1. Introduction
Although grains normally are part of a balanced diet, an increasing number of people are becoming more sensitive to them, developing clinical conditions like wheat allergies, gluten sensitivity, wheat-dependent exercise induced anaphylaxis (WDEIA), and celiac disease.
Gluten—the main group of proteins in grains such as wheat, rye, barley, and to a lesser extent oats—play a crucial role in the development of celiac disease. This auto-immune disorder of genetically predisposed individuals, affects about 1% of the world’s population and is caused by the ingestion of gluten proteins [
1]. The resistance of some gluten proteins, especially prolamins, to proteases and peptidases in the stomach and gut, and further deamidation of those proteins leads to the development of T-cell epitopes, stimulating the immune system. The consequence is inflammation and destruction of villi in the small intestine, which in the long-term leads to malnutrition. Sensitive individuals need to stick to a strict gluten-free diet due to the fact that there is no cure to celiac disease at the moment [
2].
Compliance with this gluten-free diet can often be challenging for the affected persons. In addition to the learning process involved in determining which products should be avoided, celiac patients are highly reliant on the correct labeling of products. The fact that gluten is used extensively in the food industry due to its functional properties, adds further complexity to this issue. Without an adequate allergen management plan incorporated to the HACCP (hazard analysis critical control points), the unintentional contamination of products during the production process can lead to a misbranding of products and thus threaten any gluten-sensitive people. According to Codex Alimentarius (CODEX STAN 118-1979) [
3], European Commission Regulation (EC 41/2009) [
4] and the U.S. Food and Drug Administration (FR Doc. 2013-18813) [
5], products labeled as “gluten-free” are not allowed to contain more than 20 mg/kg of gluten.
Immunoassays like ELISAs (enzyme-linked immunosorbent assays) and LFDs (lateral flow devices) have many advantages, and are therefore frequently used in the food industry to track allergen contamination. However, one big issue in allergen quantification is that results vary from kit to kit. The lack of an official reference method and official reference materials are factors contributing to this issue. Various technical differences such as antibody specificity, target analytes, sample extraction buffers, extraction time and temperature, calibration standards, and the unit of measurement are the other main factors. Furthermore, different matrixes and their different processing statuses add to the complexity of allergen testing. In particular, thermal processing during production can lead to altered protein extraction efficiencies and antibody binding affinities [
6].
The current Type I Codex method for gluten analysis is the ELISA R5 Mendez method, which is calibrated against the Prolamin Working Group (PWG) Gliadin standard and used by official control systems throughout Europe [
7]. The aim of this study was to compare the suitability of an ELISA G12 method on routine samples entering the official laboratory of AGES (Agentur für Gesundheit und Ernährungssicherheit GmbH)—part of the official food control system in Austria. The different antibodies (R5: raised against omega-Secalin from rye/G12: raised against alpha 2-gliadin 33 mer from wheat) used in these assays are not the only differences. The ELISA G12 method is calibrated with vital wheat gluten (VWG) solution from Roquette, Corby, Northants, UK. Furthermore, the kits differ in their limits of detection (LOD, 2 mg/kg gluten for the G12 assay and 3 mg/kg gluten for the R5 assay), as well as in their quantification ranges (4–200 mg/kg gluten for the G12 assay and 5–80 mg/kg gluten for the R5 assay).
2. Experimental Section
2.1. Samples
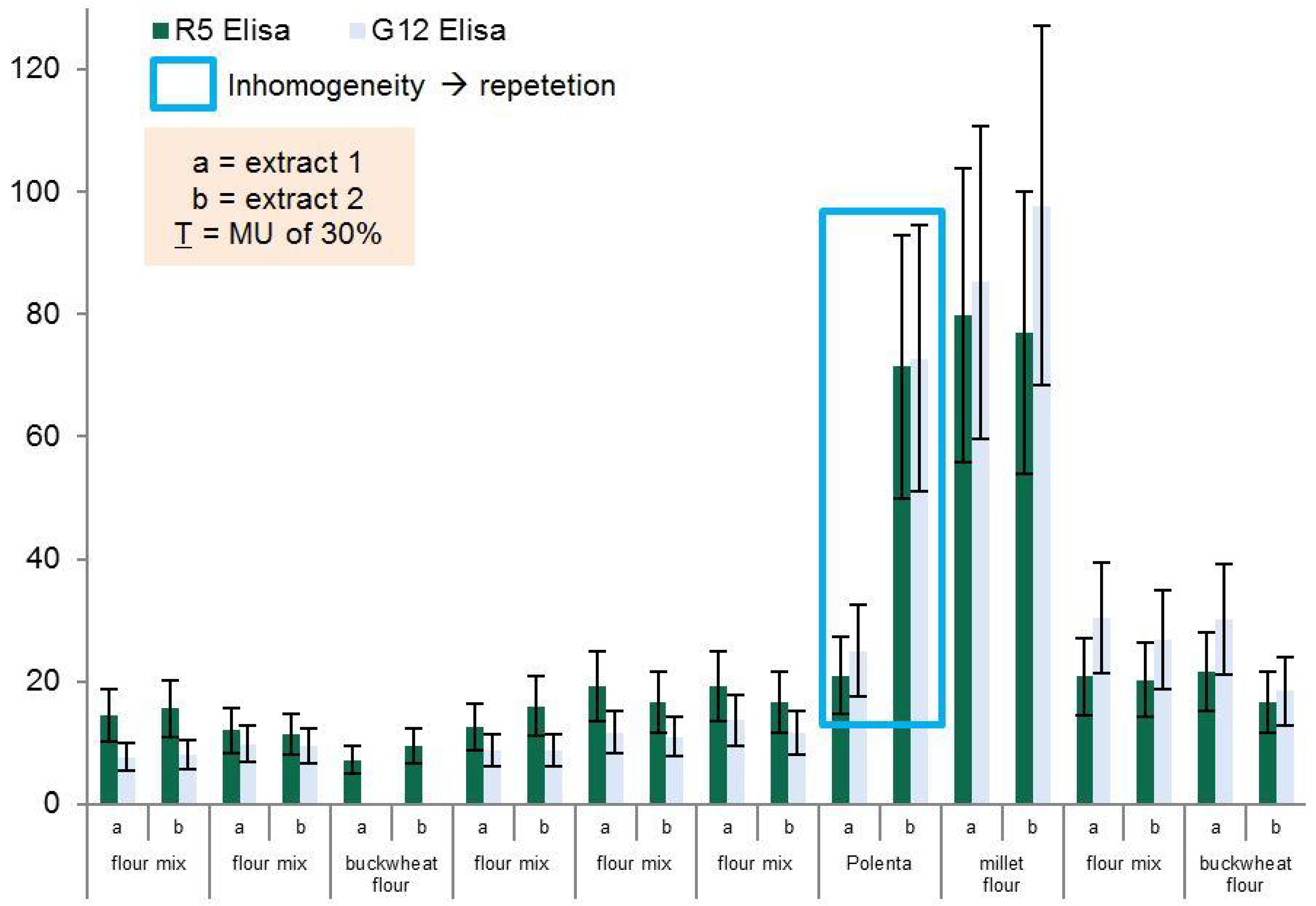
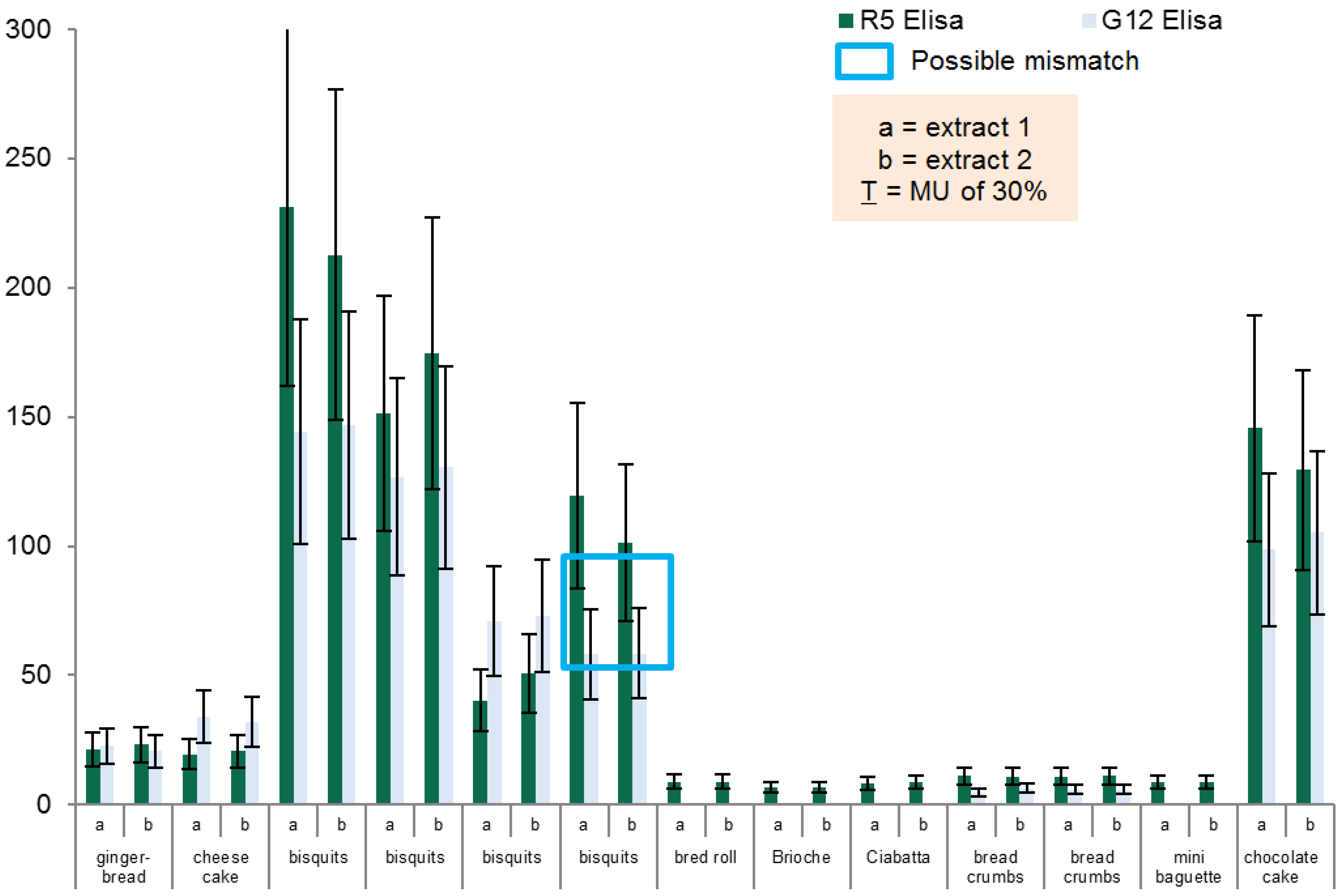
Routine samples from different types of food, all labelled as gluten-free but with a gluten content above LOQ (LOQ = 2.5 mg/kg Gliadin, R5 ELISA) were collected from Austrian supermarkets, retailers, and producers, or received from manufacturers. Samples were extracted and analyzed according to the standard procedure described below. All samples were analyzed in duplicates.
2.2. Sample Extraction Procedure
For the extraction of samples, no commercial kit procedure was used. Instead, an in-house modified extraction method was applied that combines the ethanol and cocktail extraction, described below:
To overcome problems frequently observed when testing real life samples, 5 g of homogenized material were used for the extraction procedure to reduce the impact of possible heterogeneity in our samples. Therefore, an ethanol extraction step was carried out prior to the treatment with the cocktail solution as suggested by the kit provider. After homogenization 25 mL of ethanol 60% is added to 5 g of each sample. After 3 h of incubation at 40 °C (Unihood 750, Uniequip, Martinsried, Germany) the samples are centrifuged at 460 g for 10 min (model 5810 R, Eppendorf, Hamburg, Germany) and the supernatants stored in screw top vials. The precipitates are further diluted by 6.25 mL of the cocktail solution and incubated for 40 min at 50 °C. Finally, 18.75 mL of ethanol 80% are added to the sample and incubated at room temperature for an hour while shaking every 10–15 min. After centrifugation at 3220 g for 10 min both supernatants are pooled. The cocktail was prepared according to the recipe of the patented Mendez Cocktail Solution (EP 2003448 A1), Dithiothreitol (DTT) was used as reducing agent.
2.3. Test Design
The aim of this study was to evaluate the suitability of a G12 antibody based gluten detection kit for the use in an official control system, by testing a number of routine samples. Therefore, samples entering the official laboratory were assayed with two different test kits in parallel—the G12 antibody test kit (Romer Labs) and the current R5 Codex Alimentarius type 1 method (R-Biopharm), which is accredited and has run for more than 10 years at AGES. Since the extraction of proteins from samples is the crucial step in this kind of analysis, we decided to use an identical extraction procedure (described above) prior to assay runs to eliminate side errors. Protein extracts were used immediately after extraction followed by the two individual assay procedures (a comparison of the two different assay procedures can be found in
Table 1). Each assay run was accompanied by proven material (
i.e., proficiency test material, the extraction step was included here) and a positive (PWG Gliadin standard) and a negative control (blank). The measurement uncertainty was determined with 30%.
Table 1.
Comparison of the two different assay procedures.
Table 1.
Comparison of the two different assay procedures.
| | R5 Method | G12 Method |
|---|
| Addition of standard/samples to wells | 100 µL of each standard solution or sample | Add 100 µL of each standard solution or sample |
| 1st incubation | 30 min at room temperature (RT) | 20 min at room temperature (RT) |
| Washing | Empty the contents of the microwell strips | Empty the contents of the microwell strips |
| Wash by filling each microwell with 250 μL diluted wash buffer, and then emptying the buffer from the microwell strips. | Wash by filling each microwell with 300 µL diluted wash buffer, and then emptying the buffer from the microwell strips. |
| Repeat this step a total of 3 times. | Repeat this step a total of 5 times. |
| Addition of conjugate and 2nd incubation | Add 100 μL of the diluted enzyme conjugate to each well and incubate for 30 min at room temperature (20–25 °C/68–77 °F). | Add 100 μL of the diluted enzyme conjugate to each well and incubate for 20 min at room temperature. |
| Washing | Repeat washing step as described above | Repeat washing step as described above |
| Addition of substrate and 3rd incubation | Add 50 μL of substrate and 50 μL of chromogen to each well. Mix gently by shaking the plate manually and incubate for 30 min at room temperature (20–25 °C/68–77 °F) in the dark. | Pipette 100 µL of the Substrate into each microwell and incubate at room temperature for 20 min in the dark. |
| Additions of stop solution and measurement | Add 100 μL of the stop reagent to each well. Mix gently by shaking the plate manually and measure the absorbance at 450 nm. Read within 30 min after addition of stop solution. | Pipette 100 µL of Stop Solution into each microwell. The color should change from blue to yellow. Read the strips with a microwell reader using a 450 nm filter. Record OD readings for each microwell. |