In Silico Prediction of the Toxicity of Nitroaromatic Compounds: Application of Ensemble Learning QSAR Approach
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Data Collection
2.2. Generation of Descriptors
2.3. QSAR Modeling and Validation
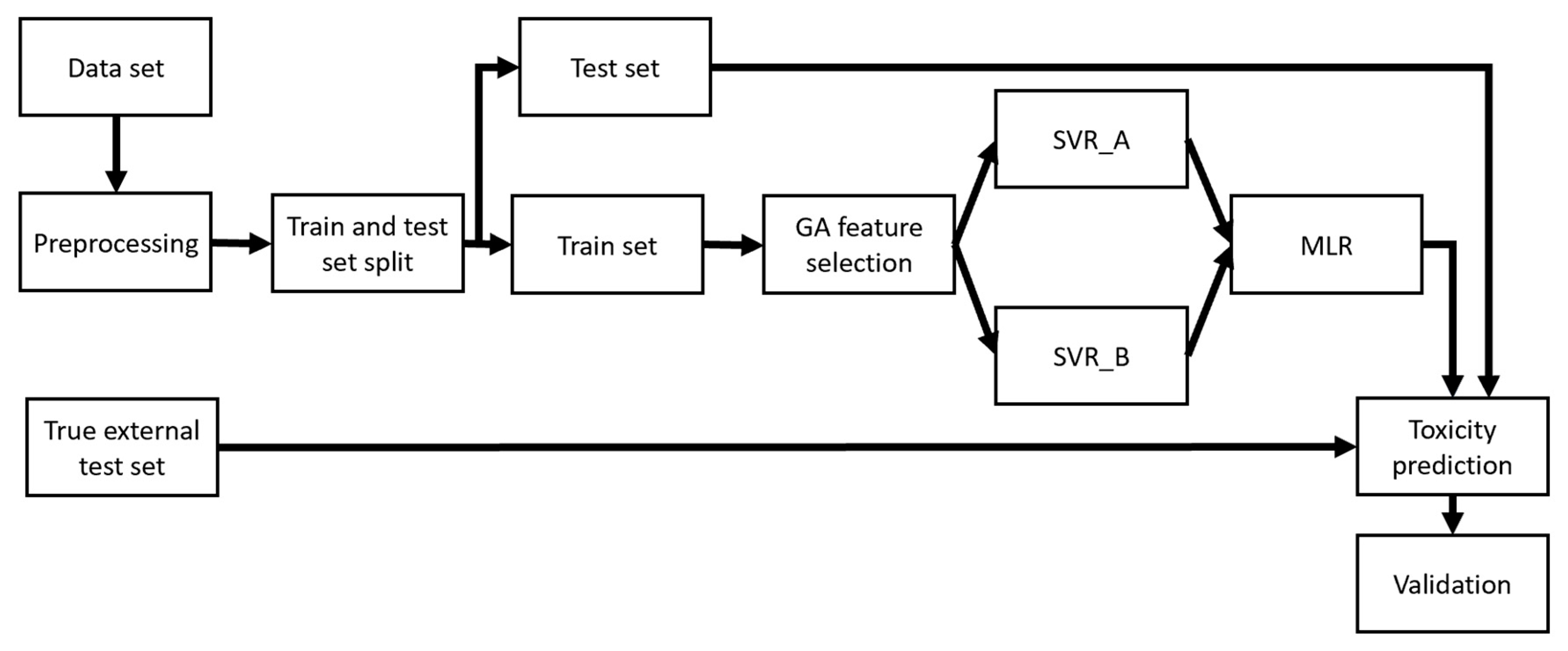
Support Vector Regression and Ensemble Model
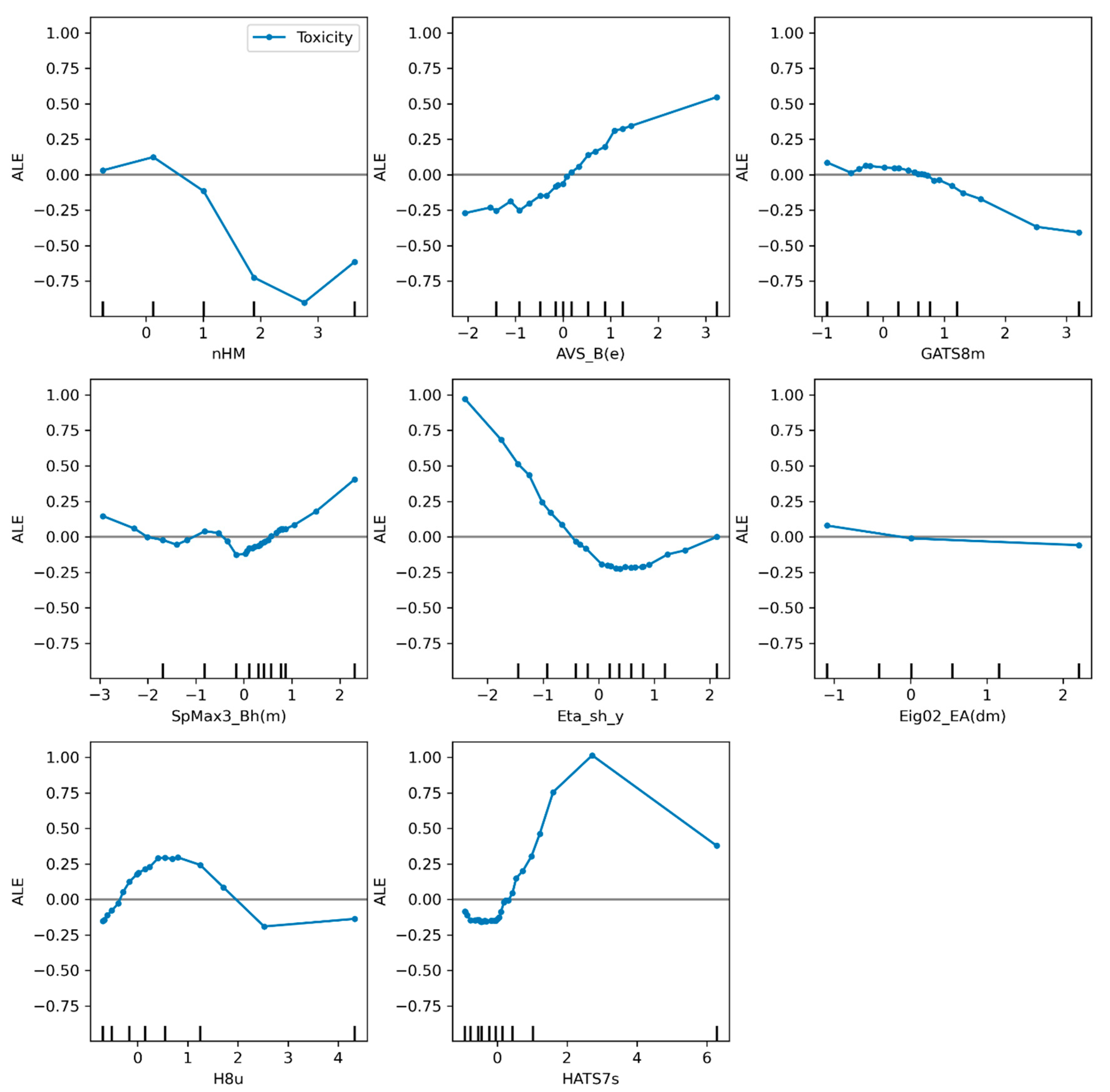
2.4. Analysis of Descriptors in Models
3. Results
3.1. Distribution of Molecular Weights and Toxicity
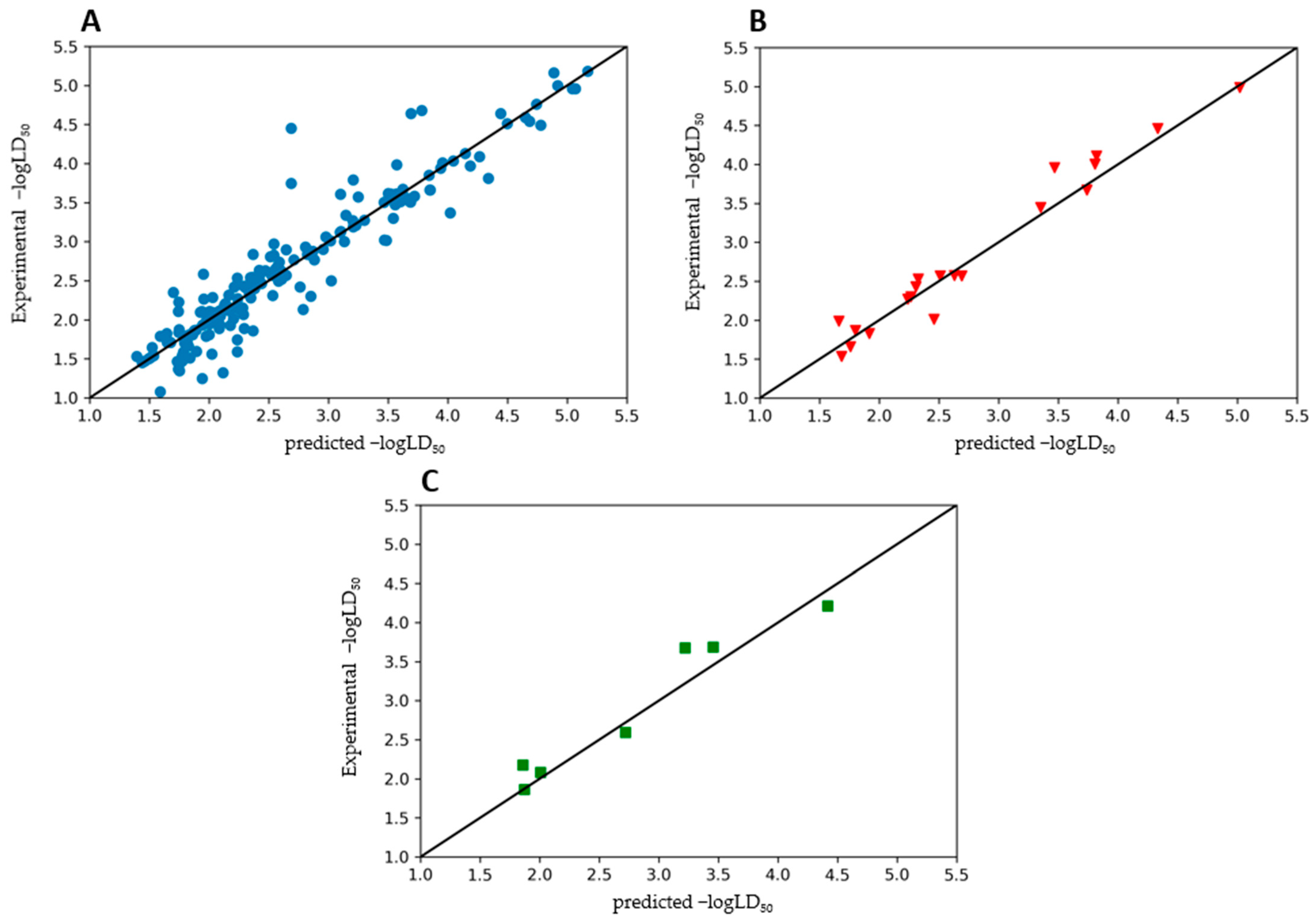
3.2. Ensemble Model
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bilal, M.; Bagheri, A.R.; Bhatt, P.; Chen, S. Environmental occurrence, toxicity concerns, and remediation of recalcitrant nitroaromatic compounds. J. Environ. Manag. 2021, 291, 112685. [Google Scholar] [CrossRef] [PubMed]
- Kovacic, P.; Somanathan, R. Nitroaromatic compounds: Environmental toxicity, carcinogenicity, mutagenicity, therapy and mechanism. J. Appl. Toxicol. 2014, 34, 810–824. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, J.; Tarale, P.; Sivanesan, S.; Bafana, A. Environmental persistence, hazard, and mitigation challenges of nitroaromatic compounds. Environ. Sci. Pollut. Res. 2019, 26, 28650–28667. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, M.; Chaudhari, A. Microbial remediation of nitro-aromatic compounds: An overview. J. Environ. Manag. 2007, 85, 496–512. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.-L.; Yu, Y.-Y.; Fang, Z.; Naraginti, S.; Zhang, Y.; Yong, Y.-C. Recent advances in nitroaromatic pollutants bioreduction by electroactive bacteria. Process Biochem. 2018, 70, 129–135. [Google Scholar] [CrossRef]
- Deng, K.; Wong, T.Y.; Wang, Y.; Leung, E.M.K.; Chan, W. Combination of precolumn nitro-reduction and ultraperformance liquid chromatography with fluorescence detection for the sensitive quantification of 1-nitronaphthalene, 2-nitrofluorene, and 1-nitropyrene in meat products. J. Agric. Food Chem. 2015, 63, 3161–3167. [Google Scholar] [CrossRef]
- Slater, E.C. Mechanism of uncoupling of oxidative phosphorylation by nitrophenols. Comp. Biochem. Physiol. 1962, 4, 281–301. [Google Scholar] [CrossRef]
- Strauss, M.J. The Nitroaromatic Group in Drug Design. Pharmacology and Toxicology (for Nonpharmacologists). Ind. Eng. Chem. Prod. Res. Dev. 1979, 18, 158–166. [Google Scholar] [CrossRef]
- Hao, Y.; Sun, G.; Fan, T.; Tang, X.; Zhang, J.; Liu, Y.; Zhang, N.; Zhao, L.; Zhong, R.; Peng, Y. In vivo toxicity of nitroaromatic compounds to rats: QSTR modelling and interspecies toxicity relationship with mouse. J. Hazard. Mater. 2020, 399, 122981. [Google Scholar] [CrossRef]
- Khan, K.; Roy, K.; Benfenati, E. Ecotoxicological QSAR modeling of endocrine disruptor chemicals. J. Hazard. Mater. 2019, 369, 707–718. [Google Scholar] [CrossRef]
- Isayev, O.; Rasulev, B.; Gorb, L.; Leszczynski, J. Structure-toxicity relationships of nitroaromatic compounds. Mol. Divers. 2006, 10, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.L.; Lyu, Y.C.; Leong, M.K. In silico prediction of the mutagenicity of nitroaromatic compounds using a novel two-QSAR approach. Toxicol. Vitr. 2017, 40, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Cassani, S.; Kovarich, S.; Papa, E.; Roy, P.P.; van der Wal, L.; Gramatica, P. Daphnia and fish toxicity of (benzo)triazoles: Validated QSAR models, and interspecies quantitative activity-activity modelling. J. Hazard. Mater. 2013, 258–259, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Tropsha, A. Best practices for QSAR model development, validation, and exploitation. Mol. Inform. 2010, 29, 476–488. [Google Scholar] [CrossRef] [PubMed]
- Katritzky, A.R.; Oliferenko, P.; Oliferenko, A.; Lomaka, A.; Karelson, M. Nitrobenzene toxicity: QSAR correlations and mechanistic interpretations. J. Phys. Org. Chem. 2003, 16, 811–817. [Google Scholar] [CrossRef]
- Casañola-Martin, G.M.; Le-Thi-Thu, H.; Pérez-Giménez, F.; Marrero-Ponce, Y.; Merino-Sanjuán, M.; Abad, C.; González-Díaz, H. Multi-output model with Box–Jenkins operators of linear indices to predict multi-target inhibitors of ubiquitin–proteasome pathway. Mol. Divers. 2015, 19, 347–356. [Google Scholar] [CrossRef]
- Bediaga, H.; Moreno, M.I.; Arrasate, S.; Vilas, J.L.; Orbe, L.; Unzueta, E.; Mercader, J.P.; González-Díaz, H. Multi-output chemometrics model for gasoline compounding. Fuel 2022, 310, 122274. [Google Scholar] [CrossRef]
- Litter, M.I. A short review on the preparation and use of iron nanomaterials for the treatment of pollutants in water and soil. Emergent Mater. 2022, 5, 391–400. [Google Scholar] [CrossRef]
- Chen, S.; Liu, H. Self-reductive palladium nanoparticles loaded on polydopamine-modified MXene for highly efficient and quickly catalytic reduction of nitroaromatics and dyes. Colloids Surf. A Physicochem. Eng. Asp. 2022, 635, 128038. [Google Scholar] [CrossRef]
- Kumunda, C.; Adekunle, A.S.; Mamba, B.B.; Hlongwa, N.W.; Nkambule, T.T.I. Electrochemical Detection of Environmental Pollutants Based on Graphene Derivatives: A Review. Front. Mater. 2020, 7, 616787. [Google Scholar] [CrossRef]
- Tiwari, J.; Gandhi, D.; Sivanesan, S.; Naoghare, P.; Bafana, A. Remediation of different nitroaromatic pollutants by a promising agent of Cupriavidus sp. strain a3. Ecotoxicol. Environ. Saf. 2020, 205, 111138. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Chen, J.; Liu, Z.; Xu, Y. CO Activation Using Nitrogen-Doped Carbon Nanotubes for Reductive Carbonylation of Nitroaromatics to Benzimidazolinone and Phenyl Urea. ACS Appl. Mater. Interfaces 2020, 12, 48700–48711. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Xiao, K.; Zhou, C.; Li, G.; Yang, H.; Li, Z.; Cheng, J. Insights into pesticide toxicity against aquatic organism: QSTR models on Daphnia Magna. Ecotoxicol. Environ. Saf. 2019, 173, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Tugcu, G.; Ertürk, M.D.; Saçan, M.T. On the aquatic toxicity of substituted phenols to Chlorella vulgaris: QSTR with an extended novel data set and interspecies models. J. Hazard. Mater. 2017, 339, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Mondal, D.; Ghosh, K.; Baidya, A.T.K.; Gantait, A.M.; Gayen, S. Identification of structural fingerprints for in vivo toxicity by using Monte Carlo based QSTR modeling of nitroaromatics. Toxicol. Mech. Methods 2020, 30, 257–265. [Google Scholar] [CrossRef]
- Kuz’min, V.E.; Muratov, E.N.; Artemenko, A.G.; Gorb, L.; Qasim, M.; Leszczynski, J. The effects of characteristics of substituents on toxicity of the nitroaromatics: HiT QSAR study. J. Comput.-Aided Mol. Des. 2008, 22, 747. [Google Scholar] [CrossRef]
- Gooch, A.; Sizochenko, N.; Rasulev, B.; Gorb, L.; Leszczynski, J. In vivo toxicity of nitroaromatics: A comprehensive quantitative structure–activity relationship study. Environ. Toxicol. Chem. 2017, 36, 2227–2233. [Google Scholar] [CrossRef]
- Toddeschini, R.; Consonni, V.; Mauri, A.; Pavan, M. Dragon Software for the Calculation of Molecular Descriptors, Version 6 for Windows; Talete SRL: Milan, Italy, 2014. [Google Scholar]
- Yap, C.W. PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints. J. Comput. Chem. 2011, 32, 1466–1474. [Google Scholar] [CrossRef]
- Kuz’min, V.E.; Artemenko, A.G.; Muratov, E.N.; Polischuk, P.G.; Ognichenko, L.N.; Liahovsky, A.V.; Hromov, A.I.; Varlamova, E.V. Virtual Screening and Molecular Design Based on Hierarchical Qsar Technology. Recent Adv. QSAR Stud. 2010, 8, 127–176. [Google Scholar] [CrossRef]
- Keshavarz, M.H.; Akbarzadeh, A.R. A simple approach for assessment of toxicity of nitroaromatic compounds without using complex descriptors and computer codes. SAR QSAR Environ. Res. 2019, 30, 347–361. [Google Scholar] [CrossRef]
- Apley, D.W.; Zhu, J. Visualizing the effects of predictor variables in black box supervised learning models. J. R. Stat. Society. Ser. B Stat. Methodol. 2020, 82, 1059–1086. [Google Scholar] [CrossRef]
- ChemIDplus: A Web-Based Chemical Search System, Mar-Apr 2000, NLM Technical Bulletin. Available online: https://www.nlm.nih.gov/pubs/techbull/ma00/ma00_chemid.html (accessed on 17 June 2022).
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem in 2021: New data content and improved web interfaces. Nucleic Acids Res. 2021, 49, D1388–D1395. [Google Scholar] [CrossRef] [PubMed]
- Hypercube Inc., N.t.S., Gainesville, Florida 32601, USA. HyperChem(TM) Professional 8.0. 2019. Available online: http://www.hypercubeusa.com/ (accessed on 13 October 2021).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- OECD. Guidance Document on the Validation of (Quantitative) Structure-Activity Relationship (Q)SAR Models; OECD: Paris, France, 2014. [Google Scholar] [CrossRef]
- Gramatica, P.; Sangion, A. A Historical Excursus on the Statistical Validation Parameters for QSAR Models: A Clarification Concerning Metrics and Terminology. J. Chem. Inf. Model. 2016, 56, 1127–1131. [Google Scholar] [CrossRef]
- Roy, K. On some aspects of validation of predictive quantitative structure-activity relationship models. Expert Opin. Drug Discov. 2007, 2, 1567–1577. [Google Scholar] [CrossRef] [PubMed]
- Golbraikh, A.; Tropsha, A. Beware of q2! J. Mol. Graph. Model 2002, 20, 269–276. [Google Scholar] [CrossRef]
- Pratim, R.P.; Paul, S.; Mitra, I.; Roy, K. On two novel parameters for validation of predictive QSAR models. Molecules 2009, 14, 1660–1701. [Google Scholar] [CrossRef]
- Erickson, M.E.; Ngongang, M.; Rasulev, B. A refractive index study of a diverse set of polymeric materials by QSPR with quantum-chemical and additive descriptors. Molecules 2020, 25, 3772. [Google Scholar] [CrossRef]
- Gramatica, P.; Chirico, N.; Papa, E.; Cassani, S.; Kovarich, S. QSARINS: A new software for the development, analysis, and validation of QSAR MLR models. J. Comput. Chem. 2013, 34, 2121–2132. [Google Scholar] [CrossRef]
- Freund, Y.; Schapire, R.E.; Singer, Y.; Warmuth, M.K. Using and combining predictors that specialize. In Proceedings of the Proceedings of the Twenty-Ninth Annual ACM Symposium on the Theory of Computing; 1997. [Google Scholar]
- Van Der Laan, M.J.; Polley, E.C.; Hubbard, A.E. Super learner. Stat. Appl. Genet. Mol. Biol. 2007, 6, 25. [Google Scholar] [CrossRef]
- Molnar, C. Interpretable Machine Learning: A Guide for Making Black Box Models Explainable. Independently Published-Amazon: Seattle, WA, USA, 2022; ISBN -13 979-841-146-333-0. [Google Scholar]
- Hao, Y.; Sun, G.; Fan, T.; Sun, X.; Liu, Y.; Zhang, N.; Zhao, L.; Zhong, R.; Peng, Y. Prediction on the mutagenicity of nitroaromatic compounds using quantum chemistry descriptors based QSAR and machine learning derived classification methods. Ecotoxicol. Environ. Saf. 2019, 186, 109822. [Google Scholar] [CrossRef] [PubMed]
- Lovrić, M.; Pavlović, K.; Žuvela, P.; Spataru, A.; Lučić, B.; Kern, R.; Wong, M.W. Machine learning in prediction of intrinsic aqueous solubility of drug-like compounds: Generalization, complexity, or predictive ability? J. Chemom. 2021, 35, e3349. [Google Scholar] [CrossRef]
- Lučić, B.; Batista, J.; Bojović, V.; Lovrić, M.; Sović Kržić, A.; Bešlo, D.; Nadramija, D.; Vikić-Topić, D. Estimation of Random Accuracy and Its Use in Validation of Predictive Quality of Classification Models within Predictive Challenges. Croat. Chem. Acta 2019, 92, 379–391. [Google Scholar] [CrossRef]
- Cronin, M.T.D.; Gregory, B.W.; Schultz, T.W. Quantitative Structure−Activity Analyses of Nitrobenzene Toxicity to Tetrahymena pyriformis. Chem. Res. Toxicol. 1998, 11, 902–908. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, H.; Altenburger, R.; Jastorff, B.; Schüürmann, G. Quantitative Structure−Activity Analysis of the Algae Toxicity of Nitroaromatic Compounds. Chem. Res. Toxicol. 2000, 13, 441–450. [Google Scholar] [CrossRef]
- Ukić, Š.; Sigurnjak, M.; Cvetnić, M.; Markić, M.; Stankov, M.N.; Rogošić, M.; Rasulev, B.; Lončarić Božić, A.; Kušić, H.; Bolanča, T. Toxicity of pharmaceuticals in binary mixtures: Assessment by additive and non-additive toxicity models. Ecotoxicol. Environ. Saf. 2019, 185, 109696. [Google Scholar] [CrossRef]
- Cvetnic, M.; Juretic Perisic, D.; Kovacic, M.; Ukic, S.; Bolanca, T.; Rasulev, B.; Kusic, H.; Loncaric Bozic, A. Toxicity of aromatic pollutants and photooxidative intermediates in water: A QSAR study. Ecotoxicol. Environ. Saf. 2019, 169, 918–927. [Google Scholar] [CrossRef]
- Sizochenko, N.; Mikolajczyk, A.; Jagiello, K.; Puzyn, T.; Leszczynski, J.; Rasulev, B. How the toxicity of nanomaterials towards different species could be simultaneously evaluated: A novel multi-nano-read-across approach. Nanoscale 2018, 10, 582–591. [Google Scholar] [CrossRef]
- Toropov, A.A.; Rasulev, B.F.; Leszczynski, J. QSAR modeling of acute toxicity by balance of correlations. Bioorganic Med. Chem. 2008, 16, 5999–6008. [Google Scholar] [CrossRef]
- Klein, D.J. Topological Indices and Related Descriptors in QSAR and QSPR. J. Chem. Inf. Comput. Sci. 2002, 42, 1507. [Google Scholar] [CrossRef]



| Method/Model | Runtime Parameters |
|---|---|
| SVR_A and SVR_B | Kernel = ’rbf’, degree = 3, gamma = ’auto’, coef 0 = 0.0, tol = 0.001, C = 5.0, epsilon = 0.1, shrinking = Ture, cache_size = 200, verbose = False, max_iter = −1 |
| MLR | Fir_intercept = True, normalize = ’False’, copy_X = True, n_jobs = −1, positive = False |
| Parameters | Regression Model | Ensemble Model | |
|---|---|---|---|
| SVR_A | SVR_B | ||
| No. of descriptors | 11 | 8 | _ |
| R2 (training) | 0.83 | 0.81 | 0.88 |
| RMSE (training) | 0.111 | 0.127 | 0.093 |
| MAE (training) | 0.221 | 0.226 | 0.199 |
| MAECV(5-Fold) | 0.484 | 0.486 | 0.480 |
| R2 (test) | 0.92 | 0.85 | 0.95 |
| RMSE (test) | 0.056 | 0.096 | 0.041 |
| MAE (test) | 0.191 | 0.250 | 0.155 |
| CCC (test) | 0.968 | 0.946 | 0.978 |
| R2 (external test) | 0.74 | 0.88 | 0.92 |
| RMSE (external test) | 0.132 | 0.123 | 0.061 |
| MAE (external test) | 0.320 | 0.319 | 0.202 |
| CCC (external test) | 0.898 | 0.931 | 0.961 |
| 0.945 | 0.906 | 0.960 | |
| 0.943 | 0.903 | 0.958 | |
| 0.510 | 0.536 | 0.560 | |
| 0.955 | 0.981 | 0.975 | |
| 1.041 | 1.007 | 1.021 | |
| Descriptor | SVR_A | SVR_B | Definition and Scope | Descriptor Type |
|---|---|---|---|---|
| AVS_B(e) | X | X | average vertex sum from Burden matrix weighted by Sanderson electronegativity | 2D matrix-based descriptors |
| HATS7s | X | X | leverage-weighted autocorrelation of lag 7/weighted by I-state | GETAWAY descriptors |
| Eta_sh_y | X | X | Eta y shape index | ETA indices |
| GATS2v | X | Geary autocorrelation of lag 2 weighted by van der Waals volume | 2D autocorrelations | |
| GATS8m | X | Geary autocorrelation of lag 8 weighted by mass | 2D autocorrelations | |
| P_VSA_LogP_3 | X | P_VSA-like on LogP, bin 3 | P_VSA-like descriptors | |
| nHM | X | number of heavy atoms | Constitutional indices | |
| RDF060s | X | Radial Distribution Function—060/weighted by I-state | RDF descriptors | |
| Dm | X | D total accessibility index/weighted by mass | WHIM descriptors | |
| H8u | X | H autocorrelation of lag 8/unweighted | GETAWAY descriptors | |
| O-059 | X | Al-O-Al | Atom-centred fragments | |
| B09[C-C] | X | Presence/absence of C—C at topological distance 9 | 2D Atom Pairs | |
| SpMax3_Bh(m) | X | largest eigenvalue n. 3 of Burden matrix weighted by mass | Burden eigenvalues | |
| CATS2D_05_NL | X | CATS2D Negative-Lipophilic at lag 05 | CATS 2D | |
| Eig02_EA(dm) | X | eigenvalue n. 2 from edge adjacency mat. weighted by dipole moment | Edge adjacency indices | |
| C-043 | X | X--CR.X | Atom-centred fragments |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daghighi, A.; Casanola-Martin, G.M.; Timmerman, T.; Milenković, D.; Lučić, B.; Rasulev, B. In Silico Prediction of the Toxicity of Nitroaromatic Compounds: Application of Ensemble Learning QSAR Approach. Toxics 2022, 10, 746. https://doi.org/10.3390/toxics10120746
Daghighi A, Casanola-Martin GM, Timmerman T, Milenković D, Lučić B, Rasulev B. In Silico Prediction of the Toxicity of Nitroaromatic Compounds: Application of Ensemble Learning QSAR Approach. Toxics. 2022; 10(12):746. https://doi.org/10.3390/toxics10120746
Chicago/Turabian StyleDaghighi, Amirreza, Gerardo M. Casanola-Martin, Troy Timmerman, Dejan Milenković, Bono Lučić, and Bakhtiyor Rasulev. 2022. "In Silico Prediction of the Toxicity of Nitroaromatic Compounds: Application of Ensemble Learning QSAR Approach" Toxics 10, no. 12: 746. https://doi.org/10.3390/toxics10120746
APA StyleDaghighi, A., Casanola-Martin, G. M., Timmerman, T., Milenković, D., Lučić, B., & Rasulev, B. (2022). In Silico Prediction of the Toxicity of Nitroaromatic Compounds: Application of Ensemble Learning QSAR Approach. Toxics, 10(12), 746. https://doi.org/10.3390/toxics10120746








