Authentication of Meat and Meat Products Using Triacylglycerols Profiling and by DNA Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples
2.2. Direct Analysis in Real Time Coupled with High-Resolution Mass Spectrometry (DART–HRMS) Analysis
2.2.1. Sample Preparation for Instrumental Analysis
2.2.2. Conditions of DART–HRMS Analysis
2.2.3. Data Analysis
2.2.4. Quality Control
2.3. Analysis by Polymerase Chain Reaction (PCR)
2.3.1. DNA Isolation
2.3.2. Primers and Probes
2.3.3. Multiplex mPCR
2.3.4. Multiplex mqPCRs
2.3.5. Data Analysis
2.3.6. Quality Control
3. Results and Discussion
3.1. Results of DART–HRMS Analysis
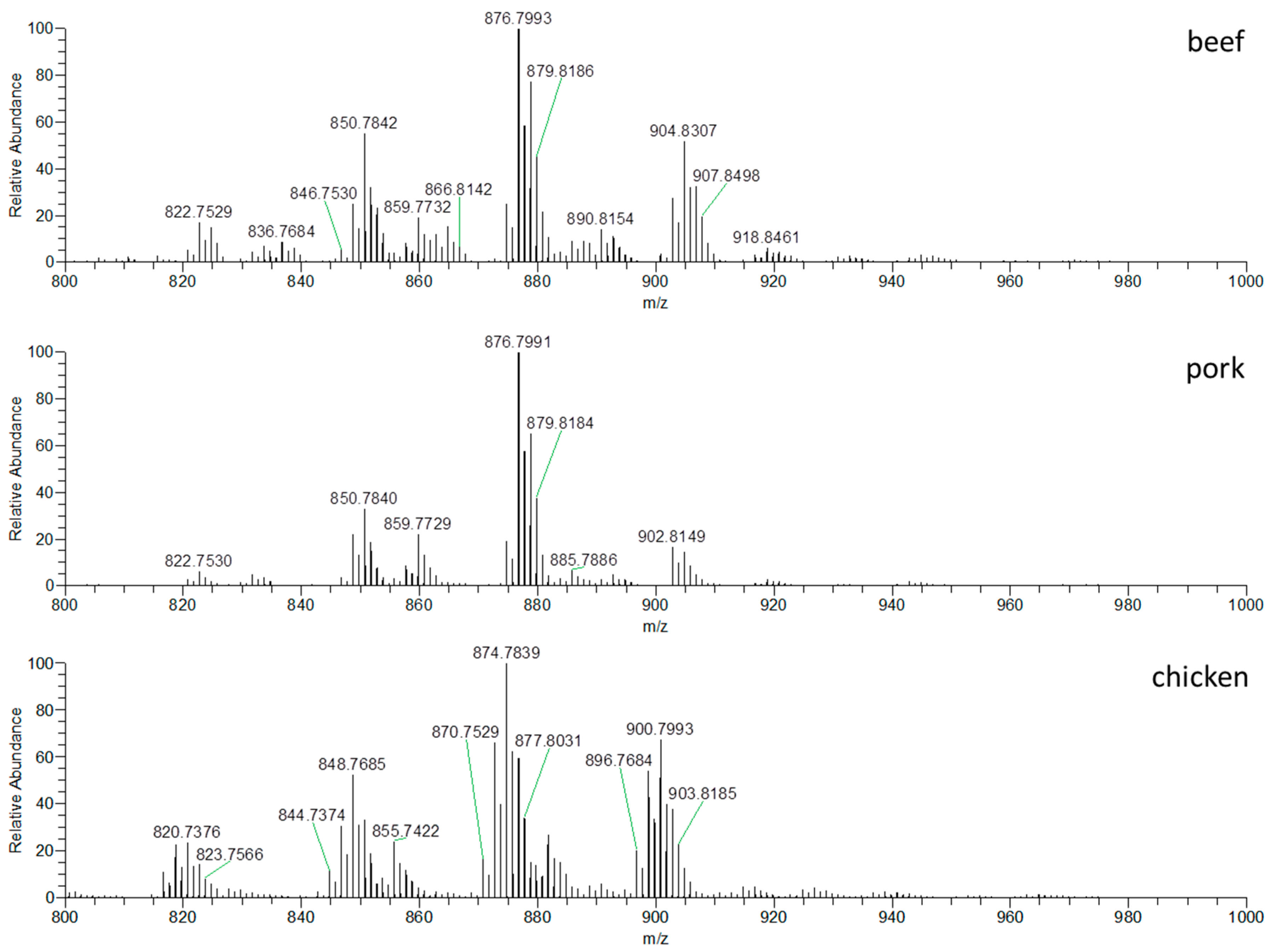
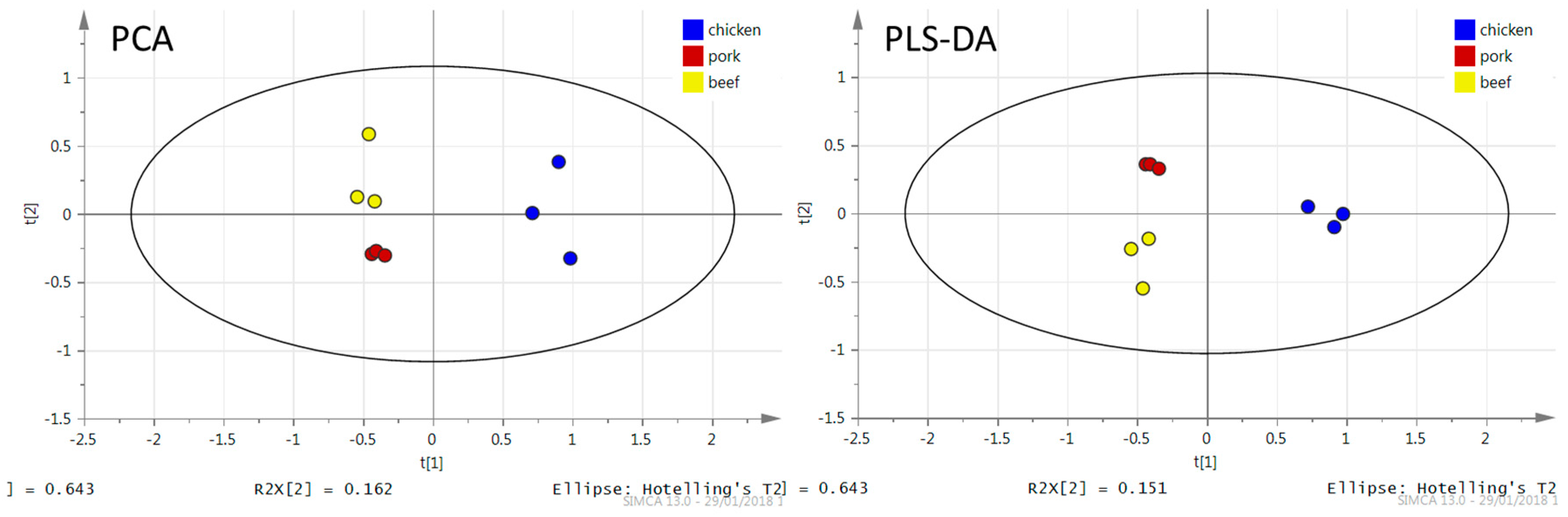
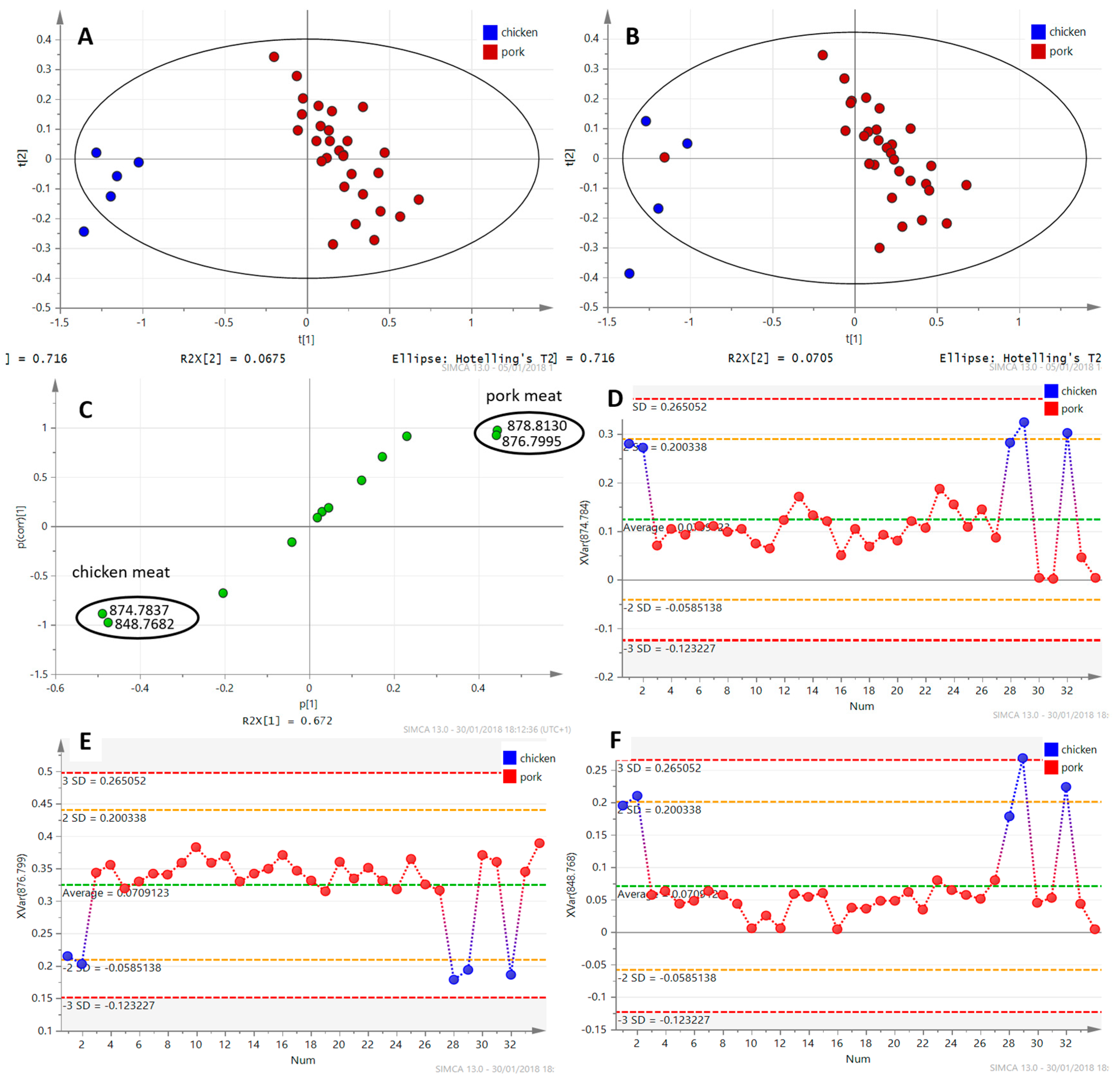
3.1.1. DART–HRMS Fingerprints of Different Meat Types
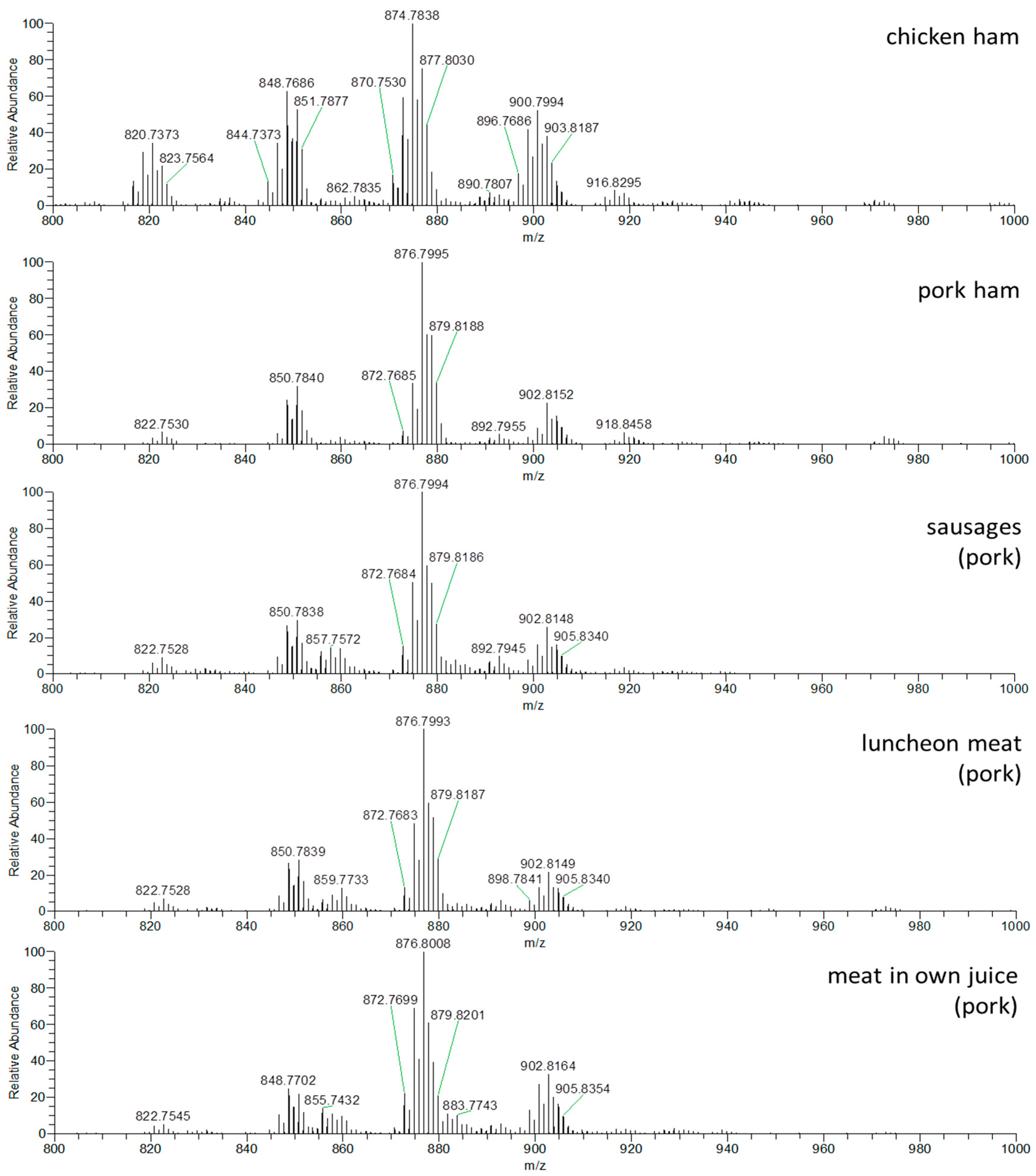
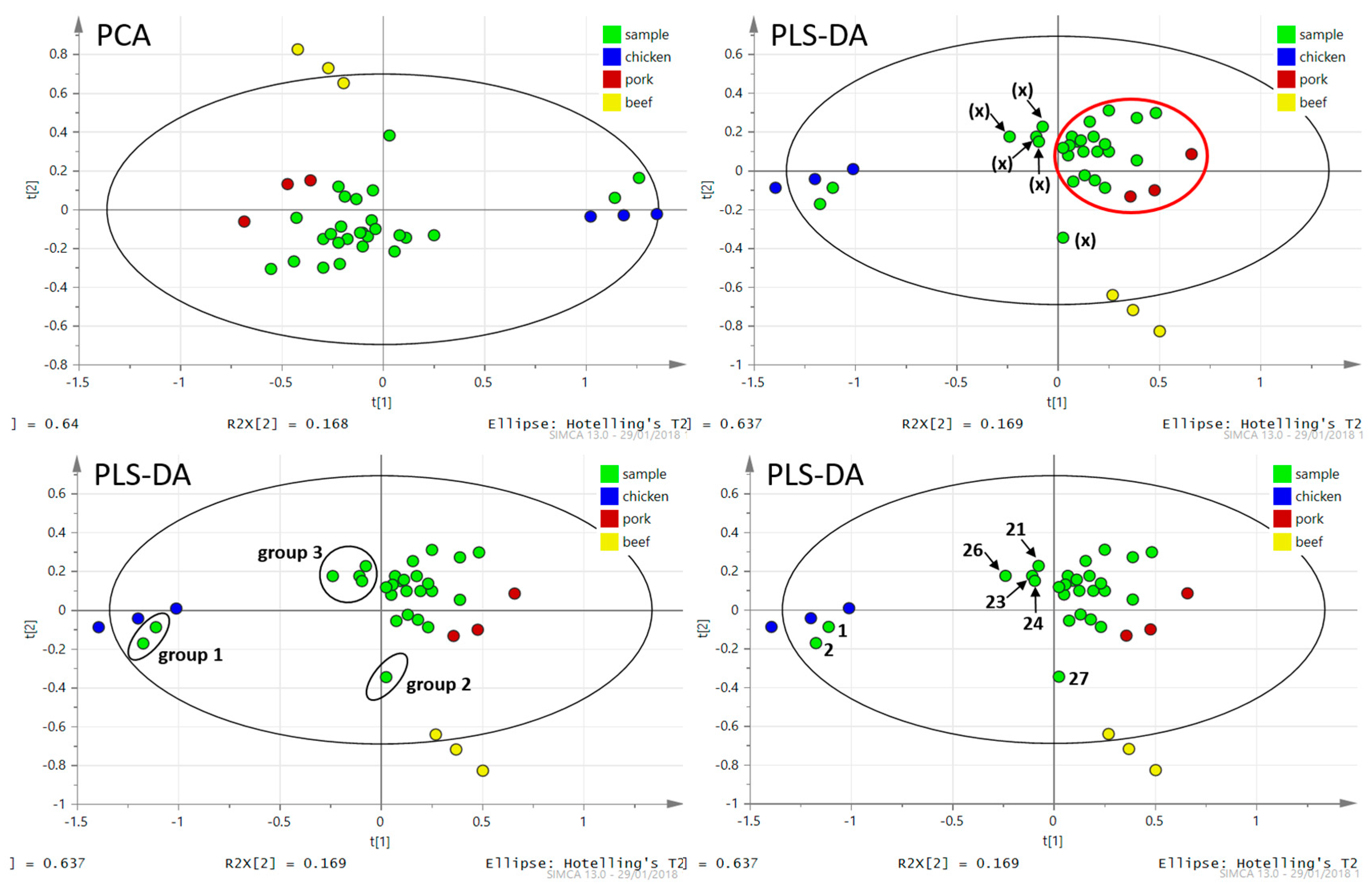
3.1.2. DART–HRMS Analysis of Real Meat Products
3.1.3. Confirmation of Isolated DNA Quality and Quantity
3.1.4. DNA Analysis of Meat and Meat Products
3.1.5. Comparison of Methods
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Hsieh, Y.P.; Woodward, B.B.; Ho, S.H. Detection of Species Substitution in Raw and Cooked Meats Using Immunoassays. J. Food Prot. 1995, 58, 555–559. [Google Scholar] [CrossRef] [PubMed]
- Ballin, N.Z.; Vogensen, F.K.; Karlsson, A.H. Species determination—Can we detect and quantify meat adulteration? Meat Sci. 2009, 83, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, V.; González, I.; Rojas, M.; García, T.; Martín, R. A review of current PCR-based methodologies for the authentication of meats from game animal species. Trends Food Sci. Technol. 2010, 21, 408–421. [Google Scholar] [CrossRef]
- Köppel, R.; Daniels, M.; Felderer, N.; Brünen-Nieweler, C. Multiplex real-time PCR for the detection and quantification of DNA from duck, goose, chicken, turkey and pork. Eur. Food Res. Technol. 2013, 236, 1093–1098. [Google Scholar] [CrossRef]
- Iwobi, A.; Sebah, D.; Spielmann, G.; Maggipinto, M.; Schrempp, M.; Kraemer, I.; Gerdes, L.; Busch, U.; Huber, I. A multiplex real-time PCR method for the quantitative determination of equine (horse) fractions in meat products. Food Control 2017, 74, 89–97. [Google Scholar] [CrossRef]
- Griffiths, A.M.; Sotelo, C.G.; Mendes, R.; Pérez-Martín, R.I.; Schröder, U.; Shorten, M.; Silva, H.A.; Verrez-Bagnis, V.; Mariani, S. Current methods for seafood authenticity testing in Europe: Is there a need for harmonisation? Food Control 2014, 45, 95–100. [Google Scholar] [CrossRef] [Green Version]
- Murugaiah, C.; Al-Talib, H.; Radu, S. Forensics: Food Authentication Using MtDNA. J. Nutr. Health Food Sci. 2015, 3, 1–10. [Google Scholar] [CrossRef]
- Mozola, M.A.; Peng, X.; Wendorf, M.; Artiga, L. Evaluation of the GeneQuence® DNA hybridization method in conjunction with 24-h enrichment protocols for detection of Salmonella spp. in select foods: Collaborative study. J. AOAC Int. 2007, 90, 738–755. [Google Scholar] [CrossRef] [Green Version]
- Tichoniuk, M.; Ligaj, M.; Filipiak, M. Application of DNA hybridization biosensor as a screening method for the detection of genetically modified food components. Sensors 2008, 8, 2118–2135. [Google Scholar] [CrossRef] [Green Version]
- Chapela, M.J.; Sotelo, C.G.; Calo-Mata, P.; Pérez-Martín, R.I.; Rehbein, H.; Hold, G.L.; Quinteiro, J.; Rey-Méndez, M.; Rosa, C.; Santos, A.T. Identification of Cephalopod Species (Ommastrephidae and Loliginidae) in Seafood Products by Forensically Informative Nucleotide Sequencing (FINS). J. Food Sci. 2002, 67, 1672–1676. [Google Scholar] [CrossRef]
- Iwobi, A.; Sebah, D.; Kraemer, I.; Losher, C.; Fischer, G.; Busch, U.; Huber, I. A multiplex real-time PCR method for the quantification of beef and pork fractions in minced meat. Food Chem. 2015, 169, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Chikuni, K.; Tanabe, R.; Muroya, S.; Shibata, K.; Yamada, J.; Shinmura, Y. A quick and simple method for the identification of meat species and meat products by PCR assay. Meat Sci. 1999, 51, 143–148. [Google Scholar] [CrossRef]
- Barbuto, M.; Galimberti, A.; Ferri, E.; Labra, M.; Malandra, R.; Galli, P.; Casiraghi, M. DNA barcoding reveals fraudulent substitutions in shark seafood products: The Italian case of “palombo” (Mustelus spp.). Food Res. Int. 2010, 43, 376–381. [Google Scholar] [CrossRef]
- Thakur, M.; Singh, S.; Shukla, M.; Sharma, L.; Agarwal, N.; Goyal, S.; Sambandam, S. Identification of Galliformes through Forensically Informative Nucleotide Sequencing (FINS) and its Implication in Wildlife Forensics. J. Forensic Res. 2012, 4. [Google Scholar] [CrossRef]
- Jírů, M.; Stranska, M.; Kocourek, V.; Krmela, A.; Tomaniová, M.; Rosmus, J.; Hajslova, J. Authentication of meat species and net muscle proteins: Updating of an old concept. Czech J. Food Sci. 2019, 37, 205–211. [Google Scholar] [CrossRef] [Green Version]
- Ruiz Orduna, A.; Husby, E.; Yang, C.T.; Ghosh, D.; Beaudry, F. Assessment of meat authenticity using bioinformatics, targeted peptide biomarkers and high-resolution mass spectrometry. Food Addit. Contam. Part A 2015, 32, 1709–1717. [Google Scholar] [CrossRef]
- Cody, R.B.; Laramée, J.A.; Durst, H.D. Versatile new ion source for the analysis of materials in open air under ambient conditions. Anal. Chem. 2005, 77, 2297–2302. [Google Scholar] [CrossRef]
- Hajslova, J.; Cajka, T.; Vaclavik, L. Challenging applications offered by direct analysis in real time (DART) in food-quality and safety analysis. TrAC Trends Anal. Chem. 2011, 30, 204–218. [Google Scholar] [CrossRef]
- Worley, B.; Powers, R. Multivariate Analysis in Metabolomics. Curr. Metab. 2013, 1, 92–107. [Google Scholar] [CrossRef]
- Blasco, H.; Błaszczyński, J.; Billaut, J.C.; Nadal-Desbarats, L.; Pradat, P.F.; Devos, D.; Moreau, C.; Andres, C.R.; Emond, P.; Corcia, P.; et al. Comparative analysis of targeted metabolomics: Dominance-based rough set approach versus orthogonal partial least square-discriminant analysis. J. Biomed. Inform. 2015, 53, 291–299. [Google Scholar] [CrossRef]
- Berrueta, L.A.; Alonso-Salces, R.M.; Héberger, K. Supervised pattern recognition in food analysis. J. Chromatogr. A 2007, 1158, 196–214. [Google Scholar] [CrossRef]
- Triba, M.N.; Le Moyec, L.; Amathieu, R.; Goossens, C.; Bouchemal, N.; Nahon, P.; Rutledge, D.N.; Savarin, P. PLS/OPLS models in metabolomics: The impact of permutation of dataset rows on the K-fold cross-validation quality parameters. Mol. Biosyst. 2015, 11, 13–19. [Google Scholar] [CrossRef]
- ISO. Foodstuffs-Methods of Analysis for the Detection of Genetically Modified Organisms and Derived Products-Nucleic Acid Extraction; ANSI: New York, NY, USA, 2005. [Google Scholar]
- Zdeňková, K.; Akhatova, D.; Fialová, E.; Krupa, O.; Kubica, L.; Lencová, S.; Demnerová, K. Detection of meat adulteration: Use of efficient and routine-suited multiplex polymerase chain reaction-based methods for species authentication and quantification in meat products. J. Food Nutr. Res. 2018, 57, 351–362. [Google Scholar]
- Laube, I.; Zagon, J.; Spiegelberg, A.; Butschke, A.; Kroh, L.W.; Broll, H. Development and design of a ‘ready-to-use’ reaction plate for a PCR-based simultaneous detection of animal species used in foods. Int. J. Food Sci. Technol. 2007, 42, 9–17. [Google Scholar] [CrossRef]
- Köppel, R.; Ruf, J.; Zimmerli, F.; Breitenmoser, A. Multiplex real-time PCR for the detection and quantification of DNA from beef, pork, chicken and turkey. Eur. Food Res. Technol. 2008, 227, 1199–1203. [Google Scholar] [CrossRef]
- Laube, I.; Spiegelberg, A.; Butschke, A.; Zagon, J.; Schauzu, M.; Kroh, L.; Broll, H. Methods for the detection of beef and pork in foods using real-time polymerase chain reaction. Int. J. Food Sci. Technol. 2003, 38, 111–118. [Google Scholar] [CrossRef]
- Hurkova, K.; Uttl, L.; Rubert, J.; Navratilova, K.; Kocourek, V.; Stranska-Zachariasova, M.; Paprstein, F.; Hajslova, J. Cranberries versus lingonberries: A challenging authentication of similar Vaccinium fruit. Food Chem. 2019, 284, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Hrbek, V.; Rektorisova, M.; Chmelarova, H.; Ovesna, J.; Hajslova, J. Authenticity assessment of garlic using a metabolomic approach based on high resolution mass spectrometry. J. Food Compos. Anal. 2018, 67, 19–28. [Google Scholar] [CrossRef]

| Sample | Product | Pork Meat/Lard (%) | Chicken Meat (%) | Beef Meat (%) | Max Fat Content (%) |
|---|---|---|---|---|---|
| 1 | Chicken ham | – | 92 | – | 1.5 |
| 2 | Poultry ham | – | 60 | – | 1.5 |
| 3 | Pork ham | 95 | – | – | 10 |
| 4 | Sausage | 16/Y | – | 35 | 40 |
| 5 | Sausage | 40/Y | – | 10 | 40 |
| 6 | Sausage | 62/Y | – | 23 | 40 |
| 7 | Sausage | 40/25 | – | 10 | 34 |
| 8 | Sausage | 54/Y | – | 26 | 44 |
| 9 | Luncheon meat | N/Y | – | Y | 40 |
| 10 | Sausage | 16/Y | – | 35 | 45 |
| 11 | Sausage | 17/Y | – | 26 | N |
| 12 | Sausage | 17.5/Y | – | 38.5 | 45 |
| 13 | Meat in natural juices | 70 | – | – | 33 |
| 14 | Luncheon meat | 79 | – | – | 30 |
| 15 | Meat in natural juices | 92 | – | – | N |
| 16 | Meat in natural juices | 70 | – | – | 40 |
| 17 | Meat in natural juices | 70 | – | – | 30 |
| 18 | Sausage | 33/30 | – | 22 | N |
| 19 | Sausage | 71/Y | – | 16 | 45 |
| 20 | Sausage | 43/30 | – | 17 | 45 |
| 21 | Luncheon meat | 48 | Y | – | 40 |
| 22 | Meat in natural juices | 30 + MSM/Y | – | N | |
| 23 | Luncheon meat | 18 | 32 | – | 30 |
| 24 | Luncheon meat | 35 | 30 | – | 25 |
| 25 | Luncheon meat | 71 | – | 40 | |
| 26 | Luncheon meat | 31/Y | 39 | – | 26 |
| 27 | Meat in natural juices | Y | Y | 70 | 27 |
| Meat Species | Name of Primer | Target Sequence | Sequence of Primer [5′-3′] | Product [bp] | References |
|---|---|---|---|---|---|
| Universal F | SIM | Cytochrome b (mtDNA) | GACCTCCCAGCTCCATCAAACATCTCATCTTGATGAAA | [12] | |
| Beef R | B | CTAGAAAAGTGTAAGACCCGTAATATAAG | 274 | [12] | |
| Pork R | P | GCTGATAGTAGATTTGTGATGACCGTA | 398 | [12] | |
| Chicken, turkey R | C | CGTATTGTACGTTCCGGCAAG | 169 | [24] | |
| Horse R | H | CTCAGATTCACTCGACGAGGGTAGTA | 439 | [12] | |
| Beef | Bos-PDE-f | Cyclic-GMP-phospho-diesterase (gDNA) | ACTCCTACCCATCATGCAGAT | 104 | [11,25] |
| Bos-PDE-r | TGTTTTTAAATATTTCAGCTAAGAAAAA | ||||
| Bos-PDE-p | TexasRed: AACATCAGGATTTTTGCTGCATTTGC:BHQ2 | ||||
| Pork | Sus1-F | Beta-actin (gDNA) | CGAGAGGCTGCCGTAAAGG | 107 | [11,26] |
| Sus1-R | TGCAAGGAACACGGCTAAGTG | ||||
| Sus1-p | HEX:TCTGACGTGACTCCCCGACCTGG:BHQ1 | ||||
| Mammals and poultry | MY-F | Myostatin (gDNA) | TTGTGCAAATCCTGAGACTCAT | 97 | [11,27] |
| MY-R | ATACCAGTGCCTGGGTTCAT | ||||
| My-p | FAM:CCCATGAAAGACGGTACAAGGTATACTG:BHQ1 | ||||
| Chicken | ChIn-F | Interleukin-2 (gDNA) | TGTTACCTGGGAGAAGTGGTTACT | 135 | [25] |
| ChIn-R | CTGACCATAAAGAATACCTACCG | [24] | |||
| ChIn-p | TAMRA:TGAAGAAAGAAACTGAAGATGACACTGAAATTAAAG:BHQ2 | [25] |
| m/z | Δppm | Formula | Identification | Significant Ions for |
|---|---|---|---|---|
| 846.7535 | 1.259 | C53H100NO6 | C 50:3 | chicken |
| 848.7682 | 1.339 | C53H102NO6 | C 50:2 | chicken/pork |
| 850.7836 | 1.559 | C53H104NO6 | C 50:1 | chicken/pork |
| 852.7975 | 4.820 | C53H106NO6 | C 50:0 | chicken/pork |
| 872.7683 | 2.104 | C55H102NO6 | C 52:4 | chicken |
| 874.7837 | 1.562 | C55H104NO6 | C 52:3 | chicken |
| 876.7995 | 1.672 | C55H106NO6 | C 52:2 | pork/beef |
| 878.8130 | 2.522 | C55H108NO6 | C 52:1 | pork/beef |
| 880.8246 | 4.112 | C55H110NO6 | C 52:0 | pork/beef |
| 896.7689 | 1.412 | C57H102NO6 | C 54:6 | chicken |
| 898.7841 | 1.898 | C57H104NO6 | C 54:5 | chicken |
| 900.7996 | 2.050 | C57H106NO6 | C 54:4 | chicken |
| 902.8148 | 3.636 | C57H108NO6 | C 54:3 | pork/beef |
| 904.8306 | 1.709 | C57H110NO6 | C 54:2 | beef |
| 906.8457 | 2.974 | C57H112NO6 | C 54:1 | beef |
| Product | Declared Composition | mPCR | mqPCR | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Pork | Chicken | Beef | Pork | Chicken/Turkey | Beef | Pork | Chicken | Beef | ||
| 1 | Chicken ham | - | + | - | - | + | - | - | + | - |
| 2 | Poultry ham | - | + | - | - | + | - | - | + | - |
| 3 | Pork ham | + | - | - | + | - | - | + | - | - |
| 4 | Sausage | + | - | + | + | - | + | + | - | + |
| 5 | Sausage | + | - | + | + | - | + | + | - | + |
| 6 | Sausage | + | - | + | + | - | + | + | - | + |
| 7 | Sausage | + | - | + | + | - | + | + | - | + |
| 8 | Sausage | + | - | + | + | - | + | + | - | + |
| 9 | Luncheon meat | + | - | + | + | - | + | + | - | + |
| 10 | Sausage | + | - | + | + | - | + | + | - | + |
| 11 | Sausage | + | - | + | + | - | + | + | - | + |
| 12 | Sausage | + | - | + | + | - | + | + | - | + |
| 13 | Meat in natural juices | + | - | - | + | - | - | + | - | - |
| 14 | Luncheon meat | + | - | - | + | - | - | + | - | - |
| 15 | Meat in natural juices | + | - | - | + | - | - | + | - | - |
| 16 | Meat in natural juices | + | - | - | + | - | - | + | - | - |
| 17 | Meat in natural juices | + | - | - | + | - | - | + | - | - |
| 18 | Sausage | + | - | + | + | - | + | + | - | + |
| 19 | Sausage | + | - | + | + | - | + | + | - | + |
| 20 | Sausage | + | - | + | + | - | + | + | - | + |
| 21 | Luncheon meat | + | + | - | -ᵻ | + | - | + | + | - |
| 22 | Meat in natural juices | + | - | - | + | - | - | + | - | - |
| 23 | Luncheon meat | + | + | - | + | + | +ᵻ | + | + | - |
| 24 | Luncheon meat | + | + | - | -ᵻ | + | - | + | + | - |
| 25 | Luncheon meat | + | - | - | + | - | - | + | - | - |
| 26 | Luncheon meat | + | + | - | + | + | +ᵻ | + | + | - |
| 27 | Meat in natural juices | + | + | + | + | + | -ᵻ | + | + | + |
| Parameters | DART–HRMS | PCR |
|---|---|---|
| Target molecule | Triacylglycerols | DNA |
| Preparation step | Hexane extract–lipophilic fraction containing triacylglycerols | DNA isolation—many methods available |
| Capacity of the machine | High-throughput method (+++) | Mainly 96 reactions in one run (++) |
| Cost of the analysis (only the retail price of chemicals is included) | Very low (+++) | Low (++) |
| Duration | Extraction: moderate (+++) Analysis: quick (+++) Evaluation: long | Extraction: moderate (++) Analysis: moderate (++) Evaluation: quick (+++) |
| Price of the required instrumentation | High (-) | Low for classical PCR instrument, moderate for qPCR device (+) |
| Feasibility for analysis of raw products | Yes (+++) | Yes, reliable (+++) |
| Feasibility for analysis of heat-treated meat products | Yes (++) | Yes, reliable (+++) |
| Feasibility for analysis of mixtures | Yes (++) | Yes, reliable (+++) |
| Feasibility for analysis of products containing high amounts of fat | Yes (+++) | Yes, gDNA is recommended as a target (+++) |
| Conduction of the experiment | Laboratory device (-), performing the analysis (+++) | Laboratory device (++), performing the analysis (+) |
| Claims for evaluation of results | Demanding for evaluation–experience is needed, because of the statistical analysis. | Simple to evaluate (+++). The PCR amplicon or fluorescence curve is or is not there, which is clearly visible from the primary results |
| Usage | Screening method | Confirmatory method |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hrbek, V.; Zdenkova, K.; Jilkova, D.; Cermakova, E.; Jiru, M.; Demnerova, K.; Pulkrabova, J.; Hajslova, J. Authentication of Meat and Meat Products Using Triacylglycerols Profiling and by DNA Analysis. Foods 2020, 9, 1269. https://doi.org/10.3390/foods9091269
Hrbek V, Zdenkova K, Jilkova D, Cermakova E, Jiru M, Demnerova K, Pulkrabova J, Hajslova J. Authentication of Meat and Meat Products Using Triacylglycerols Profiling and by DNA Analysis. Foods. 2020; 9(9):1269. https://doi.org/10.3390/foods9091269
Chicago/Turabian StyleHrbek, Vojtech, Kamila Zdenkova, Diliara Jilkova, Eliska Cermakova, Monika Jiru, Katerina Demnerova, Jana Pulkrabova, and Jana Hajslova. 2020. "Authentication of Meat and Meat Products Using Triacylglycerols Profiling and by DNA Analysis" Foods 9, no. 9: 1269. https://doi.org/10.3390/foods9091269
APA StyleHrbek, V., Zdenkova, K., Jilkova, D., Cermakova, E., Jiru, M., Demnerova, K., Pulkrabova, J., & Hajslova, J. (2020). Authentication of Meat and Meat Products Using Triacylglycerols Profiling and by DNA Analysis. Foods, 9(9), 1269. https://doi.org/10.3390/foods9091269




