Changes in miRNA Expression Profiling during Neuronal Differentiation and Methyl Mercury-Induced Toxicity in Human in Vitro Models
Abstract
:1. Introduction
2. Materials and Methods
2.1. H9 and NT-2 Cell Maintenance and Differentiation towards Neuronal Phenotype
2.2. Immunocytochemistry
2.3. Cell Viability Assay Using Alamar Blue (AB)
2.4. Exposure to Methyl Mercury Chloride
2.5. RNA Extraction and miRNAs Expression Analysis
2.6. Analysis of Stemness Marker Expression
2.7. Statistical Analysis of miRNA Expression
2.8. Gene Ontology (GO) Term Enrichment Analysis
3. Results
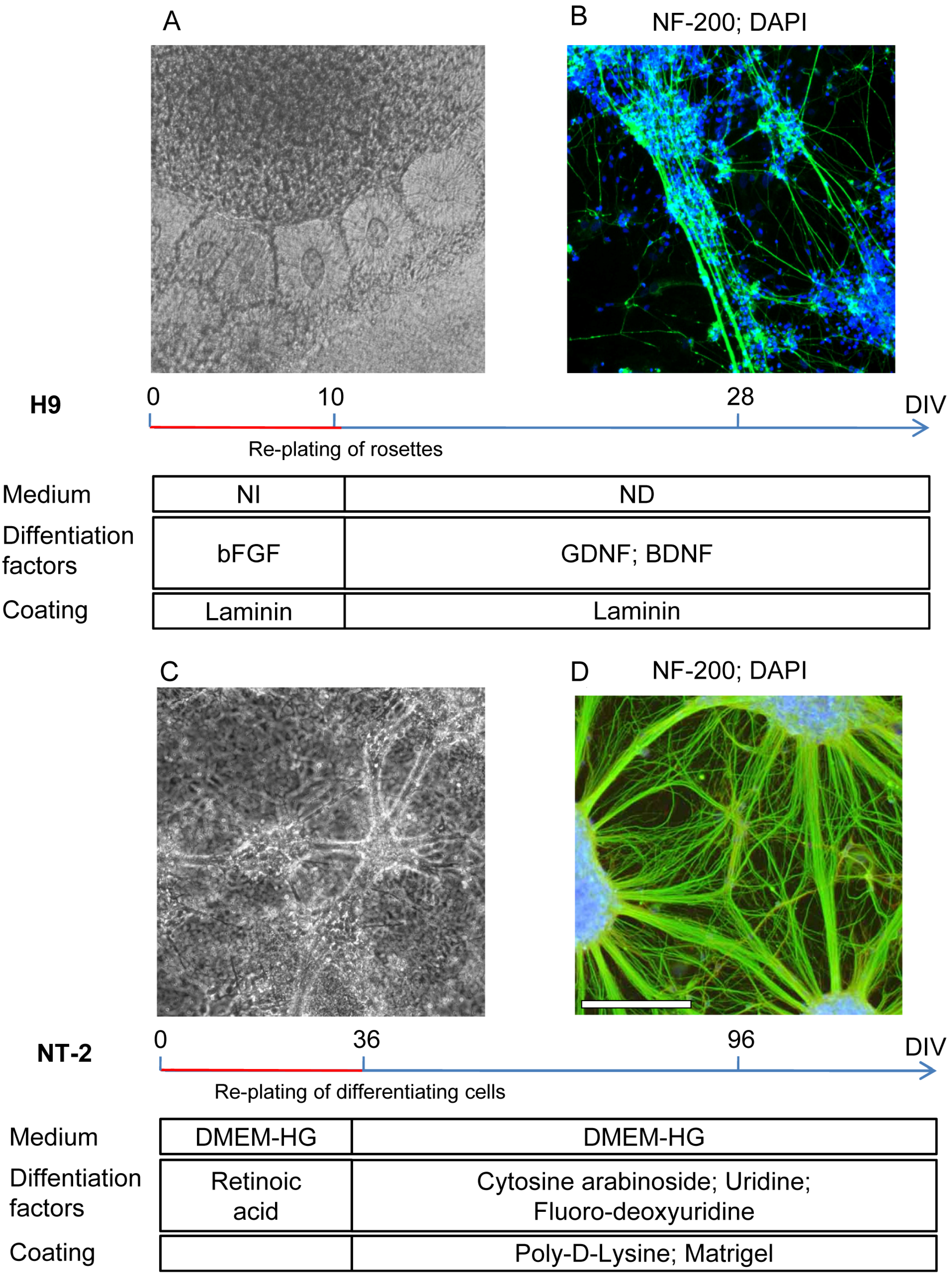
3.1. Characterization of H9 and NT2 Cells Differentiation towards Neuronal Phenotype
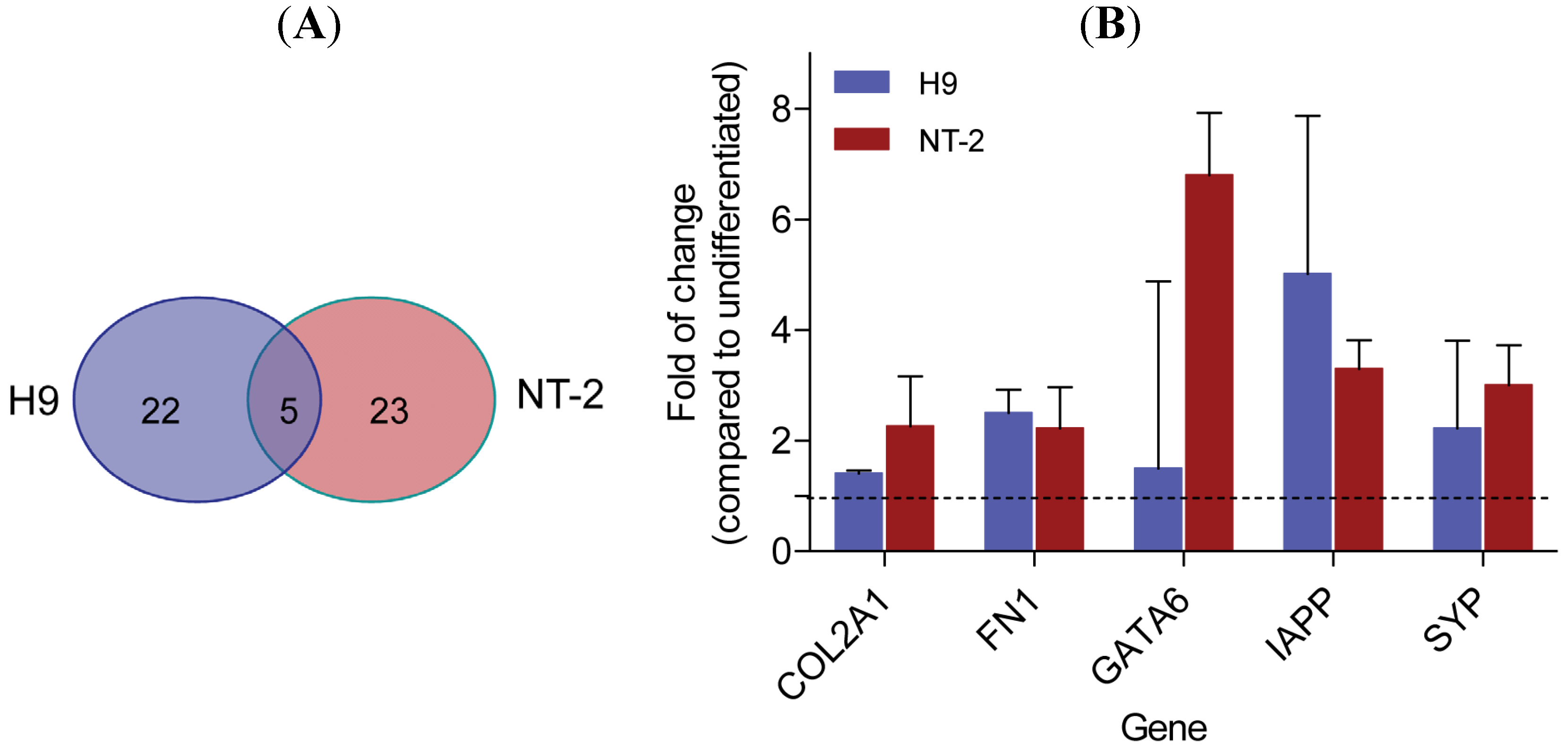
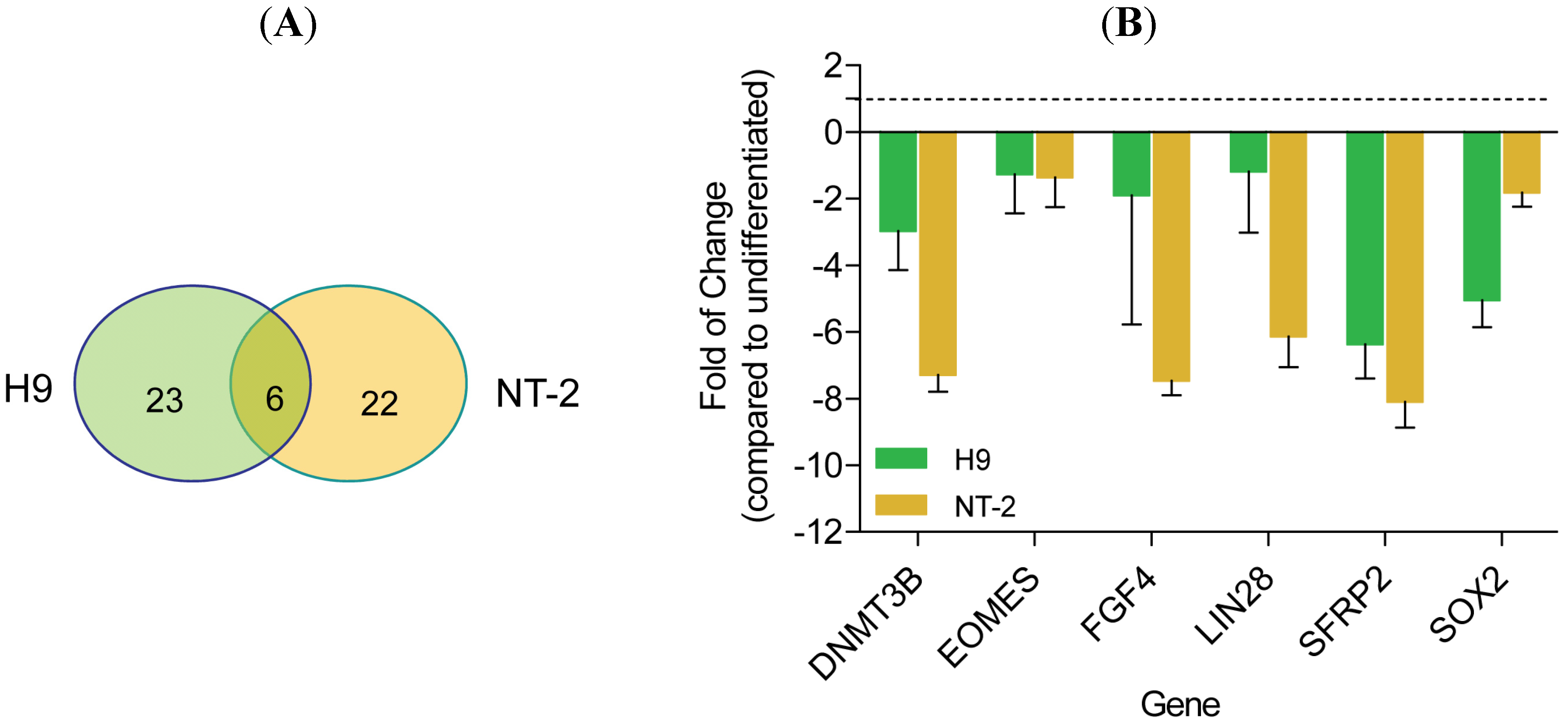
3.2. Gene Expression Analysis Comparing Neuronal Committed H9 and Neuronal Committed NT-2 Cells
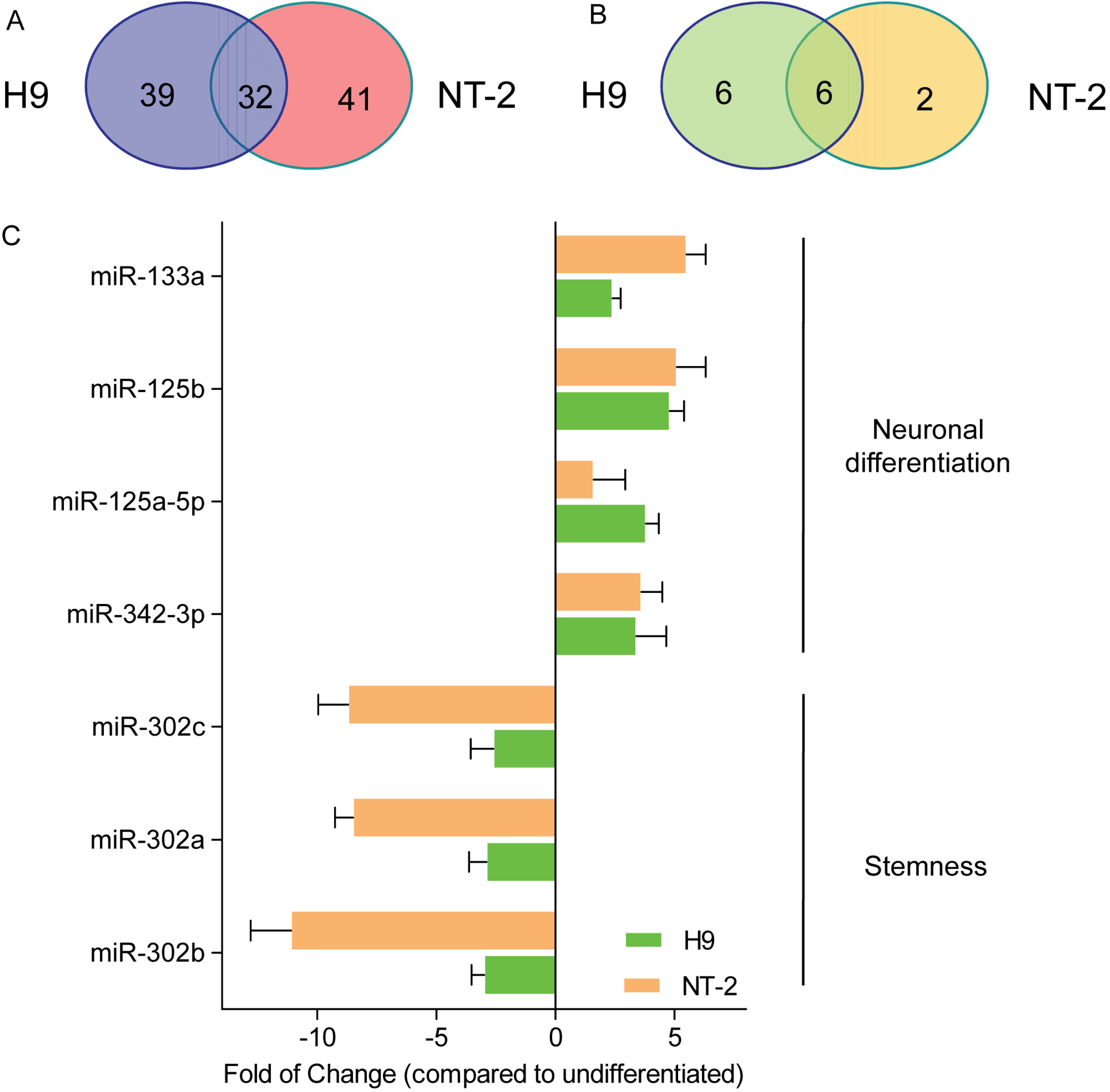
3.3. Comparison of miRNA Expression Profiles between Neuronal Committed H9 and Neuronal Committed NT-2 Cells
3.4. Analysis of miRNA Expression Profile Perturbations Comparing Neuronal Committed H9 and Neuronal Committed NT-2 Cells Exposed to MeHgCl
| Cell Line | MeHgCl Treatment | miRNA | FC | p-value |
|---|---|---|---|---|
| H9 | Time: DIV 1–10; Concentration: 25 nM | miR-183 | 2.82 | <0.05 |
| miR-15a | 19 | <0.05 | ||
| miR-526b* | 6.5 | <0.05 | ||
| NT-2 | Time: DIV 2–36 Concentration: 400 nM | miRNA-141 | 2.5 | <0.05 |
| miRNA-196b | 3.2 | <0.05 | ||
| miRNA-367 | 6.1 | <0.05 | ||
| miRNA-302b | 5.3 | <0.05 | ||
| miRNA-372 | 9.5 | <0.05 | ||
| miR-302a | 3.7 | <0.05 | ||
| miR-302c | 3.1 | <0.05 | ||
| miR-142 | 2.4 | <0.05 | ||
| miR-200c | 2.3 | <0.05 | ||
| miR-296 | −1.5 | <0.05 | ||
| miR-217* | −4.4 | <0.05 | ||
| miR-655 | −2.5 | <0.05 | ||
| miR-365 | −1.3 | <0.05 | ||
| miR-181a | −1.5 | <0.05 |
3.5. Gene ontology (GO) Term Enrichment Analysis of Deregulated miRNAs Induced by the Exposure to MeHgCl
| Upregulated miRNA-Biological Process GO-terms in NT-2 Cell Line | p-value |
|---|---|
| regulation of cellular macromolecule biosynthetic process (GO:2000112) | 7.70 × 10−30 |
| regulation of cellular metabolic process (GO:0031323) | 1.02 × 10−29 |
| regulation of cellular process (GO:0050794) | 1.27 × 10−29 |
| regulation of macromolecule biosynthetic process (GO:0010556) | 3.83 × 10−30 |
| regulation of macromolecule metabolic process (GO:0060255) | 3.78 × 10−29 |
| regulation of nucleobase-containing compound metabolic process (GO:0019219) | 7.70 × 10−30 |
| regulation of RNA metabolic process (GO:0051252) | 5.38 × 10−29 |
| regulation of transcription from RNA polymerase II promoter (GO:0006357) | 4.63 × 10−30 |
| regulation of transcription, DNA-dependent (GO:0006355) | 4.36 × 10−29 |
| transcription from RNA polymerase II promoter (GO:0006366) | 3.83 × 10−30 |
| Upregulated miRNA-Biological Process GO-terms in H9 Cell Line | p-value |
| anatomical structure morphogenesis (GO:0009653) | 2.45 × 10−7 |
| cellular protein modification process (GO:0006464) | 2.92 × 10−7 |
| generation of neurons (GO:0048699) | 1.58 × 10−8 |
| nervous system development (GO:0007399) | 1.58 × 10−8 |
| neurogenesis (GO:0022008) | 1.58 × 10−8 |
| neuron differentiation (GO:0030182) | 2.53 × 10−7 |
| protein modification process (GO:0036211) | 2.92 × 10−7 |
| regulation of cell development (GO:0060284) | 1.07 × 10−7 |
| regulation of cell differentiation (GO:0045595) | 2.69 × 10−7 |
| regulation of developmental process (GO:0050793) | 2.69 × 10−7 |
| Upregulated miRNA-Pathways in NT-2 Cell Line | p-value | Upregulated miRNA-Pathways in H9 Cell Line | p-value |
|---|---|---|---|
| Axon guidance | 7.13 × 10−10 | Cell cycle | 4.36 × 10−6 |
| Pathways in cancer | 4.86 × 10−15 | Axon guidance | 2.71 × 10−5 |
| Oocyte meiosis | 3.85 × 10−10 | TGF-beta signaling pathway | 2.46 × 10−5 |
| MAPK signaling pathway | 3.73 × 10−11 | p53 signaling pathway | 2.00 × 10−4 |
| Neurotrophin signaling pathway | 3.38 × 10−14 | Glycerophospholipid metabolism | 1.00 × 10−4 |
| Renal cell carcinoma | 1.89 × 10−8 | Oocyte meiosis | 8.25 × 10−6 |
| Wnt signaling pathway | 1.70 × 10−9 | Pathways in cancer | 1.75 × 10−10 |
| Focal adhesion | 9.74 × 10−10 | Neurotrophin signaling pathway | 1.22 × 10−6 |
| Ubiquitin mediated proteolysis | 6.55 × 10−9 | Wnt signaling pathway | 1.22 × 10−6 |
| TGF-beta signaling pathway | 4.94 × 10−9 | Small cell lung cancer | 1.00 × 10−4 |
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Christensen, M.; Schratt, G.M. microRNA involvement in developmental and functional aspects of the nervous system and in neurological diseases. Neurosci. Lett. 2009, 466, 55–62. [Google Scholar]
- Krichevsky, A.M.; King, K.S.; Donahue, C.P.; Khrapko, K.; Kosik, K.S. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 2003, 9, 1274–1281. [Google Scholar]
- Miska, E.A.; Alvarez-Saavedra, E.; Townsend, M.; Yoshii, A.; Sestan, N.; Rakic, P.; Constantine-Paton, M.; Horvitz, H.R. Microarray analysis of microRNA expression in the developing mammalian brain. Genome Biol. 2004, 5, R68. [Google Scholar]
- Krek, A.; Grun, D.; Poy, M.N.; Wolf, R.; Rosenberg, L.; Epstein, E.J.; MacMenamin, P.; da Piedade, I.; Gunsalus, K.C.; Stoffel, M.; et al. Combinatorial microRNA target predictions. Nat. Genet. 2005, 37, 495–500. [Google Scholar]
- Kim, J.; Inoue, K.; Ishii, J.; Vanti, W.B.; Voronov, S.V.; Murchison, E.; Hannon, G.; Abeliovich, A. A MicroRNA feedback circuit in midbrain dopamine neurons. Science 2007, 317, 1220–1224. [Google Scholar]
- De Pietri Tonelli, D.; Pulvers, J.N.; Haffner, C.; Murchison, E.P.; Hannon, G.J.; Huttner, W.B. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development 2008, 135, 3911–3921. [Google Scholar]
- Giraldez, A.J.; Cinalli, R.M.; Glasner, M.E.; Enright, A.J.; Thomson, J.M.; Baskerville, S.; Hammond, S.M.; Bartel, D.P.; Schier, A.F. MicroRNAs regulate brain morphogenesis in zebrafish. Science 2005, 308, 833–838. [Google Scholar]
- Delaloy, C.; Liu, L.; Lee, J.A.; Su, H.; Shen, F.; Yang, G.Y.; Young, W.L.; Ivey, K.N.; Gao, F.B. MicroRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Stem Cell 2010, 6, 323–335. [Google Scholar]
- Maller Schulman, B.R.; Liang, X.; Stahlhut, C.; DelConte, C.; Stefani, G.; Slack, F.J. The let-7 microRNA target gene, Mlin41/Trim71 is required for mouse embryonic survival and neural tube closure. Cell Cycle 2008, 7, 3935–3942. [Google Scholar]
- Makeyev, E.V.; Zhang, J.; Carrasco, M.A.; Maniatis, T. The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 2007, 27, 435–448. [Google Scholar]
- Schratt, G. Fine-tuning neural gene expression with microRNAs. Curr. Opin. Neurobiol. 2009, 19, 213–219. [Google Scholar]
- Vo, N.; Klein, M.E.; Varlamova, O.; Keller, D.M.; Yamamoto, T.; Goodman, R.H.; Impey, S. A cAMP-response element binding protein-induced microRNA regulates neuronal morphogenesis. Proc. Natl. Acad. Sci. USA 2005, 102, 16426–16431. [Google Scholar]
- Konopka, W.; Kiryk, A.; Novak, M.; Herwerth, M.; Parkitna, J.R.; Wawrzyniak, M.; Kowarsch, A.; Michaluk, P.; Dzwonek, J.; Arnsperger, T.; et al. MicroRNA loss enhances learning and memory in mice. J. Neurosci. 2010, 30, 14835–14842. [Google Scholar]
- Wang, J.; Duncan, D.; Shi, Z.; Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): Update 2013. Nucleic Acids Res. 2013, 41, W77–W83. [Google Scholar]
- Kaur, P.; Armugam, A.; Jeyaseelan, K. MicroRNAs in neurotoxicity. J. Toxicol. 2012, 2012, 870150. [Google Scholar] [CrossRef]
- Lee, Y.; Samaco, R.C.; Gatchel, J.R.; Thaller, C.; Orr, H.T.; Zoghbi, H.Y. miR-19, miR-101 and miR-130 co-regulate ATXN1 levels to potentially modulate SCA1 pathogenesis. Nat. Neurosci. 2008, 11, 1137–1139. [Google Scholar]
- Nelson, P.T.; Wang, W.X. MiR-107 is reduced in Alzheimer’s disease brain neocortex: Validation study. J. Alzheimer’s Dis. 2010, 21, 75–79. [Google Scholar]
- Grandjean, P.; Landrigan, P.J. Developmental neurotoxicity of industrial chemicals. Lancet 2006, 368, 2167–2178. [Google Scholar]
- Hogberg, H.T.; Kinsner-Ovaskainen, A.; Coecke, S.; Hartung, T.; Bal-Price, A.K. mRNA expression is a relevant tool to identify developmental neurotoxicants using an in vitro approach. Toxicol. Sci. 2010, 113, 95–115. [Google Scholar]
- Hogberg, H.T.; Kinsner-Ovaskainen, A.; Hartung, T.; Coecke, S.; Bal-Price, A.K. Gene expression as a sensitive endpoint to evaluate cell differentiation and maturation of the developing central nervous system in primary cultures of rat cerebellar granule cells (CGCs) exposed to pesticides. Toxicol. Appl. Pharmacol. 2009, 235, 268–286. [Google Scholar]
- Claudio, L.; Kwa, W.C.; Russell, A.L.; Wallinga, D. Testing methods for developmental neurotoxicity of environmental chemicals. Toxicol. Appl. Pharmacol. 2000, 164, 1–14. [Google Scholar]
- Eriksson, P. Developmental neurotoxicity of environmental agents in the neonate. Neurotoxicology 1997, 18, 719–726. [Google Scholar]
- Rodier, P.M. Developing brain as a target of toxicity. Environ. Health Perspect. 1995, 103, 73–76. [Google Scholar]
- Tilson, H.A. Neurotoxicology risk assessment guidelines: Developmental neurotoxicology. Neurotoxicology 2000, 21, 189–194. [Google Scholar]
- Buzanska, L.; Sypecka, J.; Nerini-Molteni, S.; Compagnoni, A.; Hogberg, H.T.; del Torchio, R.; Domanska-Janik, K.; Zimmer, J.; Coecke, S. A human stem cell-based model for identifying adverse effects of organic and inorganic chemicals on the developing nervous system. Stem Cells 2009, 27, 2591–2601. [Google Scholar]
- Stummann, T.C.; Hareng, L.; Bremer, S. Hazard assessment of methylmercury toxicity to neuronal induction in embryogenesis using human embryonic stem cells. Toxicology 2009, 257, 117–126. [Google Scholar]
- Balmer, N.V.; Weng, M.K.; Zimmer, B.; Ivanova, V.N.; Chambers, S.M.; Nikolaeva, E.; Jagtap, S.; Sachinidis, A.; Hescheler, J.; Waldmann, T.; et al. Epigenetic changes and disturbed neural development in a human embryonic stem cell-based model relating to the fetal valproate syndrome. Hum. Mol. Genet. 2012, 21, 4104–4114. [Google Scholar]
- Klaric, M.; Winkler, J.; Vojnits, K.; Meganathan, K.; Jagtap, S.; Ensenat-Waser, R.; Hescheler, J.; Sachinidis, A.; Bremer-Hoffmann, S. Current status of human pluripotent stem cell based in vitro toxicity tests. Front. Biosci. Schol. Ed. 2013, 5, 118–133. [Google Scholar]
- Pistollato, F.; Bremer-Hoffmann, S.; Healy, L.; Young, L.; Stacey, G. Standardization of pluripotent stem cell cultures for toxicity testing. Expert Opin. Drug Metab. Toxicol. 2012, 8, 239–257. [Google Scholar]
- Scott, C.W.; Peters, M.F.; Dragan, Y.P. Human induced pluripotent stem cells and their use in drug discovery for toxicity testing. Toxicol. Lett. 2013, 219, 49–58. [Google Scholar]
- Bosnjak, Z.J. Developmental neurotoxicity screening using human embryonic stem cells. Exp. Neurol. 2012, 237, 207–210. [Google Scholar]
- Laurenza, I.; Pallocca, G.; Mennecozzi, M.; Scelfo, B.; Pamies, D.; Bal-Price, A. A human pluripotent carcinoma stem cell-based model for in vitro developmental neurotoxicity testing: Effects of methylmercury, lead and aluminum evaluated by gene expression studies. Int. J. Dev. Neurosci. 2013, 31, 679–691. [Google Scholar]
- Hartley, R.S.; Margulis, M.; Fishman, P.S.; Lee, V.M.; Tang, C.M. Functional synapses are formed between human NTera2 (NT-2N, hNT) neurons grown on astrocytes. J. Comp. Neurol. 1999, 407, 1–10. [Google Scholar]
- Serra, M.; Leite, S.B.; Brito, C.; Costa, J.; Carrondo, M.J.; Alves, P.M. Novel culture strategy for human stem cell proliferation and neuronal differentiation. J. Neurosci. Res. 2007, 85, 3557–3566. [Google Scholar]
- Pistollato, F.; Louisse, J.; Scelfo, B.; Mennecozzi, M.; Accordi, B.; Basso, G.; Gaspar, A.J.; Zagoura, D.; Barilari, M.; Palosaari, T.; et al. Development of a pluripotent stem cell derived neuronal model to identify chemically induced pathway perturbations in relation to neurotoxicity: Effects of CREB pathway inhibition. Toxicol. Appl. Pharmacol. 2014. [Google Scholar] [CrossRef]
- O’Brien, J.; Wilson, I.; Orton, T.; Pognan, F. Investigation of the Alamar Blue (resazurin) fluorescent dye for the assessment of mammalian cell cytotoxicity. Eur. J. Biochem. 2000, 267, 5421–5426. [Google Scholar]
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J.; et al. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar]
- Dvinge, H.; Bertone, P. HTqPCR: High-throughput analysis and visualization of quantitative real-time PCR data in R. Bioinformatics 2009, 25, 3325–3326. [Google Scholar]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3. [Google Scholar] [CrossRef]
- Xiao, F.; Zuo, Z.; Cai, G.; Kang, S.; Gao, X.; Li, T. miRecords: An integrated resource or microRNA-target interactions. Nucleic Acids Res. 2009, 37, D105–D110. [Google Scholar]
- Nichols, J.; Zevnik, B.; Anastassiadis, K.; Niwa, H.; Klewe-Nebenius, D.; Chambers, I.; Schöler, H.; Smith, A. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 1998, 95, 379–391. [Google Scholar]
- Richards, M.; Tan, S.P.; Tan, J.H.; Chan, W.K.; Bongso, A. The transcriptome profile of human embryonic stem cells as defined by SAGE. Stem Cells 2004, 22, 51–64. [Google Scholar]
- Lavon, I.; Zrihan, D.; Granit, A.; Einstein, O.; Fainstein, N.; Cohen, M.A.; Zelikovitch, B.; Shoshan, Y.; Spektor, S.; Reubinoff, B.E.; et al. Gliomas display a microRNA expression profile reminiscent of neural precursor cells. Neuro-oncology 2010, 12, 422–433. [Google Scholar]
- Porrello, E.R.; Johnson, B.A.; Aurora, A.B.; Simpson, E.; Nam, Y.J.; Matkovich, S.J.; Dorn, G.W., 2nd; van Rooij, E.; Olson, E.N. MiR-15 family regulates postnatal mitotic arrest of cardiomyocytes. Circ. Res. 2011, 109, 670–679. [Google Scholar]
- Tay, Y.; Zhang, J.; Thomson, A.M.; Lim, B.; Rigoutsos, I. MicroRNAs to Nanog, Oct4 and SOX2 coding regions modulate embryonic stem cell differentiation. Nature 2008, 455, 1124–1128. [Google Scholar]
- Pallocca, G.; Fabbri, M.; Sacco, M.G.; Gribaldo, L.; Pamies, D.; Laurenza, I.; Bal-Price, A. miRNA expression profiling in a human stem cell-based model as a tool for developmental neurotoxicity testing. Cell Biol. Toxicol. 2013, 29, 239–257. [Google Scholar]
- Bal-Price, A.; Coecke, S.; Costa, L.; Crofton, K.M.; Fritsche, E.; Goldberg, A.; Grandjean, P.; Lein, P.J.; Li, A.; Lucchini, R.; et al. Advancing the science of developmental neurotoxicity (DNT): Testing for better safety evaluation. Altex 2012, 29, 202–215. [Google Scholar]
- Leist, M.; Hartung, T. Inflammatory findings on species extrapolations: Humans are definitely no 70-kg mice. Altex 2013, 30, 227–230. [Google Scholar]
- Paparella, M.; Daneshian, M.; Hornek-Gausterer, R.; Kinzl, M.; Mauritz, I.; Muhlegger, S. Uncertainty of testing methods—What do we (want to) know? Altex 2013, 30, 131–144. [Google Scholar] [CrossRef]
- Bal-Price, A.; Crofton, K.M.; Shafer, T.; Sachana, M.; Behl, M.; Forsby, A.; Hargreaves, A.; Landesmann, B.; Lein, P.J.; Louisse, J.; et al. Workshop report: Adverse outcome pathways (AOP) relevant to neurotoxicity critical rev in toxicology. Crit. Rev. Toxicol. 2014. submitted for publication. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Pallocca, G.; Fabbri, M.; Nerini-Molteni, S.; Pistollato, F.; Zagoura, D.; Sacco, M.G.; Gribaldo, L.; Bremer-Hoffmann, S.; Bal-Price, A. Changes in miRNA Expression Profiling during Neuronal Differentiation and Methyl Mercury-Induced Toxicity in Human in Vitro Models. Toxics 2014, 2, 443-463. https://doi.org/10.3390/toxics2030443
Pallocca G, Fabbri M, Nerini-Molteni S, Pistollato F, Zagoura D, Sacco MG, Gribaldo L, Bremer-Hoffmann S, Bal-Price A. Changes in miRNA Expression Profiling during Neuronal Differentiation and Methyl Mercury-Induced Toxicity in Human in Vitro Models. Toxics. 2014; 2(3):443-463. https://doi.org/10.3390/toxics2030443
Chicago/Turabian StylePallocca, Giorgia, Marco Fabbri, Silvia Nerini-Molteni, Francesca Pistollato, Dimitra Zagoura, Maria Grazia Sacco, Laura Gribaldo, Susanne Bremer-Hoffmann, and Anna Bal-Price. 2014. "Changes in miRNA Expression Profiling during Neuronal Differentiation and Methyl Mercury-Induced Toxicity in Human in Vitro Models" Toxics 2, no. 3: 443-463. https://doi.org/10.3390/toxics2030443
APA StylePallocca, G., Fabbri, M., Nerini-Molteni, S., Pistollato, F., Zagoura, D., Sacco, M. G., Gribaldo, L., Bremer-Hoffmann, S., & Bal-Price, A. (2014). Changes in miRNA Expression Profiling during Neuronal Differentiation and Methyl Mercury-Induced Toxicity in Human in Vitro Models. Toxics, 2(3), 443-463. https://doi.org/10.3390/toxics2030443




