Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling Design
2.2. Isolation of Suspected MRSA
2.3. MRSA Confirmation
2.4. Antibiotic Resistance Profiling of Confirmed MRSA Isolates
2.4.1. Disk Diffusion Assays
2.4.2. Minimum Inhibitory Concentration Assays
2.4.3. Antimicrobial Resistance Genes Determination
- Multiplex 1 (aacA-aphD, tetL, tetM, dfrA, dfrD): 95 °C (5 min), followed by 32 cycles of 95 °C (30 s), 45 °C (45 s), 72 °C (1 min) and a final extension at 72 °C (10 min).
- Multiplex 2 (blaZ, dfrK, vgaE): 95 °C (5 min), followed by 32 cycles of 95 °C (30 s), 47 °C (45 s), 72 °C (50 s) and a final extension at 72 °C (10 min).
- Multiplex 3 (ermT, norA, vgaA): 95 °C (5 min), followed by 31 cycles of 95 °C (30 s), 49 °C (40 s), 72 °C (1 min) and a final extension at 72 °C (10 min).
- Multiplex 4 (aadD, cfr): 95 °C (5 min), followed by 31 cycles of 95 °C (30 s), 52 °C (40 s), 72 °C (50 s) and a final extension at 72 °C (10 min).
- PCR 5 (fexA): 95 °C (5 min), followed by 31 cycles of 95 °C (30 s), 56 °C (40 s), 72 °C (1 min) and a final extension at 72 °C (10 min).
2.5. Enterotoxin Gene Determination
2.6. MRSA Molecular Subtyping and Phylogenetic Analysis
2.7. Quantification of Biofilm Production
2.8. Statistical Analysis
3. Results
3.1. MRSA Detection
3.2. Antibiotic Resistance Profiles of MRSA Isolates
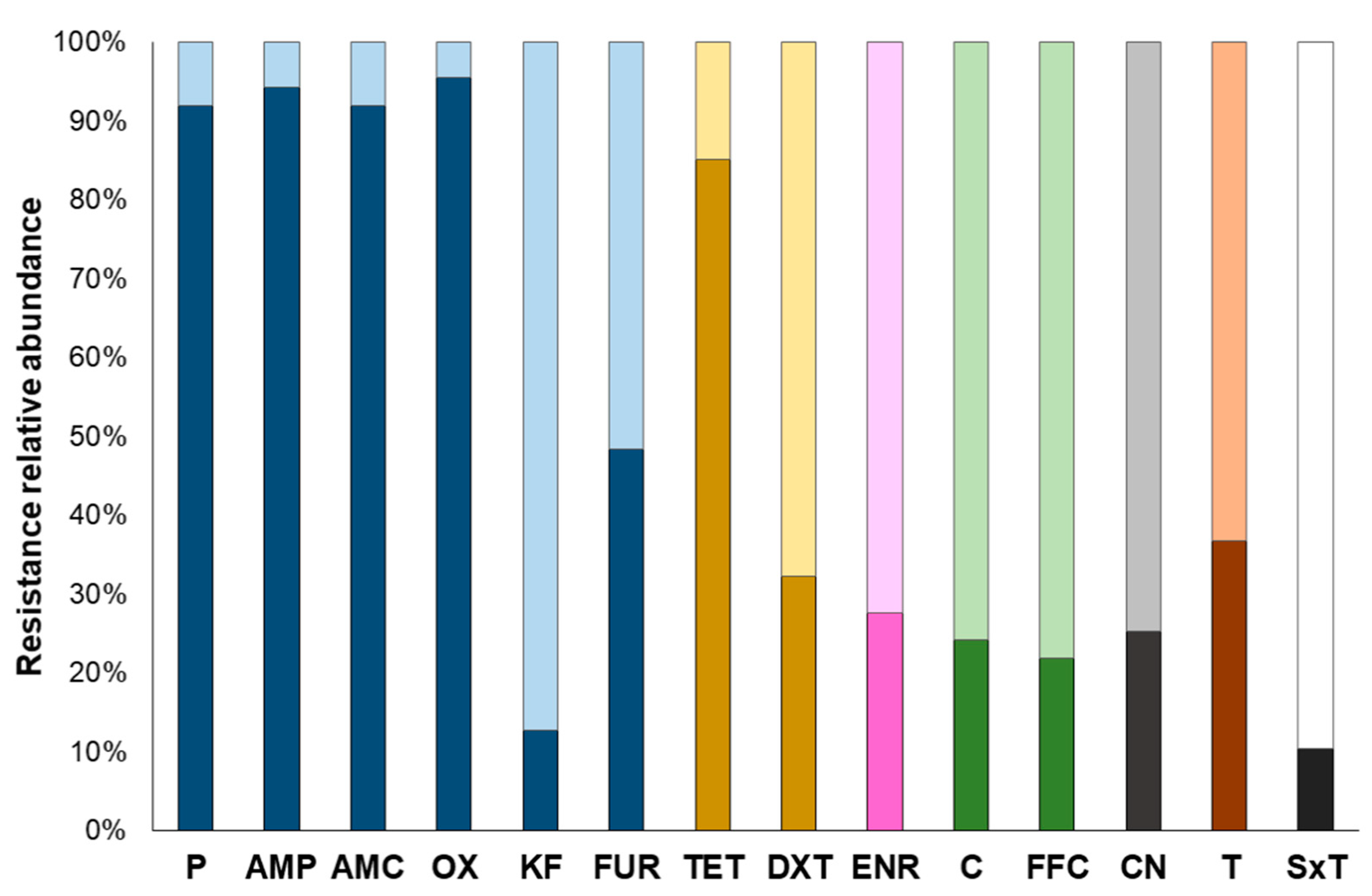
3.2.1. Phenotypic Resistance
3.2.2. Minimum Inhibitory Concentration (MIC) Assays
3.2.3. Presence of Antibiotic Resistance Genes
3.3. Presence of Enterotoxin Genes
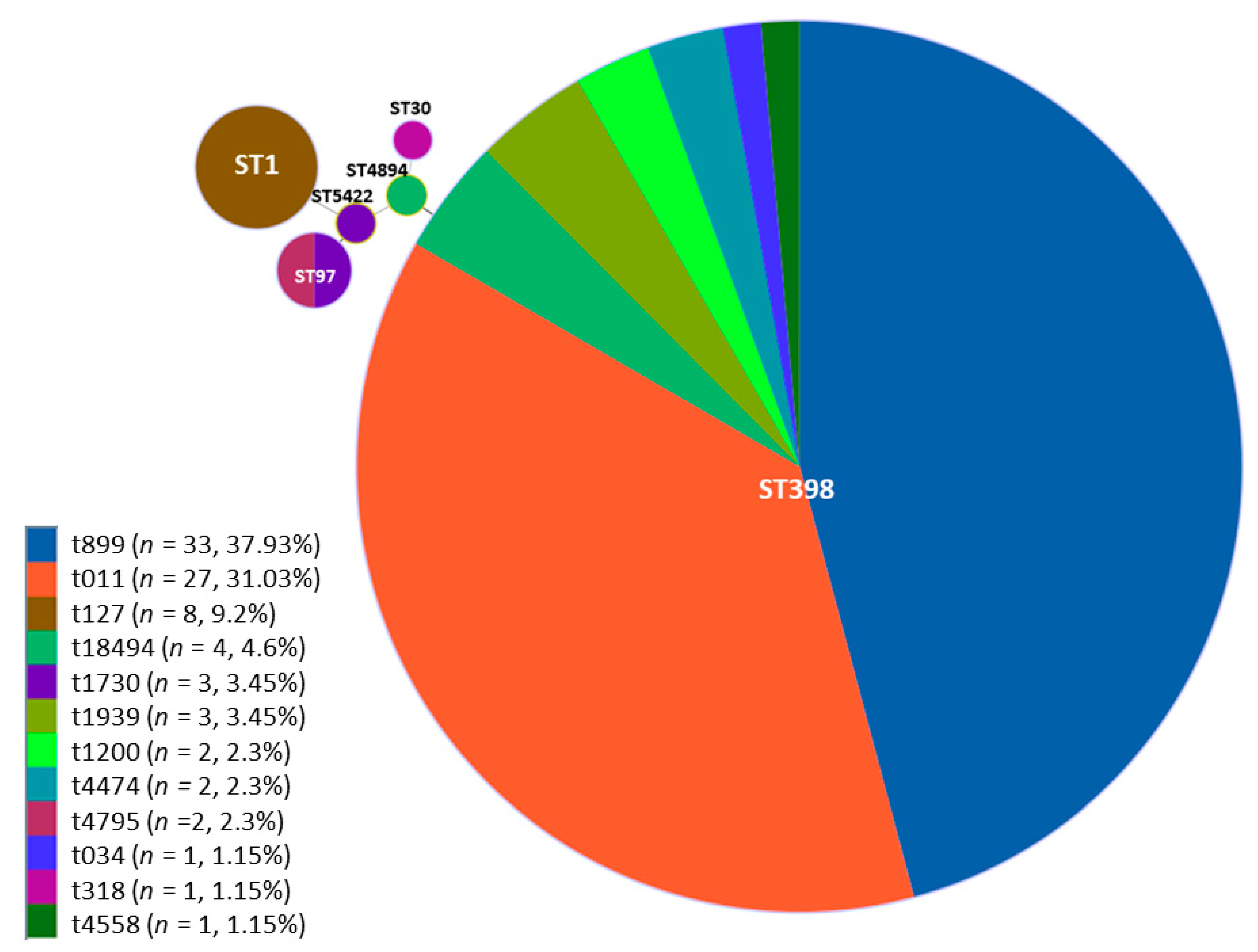
3.4. Molecular Subtyping of MRSA Isolates
3.5. Biofilm Production Assays
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Landers, T.F.; Cohen, B.; Wittum, T.E.; Larson, E.L. A review of antibiotic use in food animals: Perspective, policy, and potential. Public Health Rep. 2012, 127, 4–22. [Google Scholar] [CrossRef] [PubMed]
- Oniciuc, E.A.; Likotrafiti, E.; Alvarez-Molina, A.; Prieto, M.; López, M.; Alvarez-Ordóñez, A. Food processing as a risk factor for antimicrobial resistance spread along the food chain. Curr. Opin. Food Sci. 2019, 30, 21–26. [Google Scholar] [CrossRef]
- Guardabassi, L.; Larsen, J.; Weese, J.S.; Butaye, P.; Battisti, A.; Kluytmans, J.; Lloyd, D.H.; Skov, R.L. Public health impact and antimicrobial selection of meticillin-resistant staphylococci in animals. J. Glob. Antimicrob. Resist. 2013, 1, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Weese, J.S.; van Duijkeren, E. Methicillin-resistant Staphylococcus aureus and Staphylococcus pseudintermedius in veterinary medicine. Vet. Microbiol. 2010, 140, 418–429. [Google Scholar] [CrossRef]
- Savage, V.J.; Chopra, I.; O’Neill, A.J. Staphylococcus aureus biofilms promote horizontal transfer of antibiotic resistance. Antimicrob. Agents Chemother. 2013, 57, 1968–1970. [Google Scholar] [CrossRef]
- Grundmann, H.; Tiemersma, E. Global prevalence of meticillin-resistant Staphylococcus aureus. Lancet 2006, 368, 1866–1867. [Google Scholar] [CrossRef]
- Gajdács, M. The continuing threat of methicillin-resistant Staphylococcus aureus. Antibiotics 2019, 8, 52. [Google Scholar] [CrossRef]
- Feßler, A.T.; Olde Riekerink, R.G.M.; Rothkamp, A.; Kadlec, K.; Sampimon, O.C.; Lam, T.J.G.M.; Schwarz, S. Characterization of methicillin-resistant Staphylococcus aureus CC398 obtained from humans and animals on dairy farms. Vet. Microbiol. 2012, 160, 77–84. [Google Scholar] [CrossRef]
- Köck, R.; Loth, B.; Köksal, M.; Schulte-Wülwer, J.; Harlizius, J.; Friedrich, A.W. Persistence of nasal colonization with livestock-associated methicillin-resistant Staphylococcus aureus in pig farmers after holidays from pig exposure. Appl. Environ. Microbiol. 2012, 78, 4046–4047. [Google Scholar] [CrossRef]
- Kluytmans, J.A.J.W. Methicillin-resistant Staphylococcus aureus in food products: Cause for concern or case for complacency? Clin. Microbiol. Infect. 2010, 16, 11–15. [Google Scholar] [CrossRef]
- Voss, A.; Loeffen, F.; Bakker, J.; Klaassen, C.; Wulf, M. Methicillin-resistant Staphylococcus aureus in Pig Farming. Emerg. Infect. Dis. 2005, 11, 1965–1966. [Google Scholar] [CrossRef] [PubMed]
- Smith, T.C.; Pearson, N. The emergence of Staphylococcus aureus ST398. Vector-Borne Zoonotic Dis. 2011, 11, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Armand-Lefevre, L.; Ruimy, R.; Andremont, A. Clonal comparison of Staphylococcus from healthy pig farmers, human controls, and pigs. Emerg. Infect. Dis. 2005, 11, 711–714. [Google Scholar] [CrossRef] [PubMed]
- Hetem, D.J.; Bootsma, M.C.J.; Troelstra, A.; Bonten, M.J.M. Transmissibility of livestock-associated methicillin-resistant Staphylococcus aureus. Emerg. Infect. Dis. 2013, 19, 1797–1802. [Google Scholar] [CrossRef] [PubMed]
- Goerge, T.; Lorenz, M.B.; van Alen, S.; Hübner, N.O.; Becker, K.; Köck, R. MRSA colonization and infection among persons with occupational livestock exposure in Europe: Prevalence, preventive options and evidence. Vet. Microbiol. 2017, 200, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Boswihi, S.S.; Udo, E.E. Methicillin-resistant Staphylococcus aureus: An update on the epidemiology, treatment options and infection control. Curr. Med. Res. Pract. 2018, 8, 18–24. [Google Scholar] [CrossRef]
- Gómez-Sanz, E.; Torres, C.; Lozano, C.; Fernández-Pérez, R.; Aspiroz, C.; Ruiz-Larrea, F.; Zarazaga, M. Detection, molecular characterization, and clonal diversity of methicillin-resistant Staphylococcus aureus CC398 and CC97 in Spanish slaughter pigs of different age groups. Foodborne Pathog. Dis. 2010, 7, 1269–1277. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Caruso, M.; Normanno, G.; Latorre, L.; Miccolupo, A.; Fraccalvieri, R.; Intini, F.; Manginelli, T.; Santagada, G. MRSA in swine, farmers and abattoir workers in Southern Italy. Food Microbiol. 2019, 82, 287–293. [Google Scholar] [CrossRef]
- Filipello, V.; Di Ciccio, P.A.; Colagiorgi, A.; Tilola, M.; Romanò, A.; Vitale, N.; Losio, M.N.; Luini, M.; Zanardi, E.; Ghidini, S.; et al. Molecular characterisation and biofilm production in Staphylococcusaureus isolates from the dairy production chain in Northern Italy. Int. Dairy J. 2019, 91, 110–118. [Google Scholar] [CrossRef]
- I.Stat Consistenze Degli Allevamenti (Consistency of Farms). Available online: http://dati.istat.it/Index.aspx?DataSetCode=DCSP_CONSISTENZE# (accessed on 24 July 2020).
- European Food Safety Authority. Analysis of the baseline survey on the prevalence of methicillin-resistant Staphylococcus aureus (MRSA) in holdings with breeding pigs, in the EU, 2008—Part A: MRSA prevalence estimates. EFSA J. 2009, 7, 1376. [Google Scholar] [CrossRef]
- Battisti, A.; Franco, A.; Merialdi, G.; Hasman, H.; Iurescia, M.; Lorenzetti, R.; Feltrin, F.; Zini, M.; Aarestrup, F.M. Heterogeneity among methicillin-resistant Staphylococcus aureus from Italian pig finishing holdings. Vet. Microbiol. 2010, 142, 361–366. [Google Scholar] [CrossRef] [PubMed]
- Normanno, G.; Dambrosio, A.; Lorusso, V.; Samoilis, G.; Di Taranto, P.; Parisi, A. Methicillin-resistant Staphylococcus aureus (MRSA) in slaughtered pigs and abattoir workers in Italy. Food Microbiol. 2015, 51, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Doulgeraki, A.I.; Di Ciccio, P.; Ianieri, A.; Nychas, G.J.E. Methicillin-resistant food-related Staphylococcus aureus: A review of current knowledge and biofilm formation for future studies and applications. Res. Microbiol. 2017, 168, 1–15. [Google Scholar] [CrossRef]
- Giaouris, E.; Simões, M.; Dubois-Brissonnet, F. The Role of Biofilms in the Development and Dissemination of Microbial Resistance within the Food Industry. Foods 2020, 9, 816. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Hansen, M.F.; Røder, H.L.; Wang, N.; Burmølle, M.; He, G. Mixed-species biofilms in the food industry: Current knowledge and novel control strategies. Crit. Rev. Food Sci. Nutr. 2019, 60, 2277–2293. [Google Scholar] [CrossRef] [PubMed]
- Craft, K.M.; Nguyen, J.M.; Berg, L.J.; Townsend, S.D. Methicillin-resistant: Staphylococcus aureus (MRSA): Antibiotic-resistance and the biofilm phenotype. Medchemcomm 2019, 10, 1231–1241. [Google Scholar] [CrossRef]
- Vergara, A.; Normanno, G.; Di Ciccio, P.; Pedonese, F.; Nuvoloni, R.; Parisi, A.; Santagada, G.; Colagiorgi, A.; Zanardi, E.; Ghidini, S.; et al. Biofilm Formation and Its Relationship with the Molecular Characteristics of Food-Related Methicillin-Resistant Staphylococcus aureus (MRSA). J. Food Sci. 2017, 82, 2364–2370. [Google Scholar] [CrossRef]
- European Commission. EUROSTAT Database. Available online: https://ec.europa.eu/eurostat/web/agriculture/data/database (accessed on 29 July 2020).
- ISO. ISO 6888-2:1999-Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Coagulase-Positive Staphylococci (Staphylococcus aureus and Other Species)—Part 2: Technique Using Rabbit Plasma Fibrinogen Agar Medium. Available online: https://www.iso.org/standard/25571.html (accessed on 25 July 2020).
- Pichon, B.; Hill, R.; Laurent, F.; Larsen, A.R.; Skov, R.L.; Holmes, M.; Edwards, G.F.; Teale, C.; Kearns, A.M. Development of a real-time quadruplex PCR assay for simultaneous detection of nuc, panton-valentine leucocidin (PVL), mecA and homologue mecALGA251. J. Antimicrob. Chemother. 2012, 67, 2338–2341. [Google Scholar] [CrossRef]
- Committee for Laboratroy Standards Institute. Performance Standards for Antimicrobial Disk Susceptibility Tests, 27th ed.; Committee for Laboratroy Standards Institute: Wayne, MN, USA, 2017; Volume 37, ISBN 1-56238-1-56238-805-3. [Google Scholar]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- World Health Organization. Critically Important Antimcrobials for Human Medicine, 6th ed.; World Health Organization: Geneva, Switzerland, 2019; ISBN 978-92-4-151552-8. [Google Scholar]
- OIE List of Antimicrobials of Veterinary Importance. World Organ. Anim. Health 2007, 75, 1–9.
- European Committee on Antimicrobial Susceptibility Testing. Testing Breakpoint Tables for Interpretation of MICs and Zone Diameters (Version 10.0); EUCAST: Växjö, Sweden, 2020. [Google Scholar]
- Vali, L.; Davies, S.E.; Lai, L.L.G.; Dave, J.; Amyes, S.G.B. Frequency of biocide resistance genes, antibiotic resistance and the effect of chlorhexidine exposure on clinical methicillin-resistant Staphylococcus aureus isolates. J. Antimicrob. Chemother. 2008, 61, 524–532. [Google Scholar] [CrossRef]
- Inkster, T.; Coia, J.; Meunier, D.; Doumith, M.; Martin, K.; Pike, R.; Imrie, L.; Kane, H.; Hay, M.; Wiuff, C.; et al. First outbreak of colonization by linezolid- and glycopeptide-resistant Enterococcus faecium harbouring the cfr gene in a UK nephrology unit. J. Hosp. Infect. 2017, 97, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Kehrenberg, C.; Schwarz, S. Distribution of florfenicol resistance genes fexA and cfr among chloramphenicol-resistant Staphylococcus isolates. Antimicrob. Agents Chemother. 2006, 50, 1156–1163. [Google Scholar] [CrossRef] [PubMed]
- Lozano, C.; Aspiroz, C.; Rezusta, A.; Gómez-Sanz, E.; Simon, C.; Gómez, P.; Ortega, C.; Revillo, M.J.; Zarazaga, M.; Torres, C. Identification of novel vga(A)-carrying plasmids and a Tn5406-like transposon in meticillin-resistant Staphylococcus aureus and Staphylococcus epidermidis of human and animal origin. Int. J. Antimicrob. Agents 2012, 40, 306–312. [Google Scholar] [CrossRef] [PubMed]
- Argudín, M.A.; Vanderhaeghen, W.; Butaye, P. Diversity of antimicrobial resistance and virulence genes in methicillin-resistant non-Staphylococcus aureus staphylococci from veal calves. Res. Vet. Sci. 2015, 99, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Mehrotra, M.; Wang, G.; Johnson, W.M. Multiplex PCR for detection of genes for Staphylococcus aureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. J. Clin. Microbiol. 2000, 38, 1032–1035. [Google Scholar] [CrossRef]
- Bianchi, D.M.; Gallina, S.; Bellio, A.; Chiesa, F.; Civera, T.; Decastelli, L. Enterotoxin gene profiles of Staphylococcus aureus isolated from milk and dairy products in Italy. Lett. Appl. Microbiol. 2014, 58, 190–196. [Google Scholar] [CrossRef]
- Enright, M.C.; Day, N.P.; Davies, C.E.; Peacock, S.J.; Spratt, B.G. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 2000, 38, 1008–1015. [Google Scholar] [CrossRef]
- Staphylococcus aureus MLST. Available online: https://pubmlst.org/saureus/ (accessed on 5 December 2019).
- Shopsin, B.; Gomez, M.; Montgomery, S.O.; Smith, D.H.; Waddington, M.; Dodge, D.E.; Bost, D.A.; Riehman, M.; Naidich, S.; Kreiswirth, B.N. Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J. Clin. Microbiol. 1999, 37, 3556–3563. [Google Scholar] [CrossRef]
- Stepanović, S.; Vuković, D.; Dakić, I.; Savić, B.; Švabić-Vlahović, M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- Stepanovic, S.; Vukovic, D.; Bonaventura, G.D.I.; Djukic, S.; Ruzicka, F. Quantification of biofilm in microtiter plates: Overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 2007, 115, 891–899. [Google Scholar] [CrossRef] [PubMed]
- EFSA; ECDC. The European Union Summary Report on Antimicrobial Resistance in zoonotic and indicator bacteria from humans, animals and food in 2017/2018. EFSA J. 2020, 18, 6007. [Google Scholar] [CrossRef]
- ERSAF. Il Mercato dei Suini. Produzione e Consumo 2018 (The Swine Market. Production and Consume 2018); ERSAF: Milano, Italy, 2018. [Google Scholar]
- Pirolo, M.; Gioffrè, A.; Visaggio, D.; Gherardi, M.; Pavia, G.; Samele, P.; Ciambrone, L.; Di Natale, R.; Spatari, G.; Casalinuovo, F.; et al. Prevalence, molecular epidemiology, and antimicrobial resistance of methicillin-resistant Staphylococcus aureus from swine in southern Italy. BMC Microbiol. 2019, 19, 51. [Google Scholar] [CrossRef] [PubMed]
- Kalupahana, R.S.; Duim, B.; Verstappen, K.M.; Gamage, C.D.; Dissanayake, N.; Ranatunga, L.; Graveland, H.; Wagenaar, J.A. MRSA in Pigs and the Environment as a Risk for Employees in Pig-Dense Areas of Sri Lanka. Front. Sustain. Food Syst. 2019, 3, 25. [Google Scholar] [CrossRef]
- EFSA. Analysis of the baseline survey on the prevalence of methicillin-resistant Staphylococcus aureus (MRSA) in holdings with breeding pigs, in the EU, 2008—Part B: Factors associated with MRSA contamination of holdings. EFSA J. 2010, 8, 1597. [Google Scholar] [CrossRef]
- Alba, P.; Feltrin, F.; Cordaro, G.; Porrero, M.C.; Kraushaar, B.; Argudín, M.A.; Nykäsenoja, S.; Monaco, M.; Stegger, M.; Aarestrup, F.M.; et al. Livestock-associated methicillin resistant and methicillin susceptible Staphylococcus aureus sequence type (CC)1 in European farmed animals: High genetic relatedness of isolates from Italian cattle herds and humans. PLoS ONE 2015, 10, e0137143. [Google Scholar] [CrossRef]
- Lopes, E.; Conceição, T.; Poirel, L.; de Lencastre, H.; Aires-De-Sousa, M. Epidemiology and antimicrobial resistance of methicillin-resistant Staphylococcus aureus isolates colonizing pigs with different exposure to antibiotics. PLoS ONE 2019, 14, e0885497. [Google Scholar] [CrossRef]
- Kittl, S.; Brodard, I.; Heim, D.; Andina-Pfister, P.; Oversesch, G. Methicillin-resistant Staphylococcus aureus strains in Swiss pigs and their relation to isolates from farmers and veterinarians. Appl. Environ. Microbiol. 2020, 86, e01865-19. [Google Scholar] [CrossRef]
- Moreno-Flores, A.; Potel-Alvarellos, C.; Francisco-Tomé, M.; Constenla-Caramés, L.; Pérez-Roth, E.; López-Cotón, C.; Comesaña-Da Vila, E.; Eiroa-de la Puente, L.; Álvarez-Fernández, M. Methicillin-resistant Staphylococcus aureus in swine housed indoors in Galicia, Spain. Enferm. Infecc. Microbiol. Clin. 2020, 38, 16–20. [Google Scholar] [CrossRef]
- Witte, W. Selective pressure by antibiotic use in livestock. Int. J. Antimicrob. Agents 2000, 16, 19–24. [Google Scholar] [CrossRef]
- Chopra, I.; Roberts, M. Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Microbiol. Mol. Biol. Rev. 2001, 65, 232–260. [Google Scholar] [CrossRef] [PubMed]
- Crombé, F.; Angeles Argudfn, M.; Vanderhaeghen, W.; Hermans, K.; Haesebrouck, F.; Butaye, P. Transmission dynamics of methicillin-resistant Staphylococcus aureus in pigs. Front. Microbiol. 2013, 4, 57. [Google Scholar] [CrossRef] [PubMed]
- EFSA. Technical specifications on the harmonised monitoring and reporting of antimicrobial resistance in methicillin-resistant Staphylococcus aureus in food-producing animals and food. EFSA J. 2012, 10, 2897. [Google Scholar] [CrossRef]
- Pirolo, M.; Visaggio, D.; Gioffrè, A.; Artuso, I.; Gherardi, M.; Pavia, G.; Samele, P.; Ciambrone, L.; Di Natale, R.; Spatari, G.; et al. Unidirectional animal-to-human transmission of methicillin-resistant Staphylococcus aureus ST398 in pig farming; Evidence from a surveillance study in southern Italy. Antimicrob. Resist. Infect. Control 2019, 8, 187. [Google Scholar] [CrossRef] [PubMed]
- Cuny, C.; Nathaus, R.; Layer, F.; Strommenger, B.; Altmann, D.; Witte, W. Nasal colonization of humans with methicillin-resistant Staphylococcus aureus (MRSA) CC398 with and without exposure to pigs. PLoS ONE 2009, 4, e6800. [Google Scholar] [CrossRef] [PubMed]
- Moodley, A.; Espinosa-Gongora, C.; Nielsen, S.S.; McCarthy, A.J.; Lindsay, J.A.; Guardabassi, L. Comparative Host Specificity of Human- and Pig- Associated Staphylococcus aureus Clonal Lineages. PLoS ONE 2012, 7, e49344. [Google Scholar] [CrossRef]
- Correia, S.; Silva, V.; García-Díez, J.; Teixeira, P.; Pimenta, K.; Pereira, J.E.; Oliveira, S.; Rocha, J.; Manaia, C.M.; Igrejas, G.; et al. One Health Approach Reveals the Absence of Methicillin-Resistant Staphylococcus aureus in Autochthonous Cattle and Their Environments. Front. Microbiol. 2019, 10, 2735. [Google Scholar] [CrossRef]
- Sahibzada, S.; Abraham, S.; Coombs, G.W.; Pang, S.; Hernández-Jover, M.; Jordan, D.; Heller, J. Transmission of highly virulent community-associated MRSA ST93 and livestock-associated MRSA ST398 between humans and pigs in Australia. Sci. Rep. 2017, 7, 5273. [Google Scholar] [CrossRef]
- Pomba, C.; Hasman, H.; Cavaco, L.M.; da Fonseca, J.D.; Aarestrup, F.M. First description of meticillin-resistant Staphylococcus aureus (MRSA) CC30 and CC398 from swine in Portugal. Int. J. Antimicrob. Agents 2009, 34, 193–194. [Google Scholar] [CrossRef]
- Rachman, A.R.A.; Suhaili, Z.; Desa, M.N.M. The Evolution and Dissemination of Methicillin Resistance Determinant in Staphylococcus aureus. Available online: https://www.intechopen.com/books/the-rise-of-virulence-and-antibiotic-resistance-in-staphylococcus-aureus/the-evolution-and-dissemination-of-methicillin-resistance-determinant-in-staphylococcus-aureus (accessed on 25 July 2020).
- European Food Safety Authority. Assessment of the Public Health significance of meticillin resistant Staphylococcus aureus (MRSA) in animals and foods. EFSA J. 2009, 993, 1–73. [Google Scholar]
- Witte, W.; Strommenger, B.; Stanek, C.; Cuny, C. Methicillin-resistant Staphylococcus aureus ST398 in humans and animals, central Europe. Emerg. Infect. Dis. 2007, 13, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Larsen, J.; Petersen, A.; Sørum, M.; Stegger, M.; Van Alphen, L.; Valentiner-Branth, P.; Knudsen, L.K.; Larsen, L.S.; Feingold, B.; Price, L.B.; et al. Meticillin-resistant Staphylococcus aureus CC398 is an increasing cause of disease in people with no livestock contact in Denmark, 1999 to 2011. Eurosurveillance 2015, 20, 30021. [Google Scholar] [CrossRef] [PubMed]
- Feltrin, F.; Alba, P.; Kraushaar, B.; Ianzano, A.; Argudín, M.A.; Di Matteo, P.; Porrero, M.C.; Aarestrup, F.M.; Butaye, P.; Franco, A.; et al. A livestock-associated, multidrug-resistant, methicillin-resistant Staphylococcus aureus clonal complex 97 lineage spreading in dairy cattle and pigs in Italy. Appl. Environ. Microbiol. 2016, 82, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Nishiyama, A.; Takano, T.; Yabe, S.; Higuchi, W.; Razvina, O.; Shi, D. Community-acquired methicillin-resistant Staphylococcus aureus: Community transmission, pathogenesis, and drug resistance. J. Infect. Chemother. 2010, 16, 225–254. [Google Scholar] [CrossRef] [PubMed]
- Cuny, C.; Strommenger, B.; Witte, W.; Stanek, C. Clusters of infections in horses with MRSA ST1, ST254 and ST398 in a Veterinary Hospital. Microb. Drug Resist. 2008, 14, 307–310. [Google Scholar] [CrossRef]
- Witte, W. Community-acquired methicillin-resistant Staphylococcus aureus: What do we need to know? Clin. Microbiol. Infect. 2009, 15, 17–25. [Google Scholar] [CrossRef]
- Franco, A.; Hasman, H.; Iurescia, M.; Lorenzetti, R.; Stegger, M.; Pantosti, A.; Feltrin, F.; Ianzano, A.; Porrero, M.C.; Liapi, M.; et al. Molecular characterization of spa type t127, sequence type 1 methicillin-resistant Staphylococcus aureus from pigs. J. Antimicrob. Chemother. 2011, 66, 1231–1235. [Google Scholar] [CrossRef]
- Pinchuk, I.V.; Beswick, E.J.; Reyes, V.E. Staphylococcal Enterotoxins. Toxins 2010, 2, 2177–2197. [Google Scholar] [CrossRef]
- Hu, D.-L.; Wang, L.; Fang, R.; Okamura, M.; Ono, H.K. Staphylococcus aureus Enterotoxins. In Staphylococcus aureus; Fetsch, A., Ed.; Elsevier Academic Press: London, UK, 2018; pp. 39–55. ISBN 9780128096710. [Google Scholar]
- Schelin, J.; Wallin-Carlquist, N.; Cohn, M.T.; Lindqvist, R.; Barker, G.C.; Rådström, P. The formation of Staphylococcus aureus enterotoxin in food environments and advances in risk assessment. Virulence 2011, 2, 580–592. [Google Scholar] [CrossRef]
- European Commission. Commission Regulation (EC) N° 2073/2005 of 15th November 2005 on microbiological criteria for foodstuffs. Off. J. Eur. Union 2005, L338, 1–26. [Google Scholar]
- Vázquez-Sánchez, D.; Rodríguez-López, P. Biofilm Formation of Staphylococcus aureus. In Staphylococcus aureus; Fetsch, A., Ed.; Elsevier Academic Press: London, UK, 2018; pp. 87–103. ISBN 9780128096710. [Google Scholar]
- Balcázar, J.L.; Subirats, J.; Borrego, C.M. The role of biofilms as environmental reservoirs of antibiotic resistance. Front. Microbiol. 2015, 6, 1216. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, D.; Shi, L.; Cai, R.; Li, C.; Yan, H. Association between agr type, virulence factors, biofilm formation and antibiotic resistance of Staphylococcus aureus isolates from pork production. Front. Microbiol. 2018, 9, 1876. [Google Scholar] [CrossRef] [PubMed]
- Di Ciccio, P.; Vergara, A.; Festino, A.R.; Paludi, D.; Zanardi, E.; Ghidini, S.; Ianieri, A. Biofilm formation by Staphylococcus aureus on food contact surfaces: Relationship with temperature and cell surface hydrophobicity. Food Control 2015, 50, 930–936. [Google Scholar] [CrossRef]
- Ou, C.; Shang, D.; Yang, J.; Chen, B.; Chang, J.; Jin, F.; Shi, C. Prevalence of multidrug-resistant Staphylococcus aureus isolates with strong biofilm formation ability among animal-based food in Shanghai. Food Control 2020, 112, 107106. [Google Scholar] [CrossRef]
- Basanisi, M.G.; la Bella, G.; Xobili, G.; Tola, S.; Cafiero, M.A.; la Salandra, G. Prevalence and characterization of methicillin-resistant Staphylococcus aureus (MRSA) isolates from retail meat in South Italy. Ital. J. Food Sci. 2020, 32, 410–419. [Google Scholar] [CrossRef]
- Mama, O.M.; Morales, L.; Ruiz-Ripa, L.; Zarazaga, M.; Torres, C. High prevalence of multidrug resistant S. aureus-CC398 and frequent detection of enterotoxin genes among non-CC398 S. aureus from pig-derived food in Spain. Int. J. Food Microbiol. 2020, 320, 108510. [Google Scholar] [CrossRef]
| Antibiotic Resistance Pattern | Genotypes (No. Isolates) |
|---|---|
| P-AMC-OX-AMP-TET-T | ST398/t011 (5); ST398/t899 (1); ST398/t4474 (1) |
| P-AMC-OX-AMP-FUR-TET-T | ST398/t011 (5) |
| P-AMC-OX-AMP-FUR-TET-ENR | ST398/t899 (8); ST97/t4795 (1) |
| P-AMC-OX-AMP-FUR-TET-DXT-T | ST398/t011 (6) |
| P-AMC-OX-AMP-FUR-KF-ENR-CN | ST1/t127 (3) |
| P-AMC-OX-AMP-FUR-TET-DXT-ENR | ST5422/t1730 (1); ST398/t1939 (1) |
| P-AMC-OX-AMP-KF-ENR-CN | ST1/t127 (2) |
| P-AMC-OX-AMP-FUR-TET-DXT-C-T | ST398/t011 (2) |
| P-AMC-OX-AMP-FUR-TET-C-FFC-CN-SxT | ST398/t899 (2) |
| P-AMC-OX-AMP-FUR-TET-DXT-C-FFC-SxT | ST398/t899 (2) |
| P-AMC-OX-AMP-FUR-TET-DXT-C-FFC-T-SxT | ST398/t899 (2) |
| Isolate ID | Source | ST | spa-Type | Antibiotic Resistance Genes | Enterotoxin Genes | Biofilm Production |
|---|---|---|---|---|---|---|
| SA1 | Animal | 398 | t011 | blaZ, tetL, tetM, norA, aadD, dfrA, dfrK | n.d. | Moderate |
| SA2 | Environment | 398 | t899 | blaZ, tetL, tetM, norA, fexA, aacAaphD, aadD, vgaA, dfrA | seb, sed, see | Strong |
| SA3 | Environment | 398 | t899 | blaZ, tetL, tetM, norA, fexA, aadD, vgaA, dfrA | sea, seb, see | Weak |
| SA4 | Environment | 398 | t899 | tetL, aadD, vgaA | seb, see | Weak |
| SA5 | Animal | 97 | t4795 | blaZ, tetL, tetM, norA, aadD, vgaA, dfrA | seb, see | Moderate |
| SA6 | Animal | 97 | t4795 | n.d. | n.d. | Moderate |
| SA7 | Animal | 5422 | t1730 | blaZ, tetM, norA, dfrA, | sea, seb, sed | Strong |
| SA8 | Animal | 398 | t899 | tetM, vgaA, dfrA, dfrK | n.d. | Strong |
| SA9 | Animal | 398 | t011 | blaZ, tetM, vgaA, dfrA, dfrD | seb | Moderate |
| SA10 | Animal | 97 | t1730 | blaZ, tetL, tetM, norA, cfr, dfrA, dfrD | sea | Strong |
| SA11 | Animal | 398 | t899 | blaZ, tetL, tetM, fexA, aadD, vgaA, dfrA | sea, seb | Weak |
| SA12 | Animal | 398 | t011 | blaZ, tetL, tetM | sea, seb | Moderate |
| SA13 | Environment | 398 | t1939 | blaZ, tetM, norA, dfrA | sec | Weak |
| SA14 | Environment | 398 | t011 | tetL | sea, seb | Non-producer |
| SA15 | Environment | 30 | t318 | blaZ, tetL, tetM, fexA, cfr, aadD, dfrA | sea, seb | Strong |
| SA16 | Animal | 398 | t899 | blaZ, dfrA | see | Moderate |
| SA17 | Environment | 398 | t1939 | blaZ, tetM, norA, fexA | n.d. | Non-producing |
| SA18 | Environment | 398 | t034 | blaZ, tetL, tetM, norA, fexA, cfr, aadD, dfrA | seb, see | Strong |
| SA19 | Environment | 398 | t18494 | blaZ, tetM, norA, fexA, dfrA | sea, seb, see | Weak |
| SA20 | Environment | 398 | t1200 | tetL, fexA, cfr, aadD, dfrK | n.d | Moderate |
| SA21 | Animal | 398 | t1200 | blaZ, tetM, norA, vgaE, dfrA, dfrK | seb, sec | Weak |
| SA22 | Animal | 398 | t18494 | blaZ, tetL, tetM, vgaE, dfrA, dfrK | n.d. | Weak |
| SA23 | Animal | 398 | t011 | tetL, tetM, aadD, vgaE, dfrA, dfrK | sea, seb | Weak |
| SA24 | Animal | 4894 | t18494 | cfr | sea, sec, see | Weak |
| SA25 | Animal | 398 | t18494 | blaZ, tetL, tetM, cfr, dfrD | sea, seb, sec | Weak |
| SA26 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | see | Weak |
| SA27 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | sec, see | Weak |
| SA28 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | sec | Weak |
| SA29 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | n.d. | Moderate |
| SA30 | Animal | 398 | t899 | blaZ, tetM, norA, aadD, vgaE, dfrA, dfrK | n.d. | Weak |
| SA31 | Animal | 398 | t899 | tetM, aadD, dfrA | see | Non-producer |
| SA32 | Animal | 97 | t1730 | blaZ, tetL, tetM, norA, aacAaphD, aadD, vgaA, vgaE, dfrA, dfrK, ermT | n.d. | Weak |
| SA33 | Animal | 398 | t011 | blaZ, tetM, norA, vgaE, dfrA, dfrK | n.d. | Weak |
| SA34 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, dfrA, dfrK | sec, see | Weak |
| SA35 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE, dfrA | sec, see | Weak |
| SA36 | Animal | 398 | t899 | blaZ, tetM, norA, aadD, vgaE, dfrA, dfrD, dfrK | sea, seb, see | Non-producer |
| SA37 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaE, dfrA, dfrD, dfrK | sea, seb | Weak |
| SA38 | Animal | 398 | t011 | blaZ, tetL, tetM, norA, aadD, dfrA | n.d. | Non-producer |
| SA39 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaA, vgaE, dfrA, dfrK | sea, seb, sec, sed | Weak |
| SA40 | Animal | 398 | t011 | tetL, tetM, norA, aadD, vgaE, dfrK | seb | Weak |
| SA41 | Animal | 1 | t127 | blaZ, norA, aacAaphD, aadD, vgaE, dfrA, dfrK, ermT | sec | Weak |
| SA42 | Animal | 1 | t127 | blaZ, norA, aacAaphD, aadD, vgaE, dfrA, ermT | seb, sed | Moderate |
| SA43 | Animal | 1 | t127 | blaZ, norA, aacAaphD, aadD, vgaE, dfrA, ermT | seb, sec | Weak |
| SA44 | Animal | 1 | t127 | norA, aacAaphD, vgaA, vgaE, dfrA, dfrK, ermT | sec | Weak |
| SA45 | Animal | 1 | t127 | norA, aacAaphD, vgaA, vgaE, dfrA | seb, sec, see | Non-producer |
| SA46 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | seb | Non-producer |
| SA47 | Animal | 398 | t1939 | blaZ, tetM, fexA, cfr, aadD, vgaA, vgaE, dfrA, dfrK | seb, sec, see | Non-producer |
| SA48 | Animal | 398 | t899 | blaZ, fexA, aadD, vgaA, vgaE, dfrA, dfrK | n.d. | Non-producer |
| SA49 | Animal | 398 | t899 | blaZ, tetL, tetM, norA, aacAaphD, aadD, vgaA, vgaE, dfrA, dfrK | sec, see | Non-producer |
| SA50 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaE, dfrA | seb, see | Non-producer |
| SA51 | Animal | 398 | t899 | blaZ, norA, aadD, vgaE, dfrA, dfrD, dfrK | sea, seb | Moderate |
| SA52 | Animal | 398 | t899 | tetM, norA, aadD, vgaE, dfrA, dfrK | sec, see | Non-producer |
| SA53 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE, dfrA, dfrK | seb, sed, see | Non-producer |
| SA54 | Animal | 398 | t011 | blaZ, tetM, norA, vgaA, vgaE, dfrA, dfrK | see | Non-producer |
| SA55 | Animal | 398 | t011 | blaZ, tetM, norA, vgaA, vgaE | see | Non-producer |
| SA56 | Animal | 398 | t011 | blaZ, tetM, norA, vgaA, vgaE, dfrK | n.d. | Non-producer |
| SA57 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaE, ermT | seb | Non-producer |
| SA58 | Animal | 398 | t4474 | blaZ, tetM, norA, vgaA, vgaE, dfrA | sec, see | Non-producer |
| SA59 | Animal | 398 | t4474 | blaZ, tetM, norA, vgaE, dfrA | n.d. | Weak |
| SA60 | Animal | 398 | t011 | blaZ, tetM, norA, vgaA, vgaE, dfrA | see | Non-producer |
| SA61 | Animal | 398 | t011 | blaZ, tetM, norA, vgaA, dfrA, dfrK | seb, sec, see | Weak |
| SA62 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE, dfrA, dfrK | n.d. | Weak |
| SA63 | Animal | 398 | t899 | blaZ, tetL, tetM, aacAaphD, aadD, vgaA, vgaE, dfrA, dfrK, ermT | sec | Moderate |
| SA64 | Animal | 398 | t899 | blaZ, tetM, norA, dfrA, dfrK | seb, see | Moderate |
| SA65 | Animal | 398 | t899 | blaZ, tetL, tetM, aacAaphD, aadD, vgaA, vgaE, dfrA, dfrK, ermT | n.d. | Weak |
| SA66 | Animal | 398 | t4558 | blaZ, tetM, norA, fexA, aacAaphD, vgaA, vgaE, dfrA, dfrK | sea, sec, sed, see | Weak |
| SA67 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfrA, dfrK | sec | Weak |
| SA68 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE, dfrA, dfrK | n.d. | Weak |
| SA69 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE, dfrA, dfrK | sec | Weak |
| SA70 | Animal | 1 | t127 | blaZ, norA, aacAaphD, vgaA, vgaE, dfrA, dfrK, ermT | n.d. | Weak |
| SA71 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaA, vgaE, dfrA, dfrD | sea | Weak |
| SA72 | Animal | 398 | t011 | tetL, tetM, norA, aadD, vgaA, vgaE, dfrA, dfrK | sea | Non-producer |
| SA73 | Animal | 398 | t011 | tetM, norA, aadD, vgaA, vgaE, dfrD, dfrK | sea | Weak |
| SA74 | Animal | 398 | t011 | tetM, norA, aadD, vgaA, vgaE, dfrA, dfrD, dfrK | seb, see | Non-producer |
| SA75 | Animal | 398 | t011 | tetL, tetM, norA, aadD, vgaA, vgaE, dfrA, dfrK | sea | Weak |
| SA76 | Animal | 398 | t011 | tetM, norA, aadD, vgaE, dfrA, dfrD, dfrK | n.d. | Non-producer |
| SA77 | Animal | 1 | t127 | blaZ, tetL, norA, aacAaphD, aadD, vgaA, vgaE, dfrA, ermT | sea | Weak |
| SA78 | Animal | 1 | t127 | blaZ, tetL, norA, aacAaphD, aadD, vgaA, dfrA, dfrK, ermT | sea | Weak |
| SA79 | Animal | 398 | t899 | tetM, norA, vgaE, dfrA | sea | Non-producer |
| SA80 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfr | sea | Non-producer |
| SA81 | Animal | 398 | t899 | blaZ, tetM, norA, vgaE, dfr | sea, seb | Non-producer |
| SA82 | Animal | 398 | t899 | tetM, norA, vgaE, dfrA | sea, see | Non-producer |
| SA83 | Animal | 398 | t011 | blaZ, tetM, norA, aadD, vgaE, dfrA, dfrD | see | Non-producer |
| SA84 | Animal | 398 | t899 | blaZ, tetM, norA, vgaA, vgaE | see | Non-producer |
| SA85 | Animal | 398 | t899 | blaZ, tetM, aadD, vgaA, vgaE, dfrD | seb | Weak |
| SA86 | Animal | 398 | t011 | blaZ, tetM, norA, dfrA | sea | Non-producer |
| SA87 | Animal | 398 | t011 | blaZ, tetM, norA, vgaE, dfrA | n.d. | Non-producer |
| Genotype | Number of Isolates (%) |
|---|---|
| ST398/t899 | 33 (37.93%) |
| ST398/t011 | 27 (31.03%) |
| ST398/t1939 | 3 (3.45%) |
| ST398/t034 | 1 (1.15%) |
| ST398/t18494 | 3 (3.45%) |
| ST398/t1200 | 2 (2.30%) |
| ST398/t4474 | 2 (2.30%) |
| ST398/t4558 | 1 (1.15%) |
| ST1/t127 | 8 (9.20%) |
| ST97/t1730 | 2 (2.30%) |
| ST97/t4795 | 2 (2.30%) |
| ST4894/t18494 | 1 (1.15%) |
| ST30/t318 | 1 (1.15%) |
| ST5422/t1730 | 1 (1.15%) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodríguez-López, P.; Filipello, V.; Di Ciccio, P.A.; Pitozzi, A.; Ghidini, S.; Scali, F.; Ianieri, A.; Zanardi, E.; Losio, M.N.; Simon, A.C.; et al. Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain. Foods 2020, 9, 1141. https://doi.org/10.3390/foods9091141
Rodríguez-López P, Filipello V, Di Ciccio PA, Pitozzi A, Ghidini S, Scali F, Ianieri A, Zanardi E, Losio MN, Simon AC, et al. Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain. Foods. 2020; 9(9):1141. https://doi.org/10.3390/foods9091141
Chicago/Turabian StyleRodríguez-López, Pedro, Virginia Filipello, Pierluigi Aldo Di Ciccio, Alessandra Pitozzi, Sergio Ghidini, Federico Scali, Adriana Ianieri, Emanuela Zanardi, Marina Nadia Losio, Ancuta Cezara Simon, and et al. 2020. "Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain" Foods 9, no. 9: 1141. https://doi.org/10.3390/foods9091141
APA StyleRodríguez-López, P., Filipello, V., Di Ciccio, P. A., Pitozzi, A., Ghidini, S., Scali, F., Ianieri, A., Zanardi, E., Losio, M. N., Simon, A. C., & Alborali, G. L. (2020). Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain. Foods, 9(9), 1141. https://doi.org/10.3390/foods9091141










