Application of Near-Infrared Hyperspectral Imaging with Machine Learning Methods to Identify Geographical Origins of Dry Narrow-Leaved Oleaster (Elaeagnus angustifolia) Fruits
Abstract
1. Introduction
2. Materials and Methods
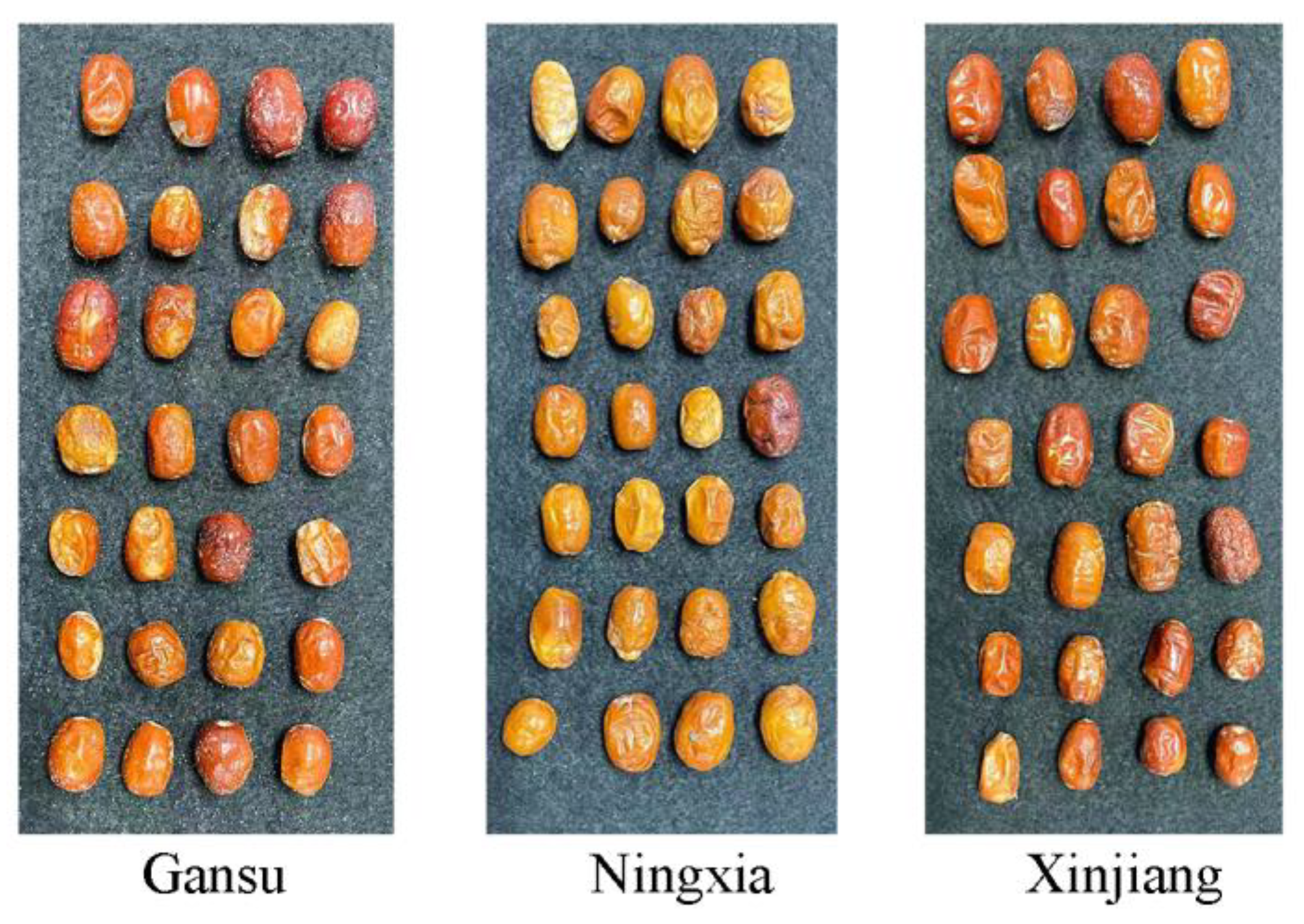
2.1. Sample Preparation
2.2. Hyperspectral Image Acquisition and Correction
2.3. Spectral Data Extraction
2.4. Data Analysis Methods
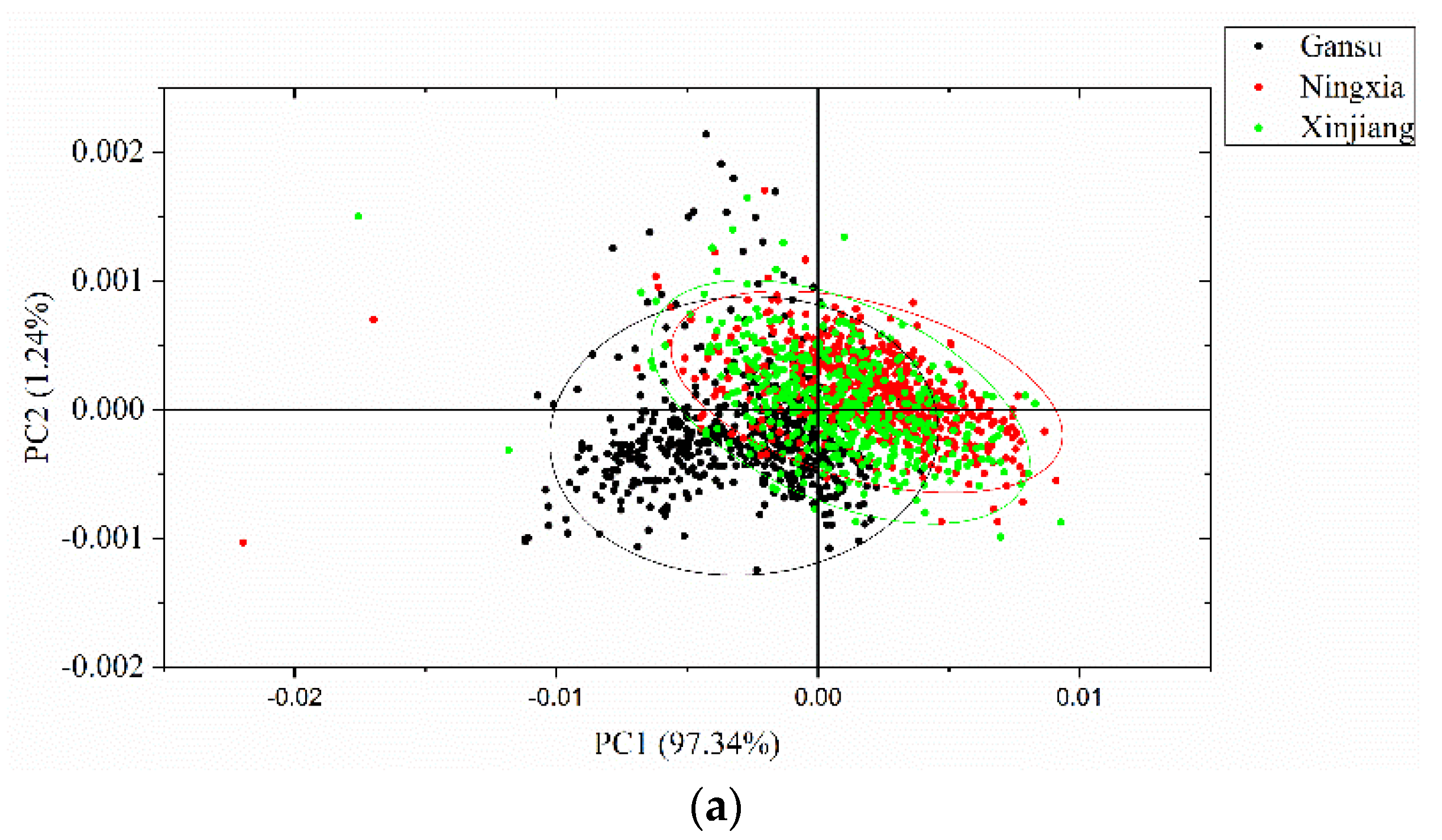
2.4.1. Principal Component Analysis
2.4.2. Partial Least Squares Discriminant Analysis
2.4.3. Support Vector Machine
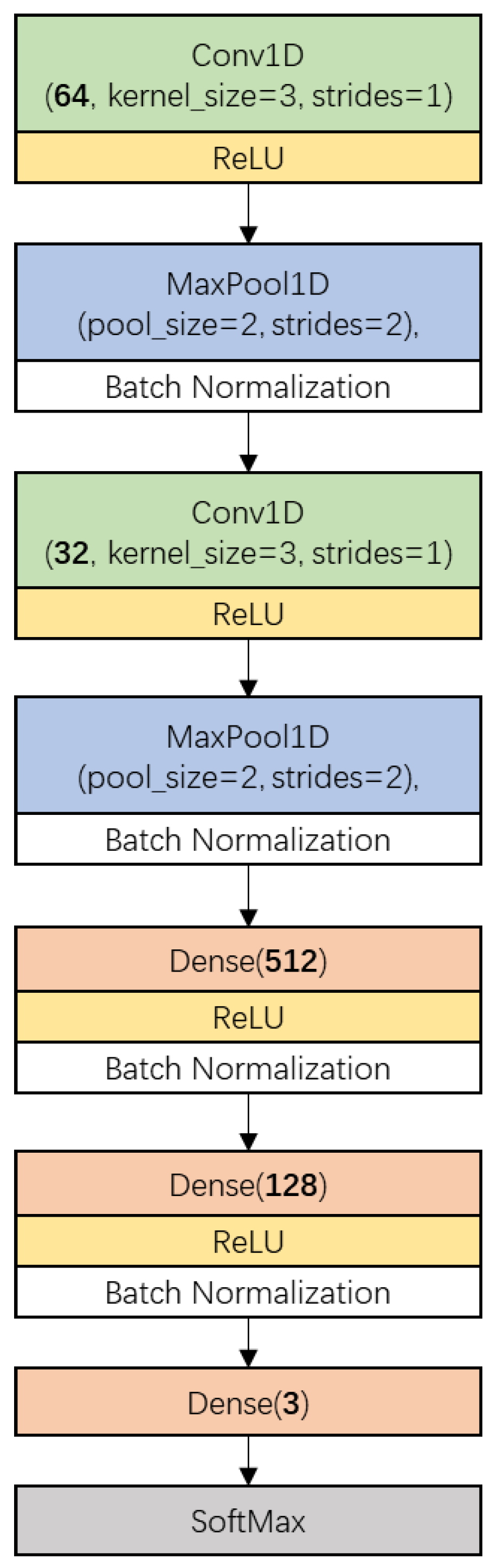
2.4.4. Convolutional Neural Network
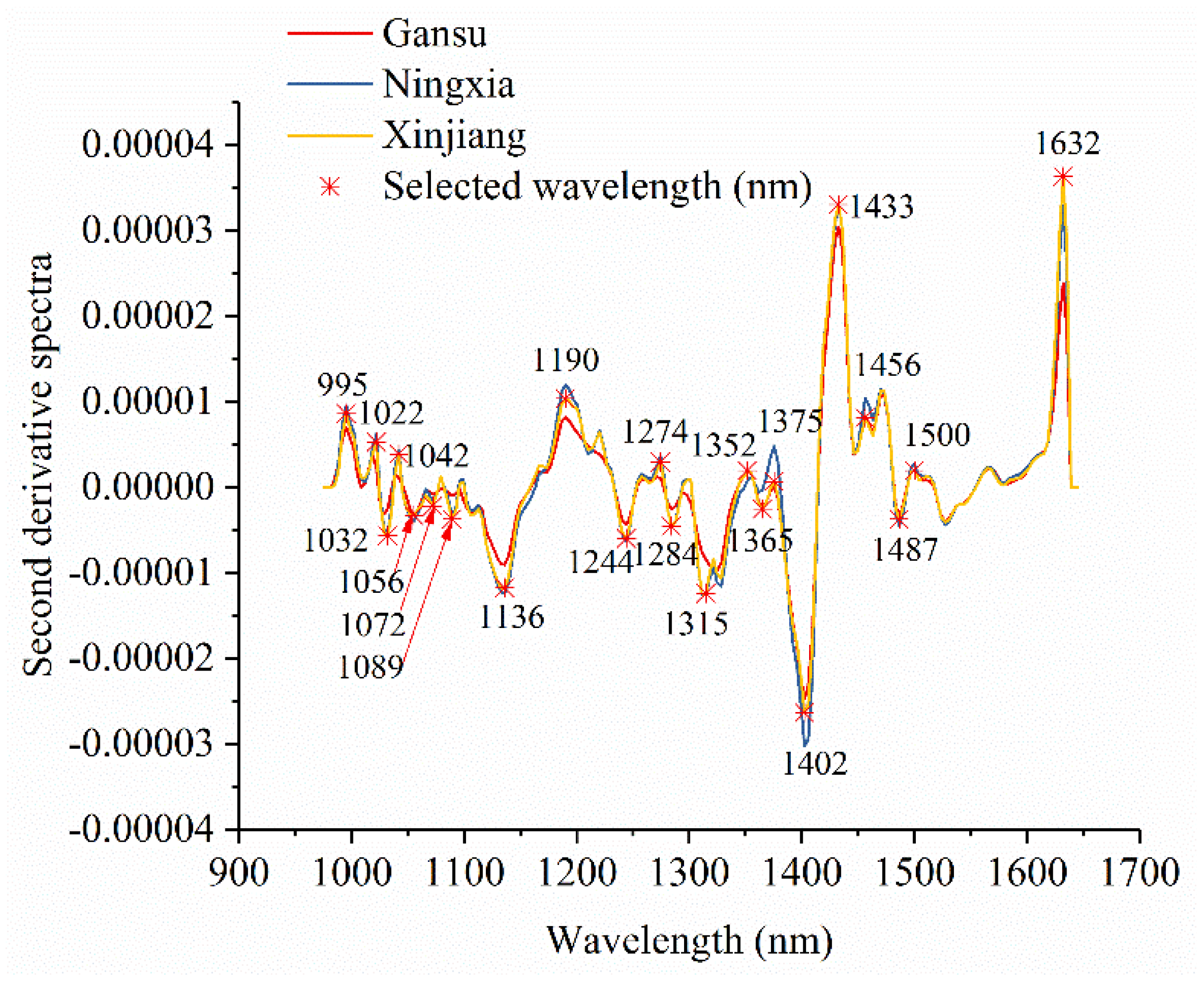
2.4.5. Optimal Wavelength Selection
2.5. Software and Model Evaluation
3. Results
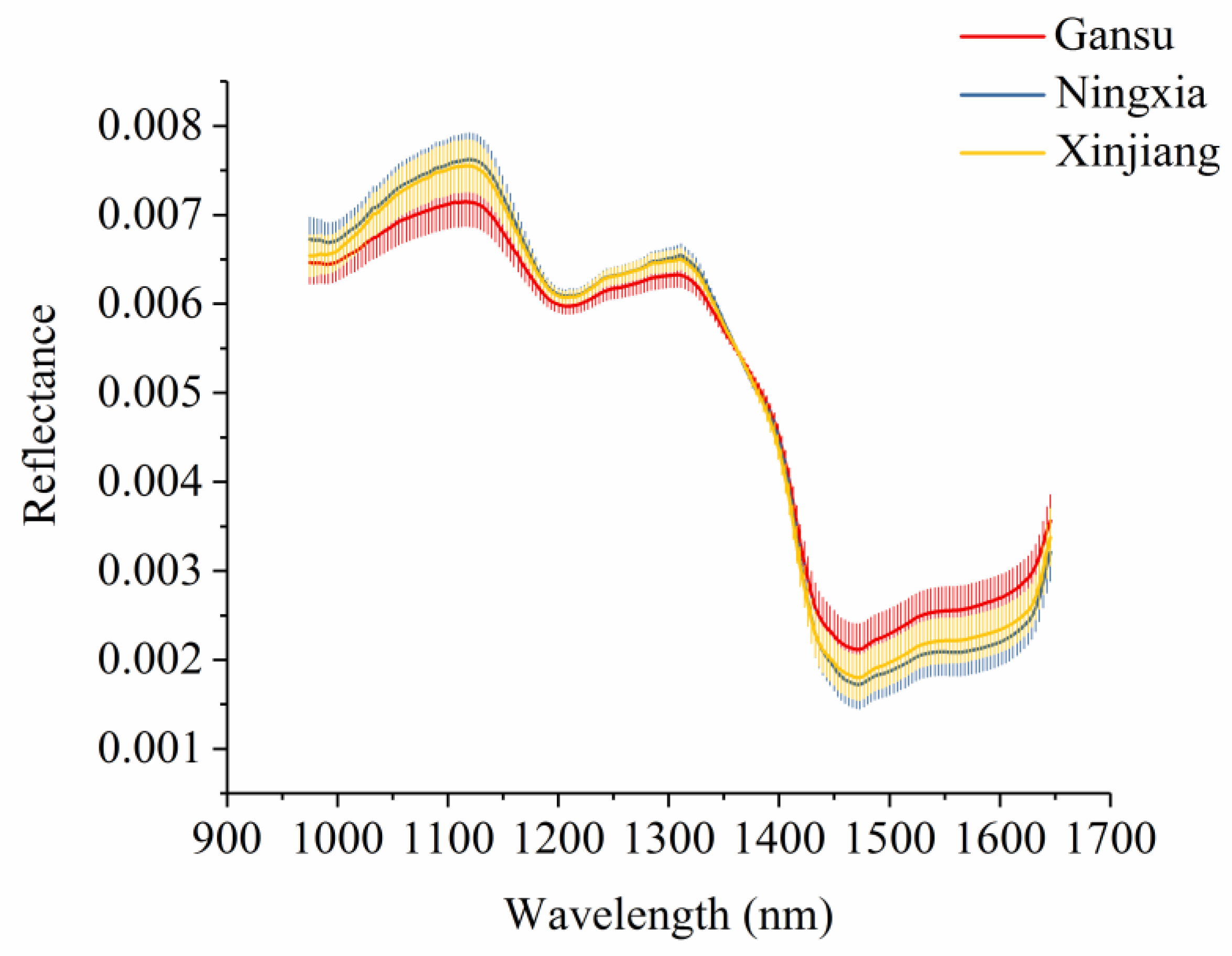
3.1. Spectral Profiles and Effective Wavelength Identification
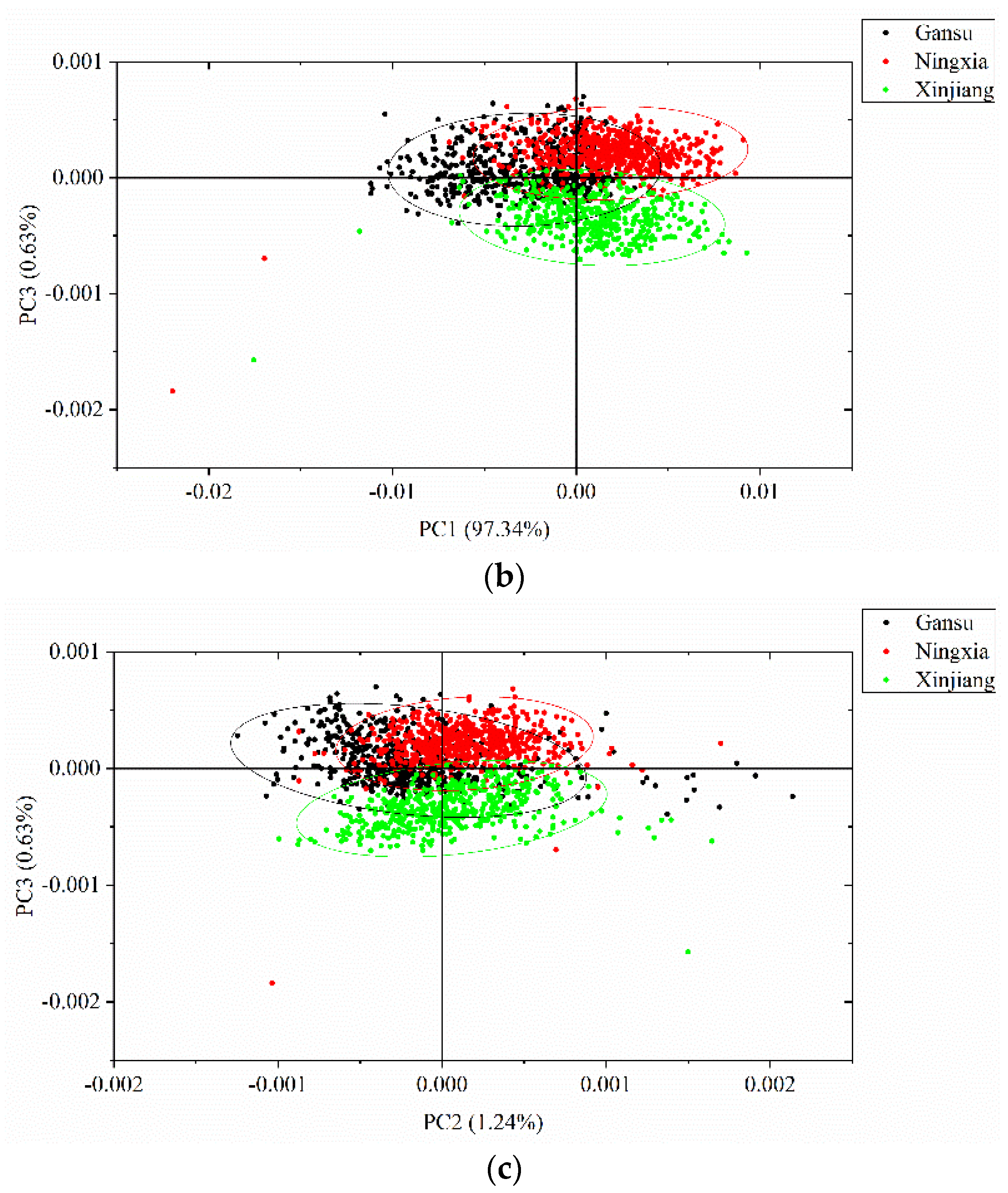
3.2. Principal Component Analysis
3.3. Classification Models Using Full Spectra
3.4. Classification Models Using Optimal Wavelengths
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhang, L.; Alvarez, L.V.; Bonthond, G.; Tian, C.; Fan, X. Cytospora elaeagnicola sp. nov. Associated with Narrow-leaved oleaster fruits Canker Disease in China. Mycobiology 2019, 47, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhanga, X.; Lia, G.; Sheng, D. Simulating the potential distribution of Elaeagnus angustifolia L. based on climatic constraints in China. Ecol. Eng. 2018, 113, 27–34. [Google Scholar] [CrossRef]
- Lin, J.; Li, J.P.; Yuan, F.; Yang, Z.; Wang, B.S.; Chen, M. Transcriptome profiling of genes involved in photosynthesis in Elaeagnus angustifolia L. under salt stress. Photosynthetica 2018, 56, 1–12. [Google Scholar] [CrossRef]
- Chen, X.; Yushuang, L.; Guangying, C.; Chi, G.; Shengge, L.; Huiming, H.; Tao, Y. Angustifolinoid A, a macrocyclic flavonoid glycoside from Elaeagnus angustifolia flowers. Tetrahedron Lett. 2018, 59, 2610–2613. [Google Scholar] [CrossRef]
- Du, H.; Chen, J.; Tian, S.; Gu, H.; Li, N.; Sun, Y.; Ru, J.; Wang, J. Extraction optimization, preliminary characterization and immunological activities in vitro of polysaccharides from Elaeagnus angustifolia L. pulp. Carbohydr. Polym. 2016, 151, 348–357. [Google Scholar] [CrossRef]
- Mcshane, R.; Auerbach, D.; Friedman, J.M.; Auble, G.T.; Shafroth, P.B.; Merigliano, M.; Scott, M.; Poff, N. Distribution of invasive and native riparian woody plants across the western USA in relation to climate, river flow, floodplain geometry and patterns of introduction. Ecography 2016, 38, 1254–1265. [Google Scholar] [CrossRef]
- Collette, L.K.D.; Pither, J. Insect assemblages associated with the exotic riparian shrub Russian olive (Elaeagnaceae), and co-occurring native shrubs in British Columbia, Canada. Can. Entomol. 2016, 148, 316–328. [Google Scholar] [CrossRef]
- Tredick, C.A.; Kelly, M.J.; Vaughan, M.R. Impacts of large-scale restoration efforts on black bear habitat use in Canyon de Chelly National Monument, Arizona, United States. J. Mammal. 2016, 97, gyw060. [Google Scholar] [CrossRef]
- Khamzina, A.; Lamers, J.P.A.; Martius, C. Above- and belowground litter stocks and decay at a multi-species afforestation site on arid, saline soil. Nutr. Cycl. Agroecosyst. 2016, 104, 187–199. [Google Scholar] [CrossRef]
- Singh, A.; Singh, N.B.; Hussain, I.; Singh, H.; Yadav, V.; Singh, S.C. Green synthesis of nano zinc oxide and evaluation of its impact on germination and metabolic activity of Solanum lycopersicum. J. Biol. 2016, 233, 84–94. [Google Scholar] [CrossRef]
- Singh, A.; Singh, N.B.; Afzal, S.; Singh, T.; Hussain, I. Zinc oxide nanoparticles: A review of their biological synthesis, antimicrobial activity, uptake, translocation and biotransformation in plants. J. Mater. Sci. 2017, 53, 185–201. [Google Scholar] [CrossRef]
- Hassanzadeh, Z.; Hassanpour, H. Evaluation of physicochemical characteristics and antioxidant properties of Elaeagnus angustifolia L. Sci. Hortic. 2018, 238, 83–90. [Google Scholar] [CrossRef]
- Waili, A.; Yili, A.; Maksimov, V.V.; Mijiti, Y.; Atamuratov, F.N.; Ziyavitdinov, Z.F.; Mamadrakhimov, A.; Asia, H.A.; Salikhov, S.I. Erratum to: Isolation of Biologically Active Constituents from Fruit of Elaeagnus angustifolia. Chem. Nat. Compd. 2016, 52, 776. [Google Scholar] [CrossRef]
- Wei, Q.; Wei, Y.; Wu, H.; Yang, X.; Zhang, H. Chemical Composition, Anti-oxidant, and Antimicrobial Activities of Four Saline-Tolerant Plant Seed Oils Extracted by SFC. J. Am. Oil Chem. Soc. 2016, 93, 1–10. [Google Scholar] [CrossRef]
- Morehart, A.L. Phomopsis canker and dieback of Elaeagnus angustifolia. Plant Dis. 2015, 64, 66. [Google Scholar] [CrossRef]
- Marena, M. Near-infrared spectroscopy and hyperspectral imaging: Non-destructive analysis of biological materials. Chem. Soc. Rev. 2014, 43, 8200–8214. [Google Scholar]
- Ru, C.; Li, Z.; Tang, R. A Hyperspectral Imaging Approach for Classifying Geographical Origins of Rhizoma Atractylodis Macrocephalae Using the Fusion of Spectrum-Image in VNIR and SWIR Ranges (VNIR-SWIR-FuSI). Sensors 2019, 19, 2045. [Google Scholar] [CrossRef]
- Noviyanto, A.; Abdulla, W.H. Honey botanical origin classification using hyperspectral imaging and machine learning. J. Food Eng. 2019, 265, 109684. [Google Scholar] [CrossRef]
- Minaei, S.; Shafiee, S.; Polder, G.; Moghadam-Charkari, N.; Van Ruth, S.; Barzegar, M.; Zahiri, J.; Alewijn, M.; Kuś, P.M. VIS/NIR imaging application for honey floral origin determination. Infrared Phys. Technol. 2017, 86, 218–225. [Google Scholar] [CrossRef]
- Puneet, M.; Alison, N.; Julius, T.; Guoping, L.; Sally, R.; Stephen, M. Near-infrared hyperspectral imaging for non-destructive classification of commercial tea products. J. Food Eng. 2018, 238, 70–77. [Google Scholar] [CrossRef]
- Zhang, C.; Liu, F.; He, Y. Identification of coffee bean varieties using hyperspectral imaging: Influence of preprocessing methods and pixel-wise spectra analysis. Sci. Rep. 2018, 8, 2166. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Zhang, C.; Zhu, H.; Zhao, Y.; He, Y. Application of near-infrared hyperspectral imaging to discriminate different geographical origins of chinese wolfberries. PLoS ONE 2017, 12, e0180534. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Zhou, L.; Gao, P.; Bao, Y.; He, Y.; Feng, L. Near-Infrared Hyperspectral Imaging Combined with Deep Learning to Identify Cotton Seed Varieties. Molecules 2019, 24, 3268. [Google Scholar] [CrossRef] [PubMed]
- Mahesh, S.; Jayas, D.S.; Paliwal, J.; White, N.D.G. Comparison of partial least squares regression (plsr) and principal components regression (pcr) methods for protein and hardness predictions using the near-infrared (nir) hyperspectral images of bulk samples of Canadian wheat. Food Bioprocess Technol. 2015, 8, 31–40. [Google Scholar] [CrossRef]
- Weinstock, B.A.; Janni, J.; Hagen, L.; Wright, S. Prediction of oil and oleic acid concentrations in individual corn (Zea mays L.) kernels using near-infrared reflectance hyperspectral imaging and multivariate analysis. Appl. Spectrosc. 2006, 60, 9. [Google Scholar] [CrossRef]
- Sun, J.; Lu, X.; Mao, H.; Wu, X.; Gao, H. Quantitative determination of rice moisture based on hyperspectral imaging technology and bcc-ls-svr algorithm. J. Food Process Eng. 2016, 40. [Google Scholar] [CrossRef]
- Zhang, N.; Liu, X.; Jin, X.; Li, C.; Wu, X.; Yang, S.; Ning, J.; Yanne, P. Determination of total iron-reactive phenolics, anthocyanins and tannins in wine grapes of skins and seeds based on near-infrared hyperspectral imaging. Food Chem. 2017, 237, 811–817. [Google Scholar] [CrossRef]
- Rodríguez-Pulido, F.J.; Hernández-Hierro, J.M.; Nogales-Bueno, J.; Gordillo, B.; González-Miret, M.L.; Heredia, F.J. A novel method for evaluating flavanols in grape seeds by near infrared hyperspectral imaging. Talanta 2014, 122, 145–150. [Google Scholar]
- Mazivila, S. Discrimination of the type of biodiesel/diesel blend (B5) using mid-infrared spectroscopy and PLS-DA. Fuel 2015, 142, 222–246. [Google Scholar] [CrossRef]
- Botelho, B.G.; Reis, N.; Oliveira, L.S.; Sena, M.M. Development and analytical validation of a screening method for simultaneous detection of five adulterants in raw milk using mid-infrared spectroscopy and PLS-DA. Food Chem. 2015, 181, 31–37. [Google Scholar] [CrossRef]
- Balage, J.M.; Amigo, J.M.; Antonelo, D.S.; Mazon, M.R.; e Silva, S.D. Shear force analysis by core location in Longissimus steaks from Nellore cattle using hyperspectral images—A feasibility study. Meat Sci. 2018, 143, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Melucci, D.; Bendini, A.; Tesini, F.; Barbieri, S.; Zappi, A.; Vichi, S.; Conte, L.; Gallina, T.T. Rapid direct analysis to discriminate geographic origin of extra virgin olive oils by flash gas chromatography electronic nose and chemometrics. Food Chem. 2016, 204, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Da Costa, G.B.; Fernandes, D.D.S.; Gomes, A.A.; De Almeida, V.E.; Veras, G. Using near infrared spectroscopy to classify soybean oil according to expiration date. Food Chem. 2016, 196, 539–543. [Google Scholar] [CrossRef]
- Du, L.; Lu, W.; Cai, Z.J.; Bao, L.; Hartmann, C.; Gao, B.; Yu, L.L. Rapid detection of milk adulteration using intact protein flow injection mass spectrometric fingerprints combined with chemometrics. Food Chem. 2017, 240, 573–578. [Google Scholar] [CrossRef]
- Schmutzler, M.; Beganovic, A.; Böhler, G.; Huck, C.W. Methods for detection of pork adulteration in veal product based on FT-NIR spectroscopy for laboratory, industrial and on-site analysis. Food Control 2015, 57, 258–267. [Google Scholar] [CrossRef]
- Yang, H.X.; Fu, H.B.; Wang, H.D.; Jia, J.W.; Sigrist, M.W.; Dong, F.Z. Laser-induced breakdown spectroscopy applied to the characterization of rock by support vector machine combined with principal component analysis. Chin. Phys. B 2016, 25, 065201. [Google Scholar] [CrossRef]
- Li, J.L.; Sun, D.W.; Pu, H.; Jayas, D.S. Determination of trace thiophanate-methyl and its metabolite carbendazim with teratogenic risk in red bell pepper (Capsicum annuum L.) by surface-enhanced Raman imaging technique. Food Chem. 2017, 218, 543–552. [Google Scholar] [CrossRef]
- Ropodi, A.I.; Panagou, E.Z.; Nychas, G.J.E. Multispectral imaging (MSI): A promising method for the detection of minced beef adulteration with horsemeat. Food Control 2017, 73, 57–63. [Google Scholar] [CrossRef]
- Wu, N.; Zhang, C.; Bai, X.; Du, X.; He, Y. Discrimination of Chrysanthemum Varieties Using Hyperspectral Imaging Combined with a Deep Convolutional Neural Network. Moleclues 2018, 23, 2831. [Google Scholar] [CrossRef]
- Qiu, Z.; Chen, J.; Zhao, Y.; Zhu, S.; He, Y.; Zhang, C. Variety Identification of Single Rice Seed Using Hyperspectral Imaging Combined with Convolutional Neural Network. Appl. Sci. 2018, 8, 212. [Google Scholar] [CrossRef]
- Zhang, C.; Feng, X.; Wang, J.; Liu, F.; He, Y.; Zhou, W. Mid-infrared spectroscopy combined with chemometrics to detect Sclerotinia stem rot on oilseed rape (Brassica napus L.) leaves. Plant Methods 2017, 13, 39. [Google Scholar] [CrossRef] [PubMed]

| Model | Category Values | Calibration | Validation | Prediction | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 2 | Total (%) | 0 | 1 | 2 | Total (%) | 0 | 1 | 2 | Total (%) | ||
| PLS | 0 * | 539 | 0 | 0 | 291 | 0 | 0 | 268 | 0 | 7 | |||
| 1 | 0 | 601 | 1 | 0 | 303 | 0 | 0 | 299 | 1 | ||||
| 2 | 0 | 0 | 481 | 0 | 0 | 241 | 0 | 0 | 240 | ||||
| Total (%) | 99.94 | 100 | 99.02 | ||||||||||
| SVM | 0 | 539 | 0 | 0 | 289 | 0 | 2 | 224 | 0 | 51 | |||
| 1 | 0 | 602 | 0 | 0 | 303 | 0 | 0 | 300 | 0 | ||||
| 2 | 0 | 0 | 481 | 0 | 0 | 241 | 0 | 0 | 240 | ||||
| Total (%) | 100 | 99.76 | 93.74 | ||||||||||
| CNN | 0 | 539 | 0 | 0 | 289 | 0 | 2 | 253 | 0 | 22 | |||
| 1 | 1 | 601 | 0 | 0 | 303 | 0 | 0 | 300 | 0 | ||||
| 2 | 6 | 0 | 475 | 4 | 0 | 237 | 0 | 0 | 240 | ||||
| Total (%) | 99.57 | 99.28 | 97.30 | ||||||||||
| Model | Category Values | Calibration | Validation | Prediction | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 2 | Total (%) | 0 | 1 | 2 | Total (%) | 0 | 1 | 2 | Total (%) | ||
| PLS | 0 * | 538 | 0 | 1 | 291 | 0 | 0 | 272 | 0 | 3 | |||
| 1 | 1 | 601 | 0 | 0 | 303 | 0 | 0 | 300 | 0 | ||||
| 2 | 1 | 0 | 480 | 0 | 0 | 241 | 0 | 0 | 240 | ||||
| Total (%) | 99.92 | 100 | 99.63 | ||||||||||
| SVM | 0 | 539 | 0 | 0 | 271 | 0 | 20 | 238 | 0 | 37 | |||
| 1 | 0 | 602 | 0 | 0 | 303 | 0 | 0 | 300 | 0 | ||||
| 2 | 2 | 0 | 479 | 1 | 0 | 240 | 1 | 0 | 239 | ||||
| Total (%) | 99.88 | 97.49 | 95.34 | ||||||||||
| CNN | 0 | 539 | 0 | 0 | 287 | 0 | 4 | 263 | 0 | 12 | |||
| 1 | 0 | 602 | 0 | 0 | 303 | 0 | 0 | 299 | 1 | ||||
| 2 | 4 | 0 | 477 | 8 | 0 | 233 | 5 | 0 | 235 | ||||
| Total (%) | 99.75 | 98.56 | 97.79 | ||||||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, P.; Xu, W.; Yan, T.; Zhang, C.; Lv, X.; He, Y. Application of Near-Infrared Hyperspectral Imaging with Machine Learning Methods to Identify Geographical Origins of Dry Narrow-Leaved Oleaster (Elaeagnus angustifolia) Fruits. Foods 2019, 8, 620. https://doi.org/10.3390/foods8120620
Gao P, Xu W, Yan T, Zhang C, Lv X, He Y. Application of Near-Infrared Hyperspectral Imaging with Machine Learning Methods to Identify Geographical Origins of Dry Narrow-Leaved Oleaster (Elaeagnus angustifolia) Fruits. Foods. 2019; 8(12):620. https://doi.org/10.3390/foods8120620
Chicago/Turabian StyleGao, Pan, Wei Xu, Tianying Yan, Chu Zhang, Xin Lv, and Yong He. 2019. "Application of Near-Infrared Hyperspectral Imaging with Machine Learning Methods to Identify Geographical Origins of Dry Narrow-Leaved Oleaster (Elaeagnus angustifolia) Fruits" Foods 8, no. 12: 620. https://doi.org/10.3390/foods8120620
APA StyleGao, P., Xu, W., Yan, T., Zhang, C., Lv, X., & He, Y. (2019). Application of Near-Infrared Hyperspectral Imaging with Machine Learning Methods to Identify Geographical Origins of Dry Narrow-Leaved Oleaster (Elaeagnus angustifolia) Fruits. Foods, 8(12), 620. https://doi.org/10.3390/foods8120620








