Estimation of Minced Pork Microbiological Spoilage through Fourier Transform Infrared and Visible Spectroscopy and Multispectral Vision Technology
Abstract
1. Introduction
2. Materials and Methods
2.1. Minced Pork Samples Preparation and Storage Conditions
2.2. Microbiological Analyses and pH Measurement
2.3. Sensors
2.3.1. FTIR Spectroscopy
2.3.2. VIS Spectroscopy
2.3.3. Image Acquisition and Pre-Processing
2.4. Data Analysis
2.4.1. Microbiological Data Analysis
2.4.2. Spectral Data Analysis and Correlation with Microbiological Data
3. Results and Discussion
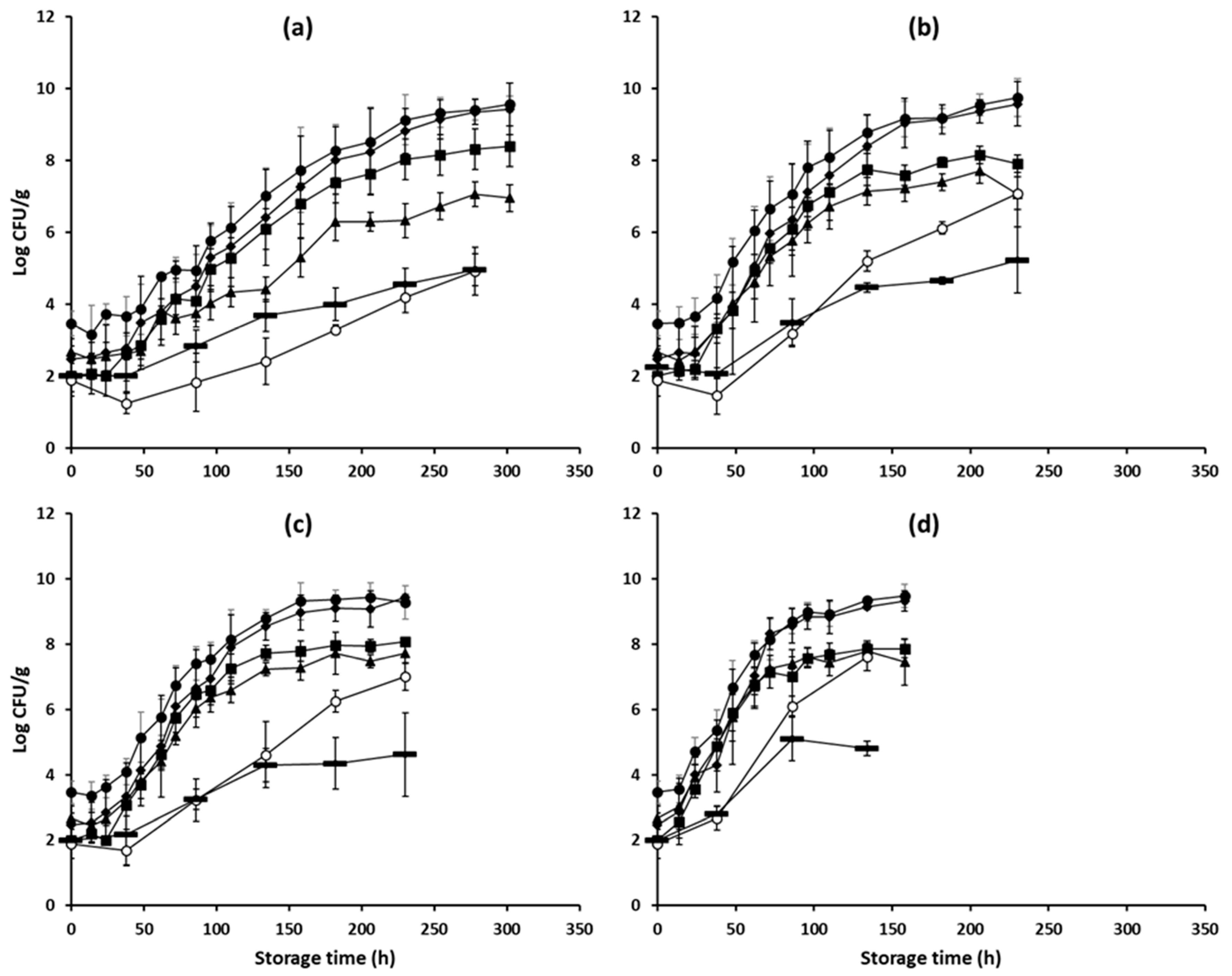
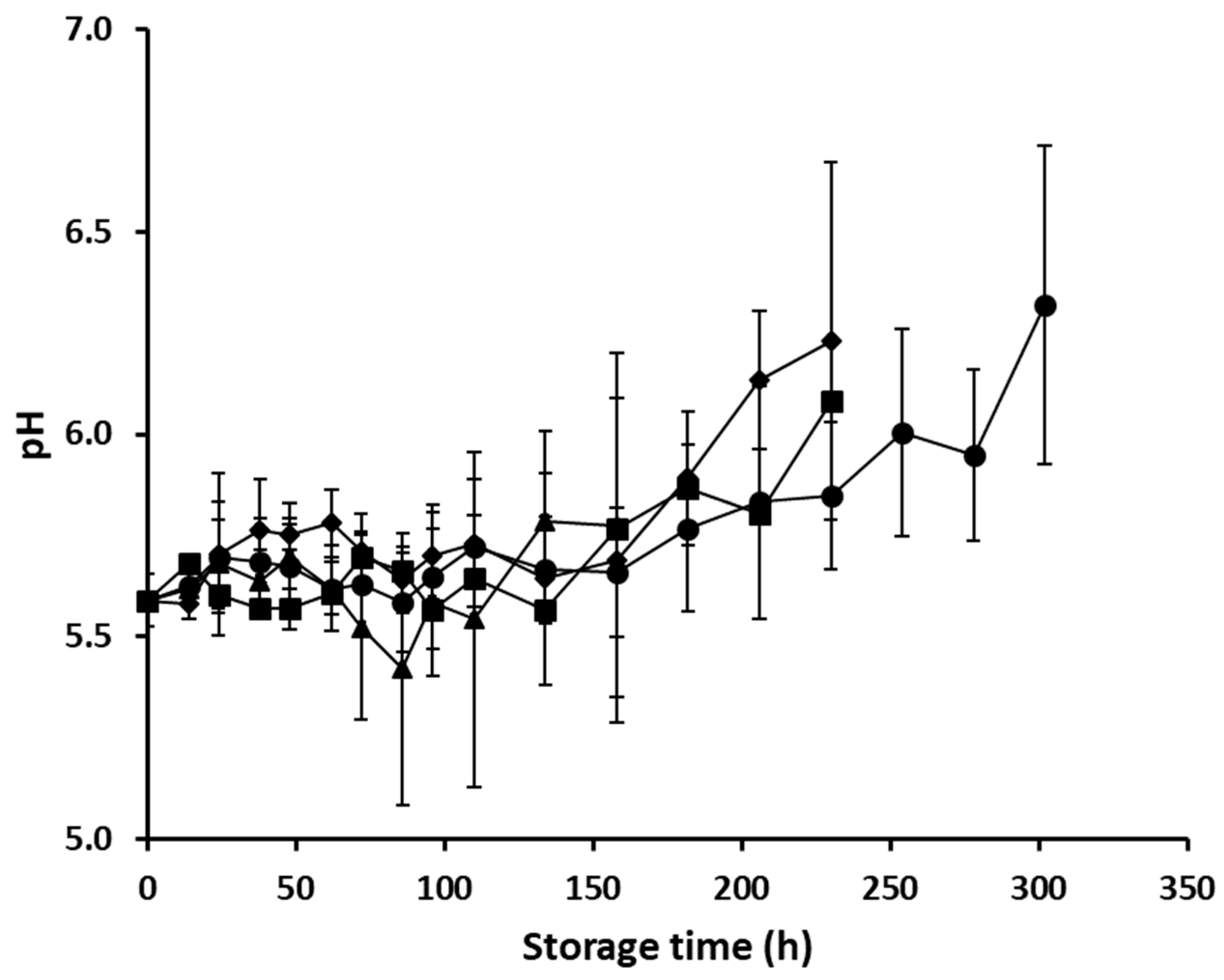
3.1. Microbiological Spoilage of Minced Pork and pH Data
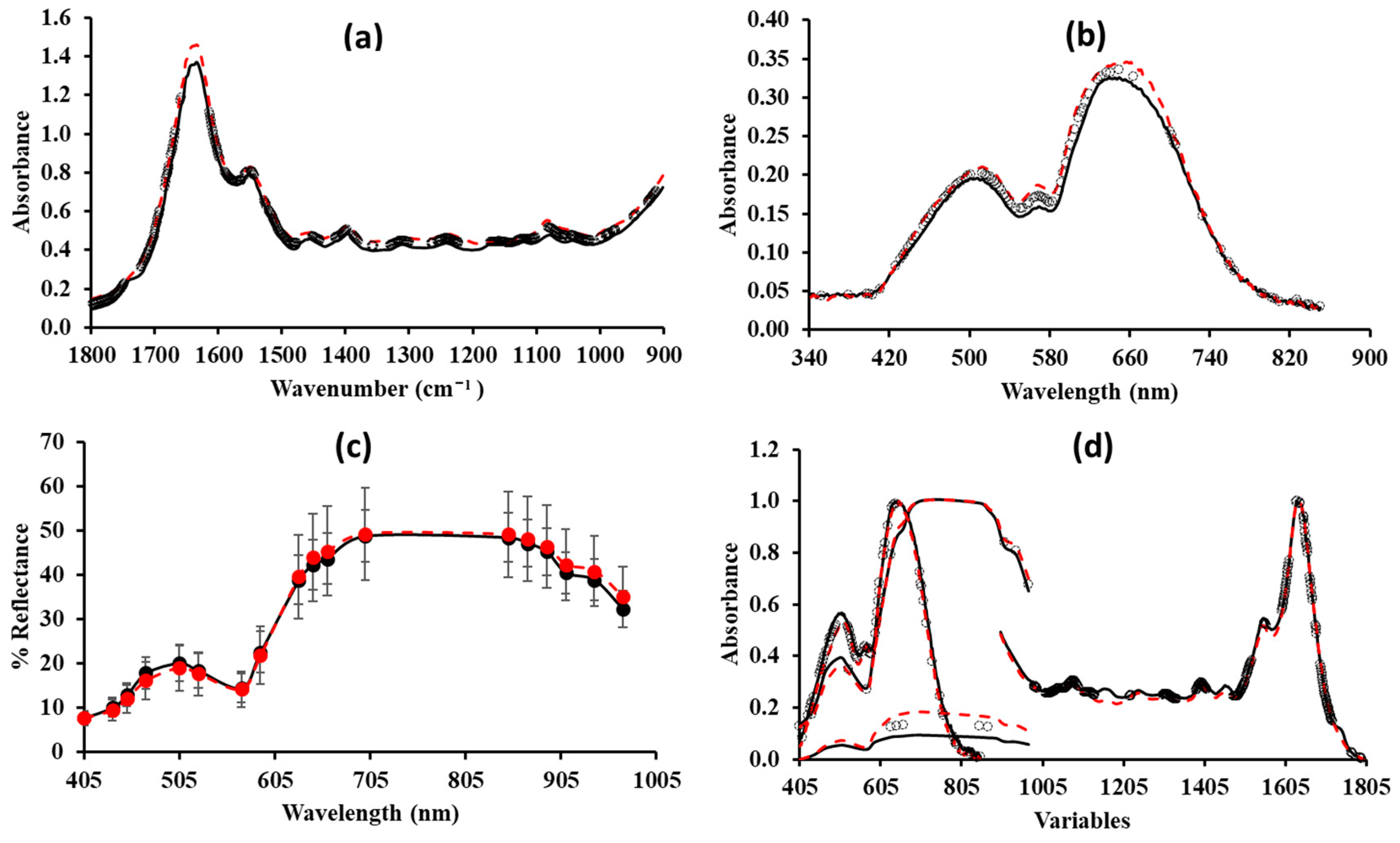
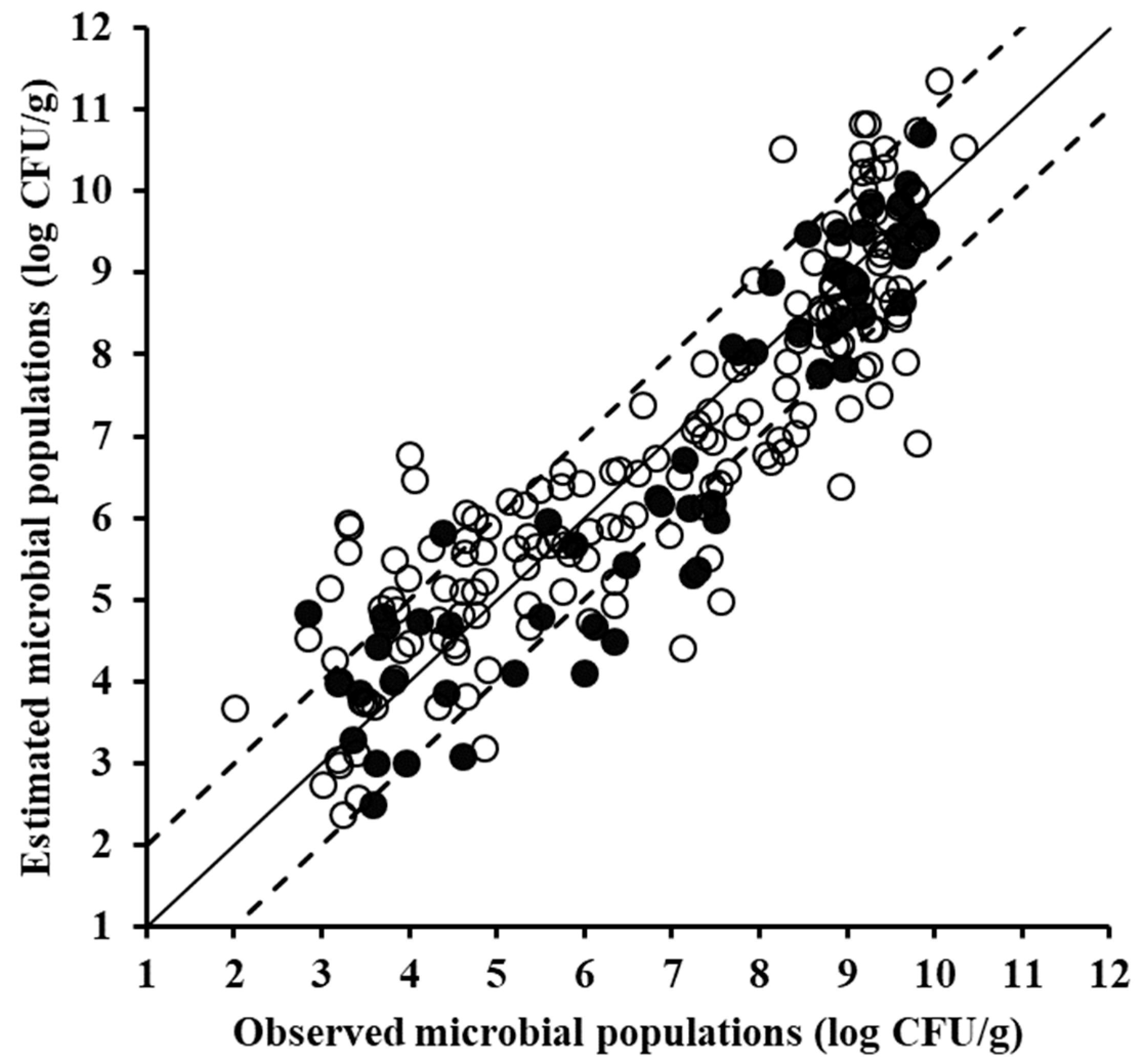
3.2. Estimation of Minced Pork Spoilage Using Spectral Data
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Nychas, G.-J.E.; Skandamis, P.N.; Tassou, C.C.; Koutsoumanis, K.P. Meat Spoilage during Distribution. Meat Sci. 2008, 78, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Nychas, G.-J.E.; Panagou, E.Z.; Mohareb, F. Novel Approaches for Food Safety Management and Communication. Curr. Opin. Food Sci. 2016, 12, 13–20. [Google Scholar] [CrossRef]
- Argyri, A.; Panagou, E.; Nychas, G. Monitoring Microbial Spoilage of Foods by Vibrational Spectroscopy (FT-IR & Raman). In Novel Food Preservation and Microbial Assessment Techniques; Boziaris, I., Ed.; CRC Press: Boca Raton, FL, USA, 2014; pp. 386–434. [Google Scholar] [CrossRef]
- Ropodi, A.I.; Panagou, E.Z.; Nychas, G.-J.E. Data Mining Derived from Food Analyses Using Non-Invasive/Non-Destructive Analytical Techniques; Determination of Food Authenticity, Quality & Safety in Tandem with Computer Science Disciplines. Trends Food Sci. Technol. 2016, 50, 11–25. [Google Scholar] [CrossRef]
- He, H.-J.; Sun, D.-W. Microbial Evaluation of Raw and Processed Food Products by Visible/Infrared, Raman and Fluorescence Spectroscopy. Trends Food Sci. Technol. 2015, 46, 199–210. [Google Scholar] [CrossRef]
- Li-Chan, E.C.Y.; Ismail, A.A.; Sedman, J.; van de Voort, F.R. Vibrational Spectroscopy of Food and Food Products. In Handbook of Vibrational Spectroscopy; Chalmers, J.M., Griffiths, P.R., Eds.; John Wiley & Sons, Ltd.: Chichester, UK, 2006. [Google Scholar] [CrossRef]
- Mohareb, F.; Papadopoulou, O.; Panagou, E.; Nychas, G.-J.; Bessant, C. Ensemble-Based Support Vector Machine Classifiers as an Efficient Tool for Quality Assessment of Beef Fillets from Electronic Nose Data. Anal. Methods 2016, 8, 3711–3721. [Google Scholar] [CrossRef]
- Zając, A.; Dymińska, L.; Lorenc, J.; Hanuza, J. Fourier Transform Infrared and Raman Spectroscopy Studies of the Time-Dependent Changes in Chicken Meat as a Tool for Recording Spoilage Processes. Food Anal. Methods 2017, 10, 640–648. [Google Scholar] [CrossRef]
- Lianou, A.; Moschonas, G.; Nychas, G.-J.E.; Panagou, E.Z. Growth of Listeria Monocytogenes in Pasteurized Vanilla Cream Pudding as Affected by Storage Temperature and the Presence of Cinnamon Extract. Food Res. Int. 2018, 106, 1114–1122. [Google Scholar] [CrossRef]
- Pavli, F.; Tassou, C.; Nychas, G.-J.; Chorianopoulos, N. Probiotic Incorporation in Edible Films and Coatings: Bioactive Solution for Functional Foods. Int. J. Mol. Sci. 2018, 19, 150. [Google Scholar] [CrossRef]
- Tsakanikas, P.; Fengou, L.-C.; Manthou, E.; Lianou, A.; Panagou, E.Z.; Nychas, G.-J.E. A Unified Spectra Analysis Workflow for the Assessment of Microbial Contamination of Ready-to-Eat Green Salads: Comparative Study and Application of Non-Invasive Sensors. Comput. Electron. Agric. 2018, 155, 212–219. [Google Scholar] [CrossRef]
- Fengou, L.-C.; Lianou, A.; Tsakanikas, P.; Gkana, E.N.; Panagou, E.Z.; Nychas, G.-J.E. Evaluation of Fourier Transform Infrared Spectroscopy and Multispectral Imaging as Means of Estimating the Microbiological Spoilage of Farmed Sea Bream. Food Microbiol. 2019, 79, 27–34. [Google Scholar] [CrossRef]
- Panagou, E.Z.; Papadopoulou, O.; Carstensen, J.M.; Nychas, G.-J.E. Potential of Multispectral Imaging Technology for Rapid and Non-Destructive Determination of the Microbiological Quality of Beef Filets during Aerobic Storage. Int. J. Food Microbiol. 2014, 174, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Koutsoumanis, K.P.; Stamatiou, A.P.; Drosinos, E.H.; Nychas, G.-J.E. Control of Spoilage Microorganisms in Minced Pork by a Self-Developed Modified Atmosphere Induced by the Respiratory Activity of Meat Microflora. Food Microbiol. 2008, 25, 915–921. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulou, O.; Panagou, E.Z.; Tassou, C.C.; Nychas, G.-J.E. Contribution of Fourier Transform Infrared (FTIR) Spectroscopy Data on the Quantitative Determination of Minced Pork Meat Spoilage. Food Res. Int. 2011, 44, 3264–3271. [Google Scholar] [CrossRef]
- Carstensen, J.M.; Hansen, J.F. An Apparatus and a Method of Recording an Image of an Object. Patent 17,198, 15 November 2003. [Google Scholar]
- Ropodi, A.I.; Panagou, E.Z.; Nychas, G.-J.E. Multispectral Imaging (MSI): A Promising Method for the Detection of Minced Beef Adulteration with Horsemeat. Food Control 2017, 73, 57–63. [Google Scholar] [CrossRef]
- Baranyi, J.; Roberts, T.A. A Dynamic Approach to Predicting Bacterial Growth in Food. Int. J. Food Microbiol. 1994, 23, 277–294. [Google Scholar] [CrossRef]
- Mehmood, T.; Liland, K.H.; Snipen, L.; Sæbø, S. A Review of Variable Selection Methods in Partial Least Squares Regression. Chemom. Intell. Lab. Syst. 2012, 118, 62–69. [Google Scholar] [CrossRef]
- Cheng, J.-H.; Sun, D.-W. Recent Applications of Spectroscopic and Hyperspectral Imaging Techniques with Chemometric Analysis for Rapid Inspection of Microbial Spoilage in Muscle Foods: Microbial Spoilage in Muscle Food. Compr. Rev. Food Sci. Food Saf. 2015, 14, 478–490. [Google Scholar] [CrossRef]
- Rinnan, Å.; van den Berg, F.; Engelsen, S.B. Review of the Most Common Pre-Processing Techniques for near-Infrared Spectra. Trends Anal. Chem. 2009, 28, 1201–1222. [Google Scholar] [CrossRef]
- Barnes, R.J.; Dhanoa, M.S.; Lister, S.J. Standard Normal Variate Transformation and De-Trending of Near-Infrared Diffuse Reflectance Spectra. Appl. Spectrosc. 1989, 43, 772–777. [Google Scholar] [CrossRef]
- Westad, F.; Martens, H. Variable Selection in near Infrared Spectroscopy Based on Significance Testing in Partial Least Squares Regression. J. Infrared Spectrosc. 2000, 8, 117–124. [Google Scholar] [CrossRef]
- Zweifel, C.; Fischer, R.; Stephan, R. Microbiological Contamination of Pig and Cattle Carcasses in Different Small-Scale Swiss Abattoirs. Meat Sci. 2008, 78, 225–231. [Google Scholar] [CrossRef] [PubMed]
- European Commission. Comission Regulation (EC) (2005); No 2073/2005 on Microbiological Criteria for Foodstuffs; OJ L 338/15. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32005R2073&from=EN (accessed on 1 June 2019).
- Dainty, R.H.; Mackey, B.M. The Relationship between the Phenotypic Properties of Bacteria from Chill-Stored Meat and Spoilage Processes. J. Appl. Bacteriol. 1992, 73, 103s–114s. [Google Scholar] [CrossRef]
- Doulgeraki, A.I.; Ercolini, D.; Villani, F.; Nychas, G.-J.E. Spoilage Microbiota Associated to the Storage of Raw Meat in Different Conditions. Int. J. Food Microbiol. 2012, 157, 130–141. [Google Scholar] [CrossRef] [PubMed]
- Iacumin, L.; Carballo, J. Microbial Ecology of Pork Meat and Pork Products. In Quantitative Microbiology in Food Processing: Modeling the Microbial Ecology; de Souza Sant’Ana, A., Ed.; John Wiley & Sons: Chichester, UK, 2016; pp. 463–482. [Google Scholar]
- Caldera, L.; Franzetti, L.; Van Coillie, E.; De Vos, P.; Stragier, P.; De Block, J.; Heyndrickx, M. Identification, Enzymatic Spoilage Characterization and Proteolytic Activity Quantification of Pseudomonas Spp. Isolated from Different Foods. Food Microbiol. 2016, 54, 142–153. [Google Scholar] [CrossRef]
- Dainty, R.H.; Hibbard, C.M. Aerobic Metabolism of Brochothrix Thermosphacta Growing on Meat Surfaces and in Laboratory Media. J. Appl. Bacteriol. 1980, 48, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Pothakos, V.; Devlieghere, F.; Villani, F.; Björkroth, J.; Ercolini, D. Lactic Acid Bacteria and Their Controversial Role in Fresh Meat Spoilage. Meat Sci. 2015, 109, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Al-Jowder, O.; Defernez, M.; Kemsley, E.K.; Wilson, R.H. Mid-Infrared Spectroscopy and Chemometrics for the Authentication of Meat Products. J. Agric. Food Chem. 1999, 47, 3210–3218. [Google Scholar] [CrossRef] [PubMed]
- Ellis, D.I.; Broadhurst, D.; Kell, D.B.; Rowland, J.J.; Goodacre, R. Rapid and Quantitative Detection of the Microbial Spoilage of Meat by Fourier Transform Infrared Spectroscopy and Machine Learning. Appl. Environ. Microbiol. 2002, 68, 2822–2828. [Google Scholar] [CrossRef] [PubMed]
- Ellis, D.I.; Broadhurst, D.; Goodacre, R. Rapid and Quantitative Detection of the Microbial Spoilage of Beef by Fourier Transform Infrared Spectroscopy and Machine Learning. Anal. Chim. Acta 2004, 514, 193–201. [Google Scholar] [CrossRef]
- Böcker, U.; Ofstad, R.; Wu, Z.; Bertram, H.C.; Sockalingum, G.D.; Manfait, M.; Egelandsdal, B.; Kohler, A. Revealing Covariance Structures in Fourier Transform Infrared and Raman Microspectroscopy Spectra: A Study on Pork Muscle Fiber Tissue Subjected to Different Processing Parameters. Appl. Spectrosc. 2007, 61, 1032–1039. [Google Scholar] [CrossRef]
- Socrates, G. Infrared and Raman Characteristic Group Frequencies: Tables and Charts, 3rd ed.; Chichester Wiley: New York, NY, USA, 2004. [Google Scholar]
- Nychas, G.J.E.; Drosinos, E.H.; Board, R.G. Chemical Changes in Stored Meat; Blackie Academic & Professional: London, UK, 1998. [Google Scholar]
- Sone, I.; Olsen, R.L.; Heia, K. Spectral Changes of Atlantic Salmon (Salmo Salar L.) Muscle during Cold Storage As Affected by the Oxidation State of Heme. J. Agric. Food Chem. 2012, 60, 9719–9726. [Google Scholar] [CrossRef] [PubMed]
- Andrés, S.; Silva, A.; Soares-Pereira, A.L.; Martins, C.; Bruno-Soares, A.M.; Murray, I. The Use of Visible and near Infrared Reflectance Spectroscopy to Predict Beef M. Longissimus Thoracis et Lumborum Quality Attributes. Meat Sci. 2008, 78, 217–224. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, Y.-R.; Huffman, R.W.; Park, B. Changes in the Visible/Near-Infrared Spectra of Chicken Carcasses in Storage. J. Food Process Eng. 1996, 19, 121–134. [Google Scholar] [CrossRef]
- Grau, R.; Sánchez, A.J.; Girón, J.; Iborra, E.; Fuentes, A.; Barat, J.M. Nondestructive Assessment of Freshness in Packaged Sliced Chicken Breasts Using SW-NIR Spectroscopy. Food Res. Int. 2011, 44, 331–337. [Google Scholar] [CrossRef]
- Dissing, B.S.; Papadopoulou, O.S.; Tassou, C.; Ersbøll, B.K.; Carstensen, J.M.; Panagou, E.Z.; Nychas, G.-J. Using Multispectral Imaging for Spoilage Detection of Pork Meat. Food Bioprocess Technol. 2013, 6, 2268–2279. [Google Scholar] [CrossRef]
- Tsakanikas, P.; Pavlidis, D.; Panagou, E.; Nychas, G.-J. Exploiting Multispectral Imaging for Non-Invasive Contamination Assessment and Mapping of Meat Samples. Talanta 2016, 161, 606–614. [Google Scholar] [CrossRef] [PubMed]
- Tao, F.; Peng, Y.; Gomes, C.L.; Chao, K.; Qin, J. A Comparative Study for Improving Prediction of Total Viable Count in Beef Based on Hyperspectral Scattering Characteristics. J. Food Eng. 2015, 162, 38–47. [Google Scholar] [CrossRef]
- Ropodi, A.I.; Panagou, E.Z.; Nychas, G.J.E. Rapid Detection of Frozen-Then-Thawed Minced Beef Using Multispectral Imaging and Fourier Transform Infrared Spectroscopy. Meat Sci. 2018, 135, 142–147. [Google Scholar] [CrossRef]
- Barbin, D.F.; ElMasry, G.; Sun, D.W.; Allen, P. Non-Destructive Determination of Chemical Composition in Intact and Minced Pork Using near-Infrared Hyperspectral Imaging. Food Chem. 2013, 138, 1162–1171. [Google Scholar] [CrossRef]
- Venel, C.; Mullen, A.M.; Downey, G.; Troy, D.J. Prediction of Tenderness and Other Quality Attributes of Beef by near Infrared Reflectance Spectroscopy between 750 and 1100 Nm; Further Studies. J. Infrared Spectrosc. 2001, 9, 185–198. [Google Scholar] [CrossRef]
- Borràs, E.; Ferré, J.; Boqué, R.; Mestres, M.; Aceña, L.; Busto, O. Data Fusion Methodologies for Food and Beverage Authentication and Quality Assessment—A Review. Anal. Chim. Acta 2015, 891, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Márquez, C.; López, M.I.; Ruisánchez, I.; Callao, M.P. FT-Raman and NIR Spectroscopy Data Fusion Strategy for Multivariate Qualitative Analysis of Food Fraud. Talanta 2016, 161, 80–86. [Google Scholar] [CrossRef] [PubMed]
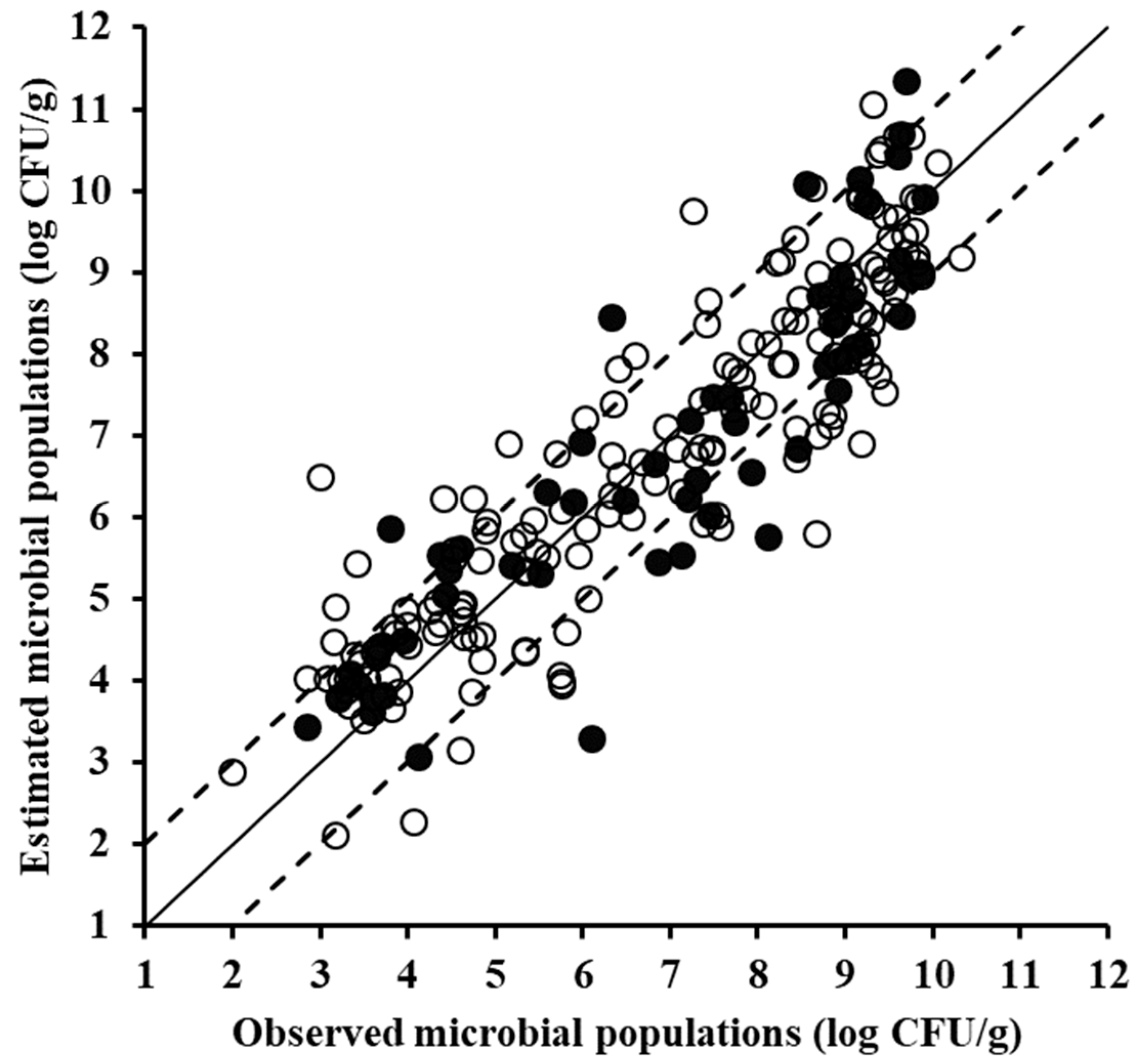
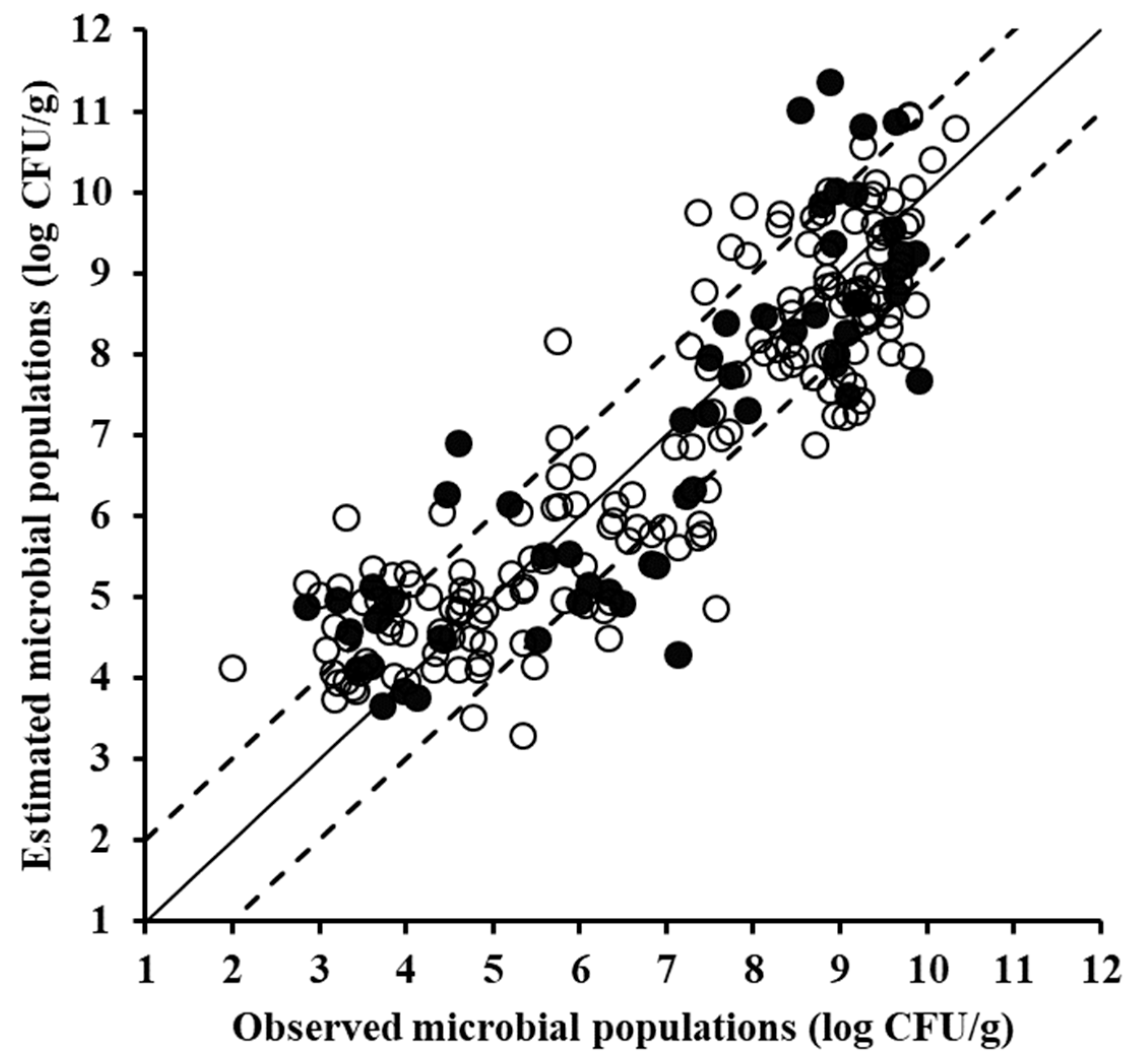
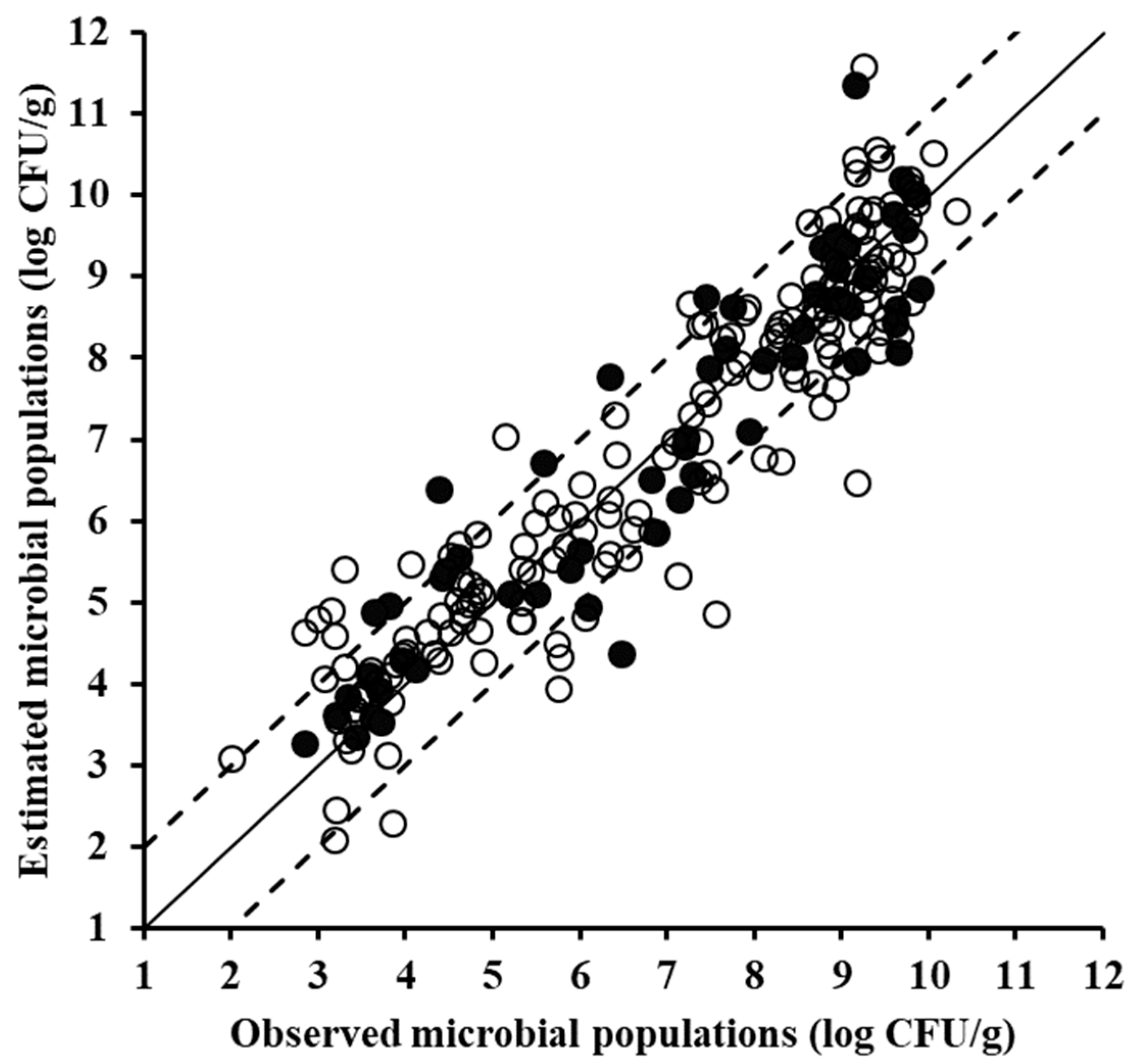
| Type of Data | Data Set | Slope | Offset | R2 | RMSE (Log CFU/g) |
|---|---|---|---|---|---|
| FTIR data | Training (n = 170) | 0.844 | 1.068 | 0.844 | 0.886 |
| Cross-validation * | 0.826 | 1.191 | 0.793 | 1.028 | |
| Testing (n = 58) | 0.951 | 0.099 | 0.834 | 0.915 | |
| VIS data | Training (n = 170) | 0.860 | 0.960 | 0.860 | 0.840 |
| Cross-validation * | 0.839 | 1.100 | 0.800 | 1.009 | |
| Testing (n = 58) | 0.843 | 0.992 | 0.788 | 1.034 | |
| MSI data | Training (n = 170) | 0.841 | 1.092 | 0.841 | 0.896 |
| Cross-validation * | 0.817 | 1.261 | 0.794 | 1.025 | |
| Testing (n = 58) | 0.834 | 1.201 | 0.727 | 1.173 | |
| Data fusion | Training (n = 170) | 0.909 | 0.622 | 0.909 | 0.676 |
| Cross-validation * | 0.893 | 0.720 | 0.864 | 0.834 | |
| Testing (n = 58) | 0.883 | 0.861 | 0.862 | 0.836 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fengou, L.-C.; Spyrelli, E.; Lianou, A.; Tsakanikas, P.; Panagou, E.Z.; Nychas, G.-J.E. Estimation of Minced Pork Microbiological Spoilage through Fourier Transform Infrared and Visible Spectroscopy and Multispectral Vision Technology. Foods 2019, 8, 238. https://doi.org/10.3390/foods8070238
Fengou L-C, Spyrelli E, Lianou A, Tsakanikas P, Panagou EZ, Nychas G-JE. Estimation of Minced Pork Microbiological Spoilage through Fourier Transform Infrared and Visible Spectroscopy and Multispectral Vision Technology. Foods. 2019; 8(7):238. https://doi.org/10.3390/foods8070238
Chicago/Turabian StyleFengou, Lemonia-Christina, Evgenia Spyrelli, Alexandra Lianou, Panagiotis Tsakanikas, Efstathios Z. Panagou, and George-John E. Nychas. 2019. "Estimation of Minced Pork Microbiological Spoilage through Fourier Transform Infrared and Visible Spectroscopy and Multispectral Vision Technology" Foods 8, no. 7: 238. https://doi.org/10.3390/foods8070238
APA StyleFengou, L.-C., Spyrelli, E., Lianou, A., Tsakanikas, P., Panagou, E. Z., & Nychas, G.-J. E. (2019). Estimation of Minced Pork Microbiological Spoilage through Fourier Transform Infrared and Visible Spectroscopy and Multispectral Vision Technology. Foods, 8(7), 238. https://doi.org/10.3390/foods8070238







