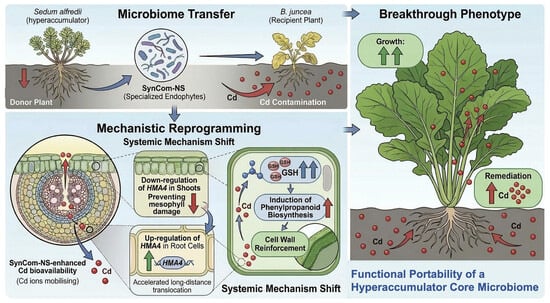
Functional Portability of a Hyperaccumulator-Derived Core Microbiome: Enhancing Cadmium Phytoextraction in Brassica juncea L. Through Molecular Reprogramming
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Synthetic Microbial Community Assembly
2.3. Greenhouse Pot Experiment and Experimental Design
2.4. Quantification of Heavy Metal Concentrations
2.5. Biochemical Evaluation of the Plant Antioxidant Defense System
2.6. RNA Extraction and Comparative Transcriptomics
2.7. Quantitative Real Time PCR Validation
2.8. Statistical Analysis and Data Visualization
3. Results
3.1. Effects of SynCom-NS Inoculation on Host Plant Growth
3.2. Influence of SynCom-NS on Cadmium Uptake and Systemic Translocation
3.3. Modulation of the Host Antioxidant Defense System
3.4. Comparative Transcriptomics and Quantitative Real Time PCR Validation
3.5. Functional Enrichment Analysis of Differentially Expressed Genes
4. Discussion
4.1. Enhancing Non-Host Metal Tolerance Through Synthetic Community Application
4.2. Microbially Driven Reprogramming of Host Metal Homeostasis and Detoxification
4.3. Transcriptomic Signatures of the Extended Phenotype in the Non Host Recipient
4.4. Practical Implications, Remediation Targets, and Study Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hou, D.; Jia, X.; Wang, L.; McGrath, S.P.; Zhu, Y.-G.; Hu, Q.; Zhao, F.-J.; Bank, M.S.; O’Connor, D.; Nriagu, J. Global soil pollution by toxic metals threatens agriculture and human health. Science 2025, 388, 316–321. [Google Scholar] [CrossRef]
- Wang, P.; Chen, H.; Kopittke, P.M.; Zhao, F.-J. Cadmium contamination in agricultural soils of China and the impact on food safety. Environ. Pollut. 2019, 249, 1038–1048. [Google Scholar] [CrossRef]
- Durand, A.; Leglize, P.; Benizri, E. Are endophytes essential partners for plants and what are the prospects for metal phytoremediation? Plant Soil 2021, 460, 1–30. [Google Scholar] [CrossRef]
- Chi, Y.; Ma, X.; Chu, S.; You, Y.; Chen, X.; Wang, J.; Wang, R.; Zhang, X.; Zhang, D.; Zhao, T.; et al. Nitrogen cycle induced by plant growth-promoting rhizobacteria drives “microbial partners” to enhance cadmium phytoremediation. Microbiome 2025, 13, 113. [Google Scholar] [CrossRef]
- Tian, Z.; Li, G.; Tang, W.; Zhu, Q.; Li, X.; Du, C.; Li, C.; Li, J.; Zhao, C.; Zhang, L. Role of Sedum alfredii and soil microbes in the remediation of ultra-high content heavy metals contaminated soil. Agric. Ecosyst. Environ. 2022, 339, 108090. [Google Scholar] [CrossRef]
- Simmer, R.A.; Schnoor, J.L. Phytoremediation, Bioaugmentation, and the Plant Microbiome. Environ. Sci. Technol. 2022, 56, 16602–16610. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, S.; Bhatt, A. Microbial Endophytes: Emerging Trends and Biotechnological Applications. Curr. Microbiol. 2023, 80, 249. [Google Scholar] [CrossRef]
- Goryluk-Salmonowicz, A.; Myczka, A.W.; Popowska, M. Antibiotic- and metal-resistant endophytes inhabit Armeria maritima hyperaccumulator. Plant Soil 2024, 495, 57–76. [Google Scholar] [CrossRef]
- Beiralas, R.; Ozer, N.; Segev, E. Abundant Sulfitobacter marine bacteria protect Emiliania huxleyi algae from pathogenic bacteria. ISME Commun. 2023, 3, 100. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Jiang, M. Aluminium-resistant bacteria boost crop yield in acidic soils. Nat. Food 2023, 4, 839–840. [Google Scholar] [CrossRef]
- Sun, Y.; Li, Y.; Chang, J.; Tian, L.; Ji, L.; Guo, L.; Gao, Q.; Veen, J.A.v.; Tian, C. Chemotaxis mediates nitrogen acquisition of maize under long-term nitrogen input. Soil Biol. Biochem. 2023, 184, 109118. [Google Scholar] [CrossRef]
- Jing, J.; Garbeva, P.; Raaijmakers, J.M.; Medema, M.H. Strategies for tailoring functional microbial synthetic communities. ISME J. 2024, 18, wrae049. [Google Scholar] [CrossRef]
- Huang, L.; Fan, Z.; Hu, Z.; Li, Z.; Fu, Y.; Wang, Q.; Lin, X.; Feng, Y. Synthetic communities derived from the core endophytic microbiome of hyperaccumulators and their role in cadmium phytoremediation. Microbiome 2024, 12, 236. [Google Scholar] [CrossRef]
- Ma, K.-W.; Niu, Y.; Jia, Y.; Ordon, J.; Copeland, C.; Emonet, A.; Geldner, N.; Guan, R.; Stolze, S.C.; Nakagami, H.; et al. Coordination of microbe-host homeostasis by crosstalk with plant innate immunity. Nat. Plants 2021, 7, 814–825. [Google Scholar] [CrossRef]
- Wippel, K.; Tao, K.; Niu, Y.; Zgadzaj, R.; Kiel, N.; Guan, R.; Dahms, E.; Zhang, P.; Jensen, D.B.; Logemann, E.; et al. Host preference and invasiveness of commensal bacteria in the Lotus and Arabidopsis root microbiota. Nat. Microbiol. 2021, 6, 1150–1162. [Google Scholar] [CrossRef]
- Singh, A.; Kumar, M.; Chakdar, H.; Pandiyan, K.; Kumar, S.C.; Zeyad, M.T.; Singh, B.N.; Ravikiran, K.T.; Mahto, A.; Srivastava, A.K.; et al. Influence of host genotype in establishing root associated microbiome of indica rice cultivars for plant growth promotion. Front. Microbiol. 2022, 13, 1033158. [Google Scholar] [CrossRef]
- Francis, J.S.; Mueller, T.G.; Vannette, R.L. Intraspecific variation in realized dispersal probability and host quality shape nectar microbiomes. New Phytol. 2023, 240, 1233–1245. [Google Scholar] [CrossRef]
- Custer, G.F.; Bresciani, L.; Dini-Andreote, F. Toward an integrative framework for microbial community coalescence. Trends Microbiol. 2024, 32, 241–251. [Google Scholar] [CrossRef]
- GB 15618-2018; Soil Environmental Quality Risk Control Standard for Soil Contamination of Agricultural Land. Ministry of Ecology and Environment of the People’s Republic of China: Beijing, China, 2018.
- Wang, Q.; Zhou, Q.; Huang, L.; Xu, S.a.; Fu, Y.; Hou, D.; Feng, Y.; Yang, X. Cadmium phytoextraction through Brassica juncea L. under different consortia of plant growth-promoting bacteria from different ecological niches. Ecotoxicol. Environ. Saf. 2022, 237, 113541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gilbert, J.A.; Liu, X.; Nie, L.; Xu, X.; Gao, G.; Lyu, L.; Ma, Y.; Fan, K.; Yang, T.; et al. SynCom-mediated herbicide degradation activates microbial carbon metabolism in soils. iMeta 2025, 4, e70058. [Google Scholar] [CrossRef] [PubMed]
- Clagnan, E.; Cucina, M.; De Nisi, P.; Dell’Orto, M.; D’Imporzano, G.; Kron-Morelli, R.; Llenas-Argelaguet, L.; Adani, F. Effects of the application of microbiologically activated bio-based fertilizers derived from manures on tomato plants and their rhizospheric communities. Sci. Rep. 2023, 13, 22478. [Google Scholar] [CrossRef]
- Ishaq, S.L.; Seipel, T.; Yeoman, C.; Menalled, F.D. Dryland Cropping Systems, Weed Communities, and Disease Status Modulate the Effect of Climate Conditions on Wheat Soil Bacterial Communities. mSphere 2020, 5, 10–1128. [Google Scholar] [CrossRef]
- Xu, Y.; Ge, Y.; Lou, Y.; Meng, J.; Shi, L.; Xia, F. Assembly strategies of the wheat root-associated microbiome in soils contaminated with phenanthrene and copper. J. Hazard. Mater. 2021, 412, 125340. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Yu, T.; Tan, W.; Zhou, Q.; Liu, L.; Nian, H.; Lian, T. GmAMT2.1/2.2-dependent ammonium nitrogen and metabolites shape rhizosphere microbiome assembly to mitigate cadmium toxicity. Npj Biofilms Microbiomes 2024, 10, 60. [Google Scholar] [CrossRef]
- Soubasakou, G.; Cavoura, O.; Damikouka, I. Phytoremediation of Cadmium-Contaminated Soils: A Review of New Cadmium Hyperaccumulators and Factors Affecting their Efficiency. Bull. Environ. Contam. Toxicol. 2022, 109, 783–787. [Google Scholar] [CrossRef]
- Ismaiel, M.M.S.; Piercey-Normore, M.D. Antioxidant enzymes of Pseudochlorella pringsheimii under two stressors: Variation of SOD Isoforms activity. J. Plant Res. 2023, 136, 755–767. [Google Scholar] [CrossRef]
- Jomova, K.; Alomar, S.Y.; Alwasel, S.H.; Nepovimova, E.; Kuca, K.; Valko, M. Several lines of antioxidant defense against oxidative stress: Antioxidant enzymes, nanomaterials with multiple enzyme-mimicking activities, and low-molecular-weight antioxidants. Arch. Toxicol. 2024, 98, 1323–1367. [Google Scholar] [CrossRef] [PubMed]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ge, X.; Peng, F.; Li, W.; Li, J.J. Exaggerated false positives by popular differential expression methods when analyzing human population samples. Genome Biol. 2022, 23, 79. [Google Scholar] [CrossRef]
- Wilcoxon, F. Individual Comparisons of Grouped Data by Ranking Methods. J. Econ. Entomol. 1946, 39, 269–270. [Google Scholar] [CrossRef]
- Islam, M.D.; Hasan, M.M.; Rahaman, A.; Haque, P.; Islam, M.S.; Rahman, M.M. Translocation and bioaccumulation of trace metals from industrial effluent to locally grown vegetables and assessment of human health risk in Bangladesh. SN Appl. Sci. 2020, 2, 1315. [Google Scholar] [CrossRef]
- Huang, L.; Wang, Q.; Zhou, Q.; Ma, L.; Wu, Y.; Liu, Q.; Wang, S.; Feng, Y. Cadmium uptake from soil and transport by leafy vegetables: A meta-analysis. Environ. Pollut. 2020, 264, 114677. [Google Scholar] [CrossRef]
- Sarver, J.; Schultz, E.; Apigo, A.; Gernandt, D.S.; Salas-Lizana, R.; Oono, R. Deep sequencing across multiple host species tests pine-endophyte specificity. Am. J. Bot. 2022, 109, 83–98. [Google Scholar] [CrossRef]
- Su, P.; Kang, H.; Peng, Q.; Peng, W.; Sun, S.; Du, X.; Zhang, C.; Lei, Z.; Bai, L.; Tang, Q.; et al. Host metabolites explain microbiome variation between different rice genotypes. Microbiome 2025, 13, 185. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Gu, S.; Guo, X.; Liu, Y.; Tao, Q.; Zhao, H.P.; Liang, Y.; Banerjee, S.; Li, T. Core Microbiota in the Rhizosphere of Heavy Metal Accumulators and Its Contribution to Plant Performance. Environ. Sci. Technol. 2022, 56, 12975–12987. [Google Scholar] [CrossRef] [PubMed]
- Hou, D.; Lin, Z.; Wang, R.; Ge, J.; Wei, S.; Xie, R.; Wang, H.; Wang, K.; Hu, Y.; Yang, X.; et al. Cadmium Exposure-Sedum alfredii Planting Interactions Shape the Bacterial Community in the Hyperaccumulator Plant Rhizosphere. Appl. Environ. Microbiol. 2018, 84, e02797-17. [Google Scholar] [CrossRef] [PubMed]
- Flores-Duarte, N.J.; Pajuelo, E.; Mateos-Naranjo, E.; Navarro-Torre, S.; Rodríguez-Llorente, I.D.; Redondo-Gómez, S.; Carrasco López, J.A. A Culturomics-Based Bacterial Synthetic Community for Improving Resilience towards Arsenic and Heavy Metals in the Nutraceutical Plant Mesembryanthemum crystallinum. Int. J. Mol. Sci. 2023, 24, 7003. [Google Scholar] [CrossRef]
- Dong, Q.; Xu, P.; Wang, Z. Differential Cadmium Distribution and Translocation in Roots and Shoots Related to Hyper-Tolerance between Tall Fescue and Kentucky Bluegrass. Front. Plant Sci. 2017, 8, 113. [Google Scholar] [CrossRef]
- Hu, Y.; Tian, S.; Foyer, C.H.; Hou, D.; Wang, H.; Zhou, W.; Liu, T.; Ge, J.; Lu, L.; Lin, X. Efficient phloem transport significantly remobilizes cadmium from old to young organs in a hyperaccumulator Sedum alfredii. J. Hazard. Mater. 2019, 365, 421–429. [Google Scholar] [CrossRef]
- Wu, Y.; Ma, L.; Liu, Q.; Vestergård, M.; Topalovic, O.; Wang, Q.; Zhou, Q.; Huang, L.; Yang, X.; Feng, Y. The plant-growth promoting bacteria promote cadmium uptake by inducing a hormonal crosstalk and lateral root formation in a hyperaccumulator plant Sedum alfredii. J. Hazard. Mater. 2020, 395, 122661. [Google Scholar] [CrossRef]
- Luo, J.; Tao, Q.; Jupa, R.; Liu, Y.; Wu, K.; Song, Y.; Li, J.; Huang, Y.; Zou, L.; Liang, Y.; et al. Role of Vertical Transmission of Shoot Endophytes in Root-Associated Microbiome Assembly and Heavy Metal Hyperaccumulation in Sedum alfredii. Environ. Sci. Technol. 2019, 53, 6954–6963. [Google Scholar] [CrossRef]
- Zulfiqar, U.; Haider, F.U.; Maqsood, M.F.; Mohy-Ud-Din, W.; Shabaan, M.; Ahmad, M.; Kaleem, M.; Ishfaq, M.; Aslam, Z.; Shahzad, B. Recent Advances in Microbial-Assisted Remediation of Cadmium-Contaminated Soil. Plants 2023, 12, 3147. [Google Scholar] [CrossRef]
- Świsłowski, P.; Rajfur, M.; Wacławek, M. Influence of Heavy Metal Concentration on Chlorophyll Content in Pleurozium schreberi Mosses. Ecol. Chem. Eng. S 2020, 27, 591–601. [Google Scholar] [CrossRef]
- Rola, K.; Majewska, E.; Chowaniec, K. Interaction effect of fungicide and chitosan on non-target lichenized fungi. Chemosphere 2023, 316, 137772. [Google Scholar] [CrossRef]
- Giraudo, M.; Cottin, G.; Esperanza, M.; Gagnon, P.; Silva, A.O.D.; Houde, M. Transcriptional and cellular effects of benzotriazole UV stabilizers UV-234 and UV-328 in the freshwater invertebrates Chlamydomonas reinhardtii and Daphnia magna. Environ. Toxicol. Chem. 2017, 36, 3333–3342. [Google Scholar] [CrossRef]
- Yue, H.; Luo, Y.; Wu, J.; Wang, N.; Wang, Y.; Chu, Z.; Tong, X.; Ma, X.; Huang, D.; Miao, H.; et al. Disposal and utilization of dead animals during breeding in livestock and poultry farming by means of synthetic microbiota. Environ. Res. 2024, 252, 119010. [Google Scholar] [CrossRef]
- Kaur, M.; Saini, K.C.; Ojah, H.; Sahoo, R.; Gupta, K.; Kumar, A.; Bast, F. Abiotic stress in algae: Response, signaling and transgenic approaches. J. Appl. Phycol. 2022, 34, 1843–1869. [Google Scholar] [CrossRef]
- Luo, J.-S.; Zhang, Z. Mechanisms of cadmium phytoremediation and detoxification in plants. Crop J. 2021, 9, 521–529. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, X.; Liu, X.; Huang, Z.; Niu, S.; Xu, T.; Zeng, J.; Li, H.; Wang, T.; Gao, Y.; et al. Physiological, biochemical and proteomic insight into integrated strategies of an endophytic bacterium Burkholderia cenocepacia strain YG-3 response to cadmium stress. Metallomics 2019, 11, 1252–1264. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.-L.; Vadiveloo, A.; Chen, D.-Z.; Gao, F. Regulation effects of indoleacetic acid on lipid production and nutrient removal of Chlorella pyrenoidosa in seawater-containing wastewater. Water Res. 2024, 248, 120864. [Google Scholar] [CrossRef]
- Finkel, O.M.; Salas-González, I.; Castrillo, G.; Conway, J.M.; Law, T.F.; Teixeira, P.; Wilson, E.D.; Fitzpatrick, C.R.; Jones, C.D.; Dangl, J.L. A single bacterial genus maintains root growth in a complex microbiome. Nature 2020, 587, 103–108. [Google Scholar] [CrossRef] [PubMed]







| Strain ID | Genus and Species | ACC Deaminase | IAA Production (μg/mL) | Cd Resistance (mM) |
|---|---|---|---|---|
| SaPI1 | Pseudomonas izuensis | + | 55.1 | 4.0 |
| SaLS1 | Leifsonia shinshuensis | + | 150.0 | 0.5 |
| SaOA1 | Ochrobactrum anthropi | − | 64.2 | 0.6 |
| SaNL1 | Novosphingobium lindaniclasticum | − | 53.9 | 2.0 |
| Gene | Forward Primers (5′-3′) | Reverse Primers (5′-3′) |
|---|---|---|
| SOD | GGTTTCCATGTCCATGCTCT | ATTGTGAAGGTGGCAGTTCC |
| CAT | TCAGCTGCCAGTTAATGCAC | GACAGCAGGTGGAGTTGGAT |
| POD | TTCGAACGGAAAAAGATGCT | AACCCTCCATGAAGGACCTC |
| GR | AAGGCAAAAGAAGGTGCTGA | AGTTCCCTTGCTGGTCTTCA |
| GST | CGTCGTCGAAGAAGAAGAGG | TTTTTGGTGGGAGTTCCAAG |
| CHLASE | GAATATCCGGTGGTGATGCT | TCCGCCGTTGATTTTATCTC |
| HMA4 | TCTGTGGCAAAGAAGTAA | ACCAAACTAGACGACCCT |
| ACTIN | CTTGCACCTAGCAGCATGAA | GGACAATGGATGGACCTGAC |
| Control Group | SynCom-NS Treatment | |
|---|---|---|
| Initial total soil Cd stock (mg/pot) | 2.0 | 2.0 |
| Final apparent soil Cd stock (mg/pot) | 1.99 | 1.81 |
| Apparent soil Cd reduction (mg/pot) | 0.01 | 0.19 |
| Total plant Cd accumulation (mg/pot) | 0.0005 | 0.0024 |
| Annotation Database | Gene Number | Annotation Percentage (%) |
|---|---|---|
| NR | 96,791 | 99.83 |
| GO | 56,740 | 58.51 |
| KEGG | 32,925 | 33.95 |
| Total | 96,958 | 99.83 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Huang, L.; Fu, S.; Du, S.; Feng, Y. Functional Portability of a Hyperaccumulator-Derived Core Microbiome: Enhancing Cadmium Phytoextraction in Brassica juncea L. Through Molecular Reprogramming. Toxics 2026, 14, 303. https://doi.org/10.3390/toxics14040303
Huang L, Fu S, Du S, Feng Y. Functional Portability of a Hyperaccumulator-Derived Core Microbiome: Enhancing Cadmium Phytoextraction in Brassica juncea L. Through Molecular Reprogramming. Toxics. 2026; 14(4):303. https://doi.org/10.3390/toxics14040303
Chicago/Turabian StyleHuang, Lukuan, Shumeng Fu, Shaoting Du, and Ying Feng. 2026. "Functional Portability of a Hyperaccumulator-Derived Core Microbiome: Enhancing Cadmium Phytoextraction in Brassica juncea L. Through Molecular Reprogramming" Toxics 14, no. 4: 303. https://doi.org/10.3390/toxics14040303
APA StyleHuang, L., Fu, S., Du, S., & Feng, Y. (2026). Functional Portability of a Hyperaccumulator-Derived Core Microbiome: Enhancing Cadmium Phytoextraction in Brassica juncea L. Through Molecular Reprogramming. Toxics, 14(4), 303. https://doi.org/10.3390/toxics14040303