Characterization of Listeria monocytogenes Isolates from Pork Production in Southern Sonora, Mexico: Serotyping, Antimicrobial Resistance, Chitosan Susceptibility, and Pathogenicity in a Chicken Embryo Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Isolation and Phenotypic Characterization of Listeria monocytogenes
2.3. Genotypic Characterization of Listeria monocytogenes Isolates
2.3.1. DNA Extraction
2.3.2. Detection of Virulence Factors by PCR
2.3.3. Molecular Serotyping of Wild Isolates of Listeria monocytogenes
2.4. Assessment of the Antibiotic Resistance Profiles of Wild-Type Listeria monocytogenes Isolates
2.5. Antimicrobial Assays with Shrimp Chitosan Against Listeria monocytogenes Isolates
2.5.1. Chitosan Preparation
2.5.2. Chitosan Antimicrobial Assay
2.6. Assessing the Virulence of Listeria monocytogenes Isolates on Chicken Embryos
2.6.1. Checking for the Dead Embryos
2.6.2. Listeria monocytogenes Inoculum
2.6.3. Infecting the Chicken Embryos with L. monocytogenes Wild Type Isolates
2.6.4. L. monocytogenes Detection in Chicken Liver Embryos
2.6.5. Ethical Declaration for the Use of Chicken Embryos
3. Results
3.1. Identification of Listeria monocytogenes Isolates
3.2. Prevalence of Listeria monocytogenes in Fresh Pork Loin and Fresh Packaged Pork Loin
3.3. Serotype Identification of Listeria monocytogenes Isolates
3.4. Detection of Virulence Factors in Listeria monocytogenes Isolates
3.5. Antibiotic Resistance of Wild-Type Listeria monocytogenes Isolates
3.6. Chitosan as an Alternative to Use Against Listeria monocytogenes in the Food Industry
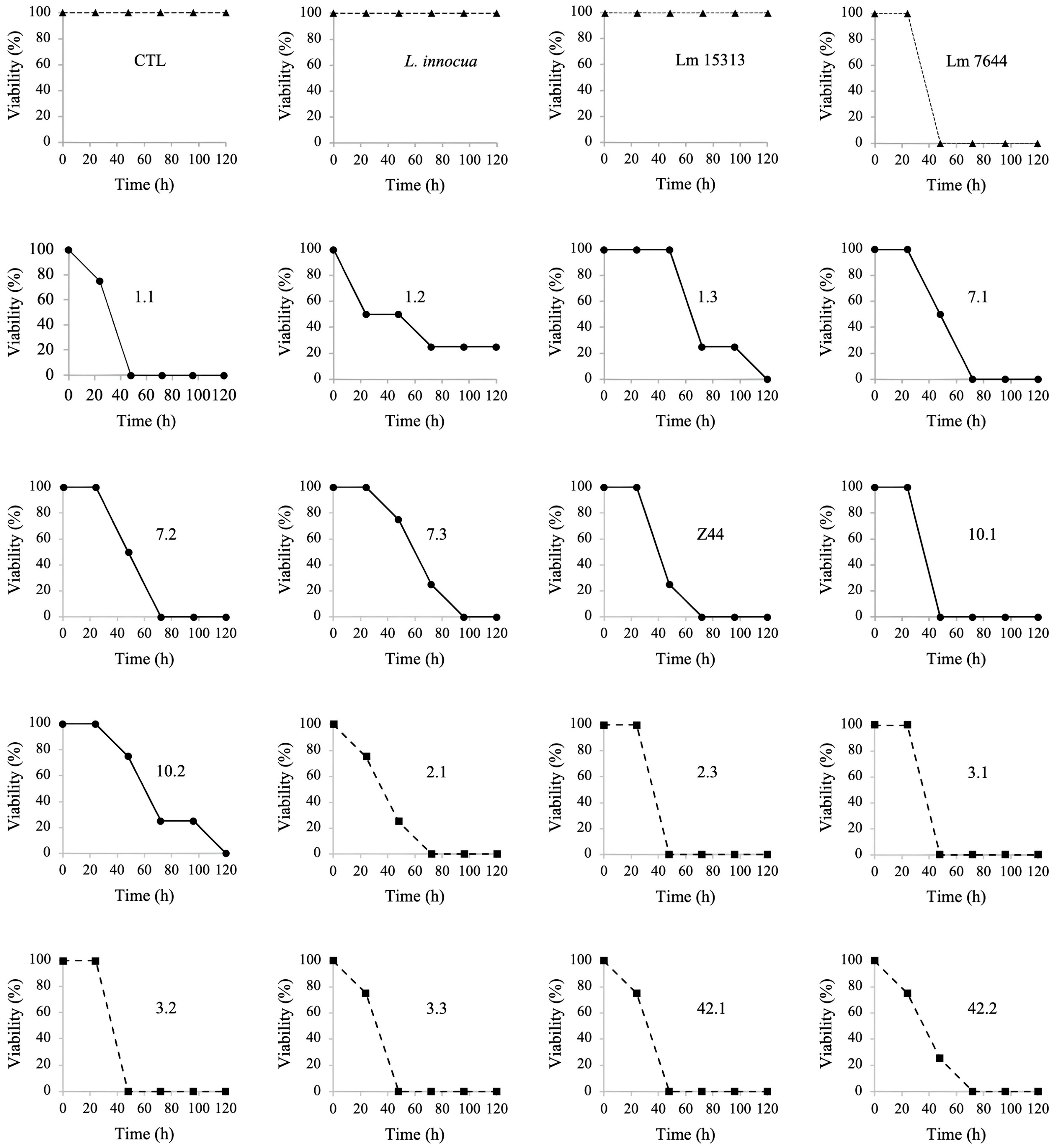
3.7. Listeria monocytogenes Infection Assays on the Chicken Embryo Model
4. Discussions
4.1. Identification of Listeria monocytogenes in Raw and Packaged Pork Samples
4.2. Prevalence of Listeria monocytogenes in Raw and Packaged Pork Samples
4.3. Serotypes of Listeria monocytogenes in Raw and Packaged Pork Samples
4.4. Virulence Factors Present in Listeria monocytogenes Isolates from Raw and Packaged Pork Samples
4.5. Antibiotic Resistance of Wild-Type Listeria monocytogenes Isolates from Raw and Packaged Pork Samples
4.6. Chitosan’s Antimicrobial Effectiveness Tested Against Listeria monocytogenes
4.7. Listeria monocytogenes Causes Infection in Chicken Embryos
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Carpentier, B.; Cerf, O. Review-Persistence of Listeria monocytogenes in Food Industry Equipment and Premises. Int. J. Food Microbiol. 2011, 145, 1–8. [Google Scholar] [CrossRef]
- Schoder, D.; Pelz, A.; Paulsen, P. Transmission Scenarios of Listeria monocytogenes on Small Ruminant on-Farm Dairies. Foods 2023, 12, 265. [Google Scholar] [CrossRef]
- Schoder, D.; Guldimann, C.; Märtlbauer, E. Asymptomatic Carriage of Listeria monocytogenes by Animals and Humans and Its Impact on the Food Chain. Foods 2022, 11, 3472. [Google Scholar] [CrossRef]
- Rossi, F.; Giaccone, V.; Colavita, G.; Amadoro, C.; Pomilio, F.; Catellani, P. Virulence Characteristics and Distribution of the Pathogen Listeria Ivanovii in the Environment and in Food. Microorganisms 2022, 10, 1679. [Google Scholar] [CrossRef]
- Muchaamba, F.; Eshwar, A.K.; Stevens, M.J.A.; Stephan, R.; Tasara, T. Different Shades of Listeria monocytogenes: Strain, Serotype, and Lineage-Based Variability in Virulence and Stress Tolerance Profiles. Front. Microbiol. 2022, 12, 792162. [Google Scholar] [CrossRef] [PubMed]
- Doumith, M.; Buchrieser, C.; Glaser, P.; Jacquet, C.; Martin, P. Differentiation of the Major Listeria monocytogenes Serovars by Multiplex PCR. J. Clin. Microbiol. 2004, 42, 3819–3822. [Google Scholar] [CrossRef]
- Ward, T.J.; Ducey, T.; Usgaard, T.; Dunn, K.A.; Bielawski, J.P. Multilocus Genotyping Assays for Single Nucleotide Polymorphism-Based Subtyping of Listeria monocytogenes Isolates. Appl. Environ. Microbiol. 2008, 74, 7629–7642. [Google Scholar] [CrossRef]
- Orsi, R.H.; den Bakker, H.C.; Wiedmann, M. Listeria monocytogenes Lineages: Genomics, Evolution, Ecology, and Phenotypic Characteristics. Int. J. Med. Microbiol. 2011, 301, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Parra-Flores, J.; Holý, O.; Bustamante, F.; Lepuschitz, S.; Pietzka, A.; Contreras-Fernández, A.; Castillo, C.; Ovalle, C.; Alarcón-Lavín, M.P.; Cruz-Córdova, A.; et al. Virulence and Antibiotic Resistance Genes in Listeria monocytogenes Strains Isolated from Ready-to-Eat Foods in Chile. Front. Microbiol. 2022, 12, 796040. [Google Scholar] [CrossRef]
- Food and Drug Administration Listeria (Listeriosis). Available online: https://www.fda.gov/food/foodborne-pathogens/listeria-listeriosis#:~:text=Past%20listeriosis%20outbreaks%20in%20the,found%20in%20raw%20pet%20food (accessed on 10 May 2025).
- Bridges, D.F.; Bilbao-Sainz, C.; Powell-Palm, M.J.; Williams, T.; Wood, D.; Sinrod, A.J.G.; Ukpai, G.; McHugh, T.H.; Rubinsky, B.; Wu, V.C.H. Viability of and Salmonella Typhimurium after Isochoric Freezing. J. Food Saf. 2020, 40, e12840. [Google Scholar] [CrossRef]
- Demaître, N.; Rasschaert, G.; De Zutter, L.; Geeraerd, A.; De Reu, K. Genetic Listeria monocytogenes Types in the Pork Processing Plant Environment: From Occasional Introduction to Plausible Persistence in Harborage Sites. Pathogens 2021, 10, 717. [Google Scholar] [CrossRef]
- Thévenot, D.; Dernburg, A.; Vernozy-Rozand, C. An Updated Review of Listeria monocytogenes in the Pork Meat Industry and Its Products. J. Appl. Microbiol. 2006, 101, 7–17. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, C.; Bassey, A.; Bai, L.; Wang, Y.; Ye, K. Quantitative Exposure Assessment of Listeria monocytogenes Cross-Contamination from Raw to Ready-to-Eat Meat under Different Food-Handling Scenarios. Food Control 2022, 137, 108972. [Google Scholar] [CrossRef]
- Pricope, L.; Nicolau, A.; Wagner, M.; Rychli, K. The Effect of Sublethal Concentrations of Benzalkonium Chloride on Invasiveness and Intracellular Proliferation of Listeria monocytogenes. Food Control 2013, 31, 230–235. [Google Scholar] [CrossRef]
- Quereda, J.J.; Morón-García, A.; Palacios-Gorba, C.; Dessaux, C.; García-del Portillo, F.; Pucciarelli, M.G.; Ortega, A.D. Pathogenicity and Virulence of Listeria monocytogenes: A Trip from Environmental to Medical Microbiology. Virulence 2021, 12, 2509–2545. [Google Scholar] [CrossRef]
- Kayode, A.J.; Igbinosa, E.O.; Okoh, A.I. Overview of Listeriosis in the Southern African Hemisphere—Review. J. Food Saf. 2020, 40, e12732. [Google Scholar] [CrossRef]
- Vázquez-Boland, J.A.; Domínguez-Bernal, G.; González-Zorn, B.; Kreft, J.; Goebel, W. Pathogenicity Islands and Virulence Evolution in Listeria. Microbes Infect. 2001, 3, 571–584. [Google Scholar] [CrossRef]
- Vázquez-Boland, J.A.; Kuhn, M.; Berche, P.; Chakraborty, T.; Domĺnguez-Bernal, G.; Goebel, W.; González-Zorn, B.; Wehland, J.; Kreft, J. Listeria Pathogenesis and Molecular Virulence Determinants. Clin. Microbiol. Rev. 2001, 14, 584–640. [Google Scholar] [CrossRef] [PubMed]
- Ntshanka, Z.; Ekundayo, T.C.; du Plessis, E.M.; Korsten, L.; Okoh, A.I. Occurrence and Molecular Characterization of Multidrug-Resistant Vegetable-Borne Listeria monocytogenes Isolates. Antibiotics 2022, 11, 1353. [Google Scholar] [CrossRef]
- EFSA. The European Union One Health 2018 Zoonoses Report. ESFA J. 2019, 17, e05926. [Google Scholar]
- Lundén, J.; Autio, T.; Markkula, A.; Hellström, S.; Korkeala, H. Adaptive and Cross-Adaptive Responses of Persistent and Non-Persistent Listeria monocytogenes Strains to Disinfectants. Int. J. Food Microbiol. 2003, 82, 265–272. [Google Scholar] [CrossRef]
- Folsom, J.P.; Frank, J.F. Chlorine Resistance of Listeria monocytogenes Biofilms and Relationship to Subtype, Cell Density, and Planktonic Cell Chlorine Resistance. J. Food Prot. 2006, 69, 1292–1296. [Google Scholar] [CrossRef] [PubMed]
- Møretrø, T.; Schirmer, B.C.T.; Heir, E.; Fagerlund, A.; Hjemli, P.; Langsrud, S. Tolerance to Quaternary Ammonium Compound Disinfectants May Enhance Growth of Listeria monocytogenes in the Food Industry. Int. J. Food Microbiol. 2017, 241, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Ardean, C.; Davidescu, C.M.; Nemeş, N.S.; Negrea, A.; Ciopec, M.; Duteanu, N.; Negrea, P.; Duda-seiman, D.; Musta, V. Factors Influencing the Antibacterial Activity of Chitosan and Chitosan Modified by Functionalization. Int. J. Mol. Sci. 2021, 22, 7449. [Google Scholar] [CrossRef] [PubMed]
- Momtaz, H.; Yadollahi, S. Molecular Characterization of Listeria monocytogenes Isolated from Fresh Seafood Samples in Iran. Diagn. Pathol. 2013, 8, 149. [Google Scholar] [CrossRef]
- CLSI. Clinical & Laboratory Standards Institute: CLSI Guidelines; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2024. [Google Scholar]
- Singh, S.; Yadav, A.S.; Singh, S.M.; Bharti, P. Prevalence of Salmonella in Chicken Eggs Collected from Poultry Farms and Marketing Channels and Their Antimicrobial Resistance. Food Res. Int. 2010, 43, 2027–2030. [Google Scholar] [CrossRef]
- Rodríguez-Núñez, J.R.; Madera-Santana, T.J.; Sánchez-Machado, D.I.; López-Cervantes, J.; Soto Valdez, H. Chitosan/Hydrophilic Plasticizer-Based Films: Preparation, Physicochemical and Antimicrobial Properties. J. Polym. Environ. 2014, 22, 41–51. [Google Scholar] [CrossRef]
- Bueno-Solano, C.; López-Cervantes, J.; Campas-Baypoli, O.N.; Lauterio-García, R.; Adan-Bante, N.P.; Sánchez-Machado, D.I. Chemical and Biological Characteristics of Protein Hydrolysates from Fermented Shrimp By-Products. Food Chem. 2009, 112, 671–675. [Google Scholar] [CrossRef]
- Weska, R.F.; Moura, J.M.; Batista, L.M.; Rizzi, J.; Pinto, L.A. Optimization of Deacetylation in the Production of Chitosan from Shrimp Wastes: Use of Response Surface Methodology. J. Food Eng. 2007, 80, 749–753. [Google Scholar] [CrossRef]
- Andersson, C.; Gripenland, J.; Johansson, J. Using the Chicken Embryo to Assess Virulence of Listeria monocytogenes and to Model Other Microbial Infections. Nat. Protoc. 2015, 10, 1155–1164. [Google Scholar] [CrossRef]
- Mateus, T.; Silva, J.; Maia, R.L.; Teixeira, P. Listeriosis during Pregnancy: A Public Health Concern. Int. Sch. Res. Not. 2013, 2013, 851712. [Google Scholar] [CrossRef]
- Yang, Y.; Kong, X.; Niu, B.; Yang, J.; Chen, Q. Differences in Biofilm Formation of Listeria monocytogenes and Their Effects on Virulence and Drug Resistance of Different Strains. Foods 2024, 13, 1076. [Google Scholar] [CrossRef] [PubMed]
- Law, J.W.-F.; Ab Mutalib, N.-S.; Chan, K.-G.; Lee, L.-H. An Insight into the Isolation, Enumeration, and Molecular Detection of Listeria monocytogenes in Food. Front. Microbiol. 2015, 6, 1227. [Google Scholar] [CrossRef] [PubMed]
- Figueroa-López, A.M.; Maldonado-Mendoza, I.E.; López-Cervantes, J.; Verdugo-Fuentes, A.A.; Ruiz-Vega, D.A.; Cantú-Soto, E.U. Prevalence and Characterization of Listeria monocytogenes Isolated from Pork Meat and on Inert Surfaces. Braz. J. Microbiol. 2019, 50, 817–824. [Google Scholar] [CrossRef] [PubMed]
- Gamboa-Marín, A.; Buitrago, M.S.; Pérez-Pérez, K.; Mercado, R.M.; Poutou-Piñales, R.; Carrascal-Camacho, A. Prevalence of Listeria monocytogenes in pork-meat and other processed products from the Colombian swine industry. Rev. MVZ Córdoba 2012, 17, 2827–2833. [Google Scholar] [CrossRef]
- Chasseignaux, E.; Gérault, P.; Toquin, M.; Salvat, G.; Colin, P.; Ermel, G. Ecology of Listeria monocytogenes in the Environment of Raw Poultry Meat and Raw Pork Meat Processing Plants. FEMS Microbiol. Lett. 2002, 210, 271–275. [Google Scholar] [CrossRef]
- Lagarde, J.; Feurer, C.; Denis, M.; Douarre, P.-E.; Piveteau, P.; Roussel, S. Listeria monocytogenes Prevalence and Genomic Diversity along the Pig and Pork Production Chain. Food Microbiol. 2024, 119, 104430. [Google Scholar] [CrossRef]
- Prencipe, V.A.; Rizzi, V.; Acciari, V.; Iannetti, L.; Giovannini, A.; Serraino, A.; Calderone, D.; Rossi, A.; Morelli, D.; Marino, L.; et al. Listeria monocytogenes Prevalence, Contamination Levels and Strains Characterization throughout the Parma Ham Processing Chain. Food Control 2012, 25, 150–158. [Google Scholar] [CrossRef]
- Capita, R.; Felices-Mercado, A.; García-Fernández, C.; Alonso-Calleja, C. Characterization of Listeria monocytogenes Originating from the Spanish Meat-Processing Chain. Foods 2019, 8, 542. [Google Scholar] [CrossRef] [PubMed]
- Coroneo, V.; Carraro, V.; Aissani, N.; Sanna, A.; Ruggeri, A.; Succa, S.; Meloni, B.; Pinna, A.; Sanna, C. Detection of virulence genes and growth potential in Listeria monocytogenes strains isolated from ricotta salata cheese. J. Food Sci. 2016, 81, M114–M120. [Google Scholar] [CrossRef]
- Duma, M.N.; Ciupescu, L.M.; Dan, S.D.; Crisan-Reget, O.L.; Tabaran, A. Virulence and Antimicrobial Resistance of Listeria monocytogenes Isolated from Ready-to-Eat Food Products in Romania. Microorganisms 2024, 12, 954. [Google Scholar] [CrossRef]
- Banović, F.; Schroten, H.; Schwerk, C. Potential Roles and Functions of Listerial Virulence Factors during Brain Entry. Toxins 2020, 12, 297. [Google Scholar] [CrossRef]
- Yu, M.; Yang, J.; Guo, M. Is the LysM Domain of Listeria monocytogenes P60 Protein Suitable for Engineering a Protein with High Peptidoglycan Binding Affinity? Bioengineered 2016, 7, 406–410. [Google Scholar] [CrossRef] [PubMed]
- Kawacka, I.; Olejnik-Schmidt, A. High Prevalence of Virulence-Associated Genes and Length Polymorphism in actA and inlB Genes Identified in Listeria monocytogenes Isolates from Meat Products and Meat-Processing Environments in Poland. Pathogens 2024, 13, 444. [Google Scholar] [CrossRef] [PubMed]
- Grossman, T.H. Tetracycline Antibiotics and Resistance. Cold Spring Harb. Perspect. Med. 2016, 6, a025387. [Google Scholar] [CrossRef] [PubMed]
- Leclercq, R. Mechanisms of Resistance to Macrolides and Lincosamides: Nature of the Resistance Elements and Their Clinical Implications. Clin. Infect. Dis. 2002, 34, 482–492. [Google Scholar] [CrossRef]
- Brenciani, A.; Morroni, G.; Schwarz, S.; Giovanetti, E. Oxazolidinones: Mechanisms of Resistance and Mobile Genetic Elements Involved. J. Antimicrob. Chemother. 2022, 77, 2596–2621. [Google Scholar] [CrossRef]
- Hershberger, E.; Donabedian, S.; Konstantinou, K.; Zervos, M.J.; Eliopoulos, G.M. Quinupristin-Dalfopristin Resistance in Gram-Positive Bacteria: Mechanism of Resistance and Epidemiology. Clin. Infect. Dis. 2004, 38, 92–98. [Google Scholar] [CrossRef]
- Rice, L.B. Mechanisms of Resistance and Clinical Relevance of Resistance to β-Lactams, Glycopeptides, and Fluoroquinolones. Mayo Clin. Proc. 2012, 87, 198–208. [Google Scholar] [CrossRef]
- Lodewijk, S.; Grauls, V. Activity of Daptomycin against Listeria monocytogenes Isolates from Cerebrospinal Fluid. Antimicrob. Agents Chemother. 2008, 52, 1850–1851. [Google Scholar] [CrossRef]
- Kettlewell, R.; Jones, C.; Felton, T.W.; Lagator, M.; Gifford, D.R. Insights into Durability against Resistance from the Antibiotic Nitrofurantoin. Npj Antimicrob. Resist. 2024, 2, 41. [Google Scholar] [CrossRef]
- Rippa, A.; Bilei, S.; Peruzy, M.F.; Marrocco, M.G.; Leggeri, P.; Bossù, T.; Murru, N. Antimicrobial Resistance of Listeria monocytogenes Strains Isolated in Food and Food-Processing Environments in Italy. Antibiotics 2024, 13, 525. [Google Scholar] [CrossRef] [PubMed]
- Morvan, A.; Moubareck, C.; Leclercq, A.; Hervé-Bazin, M.; Bremont, S.; Lecuit, M.; Courvalin, P.; Le Monnier, A. Antimicrobial Resistance of Listeria monocytogenes Strains Isolated from Humans in France. Antimicrob. Agents Chemother. 2010, 54, 2728–2731. [Google Scholar] [CrossRef]
- Şanlıbaba, P.; Tezel, B.U.; Çakmak, G.A. Prevalence and Antibiotic Resistance of Listeria monocytogenes Isolated from Ready-to-Eat Foods in Turkey. J. Food Qual. 2018, 2018, 7693782. [Google Scholar] [CrossRef]
- Godreuil, S.; Galimand, M.; Gerbaud, G.; Jacquet, C.; Courvalin, P. Efflux Pump Lde Is Associated with Fluoroquinolone Resistance in Listeria monocytogenes. Antimicrob. Agents Chemother. 2003, 47, 704–708. [Google Scholar] [CrossRef]
- Matereke, L.T.; Okoh, A.I. Listeria monocytogenes Virulence, Antimicrobial Resistance and Environmental Persistence: A Review. Pathogens 2020, 9, 528. [Google Scholar] [CrossRef]
- Barbosa, J.; Magalhães, R.; Santos, I.; Ferreira, V.; Brandão, T.R.S.; Silva, J.; Almeida, G.; Teixeira, P. Evaluation of Antibiotic Resistance Patterns of Food and Clinical Listeria monocytogenes Isolates in Portugal. Foodborne Pathog. Dis. 2013, 10, 861–866. [Google Scholar] [CrossRef]
- Kayode, A.J.; Okoh, A.I. Antibiotic Resistance Profile of Listeria monocytogenes Recovered from Ready-to-Eat Foods Surveyed in South Africa. J. Food Prot. 2022, 85, 1807–1814. [Google Scholar] [CrossRef] [PubMed]
- Almansour, A.M.; Alhadlaq, M.A.; Alzahrani, K.O.; Mukhtar, L.E.; Alharbi, A.L.; Alajel, S.M. The Silent Threat: Antimicrobial-Resistant Pathogens in Food-Producing Animals and Their Impact on Public Health. Microorganisms 2023, 11, 2127. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial Resistance: Impacts, Challenges, and Future Prospects. J. Med. Surg. Public Health 2024, 2, 100081. [Google Scholar] [CrossRef]
- Abebe, A.A.; Birhanu, A.G. Methicillin Resistant Staphylococcus aureus: Molecular Mechanisms Underlying Drug Resistance Development and Novel Strategies to Combat. IDR 2023, 16, 7641–7662. [Google Scholar] [CrossRef]
- Woodford, N.; Turton, J.F.; Livermore, D.M. Multiresistant Gram-Negative Bacteria: The Role of High-Risk Clones in the Dissemination of Antibiotic Resistance. FEMS Microbiol. Rev. 2011, 35, 736–755. [Google Scholar] [CrossRef]
- Skłodowski, K.; Chmielewska-Deptuła, S.J.; Piktel, E.; Wolak, P.; Wollny, T.; Bucki, R. Metallic Nanosystems in the Development of Antimicrobial Strategies with High Antimicrobial Activity and High Biocompatibility. Int. J. Mol. Sci. 2023, 24, 2104. [Google Scholar] [CrossRef]
- Bava, R.; Castagna, F.; Lupia, C.; Poerio, G.; Liguori, G.; Lombardi, R.; Naturale, M.D.; Mercuri, C.; Bulotta, R.M.; Britti, D.; et al. Antimicrobial Resistance in Livestock: A Serious Threat to Public Health. Antibiotics 2024, 13, 551. [Google Scholar] [CrossRef] [PubMed]
- Oleksy, M.; Dynarowicz, K.; Aebisher, D. Advances in Biodegradable Polymers and Biomaterials for Medical Applications—A Review. Molecules 2023, 28, 6213. [Google Scholar] [CrossRef] [PubMed]
- Guzman Rosellon, M.C.; Ortiz, G.D.R. Quitosano: Evaluación económica para la producción industrial de un biopolímero. Mem. Concurs. Lasallista Investig. Desarro. Innovación 2019, 6, 15–18. [Google Scholar] [CrossRef]
- Nasaj, M.; Chehelgerdi, M.; Asghari, B.; Ahmadieh-Yazdi, A.; Asgari, M.; Kabiri-Samani, S.; Sharifi, E.; Arabestani, M. Factors Influencing the Antimicrobial Mechanism of Chitosan Action and Its Derivatives: A Review. Int. J. Biol. Macromol. 2024, 277, 134321. [Google Scholar] [CrossRef] [PubMed]
- Giannoulis, N.; Karatzas, K.A.G. The Combined Effect of Chitosan and High Hydrostatic Pressure on Listeria monocytogenes and Escherichia coli. Innov. Food Sci. Emerg. Technol. 2024, 94, 103693. [Google Scholar] [CrossRef]
- Jovanović, G.D.; Klaus, A.S.; Nikšić, M.P. Antimicrobial Activity of Chitosan Coatings and Films against Listeria monocytogenes on Black Radish. Rev. Argent. Microbiol. 2016, 48, 128–136. [Google Scholar] [CrossRef] [PubMed]
- Ibañez-Peinado, D.; Ubeda-Manzanaro, M.; Martínez, A.; Rodrigo, D. Antimicrobial Effect of Insect Chitosan on Salmonella Typhimurium, Escherichia coli O157:H7 and Listeria monocytogenes Survival. PLoS ONE 2020, 15, e0244153. [Google Scholar] [CrossRef]
- Raafat, D.; Leib, N.; Wilmes, M.; François, P.; Schrenzel, J.; Sahl, H.-G. Development of in Vitro Resistance to Chitosan Is Related to Changes in Cell Envelope Structure of Staphylococcus aureus. Carbohydr. Polym. 2017, 157, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Sibanda, T.; Buys, E.M. Listeria monocytogenes Pathogenesis: The Role of Stress Adaptation. Microorganisms 2022, 10, 1522. [Google Scholar] [CrossRef]
- Severino, P.; Dussurget, O.; Vêncio, R.Z.N.; Dumas, E.; Garrido, P.; Padilla, G.; Piveteau, P.; Lemaître, J.-P.; Kunst, F.; Glaser, P.; et al. Comparative Transcriptome Analysis of Listeria monocytogenes Strains of the Two Major Lineages Reveals Differences in Virulence, Cell Wall, and Stress Response. Appl. Environ. Microbiol. 2007, 73, 6078–6088. [Google Scholar] [CrossRef]
- Yin, Y.; Tian, D.; Jiao, H.; Zhang, C.; Pan, Z.; Zhang, X.; Wang, X.; Jiao, X. Pathogenicity and Immunogenicity of a Mutant Strain of Listeria monocytogenes in the Chicken Infection Model. Clin. Vaccine Immunol. 2011, 18, 500–505. [Google Scholar] [CrossRef]
- Olier, M.; Pierre, F.; Rousseaux, S.; Lemaître, J.-P.; Rousset, A.; Piveteau, P.; Guzzo, J. Expression of Truncated Internalin A Is Involved in Impaired Internalization of Some Listeria monocytogenes Isolates Carried Asymptomatically by Humans. Infect. Immun. 2003, 71, 1217–1224. [Google Scholar] [CrossRef]
- Nishibori, T.; Cooray, K.; Xiong, H.; Kawamura, I.; Fujita, M.; Mitsuyama, M. Correlation between the Presence of Virulence-Associated Genes as Determined by PCR and Actual Virulence to Mice in Various Strains of Listeria spp. Microbiol. Immunol. 1995, 39, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Gripenland, J.; Andersson, C.; Johansson, J. Exploring the Chicken Embryo as a Possible Model for Studying Listeria monocytogenes Pathogenicity. Front. Cell. Infect. Microbiol. 2014, 4, 170. [Google Scholar] [CrossRef]
- Quereda, J.J.; Andersson, C.; Cossart, P.; Johansson, J.; Pizarro-Cerdá, J. Role in Virulence of Phospholipases, Listeriolysin O and Listeriolysin S from Epidemic Listeria monocytogenes Using the Chicken Embryo Infection Model. Vet. Res. 2018, 49, 13. [Google Scholar] [CrossRef] [PubMed]
| Serotypes | |||||||
|---|---|---|---|---|---|---|---|
| Source | Samples | Prevalence | L. monocytogenes Isolates Obtained | 1/2a | 1/2b | 4b | ND |
| Raw pork loin | 16 | 56.3% | 9 | 1 (11.1%) | 7 (77.8%) | ̶̶̶ | 1 (11.1%) |
| Raw packaged pork loin | 10 | 70% | 7 | 1 (14.3%) | 4 (57.1%) | 1 (14.3%) | 1 (14.3%) |
| TOTAL | 26 | 61.5% | 16 | 2 (12.5%) | 11 (68.75%) | 1 (6.25%) | 2 (12.5%) |
| Source | Bacterial Isolate | Virulence Factors | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| actA | hlyA | lnlA | InlB | InlC | InlJ | Iap | plcA | plcB | prfA | Serotype | ||
| Raw pork loin | L. monocytogenes ATCC 7644 * | + | + | + | + | + | + | + | + | + | + | 1/2a |
| Lm 1.1 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm 1.2 | + | + | ND | ND | + | ND | + | + | + | + | 1/2b | |
| Lm 1.3 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm 7.1 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm 7.2 | + | + | + | + | + | + | + | ND | + | + | 1/2b | |
| Lm 7.3 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm Z44 | + | + | + | + | + | + | + | + | + | + | 1/2a | |
| Lm 10.1 | + | + | + | ND | + | + | + | + | + | + | 1/2b | |
| Lm 10.2 | + | + | + | + | + | + | + | + | + | + | ND | |
| Raw packaged pork loin | Lm 2.1 | + | + | + | + | + | + | + | + | + | + | 1/2a |
| Lm 2.3 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm 3.1 | ND | + | + | + | + | + | + | ND | + | + | 4b | |
| Lm 3.2 | ND | + | + | + | + | + | + | ND | + | + | 1/2b | |
| Lm 3.3 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Lm 42.1 | + | + | + | + | + | + | + | + | + | + | ND | |
| Lm 42.2 | + | + | + | + | + | + | + | + | + | + | 1/2b | |
| Source | Isolates | Antibiotic (MIC μg/mL) * | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AMC | SAM | AMP | CRO | CIP | CLI | DAP | ERY | FOF | GEN | LVX | LZD | MXF | NIT | OXA | PEN | RIF | SYN | TET | SXT | VAN | MARI | ||
| L. monocytogenes | ATCC ♦ | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | S <32 | R >2 | S < 0.03 | S <1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 |
| Fresh pork loin | 1.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | R > 4 | R > 4 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S <1 | S < 0.5 | S <4 | S < 0.5/9.5 | S < 0.25 | 0.333 |
| 1.2 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | R > 4 | R > 4 | S < 0.5 | R > 64 | R >2 | R > 8 | S < 1 | R > 2 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.476 | |
| 1.3 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 7.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 7.2 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S <1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 7.3 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| z44 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | R > 4 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | R >8 | S < 0.5/9.5 | S < 0.25 | 0.381 | |
| 10.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | S <32 | R >2 | S < 0.03 | S <1 | S < 0.5 | S <4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 10.2 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| Fresh packaged pork loin | 2.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | R > 4 | S < 0.5 | R > 64 | R >2 | R > 8 | S < 1 | R > 2 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.429 |
| 2.3 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.286 | |
| 3.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S <1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 3.2 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.286 | |
| 3.3 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | R > 2 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.333 | |
| 42.1 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | S < 1 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.238 | |
| 42.2 | S < 4/2 | S < 8/4 | S < 2 | R > 32 | R > 2 | R > 4 | R > 4 | S < 0.5 | S | S < 4 | S < 1 | S < 1 | S < 0.5 | R > 64 | R >2 | S < 0.03 | S < 1 | S < 0.5 | S < 4 | S < 0.5/9.5 | S < 0.25 | 0.286 | |
| Log (CFU/mL) | |||||
|---|---|---|---|---|---|
| Isolate | Initial Inoculum | Control † | Chitosan (0.25%) † | Acetic Acid (1%) † | |
| Raw pork loin | Lm 1.1 | 6.36 a | 6.83 a | 0.00 b | 6.54 a |
| Lm 1.2 | 6.55 a | 6.76 a | 0.00 b | 6.13 a | |
| Lm 1.3 | 6.18 a | 6.35 a | 0.00 b | 6.27 a | |
| Lm 7.1 | 6.33 a | 6.70 a | 0.00 b | 6.12 a | |
| Lm 7.2 | 6.09 a | 6.59 a | 0.00 b | 6.48 a | |
| Lm 7.3 | 6.32 a | 6.71 a | 0.00 b | 6.21 a | |
| Lm z44 | 6.49 a | 6.58 a | 0.00 b | 6.58 a | |
| Lm 10.1 | 5.87 a | 6.58 a | 4.03 a | 6.50 a | |
| Lm 10.2 | 6.39 a | 6.61 a | 0.00 b | 6.33 a | |
| Packaged pork loin | Lm 2.1 | 5.11 a | 5.18 a | 0.00 b | 5.11 a |
| Lm 2.3 | 5.13 a | 5.43 a | 0.00 b | 5.28 a | |
| Lm 3.1 | 5.36 a | 5.53 a | 0.00 b | 5.27 a | |
| Lm 3.2 | 5.55 a | 5.77 a | 0.00 b | 5.57 a | |
| Lm 3.3 | 5.17 a | 5.24 a | 0.00 b | 5.25 a | |
| Lm 42.1 | 5.61 a | 5.75 a | 0.00 b | 5.40 a | |
| Lm 42.2 | 5.26 a | 5.63 a | 0.00 b | 5.49 a | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Figueroa-López, A.M.; Gutiérrez-Osuna, M.T.; Rodríguez-Mora, N.G.; Aragón-López, C.E.; Rodríguez-Ramírez, R.; Villa-Lerma, A.G.; Cantú-Soto, E.U. Characterization of Listeria monocytogenes Isolates from Pork Production in Southern Sonora, Mexico: Serotyping, Antimicrobial Resistance, Chitosan Susceptibility, and Pathogenicity in a Chicken Embryo Model. Foods 2025, 14, 3057. https://doi.org/10.3390/foods14173057
Figueroa-López AM, Gutiérrez-Osuna MT, Rodríguez-Mora NG, Aragón-López CE, Rodríguez-Ramírez R, Villa-Lerma AG, Cantú-Soto EU. Characterization of Listeria monocytogenes Isolates from Pork Production in Southern Sonora, Mexico: Serotyping, Antimicrobial Resistance, Chitosan Susceptibility, and Pathogenicity in a Chicken Embryo Model. Foods. 2025; 14(17):3057. https://doi.org/10.3390/foods14173057
Chicago/Turabian StyleFigueroa-López, Alejandro Miguel, Michel Teresa Gutiérrez-Osuna, Norma Gabriela Rodríguez-Mora, Carlos Eduardo Aragón-López, Roberto Rodríguez-Ramírez, Alma Guadalupe Villa-Lerma, and Ernesto Uriel Cantú-Soto. 2025. "Characterization of Listeria monocytogenes Isolates from Pork Production in Southern Sonora, Mexico: Serotyping, Antimicrobial Resistance, Chitosan Susceptibility, and Pathogenicity in a Chicken Embryo Model" Foods 14, no. 17: 3057. https://doi.org/10.3390/foods14173057
APA StyleFigueroa-López, A. M., Gutiérrez-Osuna, M. T., Rodríguez-Mora, N. G., Aragón-López, C. E., Rodríguez-Ramírez, R., Villa-Lerma, A. G., & Cantú-Soto, E. U. (2025). Characterization of Listeria monocytogenes Isolates from Pork Production in Southern Sonora, Mexico: Serotyping, Antimicrobial Resistance, Chitosan Susceptibility, and Pathogenicity in a Chicken Embryo Model. Foods, 14(17), 3057. https://doi.org/10.3390/foods14173057







