Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains
2.2. Serotyping
2.3. Antibiotic Susceptibility Testing
2.4. Growth Kinetics
2.5. Biofilm Determination
2.6. Statistical Analysis
3. Results and Discussion
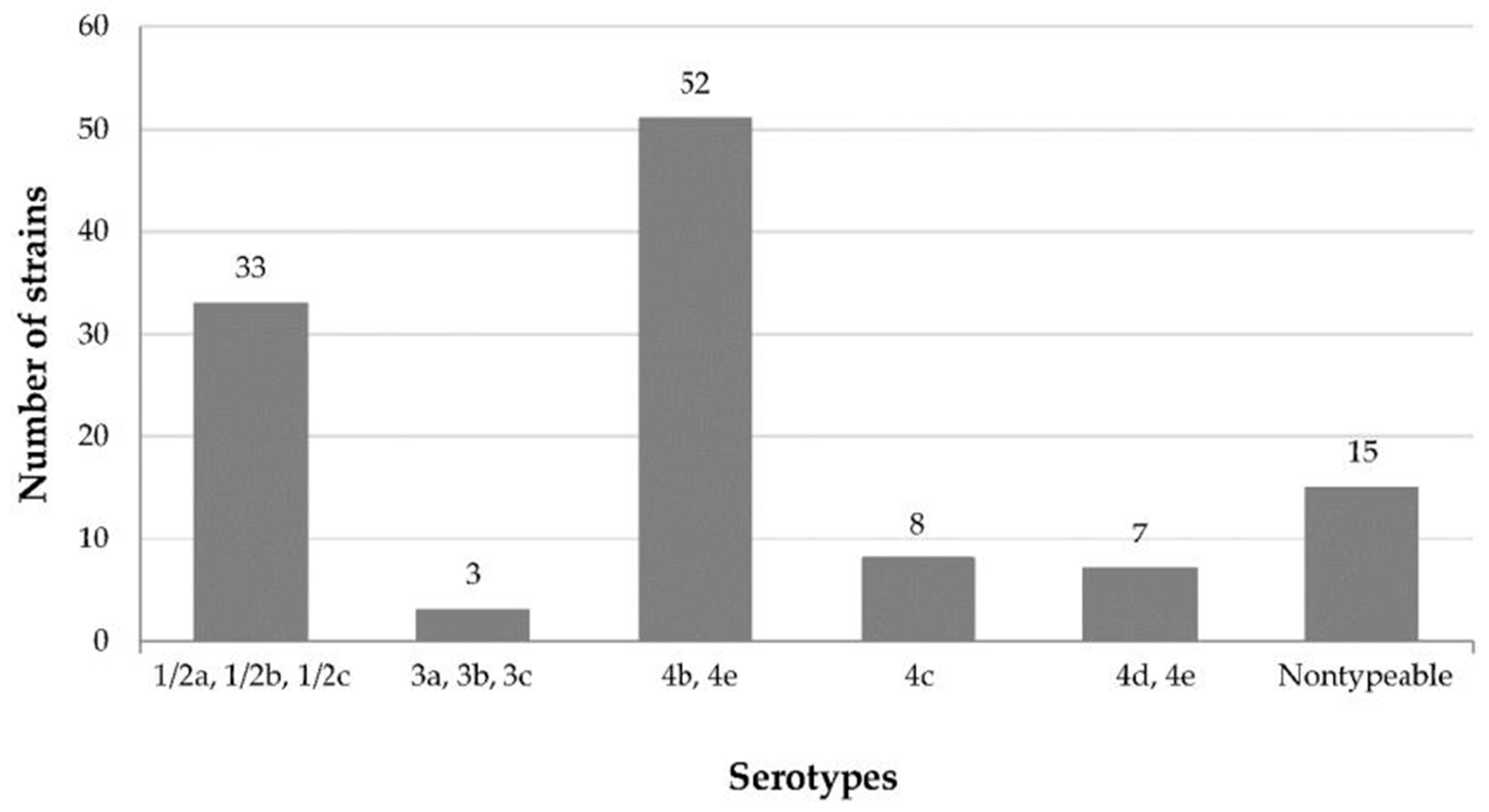
3.1. Serotyping
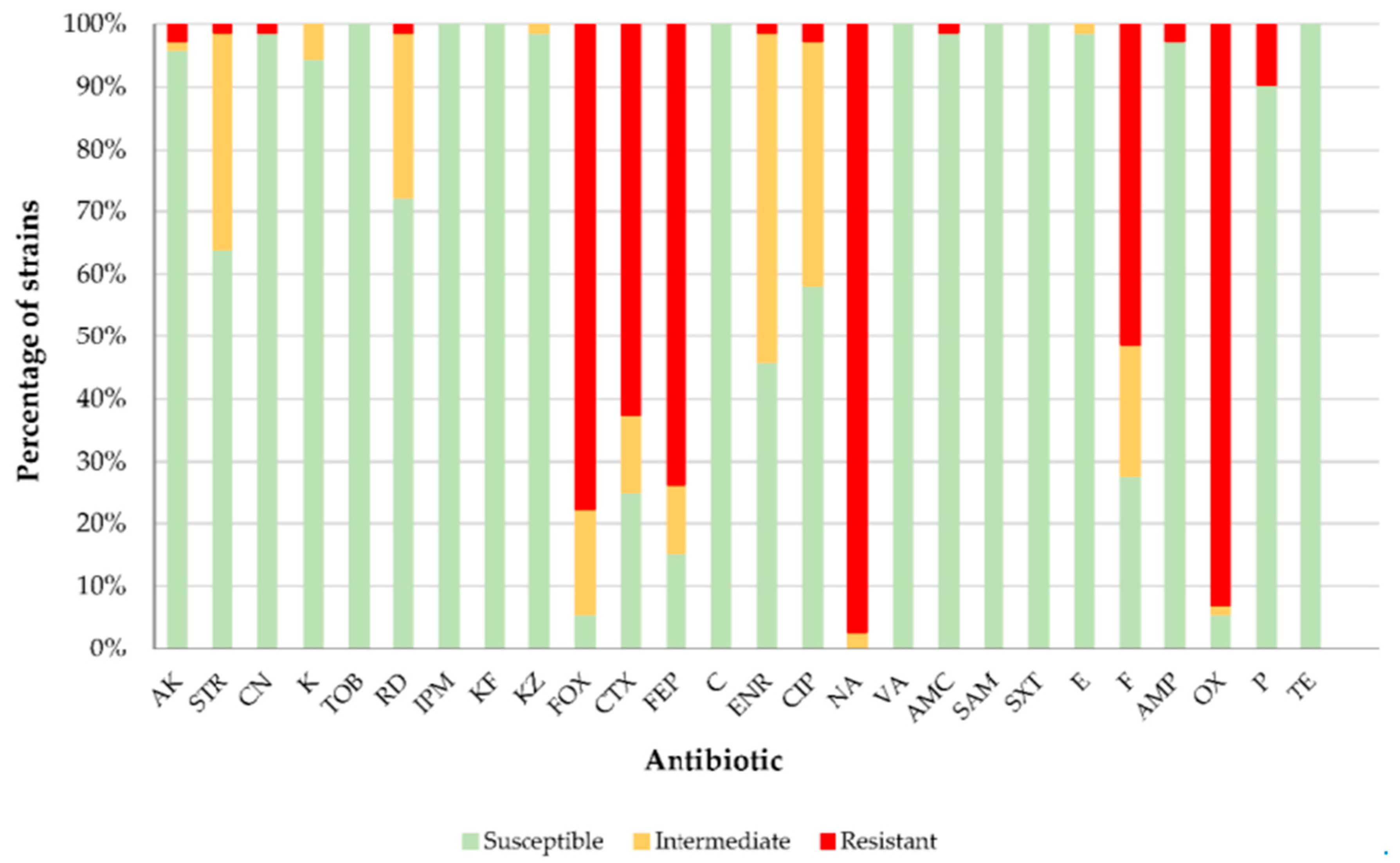
3.2. Antibiotic Resistance
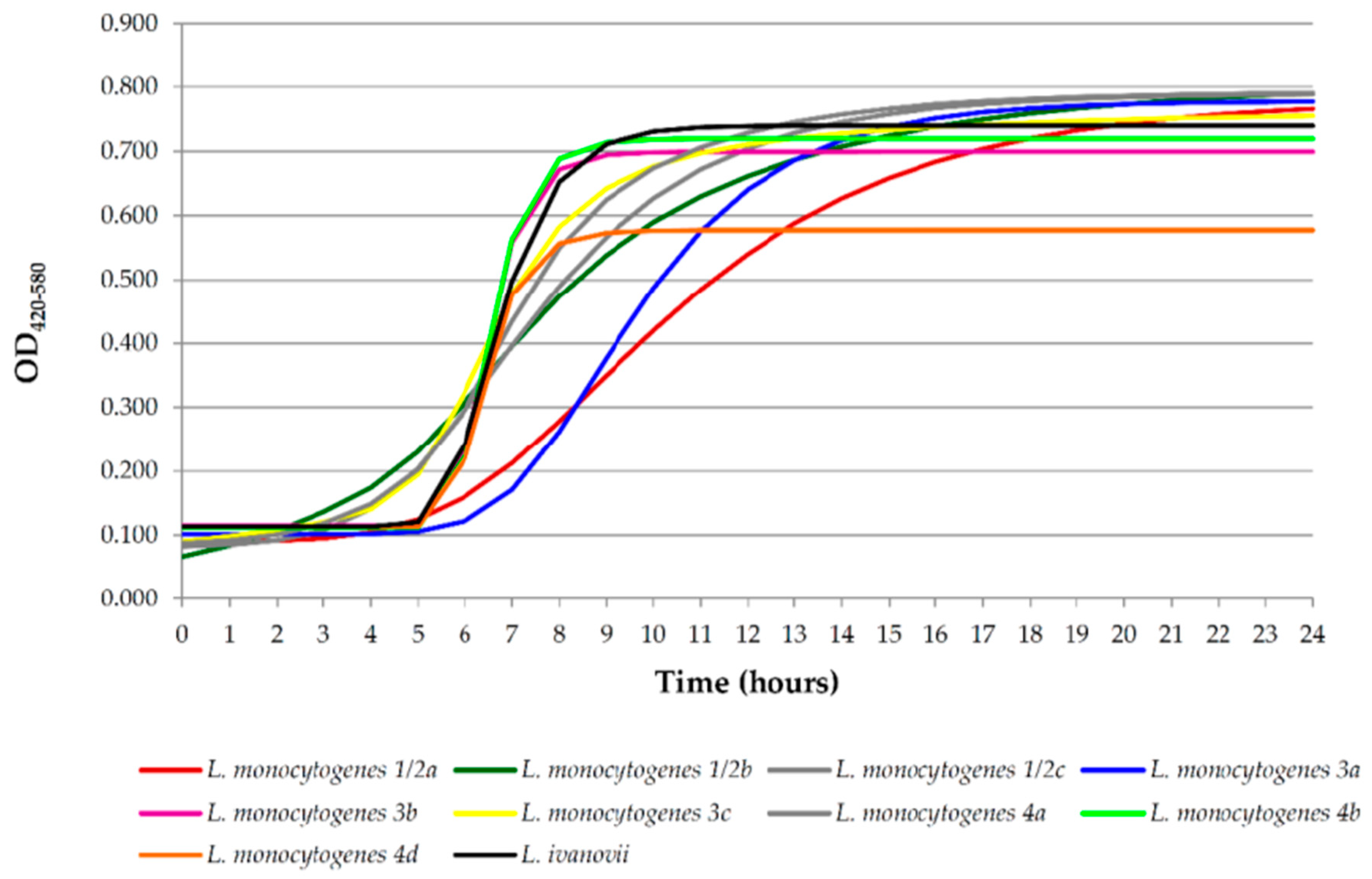
3.3. Growth Kinetics
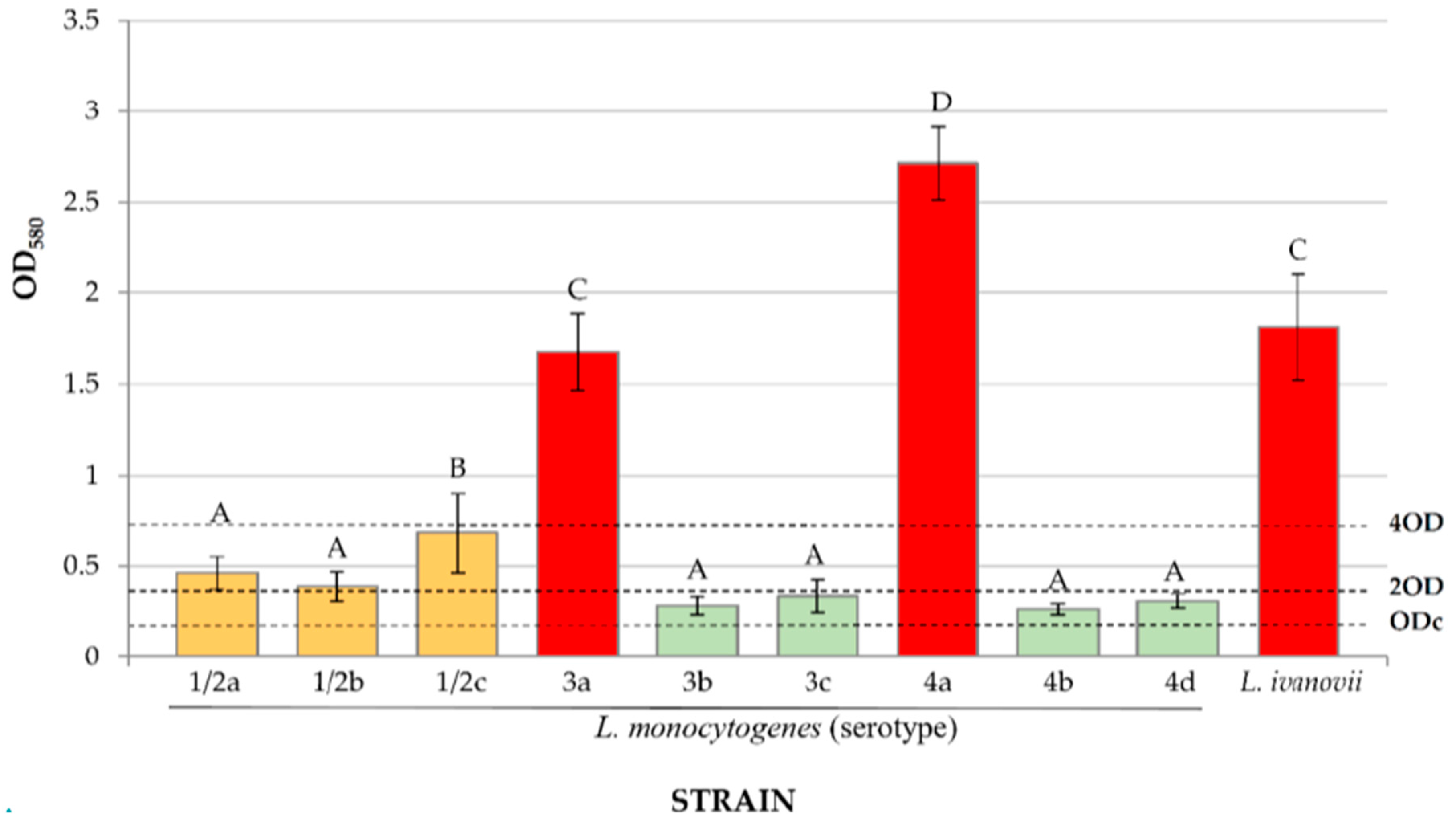
3.4. Biofilm Formation
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Leclercq, A.; Moura, A.; Vales, G.; Tessaud-Rita, N.; Aguilhon, C. Listeria thailandensis sp. nov. Int. J. Syst. Microbiol. 2019, 69, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Herrador, Z.; Gherasim, A.; López-Vélez, R.; Benito, A. Listeriosis in Spain based on hospitalisation records, 1997 to 2015: Need for greater awareness. Eurosurveillance 2019, 24, 1800271. [Google Scholar] [CrossRef] [PubMed]
- Ariza-Miguel, J.; Fernández-Natal, M.I.; Soriano, F.; Hernández, M.; Stessl, B.; Rodríguez-Lázaro, D. Molecular epidemiology of invasive listeriosis due to Listeria monocytogenes in a Spanish hospital over a nine-year study period, 2006–2014. BioMed Res. Int. 2015, 191409. [Google Scholar] [CrossRef]
- Tan, M.F.; Siow, C.C.; Dutta, A.; Mutha, N.V.; Wee, W.Y.; Heydari, H.; Tan, S.Y.; Ang, M.Y.; Wong, G.J.; Choo, S.W. Development of ListeriaBase and comparative analysis of List. monocytogenes. BMC Genom. 2015, 16, 755–774. [Google Scholar] [CrossRef]
- Donovan, S. Listeriosis: A rare but deadly disease. Clin. Microbiol. Newsl. 2015, 37, 135–141. [Google Scholar] [CrossRef]
- Lomonaco, S.; Nucera, D.; Filipello, V. The evolution and epidemiology of Listeria monocytogenes in Europe and the United States. Infect. Genet. Evol. 2015, 35, 172–183. [Google Scholar] [CrossRef]
- Parrilla, F.; Vaqué, J. Incidence study of listeriosis in Spain. Gac. Sanit. 2014, 28, 74–76. [Google Scholar] [CrossRef]
- CDC. Listeria (Listeriosis). Centers for Disease Control and Prevention. Available online: https://www.cdc.gov/listeria/index.html (accessed on 20 September 2019).
- EFSA; ECDC. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2017. EFSA J. 2018, 16, e05500. [Google Scholar] [CrossRef]
- Cunha, S.; Komora, N.; Magalhaes, R.; Almeida, G.; Ferreira, V.; Teixeira, P. Characterization of clinical and food Listeria monocytogenes isolates with different antibiotic resistance patterns through simulated gastrointestinal tract conditions and environmental stresses. Microb. Risk Anal. 2016, 1, 40–46. [Google Scholar] [CrossRef]
- Fallah, A.A.; Saei-Dehkordi, S.S.; Mahzounieh, M. Occurrence and antibiotic resistance profiles of Listeria monocytogenes isolated from seafood products and market and processing environments in Iran. Food Control 2013, 34, 630–636. [Google Scholar] [CrossRef]
- Martín, B.; Perich, A.; Gómez, D.; Yangüela, J.; Rodríguez, A.; Garriga, M.; Aymerich, T. Diversity and distribution of Listeria monocytogenes in meat processing plants. Food Microbiol. 2014, 44, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Ragon, M.; Wirth, T.; Hollandt, F.; Lavenir, R.; Lecuit, M.; Le Monnier, A.; Brisse, S. A new perspective on Listeria monocytogenes evolution. PLoS Pathog. 2008, 4, e1000146. [Google Scholar] [CrossRef] [PubMed]
- Jamshidi, A.; Zeinali, T. Significance and characteristics of Listeria monocytogenes in poultry products. Int. J. Food Sci. 2019, 7835253. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wu, Q.; Zhang, J.; Chen, M.; Yan, Z. Prevalence, antibiotic resistance and genetic diversity of Listeria monocytogenes isolated from retail ready-to-eat foods in China. Food Control 2015, 47, 340–347. [Google Scholar] [CrossRef]
- Capita, R.; Alonso-Calleja, C. Antibiotic-resistant bacteria: A challenge for the food industry. Crit. Rev. Food Sci. Nutr. 2013, 53, 11–48. [Google Scholar] [CrossRef]
- Komora, N.; Bruschi, C.; Magalhães, R.; Ferreira, V.; Teixeira, P. Survival of Listeria monocytogenes with different antibiotic resistance patterns to food-associated stresses. Int. J. Food Microbiol. 2017, 245, 79–87. [Google Scholar] [CrossRef]
- Olaimat, A.N.; Al-Holy, M.A.; Shahbaz, H.M.; Al-Nabulsi, A.A.; Ghoush, M.H.A.; Osaili, T.M.; Ayyash, M.M.; Holley, R.A. Emergence of antibiotic resistance in Listeria monocytogenes isolated from food products: A comprehensive review. Compr. Rev. Food Sci. Food Saf. 2018, 17, 1277–1292. [Google Scholar] [CrossRef]
- Gómez, D.; Azón, E.; Marco, N.; Carramiñana, J.J.; Rota, C.; Ariño, A.; Yangüela, J. Antimicrobial resistance of Listeria monocytogenes and Listeria innocua from meat products and meat-processing environment. Food Microbiol. 2014, 42, 61–65. [Google Scholar] [CrossRef]
- Díez-García, M.; Capita, R.; Alonso-Calleja, C. Influence of serotype on the growth kinetics and the ability to form biofilms of Salmonella isolates from poultry. Food Microbiol. 2012, 31, 173–180. [Google Scholar] [CrossRef]
- Rodríguez-Melcón, C.; Riesco-Peláez, F.; Carballo, J.; García-Fernández, C.; Capita, R.; Alonso-Calleja, C. Structure and viability of 24-and 72-h-old biofilms formed by four pathogenic bacteria on polystyrene and glass contact surfaces. Food Microbiol. 2018, 76, 513–517. [Google Scholar] [CrossRef]
- Capita, R.; Riesco-Peláez, F.; Alonso-Hernando, A.; Alonso-Calleja, C. Exposure of Escherichia coli ATCC 12806 to sublethal concentrations of food-grade biocides influences its ability to form biofilm, resistance to antimicrobials, and ultrastructure. Appl. Environ. Microbiol. 2014, 80, 1268–1280. [Google Scholar] [CrossRef] [PubMed]
- Piercey, M.J.; Hingston, P.A.; Hansen, L.T. Genes involved in Listeria monocytogenes biofilm formation at a simulated food processing plant temperature of 15 °C. Int. J. Food Microbiol. 2016, 223, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Breidt, F.; Gorski, L. Synergistic effects of sodium chloride, glucose, and temperature on biofilm formation by Listeria monocytogenes serotype 1/2a and 4b strains. Appl. Environ. Microbiol. 2010, 76, 1433–1441. [Google Scholar] [CrossRef] [PubMed]
- Orsi, R.H.; Den Bakker, H.C.; Wiedmann, M. Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med Microbiol. 2011, 301, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Rothrock, M.; Micciche, A.C.; Bodie, A.R.; Ricke, S.C. Listeria occurrence and potential control strategies in alternative and conventional poultry processing and retail. Front. Sustain. Food Syst. 2019, 3, 33. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Disk Susceptibility Test; Approved Standard, 9th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2012. [Google Scholar]
- Garthright, W.E. Refinements in the prediction of microbial growth curves. Food Microbiol. 1991, 8, 239–248. [Google Scholar] [CrossRef]
- Iannetti, L.; Acciari, V.A.; Antoci, S.; Addante, N.; Bardasi, L.; Bilei, S.; Calistri, P.; Cito, F.; Cogoni, P.; D’Aurelio, R.; et al. Listeria monocytogenes in ready-to-eat foods in Italy: Prevalence of contamination at retail and characterization of strains from meat products and cheese. Food Control 2016, 68, 55–61. [Google Scholar] [CrossRef]
- Bille, J.; Rocourt, J. WHO International multicenter Listeria monocytogenes subtyping study-rationale and set-up of the study. Food Microbiol. 1996, 32, 251–262. [Google Scholar] [CrossRef]
- Kérouanton, A.; Marault, M.; Petit, L.; Grout, J.; Dao, T.T.; Brisabois, A. Evaluation of a multiplex PCR assay as an alternative method for Listeria monocytogenes serotyping. J. Microbiol. Methods 2010, 80, 134–137. [Google Scholar] [CrossRef]
- Low, J.C.; Donachie, W. Clinical and serum antibody responses of lambs to infection by Listeria monocytogenes. Res. Vet. Sci. 1991, 5, 185–192. [Google Scholar] [CrossRef]
- Palumbo, J.D.; Boruchi, M.K.; Mandrell, R.E.; Gorski, L. Serotyping of Listeria monocytogenes by enzyme-linked immunosorbent assay and identification of mixed-serotype cultures by colony immunoblotting. J. Clin. Microbiol. 2003, 41, 564–571. [Google Scholar] [CrossRef] [PubMed]
- Scönberg, A.; Bannerman, E.; Courtieu, A.L.; Kiss, R.; McLauchlin, J.; Shah, S.; Wilhelms, D. Serotyping of 80 strains from the WHO multicentre international typing study of Listeria monocytogenes. Int. J. Food Microbiol. 1996, 32, 279–287. [Google Scholar] [CrossRef]
- Leclercq, A.; Chenal-Francisque, V.; Dieye, H.; Cantinelli, T.; Drali, R.; Brisse, S.; Lecuit, M. Characterization of the novel Listeria monocytogenes PCR serogrouping profile IVb-v1. Int. J. Food Microbiol. 2011, 147, 74–77. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, A.; Zhu, R.; Lan, R.; Jin, D.; Cui, Z.; Wang, Y.; Li, Z.; Wang, Y.; Xu, J.; et al. Genetic diversity and molecular typing of Listeria monocytogenes in China. BMC Microbiol. 2012, 12, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Ciolacu, L.; Nicolau, A.I.; Wagner, M.; Rychli, K. Listeria monocytogenes isolated from food samples from a Romanian black market show distinct virulence profiles. Int. J. Food Microbiol. 2015, 209, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Ebner, R.; Stephan, R.; Althaus, D.; Brisse, S.; Maury, M.; Tasara, T. Phenotypic and genotypic characteristics of Listeria monocytogenes strains isolated during 2011-2014 from different food matrices in Switzerland. Food Control 2015, 57, 321–326. [Google Scholar] [CrossRef]
- Korsak, D.; Borek, A.; Daniluk, S.; Grabowska, A.; Pappelbaum, K. Susceptibilities of Listeria monocytogenes strains isolated from food and food processing environment in Poland. Int. J. Food Microbiol. 2012, 158, 203–208. [Google Scholar] [CrossRef]
- Kramarenko, T.; Roasto, M.; Meremae, K.; Kuningas, M.; Poltsama, P.; Elias, T. Listeria monocytogenes prevalence and serotype diversity in various foods. Food Control 2013, 30, 24–29. [Google Scholar] [CrossRef]
- Prencipe, V.A.; Rizzi, V.; Acciari, V.; Iannetti, L.; Giovannini, A.; Serraino, A.; Calderone, D.; Rossi, A.; Morelli, D.; Marino, L.; et al. Listeria monocytogenes prevalence, contamination levels and strains characterization throughout the Parma ham processing chain. Food Control 2012, 25, 150–158. [Google Scholar] [CrossRef]
- Yu, T.; Jiang, X. Prevalence and characterization of Listeria monocytogenes isolated from retail food in Henan, China. Food Control 2014, 37, 228–231. [Google Scholar] [CrossRef]
- Meloni, D.; Galluzzo, P.; Mureddu, A.; Piras, F.; Griffiths, M.; Mazette, R. Listeria monocytogenes in RTE foods marketed in Italy: Prevalence and automated EcoRI ribotyping of the isolates. Int. J. Food Microbiol. 2009, 129, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Vasilev, V.; Japheth, R.; Breuer, R.; Andom, N.; Abraham, R.B.; Yoni, Y.; Valinsky, L.; Agmon, V. A survey of Listeria monocytogenes strains, isolated from ready-to-eat foods in Israel over a period of 10 years, 1998–2007. Food Control 2010, 21, 1179–1181. [Google Scholar] [CrossRef]
- Martins, E.A.; Germano, P.M.L. Listeria monocytogenes in ready-to-eat, sliced, cooked ham and salami products, marketed in the city of São Paulo, Brazil: Occurrence, quantification, and serotyping. Food Control 2011, 22, 297–302. [Google Scholar] [CrossRef]
- Fallah, A.A.; Saei-Dehkordi, S.S.; Rahnama, M.; Tahmasby, H.; Mahzounieh, M. Prevalence and antimicrobial resistance patterns of Listeria species isolated from poultry products marketed in Iran. Food Control 2012, 28, 327–332. [Google Scholar] [CrossRef]
- Notification Details—2019.2989. Foodborne Outbreak Caused by Listeria monocytogenes (>1.5x10E4 CFU/g) in Chilled Pork Products from Spain. Available online: https://webgate.ec.europa.eu/rasff-window/portal/?event=notificationDetail&NOTIF_REFERENCE=2019.2989 (accessed on 20 September 2019).
- Junta de Andalucía. Consejería de Salud y Familias. Available online: https://www.juntadeandalucia.es/organismos/saludyfamilias/actualidad/noticias/detalle/220344.html (accessed on 20 September 2019).
- Alonso-Hernando, A.; Prieto, M.; García-Fernández, C.; Alonso-Calleja, C.; Capita, R. Increase over time in the prevalence of multiple antibiotic resistance among isolates of Listeria monocytogenes from poultry in Spain. Food Control 2012, 23, 37–41. [Google Scholar] [CrossRef]
- Adzitey, F.; Ali, G.R.R.; Huda, N.; Cogan, T.; Corry, J. Prevalence, antibiotic resistance and genetic diversity of Listeria monocytogenes isolated from ducks, their rearing and processing environments in Penang, Malaysia. Food Control 2013, 32, 607–614. [Google Scholar] [CrossRef]
- Bae, D.; Mezal, E.H.; Smiley, R.D.; Cheng, C.-M.; Khan, A.A. The sub-species characterization and antimicrobial resistance of Listeria monocytogenes isolated from domestic and imported food products from 2004 to 2011. Food Res. Int. 2014, 64, 656–663. [Google Scholar] [CrossRef]
- Doménech, E.; Jimenez-Belenguer, A.; Amoros, J.A.; Ferrus, M.A.; Escriche, I. Prevalence and antimicrobial resistance of Listeria monocytogenes and Salmonella strains isolated in ready-to-eat foods in Eastern Spain. Food Control 2015, 47, 120–125. [Google Scholar] [CrossRef]
- Wang, X.-M.; Lü, X.-F.; Yin, L.; Liu, H.-F.; Zhang, W.-J.; Si, W.; Tu, S.-Y.; Shao, M.-L.; Liu, S.-G. Occurrence and antimicrobial susceptibility of Listeria monocytogenes isolates from retail raw foods. Food Control 2013, 32, 153–158. [Google Scholar] [CrossRef]
- WHO. Critically Important Antimicrobials for Human Medicine, 6th ed.; World Health Organization: Geneva, Switzerland, 2019. [Google Scholar]
- OIE. OIE List of Antimicrobial agents of Veterinary Importance; World Organization for Animal Health: Paris, France, 2018. [Google Scholar]
- Romanova, N.A.; Wolffs, P.F.G.; Brovko, L.Y.; Griffiths, M.W. Role of efflux pumps in adaptation and resistance of Listeria monocytogenes to benzalkonium chloride. Appl. Environ. Microbiol. 2006, 72, 3498–3503. [Google Scholar] [CrossRef]
- Mata, M.T.; Baquero, F.; Pérez-Díaz, J.C. A multidrug efflux transporter in Listeria monocytogenes. FEMS Microbiol. Lett. 2000, 187, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Hernando-Amado, S.; Blanco, P.; Alcalde-Rico, M.; Corona, F.; Reales-Calderón, J.A.; Sánchez, M.B.; Martínez, J.L. Multidrug efflux pumps as main players in intrinsic and acquired resistance to antimcirobials. Drug Resist. Updates 2016, 28, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Marian, M.N.; Aminah, S.M.; Zuraini, M.I.; Son, R.; Maimunah, M.; Lee, H.Y.; Wong, W.C.; Elexson, N. MPN-PCR detection and antimicrobial resistance of Listeria monocytogenes isolated from raw and ready-to-eat foods in Malaysia. Food Control 2012, 28, 309–314. [Google Scholar] [CrossRef]
- Augustin, J.-C.; Rosso, L.; Carlier, V. Estimation of temperature dependent growth rate and lag time of Listeria monocytogenes by optical density measurements. J. Microbiol. Methods 1999, 38, 137–146. [Google Scholar] [CrossRef]
- Magalhães, R.; Ferreira, V.; Brandão, T.R.S.; Palencia, R.C.; Almeida, G.; Teixeira, P. Persistent and non-persistent strains of Listeria monocytogenes: A focus on growth kinetics under different temperature, salt, and pH conditions and their sensitivity to sanitizers. Food Microbiol. 2016, 57, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Robinson, T.P.; Aboaba, O.O.; Kaloti, A.; Ocio, M.J.; Baranyi, J.; Mackey, B.M. The effect of inoculum size on the lag phase of Listeria monocytogenes. Int. J. Food Microbiol. 2001, 70, 163–173. [Google Scholar] [CrossRef]
- Tsai, H.N.; Hodgson, D.A. Development of a synthetic minimal medium for Listeria monocytogenes. Appl. Environ. Microbiol. 2003, 69, 6943–6945. [Google Scholar] [CrossRef]
- Vialette, M.; Pinon, A.; Chasseignaux, E.; Lange, M. Growths kinetics comparison of clinical and seafood Listeria monocytogenes isolates in acid and osmotic environment. Int. J. Food Microbiol. 2003, 82, 121–131. [Google Scholar] [CrossRef]
- Sant’Ana, A.S.; Franco, B.; Schaffner, D.W. Modeling the growth rate and lag time of different strains of Salmonella enterica and Listeria monocytogenes in ready-to-eat lettuce. Food Microbiol. 2012, 30, 267–273. [Google Scholar] [CrossRef]
- Xuan, X.-T.; Ding, T.; Li, J.; Ahn, J.-H.; Zhao, Y.; Chen, S.-G.; Ye, X.-Q.; Liu, D.-H. Estimation of growth parameters of Listeria monocytogenes after sublethal heat and slightly acidic electrolyzed water (SAEW) treatment. Food Control 2017, 71, 17–25. [Google Scholar] [CrossRef]
- Jarvis, N.A.; O´Bryan, C.A.; Ricke, S.C.; Johnson, M.G.; Crandall, P.G. A review of minimal and defined media for growth of Listeria monocytogenes. Food Control 2016, 66, 256–269. [Google Scholar] [CrossRef]
- Lianou, A.; Stopforth, J.D.; Yoon, Y.; Wiedmann, M.; Sofos, J.N. Growth and stress resistance variation in culture broth among Listeria monocytogenes strains of various serotypes and origins. J. Food Prot. 2006, 69, 2640–2647. [Google Scholar] [CrossRef]
- Mytilinaios, I.; Salih, M.; Schofield, H.K.; Lambert, R.J.W. Growth curve prediction from optical density data. Int. J. Food Microbiol. 2012, 154, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, J.S.; González, A.; Özçelik, N.; Rodríguez, M.R.; García de Fernando, G. Modeling the Listeria innocua micropopulation lag phase and its variability. Int. J. Food Microbiol. 2013, 164, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Melcón, C.; Capita, R.; Rodriguez-Jerez, J.J.; Martínez-Suárez, J.V.; Alonso-Calleja, C. Effect of low doses of disinfectants on the biofilm-forming ability of Listeria monocytogenes. Foodborne Pathog. Dis. 2019, 16, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Melcón, C.; Riesco-Peláez, F.; García-Fernández, C.; Alonso-Calleja, C.; Capita, R. Susceptibility of Listeria monocytogenes planktonic cultures and biofilms to sodium hypochlorite and benzalkonium chloride. Food Microbiol. 2019, 82, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Kadam, S.R.; Den Besten, H.M.W.; Dan Der Veen, S.; Zwietering, M.H.; Moezelaar, R.; Abee, T. Diversity assessment of Listeria monocytogenes biofilm formation: Impact of growth condition, serotype and strain origin. Int. J. Food Microbiol. 2013, 165, 259–264. [Google Scholar] [CrossRef]
- Nilsson, R.E.; Ross, T.; Bowman, J.P. Variability in biofilm production by Listeria monocytogenes correlated to strain origin and growth conditions. Int. J. Food Microbiol. 2011, 150, 14–24. [Google Scholar] [CrossRef]
- Nyenje, M.E.; Green, E.; Ndip, R.N. Biofilm formation and adherence characteristics of Listeria ivanovii strains isolated from ready-to-eat roods in Alice, South Africa. Sci. World J. 2012, 873909. [Google Scholar] [CrossRef]
- Eskhan, A.O.; Abu-Lail, N.I. Cellular and molecular investigations of the adhesion and mechanics of Listeria monocytogenes lineages’ I and II environmental and epidemic strains. J. Colloid Interface Sci. 2013, 394, 554–563. [Google Scholar] [CrossRef]
- Tsai, Y.-H.L.; Maron, S.B.; McGann, P.; Nightingale, K.K.; Wiedmann, M.; Orsi, R.H. Recombination and positive selection contributed to the evolution of Listeria monocytogenes lineages III and IV, two distinct and well supported uncommon L. monocytogenes lineages. Infect. Genet. Evol. 2011, 11, 1881–1890. [Google Scholar] [CrossRef] [PubMed]
- Paul, D.; Steele, C.; Donaldon, J.R.; Banes, M.M.; Kumar, R.; Bridges, S.M.; Arick, M.; Lawrence, M. Genome comparison of Listeria monocytogenes serotype 4a strain HCC23 with selected lineage I and lineage II L. monocytogenes strains and other Listeria strains. Genom. Data 2014, 2, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Franciosa, G.; Maugliani, A.; Scalfaro, C.; Floridi, F.; Aureli, P. Expression of internalin A and biofilm formation among Listeria monocytogenes clinical isolates. Int. J. Immunopathol. Pharm. 2009, 22, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Kovacevic, J.; Arguedas-Villa, C.; Wozniak, A.; Tasara, T.; Allen, K.J. Examination of food chain-derived Listeria monocytogenes strains of different serotypes reveals considerable diversity in inlA genotypes, mutability, and adaptation to cold temperatures. Appl. Environ. Microbiol. 2013, 79, 1915–1922. [Google Scholar] [CrossRef]
| Strain | Growth Kinetic Parameters | |||
|---|---|---|---|---|
| L | µ | E | T | |
| Listeria monocytogenes 1/2a | 5.319 ± 0.504 cde | 0.073 ± 0.018 a | 0.781 ± 0.015 de | 19.800 ± 0.447 d |
| Listeria monocytogenes 1/2b | 1.839 ± 2.998 a | 0.088 ± 0.049 ab | 0.805 ± 0.031 e | 18.000 ± 2.646 cd |
| Listeria monocytogenes 1/2c | 3.765 ± 0.957 bc | 0.103 ± 0.030 abc | 0.790 ± 0.038 e | 17.600 ± 2.074 cd |
| Listeria monocytogenes 3a | 6.597 ± 0.418 e | 0.118 ± 0.022 abc | 0.778 ± 0.024 de | 17.800 ± 1.304 cd |
| Listeria monocytogenes 3b | 5.717 ± 0.086 e | 0.387 ± 0.016 e | 0.699 ± 0.053 b | 9.167 ± 0.408 a |
| Listeria monocytogenes 3c | 3.479 ± 2.846 b | 0.164 ± 0.078 c | 0.761 ± 0.024 cde | 14.000 ± 3.674 b |
| Listeria monocytogenes 4a | 3.864 ± 1.656 bcd | 0.144 ± 0.081 bc | 0.793 ± 0.037 e | 15.600 ± 3.435 bc |
| Listeria monocytogenes 4b | 5.739 ± 0.111 e | 0.396 ± 0.026 e | 0.719 ± 0.033 bc | 9.167 ± 0.408 a |
| Listeria monocytogenes 4d | 5.729 ± 0.133 e | 0.345 ± 0.066 e | 0.578 ± 0.049 a | 9.000 ± 0.894 a |
| Listeria ivanovii | 5.528 ± 0.226 de | 0.272 ± 0.019 d | 0.740 ± 0.023 bcd | 11.000 ± 0.000 a |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Capita, R.; Felices-Mercado, A.; García-Fernández, C.; Alonso-Calleja, C. Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain. Foods 2019, 8, 542. https://doi.org/10.3390/foods8110542
Capita R, Felices-Mercado A, García-Fernández C, Alonso-Calleja C. Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain. Foods. 2019; 8(11):542. https://doi.org/10.3390/foods8110542
Chicago/Turabian StyleCapita, Rosa, Amanda Felices-Mercado, Camino García-Fernández, and Carlos Alonso-Calleja. 2019. "Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain" Foods 8, no. 11: 542. https://doi.org/10.3390/foods8110542
APA StyleCapita, R., Felices-Mercado, A., García-Fernández, C., & Alonso-Calleja, C. (2019). Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain. Foods, 8(11), 542. https://doi.org/10.3390/foods8110542






