. Appl. Bioinform. Chem. 2015, 8, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W. Computational methods in drug discovery. Pharmacol. Rev. 2014, 66, 334–395. [Google Scholar] [CrossRef] [PubMed]
- Rose, P.W. The RCSB Protein Data Bank: Integrative view of protein, gene and 3D structural information. Nucleic Acids Res. 2016, 45, D271–D281. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Bragina, M.E.; Daina, A.; Perez, M.A.S.; Michielin, O.; Zoete, V. SwissSimilarity 2021: Novel chemical libraries and additional methods for ligand-based virtual screening. Int. J. Mol. Sci. 2022, 23, 811. [Google Scholar] [CrossRef]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J. Cheminform. 2011, 3, 33. [Google Scholar] [CrossRef]
- Halgren, T.A. Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J. Comput. Chem. 1996, 17, 490–519. [Google Scholar] [CrossRef]
- Lim, V.T.; Hahn, D.F.; Tresadern, G.; Bayly, C.I.; Mobley, D.L. Benchmark assessment of molecular geometries and energies from small molecule force fields. F1000Research 2020, 9, 1390. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Abdullahi, S.H.; Moin, A.T.; Uzairu, A.; Umar, A.B.; Ibrahim, M.T.; Usman, M.T.; Nawal, N.; Bayil, I.; Zubair, T. Molecular docking and MD simulation studies of benzoxazole and benzothiazole derivatives as VEGFR-2 inhibitors. Intell. Pharm. 2024, 2, 232–250. [Google Scholar] [CrossRef]
- Stielow, M.; Witczyńska, A.; Kubryń, N.; Fijałkowski, Ł.; Nowaczyk, J.; Nowaczyk, A. Bioavailability of drugs: Current state of knowledge. Molecules 2023, 28, 8038. [Google Scholar] [CrossRef] [PubMed]
- Piton, J.; Petrella, S.; Delarue, M.; André-Leroux, G.; Jarlier, V.; Aubry, A.; Mayer, C. Structural insights into the quinolone resistance mechanism of Mycobacterium tuberculosis DNA gyrase. PLoS ONE 2010, 5, e12245. [Google Scholar] [CrossRef]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A.F.; Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model. 2021, 61, 3891–3898. [Google Scholar] [CrossRef]
- Sharma, S.; Sharma, A.; Gupta, U. Molecular docking studies on antifungal activity of Allium sativum (garlic) against mucormycosis. Res. Sq. 2021. preprint. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera: A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Irwin, J.J.; Shoichet, B.K. ZINC: A free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 2005, 45, 177–182. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multilevel parallelism. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Bugnon, M.; Goullieux, M.; Röhrig, U.F.; Perez, M.A.S.; Daina, A.; Michielin, O.; Zoete, V. SwissParam 2023: A web tool for small molecule parametrization. J. Chem. Inf. Model. 2023, 63, 6469–6475. [Google Scholar] [CrossRef]
- Zoete, V.; Cuendet, M.A.; Grosdidier, A.; Michielin, O. SwissParam: A fast force field generation tool for small organic molecules. J. Comput. Chem. 2011, 32, 2359–2368. [Google Scholar] [CrossRef] [PubMed]
- Brooks, B.R.; Brooks, C.L., III; MacKerell, A.D., Jr.; Nilsson, L.; Petrella, R.J.; Roux, B.; Won, Y.; Archontis, G.; Bartels, C.; Boresch, S.; et al. CHARMM: The biomolecular simulation program. J. Comput. Chem. 2009, 30, 1545–1614. [Google Scholar] [CrossRef] [PubMed]
- Abascal, J.L.F.; Vega, C. A general purpose model for the condensed phases of water: TIP4P/2005. J. Chem. Phys. 2005, 123, 234505. [Google Scholar] [CrossRef] [PubMed]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 2008, 4, 116–122. [Google Scholar] [CrossRef]
- Sittel, F.; Jain, A.; Stock, G. Principal component analysis of molecular dynamics: On the use of Cartesian vs internal coordinates. J. Chem. Phys. 2014, 141, 014111. [Google Scholar] [CrossRef]
- Papaleo, E.; Mereghetti, P.; Fantucci, P.; Grandori, R.; De Gioia, L. Free-energy landscape, PCA, and structural clustering to identify conformations from MD: Myoglobin case. J. Mol. Graph. Model. 2009, 27, 889–899. [Google Scholar] [CrossRef]
- Kumari, R.; Kumar, R.; Lynn, A.; OSDD Consortium. g_mmpbsa: A GROMACS tool for MM-PBSA calculations. J. Chem. Inf. Model. 2014, 54, 1951–1962. [Google Scholar] [CrossRef]
- Wang, E.; Sun, H.; Wang, J.; Wang, Z.; Liu, H.; Zhang, J.Z.H.; Hou, T. End-point binding free energy calculation with MM/PBSA and MM/GBSA: Strategies and applications. Chem. Rev. 2019, 119, 9478–9508. [Google Scholar] [CrossRef] [PubMed]
- Paul, L.; Shadrack, D.M.; Mudogo, C.N.; Mtei, K.M.; Machunda, R.L.; Ntie-Kang, F. Structural characterization of cassava linamarase–linamarin complex: A computational approach. J. Biomol. Struct. Dyn. 2022, 40, 9270–9278. [Google Scholar] [CrossRef] [PubMed]
- Baum, B.; Muley, L.; Smolinski, M.; Heine, A.; Hangauer, D.; Klebe, G. Non-additivity of Functional Group Contributions in Protein–Ligand Binding: A Comprehensive Study by Crystallography and Isothermal Titration Calorimetry. J. Mol. Biol. 2010, 397, 1042–1054. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.D.; Kim, M.-S. Identification of promising inhibitory heterocyclic compounds against acetylcholinesterase using QSAR, ADMET, biological activity, and molecular docking. Comput. Biol. Chem. 2023, 104, 107872. [Google Scholar] [CrossRef]
- Dias, M.V.B.; Vasconcelos, I.B.; Prado, A.M.X.; Fadel, V.; Basso, L.A.; de Azevedo, W.F.; Santos, D.S. Binding of isonicotinyl-NAD adduct to wild-type and isoniazid-resistant InhA: Crystallographic studies. J. Struct. Biol. 2007, 159, 369–380. [Google Scholar] [CrossRef]
- Sucharitha, P.; Ramesh Reddy, K.; Satyanarayana, S.V.; Garg, T. ADMET assessment of drugs using computational tools. In Computational Approaches for Novel Therapeutic and Diagnostic Designing to Mitigate SARS-CoV-2 Infection; Academic Press: Cambridge, MA, USA, 2022; pp. 335–355. [Google Scholar] [CrossRef]
- Alsenan, S.; Al-Turaiki, I.; Hafez, A. A deep learning approach to predict blood–brain barrier permeability. PeerJ Comput. Sci. 2021, 7, e515. [Google Scholar] [CrossRef]
- Veber, D.F.; Johnson, S.R.; Cheng, H.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef]
- Guterres, H.; Im, W. Improving protein–ligand docking results with high-throughput molecular dynamics simulations. J. Chem. Inf. Model. 2020, 60, 2189–2198. [Google Scholar] [CrossRef]
- Liu, K.; Watanabe, E.; Kokubo, H. Exploring stability of ligand binding modes by molecular dynamics simulations. J. Comput. Aided Mol. Des. 2017, 31, 201–211. [Google Scholar] [CrossRef]
- Cao, X.; Hummel, M.H.; Wang, Y.; Simmerling, C.; Coutsias, E.A. Exact analytical algorithm for solvent-accessible surface area and derivatives in implicit solvent simulations on GPUs. J. Chem. Theory Comput. 2024, 20, 4456–4468. [Google Scholar] [CrossRef]
- Huang, H.; Simmerling, C. Fast pairwise approximation of solvent accessible surface area for implicit solvent simulations. J. Chem. Theory Comput. 2018, 14, 5797–5814. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.; Raghavan, V. Molecular dynamics simulation of the thermal treatment of the Ara h 6 peanut protein. Processes 2025, 13, 434. [Google Scholar] [CrossRef]
- Islam, M.T.; Aktaruzzaman, M.; Barai, C.; Rafi, F.I.; Hasan, A.R.; Tasnim, T.; Sarder, P.; Albadrani, G.M.; Al-Ghadi, M.Q.; Sayed, A.A.; et al. In silico screening of dietary compounds as butyrylcholinesterase inhibitors for Alzheimer’s disease. Sci. Rep. 2025, 15, 17134. [Google Scholar] [CrossRef]
- Maruyama, Y.; Igarashi, R.; Ushiku, Y.; Mitsutake, A. Analysis of protein folding simulation with moving RMSD. J. Chem. Inf. Model. 2023, 63, 1529–1541. [Google Scholar] [CrossRef]
- Mascoli, V.; Liguori, N.; Cupellini, L.; Elias, E.; Mennucci, B.; Croce, R. Interactions driving carotenoid binding in light-harvesting complexes. Chem. Sci. 2021, 12, 5113–5122. [Google Scholar] [CrossRef]
- Ragunathan, A.; Malathi, K.; Anbarasu, A. MurB as a target in addressing Vibrio cholerae resistance through docking and MD simulation. J. Cell. Biochem. 2018, 119, 1726–1732. [Google Scholar] [CrossRef]
- Bianco, V.; Iskrov, S.; Franzese, G. Hydrogen bonds in water dynamics and protein stability. J. Biol. Phys. 2012, 38, 27–48. [Google Scholar] [CrossRef]
- Qin, X.; Zhong, J.; Wang, Y. Homology modeling, docking, and MD simulation of a mutant T1 lipase with fatty acids. J. Biotechnol. 2021, 337, 24–34. [Google Scholar] [CrossRef]
- Abdelsattar, A.S.; Mansour, Y.; Aboul-ela, F. Perturbed free-energy landscape: Linking ligand binding to biomolecular folding. ChemBioChem 2021, 22, 1499–1516. [Google Scholar] [CrossRef]















| Compound | Binding Energy (Kcal/Mol) | Structures |
|---|---|---|
| ZINC000013629834 | −10.90 |  |
| ZINC000253411694 | −10.36 |  |
| ZINC000009418955 | −10.33 |  |
| ZINC000009306325 | −10.31 |  |
| ZINC000032851106 | −10.21 |  |
| ZINC000009304208 | −10.08 |  |
| ZINC000009548024 | −9.81 |  |
| ZINC000253411693 | −9.76 |  |
| ZINC000007039726 | −9.75 | 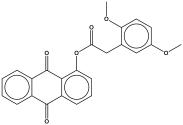 |
| Native ligand | −9.38 |  |
| Rifampicin | −9.05 |  |
| Compounds | Interactions with InhA | Distance for H Bonds (Å) |
|---|---|---|
| ZINC000013629834 | Conventional Hydrogen Bond: GLY96, SER94, THR196, ILE21 Van der Waals: ASP64, GLN66, VAL65, GLY14, ALA22, PRO193, TYR158, ILE194, MET199, PHE149, LYS165, MET147, ILE16, PHE97 Carbon Hydrogen Bond: ILE95, SER20 Pi-Sigma: PHE41, ILE122 | 3.2, 1.8, 2.9 and 3.2 |
| ZINC000253411694 | Conventional Hydrogen Bond: ILE21, THR196, SER20 Carbon Hydrogen Bond: ASP64 Pi-Sigma: ILE95 Pi-Pi Stacked: PHE41 Pi-Alkyl: ILE16, ILE122, VAL65 | 3.2, 2.8 and 3.0 |
| Rifampicin | Conventional Hydrogen Bond: MET98, SER94, ILE21 Van der Waals: ALA191, GLY192, TYR158, PHE149, LYS165, MET147, ALA22, SER20, GLY96, THR196, PHE97 Carbon Hydrogen Bond: ASP148, ILE194 Pi-Sigma: PRO193 | 3.1, 2.1 and 2.9 |
| Descriptor | ZINC000013629834 | ZINC000253411694 | Rifampicin |
|---|---|---|---|
| Total no. of atoms | 30 | 32 | 32 |
| Molecular weight (g/mol) | 406.39 | 432.00 | 437.40 |
| No. of H-bond acceptors | 7 | 5 | 8 |
| No. of H-bond donors | 3 | 3 | 2 |
| No. of rotatable bonds | 6 | 5 | 7 |
| TPSA (Å2) | 130.34 | 104.56 | 124.00 |
| LogP | 1.81 | 3.08 | 2.46 |
| LogS (ESOL) | −7.37 | −8.13 | −6.76 |
| GI absorbance | High | High | High |
| BBB permeant | No | No | No |
| LogKp (skin permeation) (cm/s) | −7.16 | −6.74 | −7.07 |
| Lipinski rule | 0 | 0 | 0 |
| Veber | 0 | 0 | 0 |
| Bioavailability score | 0.55 | 0.55 | 0.55 |
| Lead likeness | 1 | 1 | 1 |
| Synthetic accessibility | 3.55 | 4.06 | 3.91 |
| Complexes | ∆EvdW | ∆Eelect | ∆GPB | ∆GSASA | ∆Gbinding |
|---|---|---|---|---|---|
| ZINC000013629834 | −201.951 ± 0.974 | −67.275 ± 0.913 | 200.601 ± 1.300 | −21.560 ± 0.095 | −90.159 ± 1.499 |
| ZINC000253411694 | −185.814 ± 1.215 | −67.608 ± 0.878 | 183.415 ± 1.231 | −21.557 ± 0.114 | −91.646 ± 1.524 |
| Rifampicin | −216.680 ± 1.366 | −90.289 ± 1.775 | 239.704 ± 1.340 | −23.364 ± 0.092 | −90.734 ± 1.431 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Paul, L.; Paluch, A.S. Structure-Based Screening and Molecular Dynamics of Rifampicin Analogues Targeting InhA of Mycobacterium tuberculosis. ChemEngineering 2026, 10, 28. https://doi.org/10.3390/chemengineering10020028
Paul L, Paluch AS. Structure-Based Screening and Molecular Dynamics of Rifampicin Analogues Targeting InhA of Mycobacterium tuberculosis. ChemEngineering. 2026; 10(2):28. https://doi.org/10.3390/chemengineering10020028
Chicago/Turabian StylePaul, Lucas, and Andrew S. Paluch. 2026. "Structure-Based Screening and Molecular Dynamics of Rifampicin Analogues Targeting InhA of Mycobacterium tuberculosis" ChemEngineering 10, no. 2: 28. https://doi.org/10.3390/chemengineering10020028
APA StylePaul, L., & Paluch, A. S. (2026). Structure-Based Screening and Molecular Dynamics of Rifampicin Analogues Targeting InhA of Mycobacterium tuberculosis. ChemEngineering, 10(2), 28. https://doi.org/10.3390/chemengineering10020028







