What We Know and What We Need to Know about Aromatic and Cationic Biogenic Amines in the Gastrointestinal Tract
Abstract
1. Introduction
2. Histamine Biochemistry and Physiology
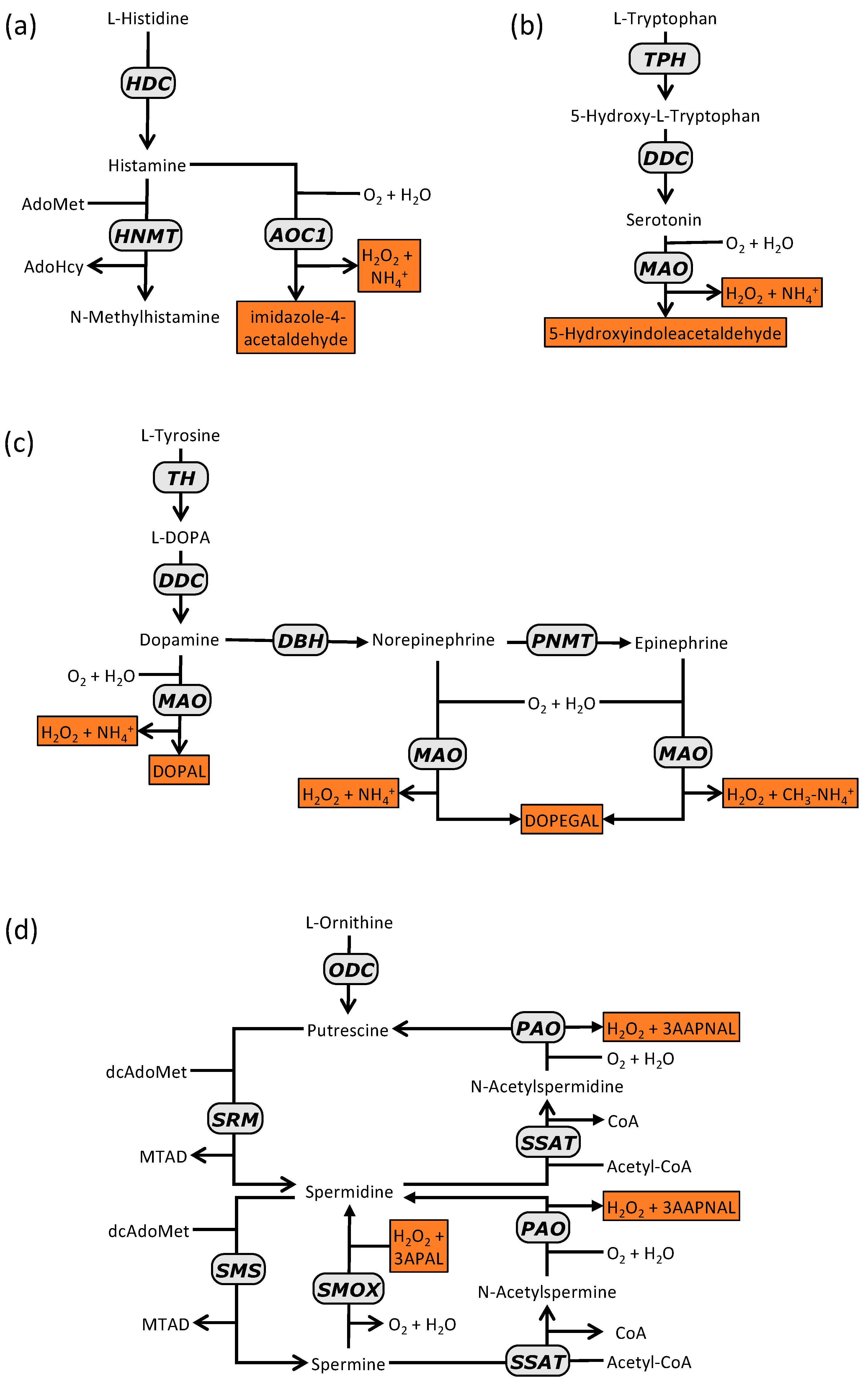
2.1. Histamine Synthesis
2.2. Exogenous Histamine Synthesis
2.3. Histamine Degradation
2.4. Histamine Transport and Storage Mechanisms
- Mast cell degranulation by immune stimuli. The presence of specific antigens induces IgE synthesis, inducing a high affinity binding between the specific IgE and IgE receptor known as FcεRI. This high affinity complex induces degranulation after further expositions to the antigen.
- Cytokines can also induce degranulation. It is mediated by vesicular trafficking events involving fusion and/or content interchange between secretory granules and vesicles driven to exocytosis.
- Constitutive HIS leakage due to non-active transport through cytosolic membranes or trans-Golgi vesicles driven to exocytosis.
2.5. Histamine Signalling and Physiological Functions
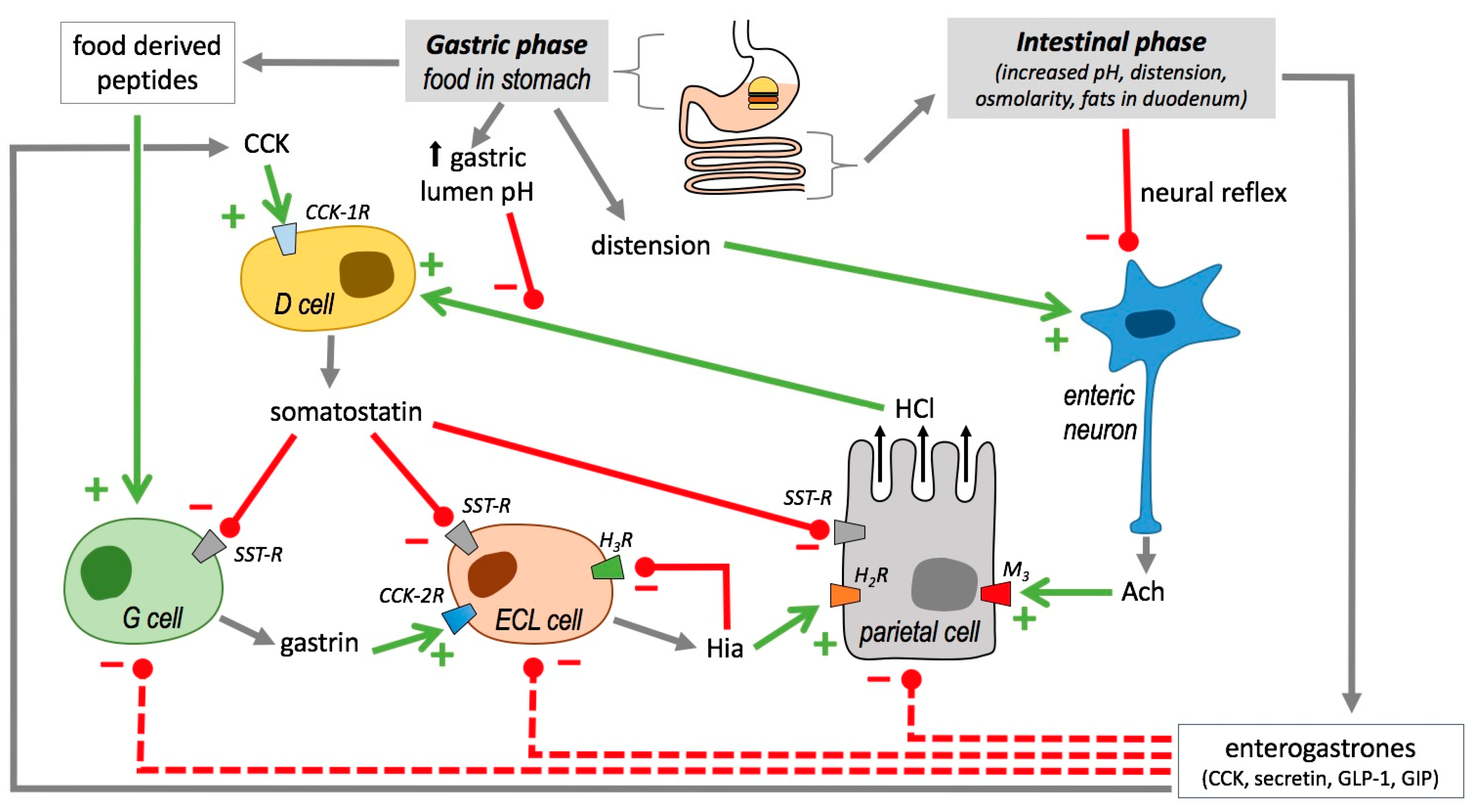
2.5.1. Histamine and Acid Gastric Secretion
2.5.2. Histamine and Immune Response in Gastrointestinal Tract
3. Serotonin Biochemistry and Physiology
3.1. Serotonin Synthesisn
3.2. Serotonin Degradation
3.3. Serotonin Transport and Storage Mechanism
3.4. Serotonin Signalling and Physiological Functions
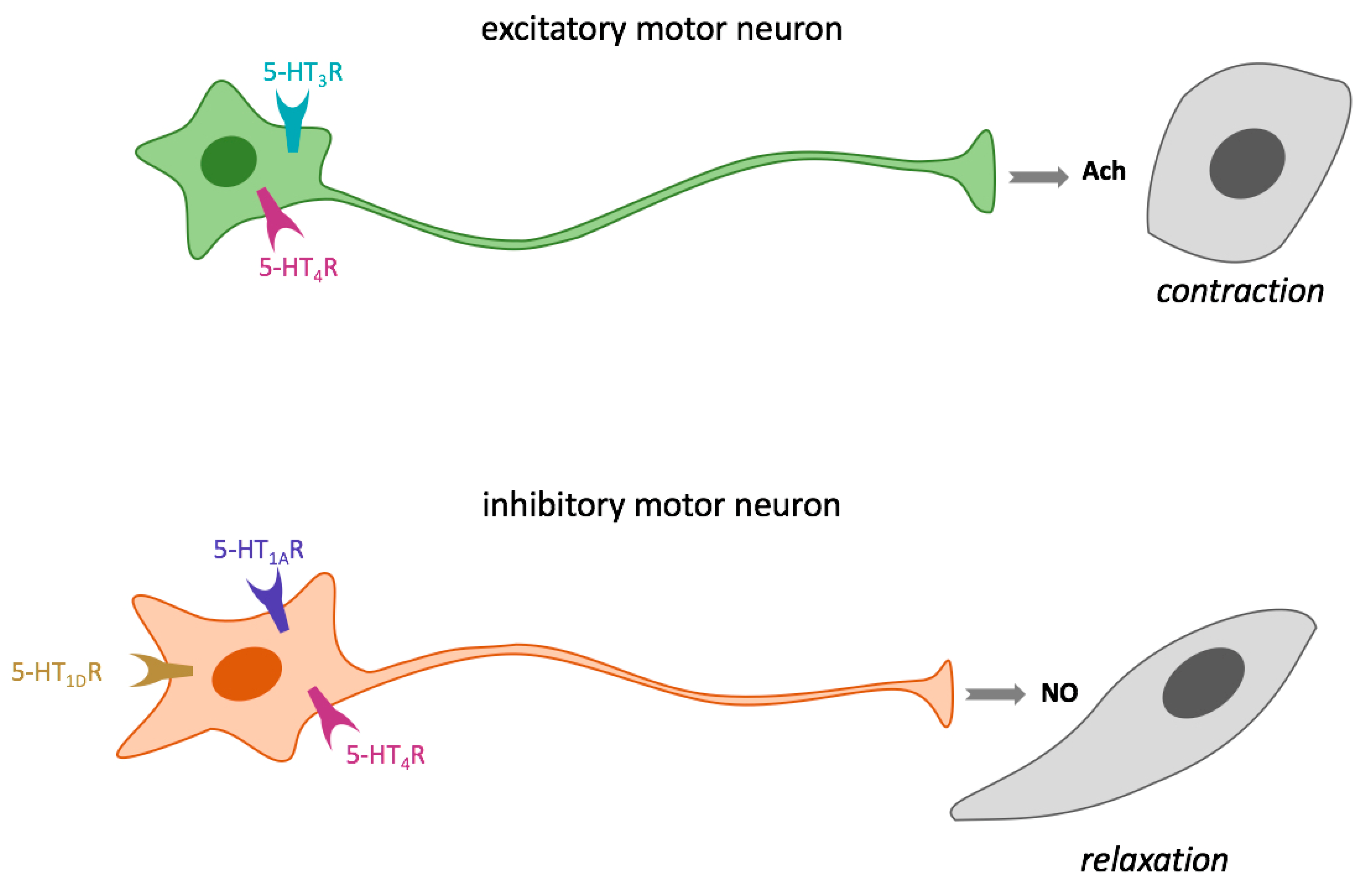
3.4.1. Regulation of GIT Smooth Muscle Contraction and Relaxation
3.4.2. Mucosal Sensory Transduction
3.4.3. Serotonin and Immune Response in GIT
4. Biochemistry and Physiology of Catecholamines
4.1. Synthesis of Catecholamines
4.2. Degradation of Catecholamines
4.3. Signalling and Physiological Functions of Catecholamines
Regulation of Intestinal Blood Flux, Immunity and Motility
- Nutrient absorption. Both epinephrine and norepinephrine play important roles in nutrient absorption regulation. Epinephrine is able to induce a hyperglycemic response acting through β-adrenergic receptors, and it increases absorption of oligopeptides when bound to α-adrenoceptors.
- Intestinal motility. CAs binding to β-adrenoreceptors induces smooth muscle relaxation that lead to a global food transit delay. On the contrary, their bindings to α-adrenoreceptors stimulate intestinal smooth muscle contraction, and consequently gut motility and food transit.
- CAs, immune system and GIT. Recently, CAs, as well as 5-HT, have been described as regulators of the innate immune system, which can be related to food intolerance. In addition, it is also reported that these amines can influence the intestinal microbiota [115].
5. Biochemistry and Physiology of Polyamines
5.1. Synthesis of Polyamines
5.2. Degradation and Recycling of Polyamines
5.3. Polyamine Transport Systems
5.4. Physiological Functions of Polyamines
5.5. The Particular Case of Agmatine
6. Biogenic Amines and Microbiota-Intestine Crosstalk
7. What Is Known about Biogenic Amines Roles in Human Gastro Intestinal Pathologies?
7.1. Gastric Diseases
7.1.1. Peptic Ulcers
7.1.2. Gastric Cancer
7.2. Intestinal Diseases
7.2.1. Irritable Bowel Syndrome
7.2.2. Inflammatory Bowel Diseases
7.2.3. Intestinal Neoplasias
8. Conclusions and Future Prospects
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sánchez-Jiménez, F.; Ruiz-Perez, M.V.; Urdiales, J.L.; Medina, M.A. Pharmacological potential of biogenic amine-polyamine interactions beyond neurotransmission. Br. J. Pharmacol. 2013, 170, 4–16. [Google Scholar] [CrossRef] [PubMed]
- Okada, K.; Hidese, R.; Fukuda, W.; Niitsu, M.; Takao, K.; Horai, Y.; Umezawa, N.; Higuchi, T.; Oshima, T.; Yoshikawa, Y.; et al. Identification of a novel aminopropyltransferase involved in the synthesis of branched-chain polyamines in hyperthermophiles. J. Bacteriol. 2014, 196, 1866–1876. [Google Scholar] [CrossRef] [PubMed]
- Suzzi, G.; Torriani, S. Editorial: Biogenic amines in foods. Front. Microbiol. 2015, 6, 472. [Google Scholar] [CrossRef] [PubMed]
- Bodmer, S.; Imark, C.; Kneubühl, M. Biogenic amines in foods: Histamine and food processing. Inflamm. Res. 1999, 48, 296–300. [Google Scholar] [CrossRef] [PubMed]
- Moya-Garcia, A.A.; Pino-Angeles, A.; Gil-Redondo, R.; Morreale, A.; Sanchez-Jimenez, F. Structural features of mammalian histidine decarboxylase reveal the basis for specific inhibition. Br. J. Pharmacol. 2009, 157, 4–13. [Google Scholar] [CrossRef] [PubMed]
- Giardina, G.; Montioli, R.; Gianni, S.; Cellini, B.; Paiardini, A.; Voltattorni, C.B.; Cutruzzolà, F. Open conformation of human DOPA decarboxylase reveals the mechanism of PLP addition to Group II decarboxylases. Proc. Natl. Acad. Sci. USA 2011, 108, 20514–20519. [Google Scholar] [CrossRef] [PubMed]
- Sandmeier, E.; Hale, T.I.; Christen, P. Multiple evolutionary origin of pyridoxal-5′-phosphate-dependent amino acid decarboxylases. Eur. J. Biochem. 1994, 221, 997–1002. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Jiménez, F.; Moya-García, A.A.; Pino-Ángeles, A. New structural insights to help in the search for selective inhibitors of mammalian pyridoxal 5′-phosphate-dependent histidine decarboxylase. Inflamm. Res. 2006, 55, S55–S56. [Google Scholar] [CrossRef] [PubMed]
- Fonda, M.L.; Eggers, D.K.; Mehta, R. Vitamin B-6 metabolism in the livers of young adult and senescent mice. Exp. Gerontol. 1980, 15, 457–463. [Google Scholar] [CrossRef]
- Jalkanen, S.; Salmi, M. Cell surface monoamine oxidases: Enzymes in search of a function. EMBO J. 2001, 20, 3893–3901. [Google Scholar] [CrossRef] [PubMed]
- Finney, J.; Moon, H.-J.; Ronnebaum, T.; Lantz, M.; Mure, M. Human copper-dependent amine oxidases. Arch. Biochem. Biophys. 2014, 546, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Edmondson, D.E.; Binda, C.; Mattevi, A. Structural insights into the mechanism of amine oxidation by monoamine oxidases A and B. Arch. Biochem. Biophys. 2007, 464, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-López, R.; Morales, M.; Sánchez-Jiménez, F. Histamine and its receptors as a module of the biogenic amine diseasome. In Histamine Receptors; Springer International Publishing: Cham, Switzerland, 2016; pp. 173–214. [Google Scholar]
- Schubert, M.L. Gastric acid secretion. Curr. Opin. Gastroenterol. 2016, 32, 452–460. [Google Scholar] [CrossRef] [PubMed]
- Barcik, W.; Wawrzyniak, M.; Akdis, C.A.; O’Mahony, L. Immune regulation by histamine and histamine-secreting bacteria. Curr. Opin. Immunol. 2017, 48, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Schneider, E.; Rolli-Derkinderen, M.; Arock, M.; Dy, M. Trends in histamine research: New functions during immune responses and hematopoiesis. Trends Immunol. 2002, 23, 255–263. [Google Scholar] [CrossRef]
- Peters, L.J.; Kovacic, J.P. Histamine: Metabolism, physiology, and pathophysiology with applications in veterinary medicine. J. Vet. Emerg. Crit. Care 2009, 19, 311–328. [Google Scholar] [CrossRef] [PubMed]
- Panula, P.; Chazot, P.L.; Cowart, M.; Gutzmer, R.; Leurs, R.; Liu, W.L.S.; Stark, H.; Thurmond, R.L.; Haas, H.L. International union of basic and clinical pharmacology. XCVIII. Histamine receptors. Pharmacol. Rev. 2015, 67, 601–655. [Google Scholar] [CrossRef] [PubMed]
- Gardini, F.; Özogul, Y.; Suzzi, G.; Tabanelli, G.; Özogul, F. Technological factors affecting biogenic amine content in foods: A review. Front. Microbiol. 2016, 7, 1218. [Google Scholar] [CrossRef] [PubMed]
- Piletz, J.E.; Aricioglu, F.; Cheng, J.-T.; Fairbanks, C.A.; Gilad, V.H.; Haenisch, B.; Halaris, A.; Hong, S.; Lee, J.E.; Li, J.; et al. Agmatine: Clinical applications after 100 years in translation. Drug Discov. Today 2013, 18, 880–893. [Google Scholar] [CrossRef] [PubMed]
- López-Contreras, A.J.; López-Garcia, C.; Jiménez-Cervantes, C.; Cremades, A.; Peñafiel, R. Mouse ornithine decarboxylase-like gene encodes an antizyme inhibitor devoid of ornithine and arginine decarboxylating activity. J. Biol. Chem. 2006, 281, 30896–30906. [Google Scholar] [CrossRef] [PubMed]
- Biji, K.B.; Ravishankar, C.N.; Venkateswarlu, R.; Mohan, C.O.; Gopal, T.K.S. Biogenic amines in seafood: A review. J. Food Sci. Technol. 2016, 53, 2210–2218. [Google Scholar] [CrossRef] [PubMed]
- Morgan, D.M.L.; White, A.; Sánchez-Jiménez, F.; Bardócz, S. COST 917—Biogenically Active Amines in Food. Volume IV, First General Workshop; Office for Official Publicationsn of European Communities: Luxemburg, 2000. [Google Scholar]
- Wallace, H.M.; Hughes, A. COST Action 922. Health Implications of Dietary Amines; Office for Official Publicationsn of European Communities: Luxemburg, 2004. [Google Scholar]
- Naila, A.; Flint, S.; Fletcher, G.; Bremer, P.; Meerdink, G. Control of biogenic amines in food-existing and emerging approaches. J. Food Sci. 2010, 75, R139–R150. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Jiménez, F.; Pino-Ángeles, A.; Rodríguez-López, R.; Morales, M.; Urdiales, J.L. Structural and functional analogies and differences between histidine decarboxylase and aromatic l-amino acid decarboxylase molecular networks: Biomedical implications. Pharmacol. Res. 2016, 114, 90–102. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-López, R.; Reyes-Palomares, A.; Sánchez-Jiménez, F.; Medina, M.Á. PhenUMA: A tool for integrating the biomedical relationships among genes and diseases. BMC Bioinform. 2014, 15, 375. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Palomares, A.; Bueno, A.; Rodríguez-López, R.; Medina, M.Á.; Sánchez-Jiménez, F.; Corpas, M.; Ranea, J.A.G. Systematic identification of phenotypically enriched loci using a patient network of genomic disorders. BMC Genom. 2016, 17, 232. [Google Scholar] [CrossRef] [PubMed]
- Pino-Ángeles, A.; Reyes-Palomares, A.; Melgarejo, E.; Sánchez-Jiménez, F. Histamine: An undercover agent in multiple rare diseases? J. Cell. Mol. Med. 2012, 16, 1947–1960. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Pérez, M.V.; Medina, M.Á.; Urdiales, J.L.; Keinänen, T.A.; Tuomo, A.; Sánchez-Jiménez, F. Polyamine metabolism is sensitive to glycolysis inhibition in human neuroblastoma cells. J. Biol. Chem. 2015, 290, 6106–6119. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Ohtsu, H. l-histidine decarboxylase as a probe in studies on histamine. Chem. Rec. 2002, 2, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Stark, H. Histamine H4 Receptor: A Novel Drug Target for Immunoregulation and Inflammation; Versita: Berlin, Germany, 2013. [Google Scholar]
- Nakanishi, T.; Kekuda, R.; Fei, Y.J.; Hatanaka, T.; Sugawara, M.; Martindale, R.G.; Leibach, F.H.; Prasad, P.D.; Ganapathy, V. Cloning and functional characterization of a new subtype of the amino acid transport system N. Am. J. Physiol. Cell Physiol. 2001, 281, C1757–C1768. [Google Scholar] [CrossRef] [PubMed]
- Moya-Garcia, A.A.; Medina, M.A.; Sánchez-Jiménez, F. Mammalian histidine decarboxylase: From structure to function. Bioessays 2005, 27, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Schwelberger, H.G.; Ahrens, F.; Fogel, W.A.; Sánchez-Jiménez, F. Histamine metabolism. In Histamine H4 Receptor: A Novel Drug Target in Immunoregulation and Inflammation; Stark, H., Ed.; Versita: Berlin, Germany, 2013; pp. 63–102. [Google Scholar]
- Metcalfe, D.D. Mast cells and mastocytosis. Blood 2008, 112, 946–956. [Google Scholar] [CrossRef] [PubMed]
- Ennis, M.; Ciz, M.; Dib, K.; Friedman, S.; Gangwar, R.S.; Gibbs, B.F.; Levi-Schaffer, F.; Lojek, A.; Migalovich-Sheikhet, H.; O’Mahony, L.; et al. Histamine receptors and inflammatory cells. In Histamine H4 Receptor: A Novel Drug Target in Immunoregulation and Inflammation; Stark, H., Ed.; Versita: London, UK, 2013; pp. 103–144. [Google Scholar]
- Dwyer, D.F.; Barrett, N.A.; Austen, K.F.; Immunological Genome Project Consortium. Expression profiling of constitutive mast cells reveals a unique identity within the immune system. Nat. Immunol. 2016, 17, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, R.; Furini, C.R.G.; Passani, M.B.; Provensi, G.; Baldi, E.; Bucherelli, C.; Izquierdo, I.; de Carvalho Myskiw, J.; Blandina, P. Memory retrieval of inhibitory avoidance requires histamine H1 receptor activation in the hippocampus. Proc. Natl. Acad. Sci. USA 2016, 113, E2714–E2720. [Google Scholar] [CrossRef] [PubMed]
- Bernsand, M.; Ericsson, P.; Bjorkqvist, M.; Zhao, C.-M.; Hakanson, R.; Norlen, P. Submucosal microinfusion of endothelin and adrenaline mobilizes ECL-cell histamine in rat stomach, and causes mucosal damage: A microdialysis study. Br. J. Pharmacol. 2003, 140, 707–717. [Google Scholar] [CrossRef] [PubMed]
- Krauth, M.-T.T.; Agis, H.; Aichberger, K.J.; Simonitsch-Klupp, I.; Müllauer, L.; Mayerhofer, M.; Böhm, A.; Horny, H.-P.P.; Valent, P. Immunohistochemical detection of histidine decarboxylase in neoplastic mast cells in patients with systemic mastocytosis. Hum. Pathol. 2006, 37, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Osefo, N.; Ito, T.; Jensen, R.T. Gastric acid hypersecretory states: Recent insights and advances. Curr. Gastroenterol. Rep. 2009, 11, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Kuramasu, A.; Saito, H.; Suzuki, S.; Watanabe, T.; Ohtsu, H. Mast cell-/basophil-specific transcriptional regulation of human l-histidine decarboxylase gene by CpG methylation in the promoter region. J. Biol. Chem. 1998, 273, 31607–31614. [Google Scholar] [CrossRef] [PubMed]
- Correa-Fiz, F.; Reyes-Palomares, A.; Fajardo, I.; Melgarejo, E.; Gutiérrez, A.; García-Ranea, J.A.; Medina, M.A.; Sánchez-Jiménez, F. Regulatory cross-talk of mouse liver polyamine and methionine metabolic pathways: A systemic approach to its physiopathological consequences. Amino Acids 2011, 42, 577–595. [Google Scholar] [CrossRef] [PubMed]
- Smolinska, S.; Jutel, M.; Crameri, R.; O’Mahony, L. Histamine and gut mucosal immune regulation. Allergy 2014, 69, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Mamune-Sato, R.; Yamauchi, K.; Tanno, Y.; Ohkawara, Y.; Ohtsu, H.; Katayose, D.; Maeyama, K.; Watanabe, T.; Shibahara, S.; Takishima, T. Functional analysis of alternatively spliced transcripts of the human histidine decarboxylase gene and its expression in human tissues and basophilic leukemia cells. Eur. J. Biochem. 1992, 209, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Abrighach, H.; Fajardo, I.; Sánchez-Jiménez, F.; Urdiales, J.L. Exploring polyamine regulation by nascent histamine in a human-transfected cell model. Amino Acids 2010, 38, 561–573. [Google Scholar] [CrossRef] [PubMed]
- Olmo, M.T.; Urdiales, J.L.; Pegg, A.E.; Medina, M.A.; Sánchez-Jiménez, F. In vitro study of proteolytic degradation of rat histidine decarboxylase. Eur. J. Biochem. 2000, 267, 1527–1531. [Google Scholar] [CrossRef] [PubMed]
- Fleming, J.V.; Fajardo, I.; Langlois, M.R.; Sanchez-Jimenez, F.; Wang, T.C. The C-terminus of rat l-histidine decarboxylase specifically inhibits enzymic activity and disrupts pyridoxal phosphate-dependent interactions with l-histidine substrate analogues. Biochem. J. 2004, 381, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Furuta, K.; Nakayama, K.; Sugimoto, Y.; Ichikawa, A.; Tanaka, S. Activation of histidine decarboxylase through post-translational cleavage by caspase-9 in a mouse mastocytoma P-815. J. Biol. Chem. 2007, 282, 13438–13446. [Google Scholar] [CrossRef] [PubMed]
- Olmo, M.T.; Rodríguez-Agudo, D.; Medina, M.A.; Sánchez-Jiménez, F. The pest regions containing C-termini of mammalian ornithine decarboxylase and histidine decarboxylase play different roles in protein degradation. Biochem. Biophys. Res. Commun. 1999, 257, 269–272. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Agudo, D.; Olmo, M.T.; Sanchez-Jimenez, F.; Medina, M.A. Rat histidine decarboxylase is a substrate for m-calpain in vitro. Biochem. Biophys. Res. Commun. 2000, 271, 777–781. [Google Scholar] [CrossRef] [PubMed]
- Pino-Angeles, A.; Morreale, A.; Negri, A.; Sánchez-Jiménez, F.; Moya-García, A.A. Substrate uptake and protein stability relationship in mammalian histidine decarboxylase. Proteins 2010, 78, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Olmo, M.T.; Sanchez-Jimenez, F.; Medina, M.A.; Hayashi, H. Spectroscopic analysis of recombinant rat histidine decarboxylase. J. Biochem. 2002, 132, 433–439. [Google Scholar] [CrossRef] [PubMed]
- Moya-García, A.A.; Ruiz-Pernía, J.; Martí, S.; Sánchez-Jiménez, F.; Tuñón, I. Analysis of the decarboxylation step in mammalian histidine decarboxylase. A computational study. J. Biol. Chem. 2008, 283, 12393–12401. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Caso, C.; Rodríguez-Agudo, D.; Moya-García, A.A.; Fajardo, I.; Medina, M.A.; Subramaniam, V.; Sánchez-Jiménez, F. Local changes in the catalytic site of mammalian histidine decarboxylase can affect its global conformation and stability. Eur. J. Biochem. 2003, 270, 4376–4387. [Google Scholar] [CrossRef] [PubMed]
- Fleming, J.V.; Sánchez-Jiménez, F.; Moya-García, A.A.; Langlois, M.R.; Wang, T.C. Mapping of catalytically important residues in the rat l-histidine decarboxylase enzyme using bioinformatic and site-directed mutagenesis approaches. Biochem. J. 2004, 379, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Morgan, D.M.L.; Milovic, V.; Krizek, M.; White, A. COST Action 917. Biogenically Active Amines in Food. Volume V. Polyamines and Tumor Growth, Biologically Active Amines in Food Processing and Amines Produced by Bacteria; European Commission: Luxemburg, 2001. [Google Scholar]
- Wallace, H.M. Health implications of dietary amines: An overview of COST Action 922 (2001–2006). Biochem. Soc. Trans. 2007, 35, 293–294. [Google Scholar] [CrossRef] [PubMed]
- Schwelberger, H.G. Histamine N-methyltransferase (HNMT) enzyme and gene. In Histamine: Biology and Medical Aspects; Falus, A., Grosman, N., Darvas, Z., Eds.; SpringMed Publishing: Budapest, Hungary, 2004; pp. 53–59. [Google Scholar]
- Fagerberg, L.; Hallström, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Duff, M.O.; Olson, S.; Wei, X.; Garrett, S.C.; Osman, A.; Bolisetty, M.; Plocik, A.; Celniker, S.E.; Graveley, B.R. Genome-wide identification of zero nucleotide recursive splicing in Drosophila. Nature 2015, 521, 376–379. [Google Scholar] [CrossRef] [PubMed]
- Prell, G.D.; Green, J.P. Measurement of histamine metabolites in brain and cerebrospinal fluid provides insights into histaminergic activity. Agents Actions 1994, 41, C5–C8. [Google Scholar] [CrossRef] [PubMed]
- Song, W.-B.; Lv, Y.-H.; Zhang, Z.-S.; Li, Y.-N.; Xiao, L.-P.; Yu, X.-P.; Wang, Y.-Y.; Ji, H.-L.; Ma, L. Soluble intercellular adhesion molecule-1, D-lactate and diamine oxidase in patients with inflammatory bowel disease. World J. Gastroenterol. 2009, 15, 3916–3919. [Google Scholar] [CrossRef] [PubMed]
- Koepsell, H.; Lips, K.; Volk, C. Polyspecific organic cation transporters: Structure, function, physiological roles, and biopharmaceutical implications. Pharm. Res. 2007, 24, 1227–1251. [Google Scholar] [CrossRef] [PubMed]
- Eiden, L.E.; Weihe, E. VMAT2: A dynamic regulator of brain monoaminergic neuronal function interacting with drugs of abuse. Ann. N. Y. Acad. Sci. 2011, 1216, 86–98. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Faroldi, G.; Rodriguez, C.E.; Urdiales, J.L.; Perez-Pomares, J.M.; Davila, J.C.; Pejler, G.; Sanchez-Jimenez, F.; Fajardo, I. Polyamines are present in mast cell secretory granules and are important for granule homeostasis. PLoS ONE 2010, 5, e15071. [Google Scholar] [CrossRef] [PubMed]
- Andersson, K.; Chen, D.; Håkanson, R.; Mattsson, H.; Sundler, F. Enterochromaffin-like cells in the rat stomach: Effect of alpha-fluoromethylhistidine-evoked histamine depletion. A chemical, histochemical and electron-microscopic study. Cell Tissue Res. 1992, 270, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Thurmond, R.L. The histamine H4 receptor: From orphan to the clinic. Front. Pharmacol. 2015, 6, 65. [Google Scholar] [CrossRef] [PubMed]
- Kusters, J.G.; van Vliet, A.H.M.; Kuipers, E.J. Pathogenesis of Helicobacter pylori infection. Clin. Microbiol. Rev. 2006, 19, 449–490. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Aihara, T.; Zhao, C.-M.; Håkanson, R.; Okabe, S. Differentiation of the Gastric Mucosa I. Role of histamine in control of function and integrity of oxyntic mucosa: Understanding gastric physiology through disruption of targeted genes. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 291, G539–G544. [Google Scholar] [CrossRef] [PubMed]
- Waldum, H.L.; Hauso, Ø.; Fossmark, R. The regulation of gastric acid secretion—Clinical perspectives. Acta Physiol. 2014, 210, 239–256. [Google Scholar] [CrossRef] [PubMed]
- Coruzzi, G.; Adami, M.; Pozzoli, C.; de Esch, I.J.P.; Smits, R.; Leurs, R. Selective histamine H3 and H4 receptor agonists exert opposite effects against the gastric lesions induced by HCl in the rat stomach. Eur. J. Pharmacol. 2011, 669, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Rydning, A.; Lyng, O.; Falkmer, S.; Grønbech, J.E. Histamine is involved in gastric vasodilation during acid back diffusion via activation of sensory neurons. Am. J. Physiol. Gastrointest. Liver Physiol. 2002, 283, G603–G611. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, J.; Misra, M.; Rizvi, W.; Kumar, A. Histamine aspects in acid peptic diseases and cell proliferation. In Biomedical Aspects of Histamine. Current Perspectives; Shahid, M., Khardori, N., Khan, R.A., Tripathi, T., Eds.; Springer: Dordrecht, The Netherlands, 2010; pp. 175–198. [Google Scholar]
- Sander, L.E.; Lorentz, A.; Sellge, G.; Coëffier, M.; Neipp, M.; Veres, T.; Frieling, T.; Meier, P.N.; Manns, M.P.; Bischoff, S.C. Selective expression of histamine receptors H1R, H2R, and H4R, but not H3R, in the human intestinal tract. Gut 2006, 55, 498–504. [Google Scholar] [CrossRef] [PubMed]
- Schneider, E.; Leite-de-moraes, M.; Dy, M. Histamine, immune cells and autoimmunity. Adv. Exp. Med. Biol. 2010, 709, 81–94. [Google Scholar] [PubMed]
- Xie, H.; He, S.-H. Roles of histamine and its receptors in allergic and inflammatory bowel diseases. World J. Gastroenterol. 2005, 11, 2851–2857. [Google Scholar] [CrossRef] [PubMed]
- Gutzmer, R.; Diestel, C.; Mommert, S.; Köther, B.; Stark, H.; Wittmann, M.; Werfel, T. Histamine H4 receptor stimulation suppresses IL-12p70 production and mediates chemotaxis in human monocyte-derived dendritic cells. J. Immunol. 2005, 174, 5224–5232. [Google Scholar] [CrossRef] [PubMed]
- Gutzmer, R.; Mommert, S.; Gschwandtner, M.; Zwingmann, K.; Stark, H.; Werfel, T. The histamine H4 receptor is functionally expressed on T(H)2 cells. J. Allergy Clin. Immunol. 2009, 123, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Coruzzi, G.; Adami, M.; Pozzoli, C. Role of histamine H4 receptors in the gastrointestinal tract. Front. Biosci. 2012, 4, 226–239. [Google Scholar] [CrossRef]
- Mittal, R.; Debs, L.H.; Patel, A.P.; Nguyen, D.; Patel, K.; O’Connor, G.; Grati, M.; Mittal, J.; Yan, D.; Eshraghi, A.A.; et al. Neurotransmitters: The critical modulators regulating gut-brain axis. J. Cell. Physiol. 2017, 232, 2359–2372. [Google Scholar] [CrossRef] [PubMed]
- Sikander, A.; Rana, S.V.; Prasad, K.K. Role of serotonin in gastrointestinal motility and irritable bowel syndrome. Clin. Chim. Acta 2009, 403, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Roberts, K.M.; Fitzpatrick, P.F. Mechanisms of tryptophan and tyrosine hydroxylase. IUBMB Life 2013, 65, 350–357. [Google Scholar] [CrossRef] [PubMed]
- O’Mahony, S.M.; Clarke, G.; Borre, Y.E.; Dinan, T.G.; Cryan, J.F. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav. Brain Res. 2015, 277, 32–48. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, H.; Tsukiyama, F.; Ishii, S.; Mizuguchi, H.; Kagamiyama, H. Acid-base chemistry of the reaction of aromatic l-amino acid decarboxylase and dopa analyzed by transient and steady-state kinetics: Preferential binding of the substrate with its amino group unprotonated. Biochemistry 1999, 38, 15615–15622. [Google Scholar] [CrossRef] [PubMed]
- Bertoldi, M.; Voltattorni, C.B. Dopa decarboxylase exhibits low pH half-transaminase and high pH oxidative deaminase activities toward serotonin (5-hydroxytryptamine). Protein Sci. 2001, 10, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Bertoldi, M.; Gonsalvi, M.; Contestabile, R.; Voltattorni, C.B. Mutation of tyrosine 332 to phenylalanine converts dopa decarboxylase into a decarboxylation-dependent oxidative deaminase. J. Biol. Chem. 2002, 277, 36357–36362. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Pérez, M.V.; Pino-Ángeles, A.; Medina, M.A.; Sánchez-Jiménez, F.; Moya-García, A.A. Structural perspective on the direct inhibition mechanism of EGCG on mammalian histidine decarboxylase and DOPA decarboxylase. J. Chem. Inf. Model. 2012, 52, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Montioli, R.; Cellini, B.; Dindo, M.; Oppici, E.; Voltattorni, C.B. Interaction of human Dopa decarboxylase with l-Dopa: Spectroscopic and kinetic studies as a function of pH. BioMed Res. Int. 2013, 2013, 161456. [Google Scholar] [CrossRef] [PubMed]
- Bortolato, M.; Chen, K.; Shih, J.C. The degradation of serotonin: Role of MAO. Handb. Behav. Neurosci. 2010, 21, 203–218. [Google Scholar] [CrossRef]
- Meiser, J.; Weindl, D.; Hiller, K. Complexity of dopamine metabolism. Cell Commun. Signal. 2013, 11, 34. [Google Scholar] [CrossRef] [PubMed]
- Nagatsu, T. Progress in monoamine oxidase (MAO) research in relation to genetic engineering. Neurotoxicology 2004, 25, 11–20. [Google Scholar] [CrossRef]
- Strege, P.R.; Knutson, K.; Eggers, S.J.; Li, J.H.; Wang, F.; Linden, D.; Szurszewski, J.H.; Milescu, L.; Leiter, A.B.; Farrugia, G.; et al. Sodium channel NaV1.3 is important for enterochromaffin cell excitability and serotonin release. Sci. Rep. 2017, 7, 15650. [Google Scholar] [CrossRef] [PubMed]
- Mawe, G.M.; Hoffman, J.M. Serotonin signalling in the gut—Functions, dysfunctions and therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 473–486. [Google Scholar] [CrossRef] [PubMed]
- Gershon, M.D. 5-HT (serotonin) physiology and related drugs. Curr. Opin. Gastroenterol. 2000, 16, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Greenwood-van Meerveld, B. Importance of 5-hydroxytryptamine receptors on intestinal afferents in the regulation of visceral sensitivity. Neurogastroenterol. Motil. 2007, 19, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Versteeg, R.I.; Serlie, M.J.; Kalsbeek, A.; la Fleur, S.E. Serotonin, a possible intermediate between disturbed circadian rhythms and metabolic disease. Neuroscience 2015, 301, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Shortall, S.E.; Spicer, C.H.; Ebling, F.J.P.; Green, A.R.; Fone, K.C.F.; King, M.V. Contribution of serotonin and dopamine to changes in core body temperature and locomotor activity in rats following repeated administration of mephedrone. Addict. Biol. 2016, 21, 1127–1139. [Google Scholar] [CrossRef] [PubMed]
- Arreola, R.; Becerril-Villanueva, E.; Cruz-Fuentes, C.; Velasco-Velázquez, M.A.; Garcés-Alvarez, M.E.; Hurtado-Alvarado, G.; Quintero-Fabian, S.; Pavón, L. Immunomodulatory Effects Mediated by Serotonin. J. Immunol. Res. 2015, 2015, 354957. [Google Scholar] [CrossRef] [PubMed]
- Maggiorani, D.; Manzella, N.; Edmondson, D.E.; Mattevi, A.; Parini, A.; Binda, C.; Mialet-Perez, J. Monoamine oxidases, oxidative stress, and altered mitochondrial dynamics in cardiac ageing. Oxid. Med. Cell. Longev. 2017, 2017, 3017947. [Google Scholar] [CrossRef] [PubMed]
- Margolis, K.G.; Gershon, M.D. Enteric neuronal regulation of intestinal inflammation. Trends Neurosci. 2016, 39, 614–624. [Google Scholar] [CrossRef] [PubMed]
- Filip, M.; Bader, M. Overview on 5-HT receptors and their role in physiology and pathology of the central nervous system. Pharmacol. Rep. 2009, 61, 761–777. [Google Scholar] [CrossRef]
- Li, Z.; Chalazonitis, A.; Huang, Y.-Y.; Mann, J.J.; Margolis, K.G.; Yang, Q.M.; Kim, D.O.; Côté, F.; Mallet, J.; Gershon, M.D. Essential roles of enteric neuronal serotonin in gastrointestinal motility and the development/survival of enteric dopaminergic neurons. J. Neurosci. 2011, 31, 8998–9009. [Google Scholar] [CrossRef] [PubMed]
- Soyer, T.; Aktuna, Z.; Reşat Aydos, T.; Osmanoğlu, G.; Korkut, O.; Akman, H.; Cakmak, M. Esophageal and gastric smooth muscle activity after carbon dioxide pneumoperitoneum. J. Surg. Res. 2010, 161, 278–281. [Google Scholar] [CrossRef] [PubMed]
- Flatmark, T. Catecholamine biosynthesis and physiological regulation in neuroendocrine cells. Acta Physiol. Scand. 2000, 168, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; McLeish, M.J. Kinetic and pH studies on human phenylethanolamine N-methyltransferase. Arch. Biochem. Biophys. 2013, 539, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Eisenhofer, G.; Kopin, I.J.; Goldstein, D.S. Catecholamine metabolism: A contemporary view with implications for physiology and medicine. Pharmacol. Rev. 2004, 56, 331–349. [Google Scholar] [CrossRef] [PubMed]
- Daubner, S.C.; Le, T.; Wang, S. Tyrosine hydroxylase and regulation of dopamine synthesis. Arch. Biochem. Biophys. 2011, 508, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Tank, A.W.; Lee Wong, D. Peripheral and Central Effects of Circulating Catecholamines. Compr. Physiol. 2014, 5, 1–15. [Google Scholar] [CrossRef]
- Kobayashi, K. Role of catecholamine signaling in brain and nervous system functions: New insights from mouse molecular genetic study. J. Investig. Dermatol. Symp. Proc. 2001, 6, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Arreola, R.; Alvarez-Herrera, S.; Pérez-Sánchez, G.; Becerril-Villanueva, E.; Cruz-Fuentes, C.; Flores-Gutierrez, E.O.; Garcés-Alvarez, M.E.; de la Cruz-Aguilera, D.L.; Medina-Rivero, E.; Hurtado-Alvarado, G.; et al. Immunomodulatory effects mediated by dopamine. J. Immunol. Res. 2016, 2016, 3160486. [Google Scholar] [CrossRef] [PubMed]
- Beaulieu, J.-M.; Espinoza, S.; Gainetdinov, R.R. Dopamine receptors—IUPHAR review 13. Br. J. Pharmacol. 2015, 172, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Elenkov, I.J.; Wilder, R.L.; Chrousos, G.P.; Vizi, E.S. The sympathetic nerve—An integrative interface between two supersystems: The brain and the immune system. Pharmacol. Rev. 2000, 52, 595–638. [Google Scholar] [PubMed]
- Rizzetto, L.; Fava, F.; Tuohy, K.M.; Selmi, C. Connecting the immune system, systemic chronic inflammation and the gut microbiome: The role of sex. J. Autoimmun. 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- Natale, G.; Ryskalin, L.; Busceti, C.L.; Biagioni, F.; Fornai, F. The nature of catecholamine-containing neurons in the enteric nervous system in relationship with organogenesis, normal human anatomy and neurodegeneration. Arch. Ital. Biol. 2017, 155, 118–130. [Google Scholar] [PubMed]
- Rodriguez-Caso, C.; Montañez, R.; Cascante, M.; Sanchez-Jimenez, F.; Medina, M.A. Mathematical modeling of polyamine metabolism in mammals. J. Biol. Chem. 2006, 281, 21799–21812. [Google Scholar] [CrossRef] [PubMed]
- Pegg, A.E. Mammalian polyamine metabolism and function. IUBMB Life 2009, 61, 880–894. [Google Scholar] [CrossRef] [PubMed]
- Pegg, A.E. Functions of Polyamines in Mammals. J. Biol. Chem. 2016, 291, 14904–14912. [Google Scholar] [CrossRef] [PubMed]
- Perez-Leal, O.; Merali, S. Regulation of polyamine metabolism by translational control. Amino Acids 2012, 42, 611–617. [Google Scholar] [CrossRef] [PubMed]
- Kahana, C. Protein degradation, the main hub in the regulation of cellular polyamines. Biochem. J. 2016, 473, 4551–4558. [Google Scholar] [CrossRef] [PubMed]
- Pegg, A.E. Regulation of ornithine decarboxylase. J. Biol. Chem. 2006, 281, 14529–14532. [Google Scholar] [CrossRef] [PubMed]
- Pegg, A.E. S-Adenosylmethionine decarboxylase. Essays Biochem. 2009, 46, 25–45. [Google Scholar] [CrossRef] [PubMed]
- Pegg, A.E.; Casero, R.A. Current status of the polyamine research field. In Polyamines. Methods and Protocols; Pegg, A.E., Casero, R.A., Eds.; Humana Press: Newyork, NK, USA, 2011; pp. 3–35. [Google Scholar]
- Casero, R.A.; Marton, L.J. Targeting polyamine metabolism and function in cancer and other hyperproliferative diseases. Nat. Rev. Drug Discov. 2007, 6, 373–390. [Google Scholar] [CrossRef] [PubMed]
- Casero, R.A.; Pegg, A.E. Polyamine catabolism and disease. Biochem. J. 2009, 421, 323–338. [Google Scholar] [CrossRef] [PubMed]
- Cecco, L.; Antoniello, S.; Auletta, M.; Cerra, M.; Bonelli, P. Pattern and concentration of free and acetylated polyamines in urine of cirrhotic patients. Int. J. Biol. Mark. 1992, 7, 52–58. [Google Scholar]
- Qian, H.; Beard, D.A. Metabolic futile cycles and their functions: A systems analysis of energy and control. Syst. Biol. 2006, 153, 192–200. [Google Scholar] [CrossRef]
- Reyes-Palomares, A.; Montañez, R.; Sánchez-Jiménez, F.; Medina, M.A.; Sanchez-Jimenez, F.; Medina, M.A. A combined model of hepatic polyamine and sulfur amino acid metabolism to analyze S-adenosyl methionine availability. Amino Acids 2012, 42, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Muth, A.; Madan, M.; Archer, J.J.; Ocampo, N.; Rodriguez, L.; Phanstiel, O. Polyamine transport inhibitors: Design, synthesis, and combination therapies with difluoromethylornithine. J. Med. Chem. 2014, 57, 348–363. [Google Scholar] [CrossRef] [PubMed]
- Uimari, A.; Keinänen, T.A.; Karppinen, A.; Woster, P.; Uimari, P.; Jänne, J.; Alhonen, L. Spermine analogue-regulated expression of spermidine/spermine N1-acetyltransferase and its effects on depletion of intracellular polyamine pools in mouse fetal fibroblasts. Biochem. J. 2009, 422, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Soulet, D.; Gagnon, B.; Rivest, S.; Audette, M.; Poulin, R. A fluorescent probe of polyamine transport accumulates into intracellular acidic vesicles via a two-step mechanism. J. Biol. Chem. 2004, 279, 49355–49366. [Google Scholar] [CrossRef] [PubMed]
- Poulin, R.; Casero, R.A.; Soulet, D. Recent advances in the molecular biology of metazoan polyamine transport. Amino Acids 2011, 42, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Belting, M.; Mani, K.; Jönsson, M.; Cheng, F.; Sandgren, S.; Jonsson, S.; Ding, K.; Delcros, J.-G.; Fransson, L.-A. Glypican-1 is a vehicle for polyamine uptake in mammalian cells: A pivital role for nitrosothiol-derived nitric oxide. J. Biol. Chem. 2003, 278, 47181–47189. [Google Scholar] [CrossRef] [PubMed]
- Abdulhussein, A.A.; Wallace, H.M. Polyamines and membrane transporters. Amino Acids 2014, 46, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Uemura, T.; Yerushalmi, H.F.; Tsaprailis, G.; Stringer, D.E.; Pastorian, K.E.; Hawel, L.; Byus, C.V.; Gerner, E.W. Identification and characterization of a diamine exporter in colon epithelial cells. J. Biol. Chem. 2008, 283, 26428–26435. [Google Scholar] [CrossRef] [PubMed]
- Uemura, T.; Gerner, E.W. Polyamine transport systems in mammalian cells and tissues. Methods Mol. Biol. 2011, 720, 339–348. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Chica, J.; Medina, M.A.; Sánchez-Jiménez, F.; Ramírez, F.J. Fourier transform Raman study of the structural specificities on the interaction between DNA and biogenic polyamines. Biophys. J. 2001, 80, 443–454. [Google Scholar] [CrossRef]
- Finger, S.; Schwieger, C.; Arouri, A.; Kerth, A.; Blume, A. Interaction of linear polyamines with negatively charged phospholipids: The effect of polyamine charge distance. Biol. Chem. 2014, 395, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Lightfoot, H.L.; Hall, J. Endogenous polyamine function—The RNA perspective. Nucleic Acids Res. 2014, 42, 11275–11290. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.J.; Tajmir-Riahi, H.A.; Thomas, T. Polyamine-DNA interactions and development of gene delivery vehicles. Amino Acids 2016, 48, 2423–2431. [Google Scholar] [CrossRef] [PubMed]
- Acosta-Andrade, C.; Artetxe, I.; Lete, M.G.; Monasterio, B.G.; Ruiz-Mirazo, K.; Goñi, F.M.; Sánchez-Jiménez, F. Polyamine-RNA-membrane interactions: From the past to the future in biology. Colloids Surf. B Biointerfaces 2017, 155, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, K.; Kashiwagi, K. Modulation of protein synthesis by polyamines. IUBMB Life 2015, 67, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Ramani, D.; De Bandt, J.P.; Cynober, L. Aliphatic polyamines in physiology and diseases. Clin. Nutr. 2014, 33, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Miller-Fleming, L.; Olin-Sandoval, V.; Campbell, K.; Ralser, M. Remaining mysteries of molecular biology: The role of polyamines in the cell. J. Mol. Biol. 2015, 427, 3389–3406. [Google Scholar] [CrossRef] [PubMed]
- Murphy, G.M. Polyamines in the human gut. Eur. J. Gastroenterol. Hepatol. 2001, 13, 1011–1014. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Q.; Ray, R.M.; Viar, M.J.; Johnson, L.R. Polyamine regulation of ornithine decarboxylase and its antizyme in intestinal epithelial cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2001, 280, G130–G138. [Google Scholar] [CrossRef] [PubMed]
- Seiler, N.; Raul, F. Polyamines and the intestinal tract. Crit. Rev. Clin. Lab. Sci. 2007, 44, 365–411. [Google Scholar] [CrossRef] [PubMed]
- Deloyer, P.; Peulen, O.; Dandrifosse, G. Dietary polyamines and non-neoplastic growth and disease. Eur. J. Gastroenterol. Hepatol. 2001, 13, 1027–1032. [Google Scholar] [CrossRef] [PubMed]
- Arndt, M.A.; Battaglia, V.; Parisi, E.; Lortie, M.J.; Isome, M.; Baskerville, C.; Pizzo, D.P.; Ientile, R.; Colombatto, S.; Toninello, A.; et al. The arginine metabolite agmatine protects mitochondrial function and confers resistance to cellular apoptosis. Am. J. Physiol. Cell Physiol. 2009, 296, C1411–C1419. [Google Scholar] [CrossRef] [PubMed]
- Satriano, J.; Isome, M.; Casero, R.A.; Thomson, S.C.; Blantz, R.C. Polyamine transport system mediates agmatine transport in mammalian cells. Am. J. Physiol. Cell Physiol. 2001, 281, C329–C334. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.C.; Reis, D.J. Agmatine selectively blocks the N-methyl-d-aspartate subclass of glutamate receptor channels in rat hippocampal neurons. J. Pharmacol. Exp. Ther. 1999, 288, 544–549. [Google Scholar] [PubMed]
- Molderings, G.J.; Heinen, A.; Menzel, S.; Lübbecke, F.; Homann, J.; Göthert, M. Gastrointestinal uptake of agmatine: Distribution in tissues and organs and pathophysiologic relevance. Ann. N. Y. Acad. Sci. 2003, 1009, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Halaris, A.; Plietz, J. Agmatine: Metabolic pathway and spectrum of activity in brain. CNS Drugs 2007, 21, 885–900. [Google Scholar] [CrossRef] [PubMed]
- Winter, T.N.; Elmquist, W.F.; Fairbanks, C.A. OCT2 and MATE1 provide bidirectional agmatine transport. Mol. Pharm. 2011, 8, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Sugiyama, Y.; Nara, M.; Sakanaka, M.; Gotoh, A.; Kitakata, A.; Okuda, S.; Kurihara, S. Comprehensive analysis of polyamine transport and biosynthesis in the dominant human gut bacteria: Potential presence of novel polyamine metabolism and transport genes. Int. J. Biochem. Cell Biol. 2017, 93, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Landete, J.M.; De las Rivas, B.; Marcobal, A.; Muñoz, R. Updated molecular knowledge about histamine biosynthesis by bacteria. Crit. Rev. Food Sci. Nutr. 2008, 48, 697–714. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, B.P.; Hall, A.; Ayyaswamy, S.; Nelson, J.W.; Fultz, R.; Major, A.; Haag, A.; Esparza, M.; Lugo, M.; Venable, S.; et al. Diacylglycerol kinase synthesized by commensal Lactobacillus reuteri diminishes protein kinase C phosphorylation and histamine-mediated signaling in the mammalian intestinal epithelium. Mucosal. Immunol. 2018, 11, 380–393. [Google Scholar] [CrossRef] [PubMed]
- Williams, B.B.; Van Benschoten, A.H.; Cimermancic, P.; Donia, M.S.; Zimmermann, M.; Taketani, M.; Ishihara, A.; Kashyap, P.C.; Fraser, J.S.; Fischbach, M.A. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe 2014, 16, 495–503. [Google Scholar] [CrossRef] [PubMed]
- Westfall, S.; Lomis, N.; Kahouli, I.; Dia, S.Y.; Singh, S.P.; Prakash, S. Microbiome, probiotics and neurodegenerative diseases: Deciphering the gut brain axis. Cell. Mol. Life Sci. 2017, 74, 3769–3787. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.; Pan, J.; Liu, Y.; Wang, H.; Zhou, W.; Wang, X. Intestinal crosstalk between microbiota and serotonin and its impact on gut motility. Curr. Pharm. Biotechnol. 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Ganesh, B.P.; Shi, Z.; Shah, R.R.; Fultz, R.; Major, A.; Venable, S.; Lugo, M.; Hoch, K.; Chen, X.; et al. Gut microbe-mediated suppression of inflammation-associated colon carcinogenesis by luminal histamine production. Am. J. Pathol. 2017, 187, 2323–2336. [Google Scholar] [CrossRef] [PubMed]
- Russo, F.; Linsalata, M.; Orlando, A. Probiotics against neoplastic transformation of gastric mucosa: Effects on cell proliferation and polyamine metabolism. World J. Gastroenterol. 2014, 20, 13258–13272. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishnan, K.; Salinas, R.C. Peptic ulcer disease. Am. Fam. Physician 2007, 76, 1005–1012. [Google Scholar] [PubMed]
- Calam, J.; Baron, J.H. ABC of the upper gastrointestinal tract: Pathophysiology of duodenal and gastric ulcer and gastric cancer. BMJ 2001, 323, 980–982. [Google Scholar] [CrossRef] [PubMed]
- Lai, L.H.; Sung, J.J.Y. Helicobacter pylori and benign upper digestive disease. Best Pract. Res. Clin. Gastroenterol. 2007, 21, 261–279. [Google Scholar] [CrossRef] [PubMed]
- Konturek, S.J.; Konturek, P.C.; Brzozowski, T.; Konturek, J.W.; Pawlik, W.W. From nerves and hormones to bacteria in the stomach; Nobel prize for achievements in gastrology during last century. J. Physiol. Pharmacol. 2005, 56, 507–530. [Google Scholar] [PubMed]
- Singh, V.; Gohil, N.; Ramírez-García, R. New insight into the control of peptic ulcer by targeting the histamine H2 receptor. J. Cell. Biochem. 2018, 119, 2003–2011. [Google Scholar] [CrossRef] [PubMed]
- Safavi, M.; Sabourian, R.; Foroumadi, A. Treatment of Helicobacter pylori infection: Current and future insights. World J. Clin. Cases 2016, 4, 5–19. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, C.A.; Marques, C.R.; dos Santos Costa, R.; da Silva, H.B.F.; Alcantara-Neves, N.M. Cytokines, cytokine gene polymorphisms and Helicobacter pylori infection: Friend or foe? World J. Gastroenterol. 2014, 20, 5235–5243. [Google Scholar] [CrossRef] [PubMed]
- Wessler, S.; Höcker, M.; Fischer, W.; Wang, T.C.; Rosewicz, S.; Haas, R.; Wiedenmann, B.; Meyer, T.F.; Naumann, M. Helicobacter pylori activates the histidine decarboxylase promoter through a mitogen-activated protein kinase pathway independent of pathogenicity island-encoded virulence factors. J. Biol. Chem. 2000, 275, 3629–3636. [Google Scholar] [CrossRef] [PubMed]
- Wadhwa, R.; Song, S.; Lee, J.-S.; Yao, Y.; Wei, Q.; Ajani, J.A. Gastric cancer-molecular and clinical dimensions. Nat. Rev. Clin. Oncol. 2013, 10, 643–655. [Google Scholar] [CrossRef] [PubMed]
- Takaishi, S.; Cui, G.; Frederick, D.M.; Carlson, J.E.; Houghton, J.; Varro, A.; Dockray, G.J.; Ge, Z.; Whary, M.T.; Rogers, A.B.; et al. Synergistic inhibitory effects of gastrin and histamine receptor antagonists on Helicobacter-induced gastric cancer. Gastroenterology 2005, 128, 1965–1983. [Google Scholar] [CrossRef] [PubMed]
- Langman, M.J.; Dunn, J.A.; Whiting, J.L.; Burton, A.; Hallissey, M.T.; Fielding, J.W.; Kerr, D.J. Prospective, double-blind, placebo-controlled randomized trial of cimetidine in gastric cancer. Br. J. Cancer 1999, 81, 1356–1362. [Google Scholar] [CrossRef] [PubMed]
- Garvin, B.; Wiley, J.W. The role of serotonin in irritable bowel syndrome: Implications for management. Curr. Gastroenterol. Rep. 2008, 10, 363–368. [Google Scholar] [CrossRef] [PubMed]
- Crowell, M.D. Role of serotonin in the pathophysiology of the irritable bowel syndrome. Br. J. Pharmacol. 2004, 141, 1285–1293. [Google Scholar] [CrossRef] [PubMed]
- Foley, S.; Garsed, K.; Singh, G.; Duroudier, N.P.; Swan, C.; Hall, I.P.; Zaitoun, A.; Bennett, A.; Marsden, C.; Holmes, G.; et al. Impaired uptake of serotonin by platelets from patients with irritable bowel syndrome correlates with duodenal immune activation. Gastroenterology 2011, 140, 1434–1443. [Google Scholar] [CrossRef] [PubMed]
- Mawe, G.M.; Coates, M.D.; Moses, P.L. Intestinal serotonin signalling in irritable bowel syndrome. Aliment. Pharmacol. Ther. 2006, 23, 1067–1076. [Google Scholar] [CrossRef] [PubMed]
- Konturek, P.C.; Brzozowski, T.; Konturek, S.J. Stress and the gut: Pathophysiology, clinical consequences, diagnostic approach and treatment options. J. Physiol. Pharmacol. 2011, 62, 591–599. [Google Scholar] [PubMed]
- Liu, Q.; Yang, Q.; Sun, W.; Vogel, P.; Heydorn, W.; Yu, X.-Q.; Hu, Z.; Yu, W.; Jonas, B.; Pineda, R.; et al. Discovery and characterization of novel tryptophan hydroxylase inhibitors that selectively inhibit serotonin synthesis in the gastrointestinal tract. J. Pharmacol. Exp. Ther. 2008, 325, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Matthes, S.; Bader, M. Peripheral Serotonin Synthesis as a New Drug Target. Trends Pharmacol. Sci. 2018, 39, 560–572. [Google Scholar] [CrossRef] [PubMed]
- Fogel, W.A.; Lewiński, A.; Jochem, J. Histamine in idiopathic inflammatory bowel diseases—Not a standby player. Folia Med. Cracov. 2005, 46, 107–118. [Google Scholar] [PubMed]
- Neumann, D.; Seifert, R. The therapeutic potential of histamine receptor ligands in inflammatory bowel disease. Biochem. Pharmacol. 2014, 91, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Deiteren, A.; De Man, J.G.; Pelckmans, P.A.; De Winter, B.Y. Histamine H4 receptors in the gastrointestinal tract. Br. J. Pharmacol. 2015, 172, 1165–1178. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Gan, H.-T. Controversies about the use of serological markers in diagnosis of inflammatory bowel disease. World J. Gastroenterol. 2010, 16, 279–280. [Google Scholar] [CrossRef] [PubMed]
- Thomas, H. IBD: Probiotics for IBD: A need for histamine? Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 62–63. [Google Scholar] [CrossRef] [PubMed]
- Milano, A.F.; Singer, R.B. The cancer mortality risk project—Cancer mortality risks by anatomic site: Part 1—Introductory overview; part II—Carcinoma of the Colon: 20-Year mortality follow-up derived from 1973-2013 (NCI) SEER*Stat Survival Database. J. Insur. Med. 2017, 47, 65–94. [Google Scholar] [CrossRef] [PubMed]
- Babbar, N.; Gerner, E.W. Targeting polyamines and inflammation for cancer prevention. Recent Results Cancer Res. 2010, 188, 49–64. [Google Scholar] [CrossRef]
- Shen, P.; Pichler, M.; Chen, M.; Calin, G.A.; Ling, H. To WNT or Lose: The Missing Non-Coding Linc in Colorectal Cancer. Int. J. Mol. Sci. 2017, 18, 2003. [Google Scholar] [CrossRef] [PubMed]
- Gerner, E.W.; Meyskens, F.L. Combination chemoprevention for colon cancer targeting polyamine synthesis and inflammation. Clin. Cancer Res. 2009, 15, 758–761. [Google Scholar] [CrossRef] [PubMed]
- Ogino, S.; Shima, K.; Baba, Y.; Nosho, K.; Irahara, N.; Kure, S.; Chen, L.; Toyoda, S.; Kirkner, G.J.; Wang, Y.L.; et al. Colorectal cancer expression of peroxisome proliferator-activated receptor gamma (PPARG, PPARgamma) is associated with good prognosis. Gastroenterology 2009, 136, 1242–1250. [Google Scholar] [CrossRef] [PubMed]
- Babbar, N.; Ignatenko, N.A.; Casero, R.A.; Gerner, E.W. Cyclooxygenase-independent induction of apoptosis by sulindac sulfone is mediated by polyamines in colon cancer. J. Biol. Chem. 2003, 278, 47762–47775. [Google Scholar] [CrossRef] [PubMed]
- Gerner, E.W.; Meyskens, F.L. Polyamines and cancer: Old molecules, new understanding. Nat. Rev. Cancer 2004, 4, 781–792. [Google Scholar] [CrossRef] [PubMed]
- Vargas, A.J.; Ashbeck, E.L.; Thomson, C.A.; Gerner, E.W.; Thompson, P.A. Dietary polyamine intake and polyamines measured in urine. Nutr. Cancer 2014, 66, 1144–1153. [Google Scholar] [CrossRef] [PubMed]
- Simon, T.; László, V.; Falus, A. Impact of histamine on dendritic cell functions. Cell Biol. Int. 2011, 35, 997–1000. [Google Scholar] [CrossRef] [PubMed]
- Martinel Lamas, D.J.; Rivera, E.S.; Medina, V.A. Histamine H4 receptor: Insights into a potential therapeutic target in breast cancer. Front. Biosci. 2015, 7, 1–9. [Google Scholar]
- Palucka, A.K.; Coussens, L.M. The Basis of Oncoimmunology. Cell 2016, 164, 1233–1247. [Google Scholar] [CrossRef] [PubMed]
- Medina, V.A.; Coruzzi, G.; Martinel Lamas, D.J.; Massari, N.; Adami, M.; Levi-Schaffer, F.; Ben-Zimra, M.; Schwelberger, H.G.; Rivera, E.S. Histamine in cancer. In Histamine H4 Receptor: A Novel Drug Target in Immunoregulation and Inflammation; Stark, H., Ed.; Versita: London, UK, 2013; pp. 259–308. [Google Scholar]
- Barilli, A.; Rotoli, B.M.; Visigalli, R.; Ingoglia, F.; Cirlini, M.; Prandi, B.; Dall’Asta, V. Gliadin-mediated production of polyamines by RAW264.7 macrophages modulates intestinal epithelial permeability in vitro. Biochim. Biophys. Acta 2015, 1852, 1779–1786. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Ruiz, M.; Moreno, E.; Moreno-Delgado, D.; Navarro, G.; Mallol, J.; Cortés, A.; Lluís, C.; Canela, E.I.; Casadó, V.; McCormick, P.J.; et al. Heteroreceptor complexes formed by dopamine d1, histamine h3, and n-methyl-d-aspartate glutamate receptors as targets to prevent neuronal death in Alzheimer’s disease. Mol. Neurobiol. 2017, 54, 4537–4550. [Google Scholar] [CrossRef] [PubMed]
- Burban, A.; Faucard, R.; Armand, V.; Bayard, C.; Vorobjev, V.; Arrang, J.-M. Histamine potentiates N-methyl-d-aspartate receptors by interacting with an allosteric site distinct from the polyamine binding site. J. Pharmacol. Exp. Ther. 2010, 332, 912–921. [Google Scholar] [CrossRef] [PubMed]
- Hornung, B.; Martins Dos Santos, V.A.P.; Smidt, H.; Schaap, P.J. Studying microbial functionality within the gut ecosystem by systems biology. Genes Nutr. 2018, 13, 5. [Google Scholar] [CrossRef] [PubMed]
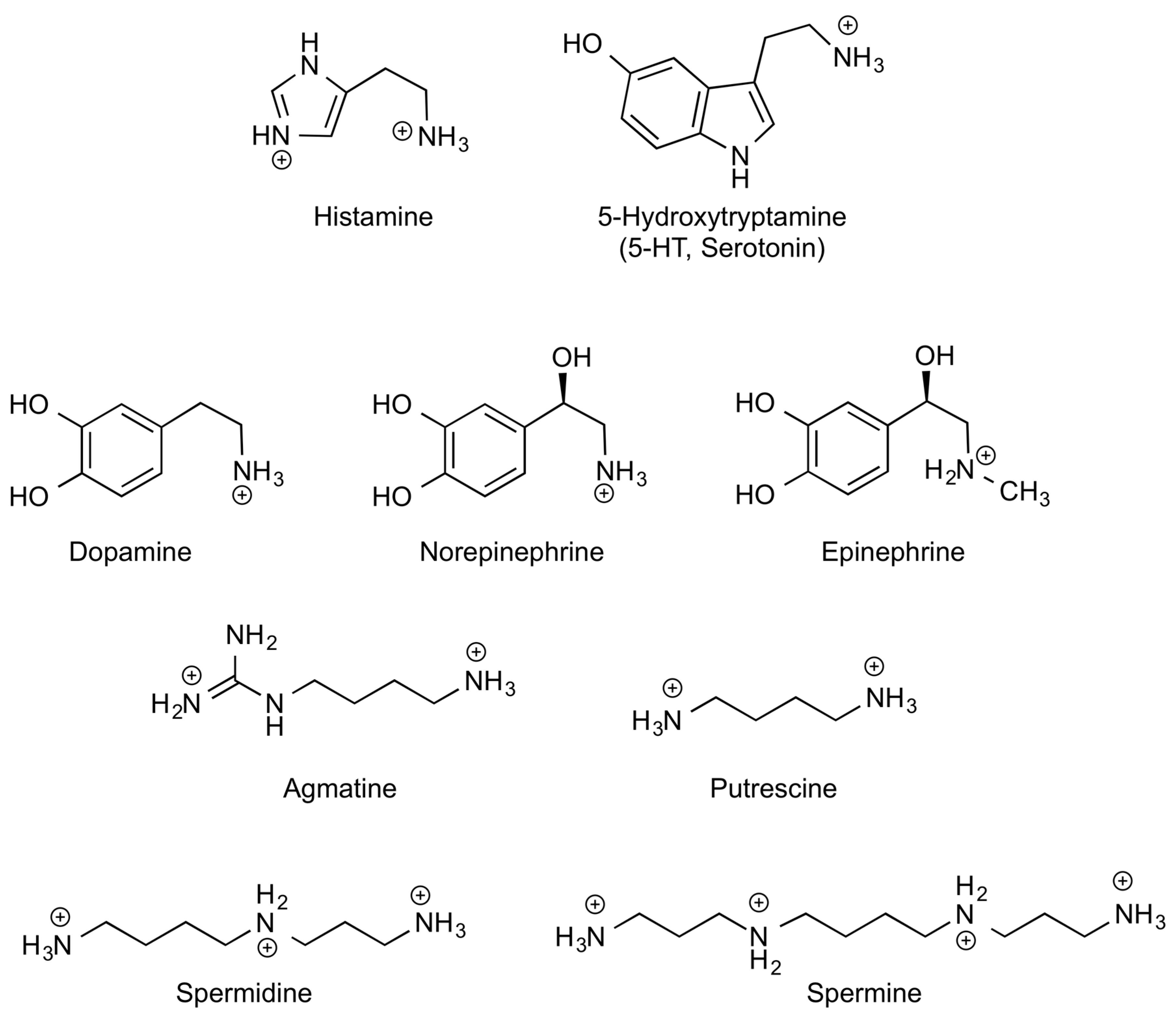
| Common Names (Abbreviations) * | IUPAC Names | Precursor l-Amino Acids | Physiological Roles |
|---|---|---|---|
| Histamine (HIS) | 2-(1H-Imidazol-4-yl)ethanamine | l-Histidine | Neurotransmitter. Immune mediator. Gastric acid secretion inducer. |
| Serotonin (5-HT) | 3-(2-Aminoethyl)-1H-indol-5-ol | l-Triptophan | Neurotransmitter related to reward motivated behaviour. Modulator of vessel constriction and intestinal motility. |
| Catecholamines (CAs): | l-Tyrosine | Blood pressure regulators. Modulators of nutrient absorption and intestinal motility. | |
| Dopamine (DA) | 4-(2-Aminoethyl)benzene-1,2-diol | ||
| Epinephrine | (R)-4-(1-Hydroxy-2-(methyl amino)ethyl)benzene-1,2-diol | ||
| Norepinephrine | (R)-4-(2-amino-1-hydroxy ethyl)benzene-1,2-diol | ||
| Polyamines (PAs): | Essential for cell viability, proliferation and correct differentiation. | ||
| Putrescine (Put) | Butane-1,4-diamine | l-Ornithine | |
| Spermidine (Spd) | N′-(3-aminopropyl)butane-1,4-diamine | l-Ornithine + l-Methionine | |
| Spermine (Spm) | N,N′-bis(3-aminopropyl)butane-1,4-diamine | ||
| Agmatine (Agm) | 2-(4-aminobutyl)guanidine | l-Arginine | Anti-apoptotic effects. Positive effects on brain, hepatic and renal functions. |
| Properties | HIS Receptor 1 (H1R) | HIS Receptor 2 (H2R) | HIS Receptor 3 (H3R) | HIS Receptor 4 (H4R) |
|---|---|---|---|---|
| Chromosome | 3 | 5 | 20 | 18 |
| Molecular weight (KDa) | 56 | 40 | 49 | 44 |
| G protein signalling | Gαq | Gαs | Gi/o | Gi/o |
| Elicited signalling | PLC activation Increase of Ca2+ Production of NOS and cGMP | PKA activation Increase of cAMP PLC activation Increase of Ca2+ | Decrease of cAMP Inhibition of Ca2+ channels | Inhibition of cAMP Stimulation of MAP kinase phosphorylation |
| Expression | Brain, smooth muscle, skin, gastrointestinal and genitourinary tract, adrenal medulla, immune system and heart | Brain, smooth muscle, skin, gastrointestinal and genitourinary tract, adrenal medulla, immune system and heart | Widely found in brain and gastric mucosa | Inflammatory cells, dendritic cells and peripheral nerves |
| Physiological effects | Smooth muscle contraction Vasodilation and increase of vascular permeability | Inhibition of chemotaxis in basophils, gastric secretion of HCl and duodenal bicarbonate secretion | Release regulation of HIS (and other neurotransmitters) release from neurons Inhibition the secretion of gastric acid | Inflammatory processes such as allergies and asthma |
| Properties | 5-HT1A Receptors (5-HT1AR) | 5-HT1D Receptors (5-HT1DR) | 5-HT2 Receptors (5-HT2R) | 5-HT3 Receptors (5-HT3R) | 5-HT4 Receptors (5-HT4R) | 5-HT7 Receptors (5-HT7R) |
|---|---|---|---|---|---|---|
| Chromosome | 5 | 6 | 13/2/X | 11 (A, B and C) and 3 (D and E) | 5 | 10 |
| Molecular weight (KDa) | 421 | 390 | 471/481/458 | Pentameric 478 (A); 441 (B); 447 (C); 279 (D); 471 (E) | 387 | 445 |
| G protein signalling | Gi/o | Gi/o | Gq/11 | Activated by ligand binding and opening channels | Gs | Gs |
| Expression | Enteric neurons, substantia nigra, hippocampus | Enteric neurons, substantia nigra, basal ganglia | Stomach, fundus, caudate nucleus, cerebellum | Enteric, sympathetic and vagus nerves, area postrema | Enteric neurons (myenteric plexus), hippocampus | Smooth muscle, thalamus, hypothalamus and hippocampus |
| Physiological effects | Neuronal inhibition | Neuronal inhibition | Muscle contraction | Neuronal depolarization Increased neurotransmitter release | Muscle contraction Positive effects on cholinergic transmission. | Muscle relaxation |
| Location | Acid Secretion | Gastric Features and Histology | Intestinal Features and Histology | Pathology |
|---|---|---|---|---|
| Stomach (pan-gastritis) | Hyposecretion | Chronic inflammation and parietal cell apoptosis Atrophy Intestinal metaplasia | Normal | Gastric ulcer Gastric cancer |
| Pylorus area | Hypersecretion | Chronic inflammation and increased gastrin released Inhibition of somatostatin Increase parietal cell stimulation | Gastric metaplasia Active chronic inflammation | Duodenal ulcer |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Reina, A.; Urdiales, J.L.; Sánchez-Jiménez, F. What We Know and What We Need to Know about Aromatic and Cationic Biogenic Amines in the Gastrointestinal Tract. Foods 2018, 7, 145. https://doi.org/10.3390/foods7090145
Fernández-Reina A, Urdiales JL, Sánchez-Jiménez F. What We Know and What We Need to Know about Aromatic and Cationic Biogenic Amines in the Gastrointestinal Tract. Foods. 2018; 7(9):145. https://doi.org/10.3390/foods7090145
Chicago/Turabian StyleFernández-Reina, Alberto, José Luis Urdiales, and Francisca Sánchez-Jiménez. 2018. "What We Know and What We Need to Know about Aromatic and Cationic Biogenic Amines in the Gastrointestinal Tract" Foods 7, no. 9: 145. https://doi.org/10.3390/foods7090145
APA StyleFernández-Reina, A., Urdiales, J. L., & Sánchez-Jiménez, F. (2018). What We Know and What We Need to Know about Aromatic and Cationic Biogenic Amines in the Gastrointestinal Tract. Foods, 7(9), 145. https://doi.org/10.3390/foods7090145





