Fermentation Process Optimization of Strawberry Wine Using the 2-Phenylethanol Tolerant Saccharomyces cerevisiae AFRC01 and Comparative Genomic Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Materials
2.2. Selection of Fermented Strawberry Wine for Single-Factor Experiments
2.3. Quantification of 2-Phenylethanol Content
2.4. Response Surface Methodology Optimization and Model Validation
2.5. Sensory Analysis
2.6. Genome Sequencing and Functional Annotation
2.7. Expression Analysis of Phenylethanol Biosynthesis-Related Genes
2.8. Statistical Analysis
3. Results and Analysis
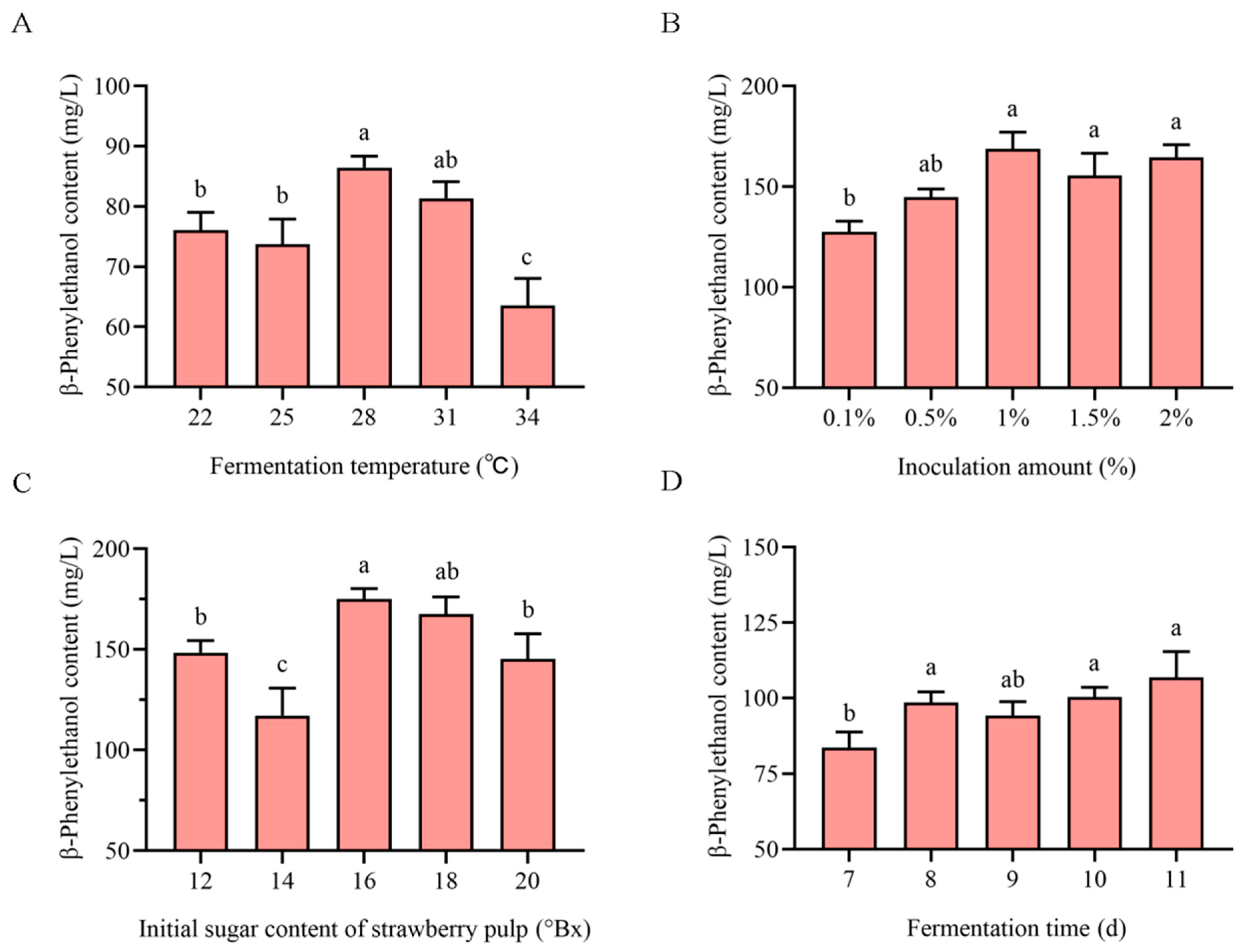
3.1. Single-Factor Optimization of Strawberry Wine Fermentation Conditions
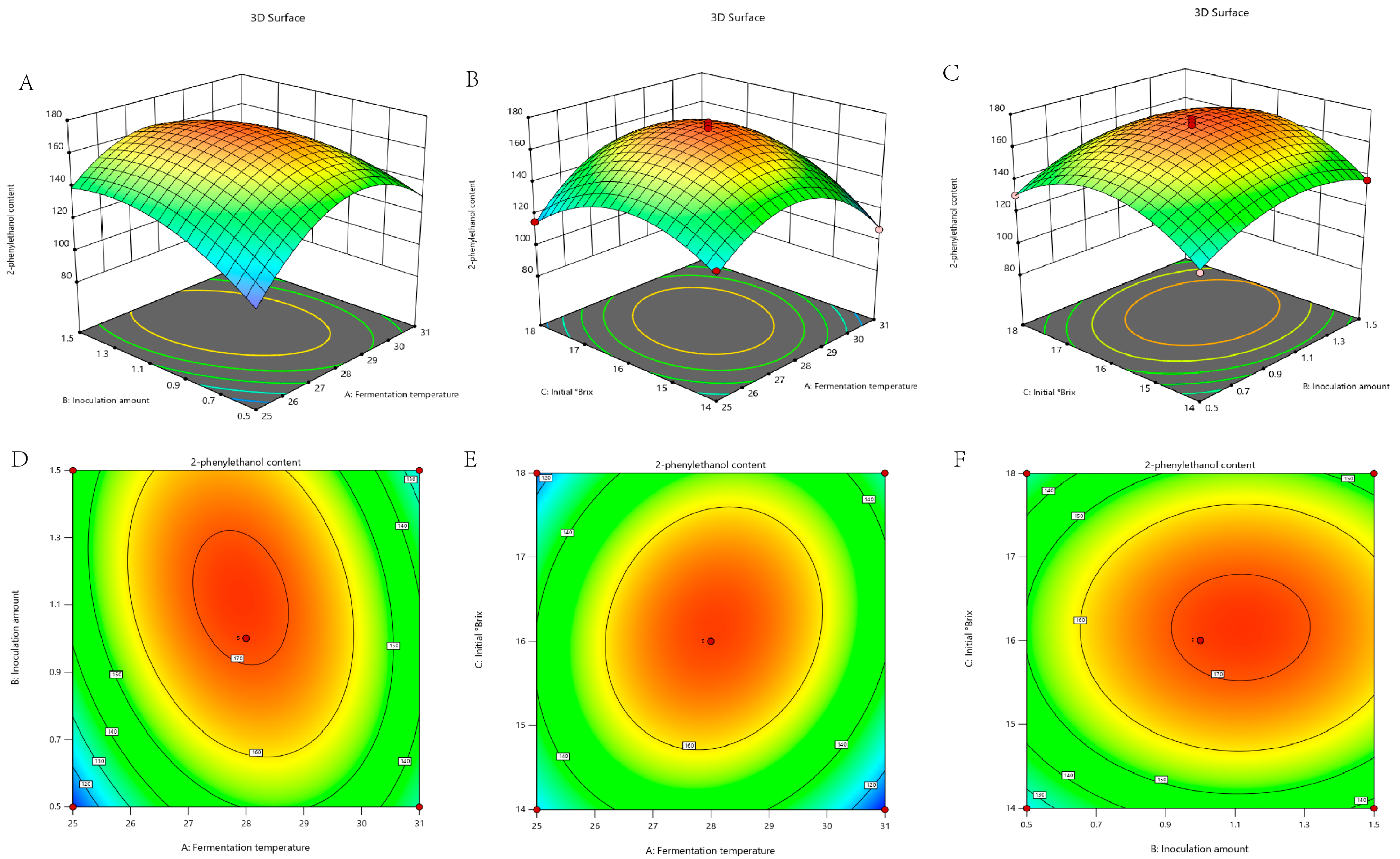
3.2. Response Surface Methodology (RSM) Optimization Experiments
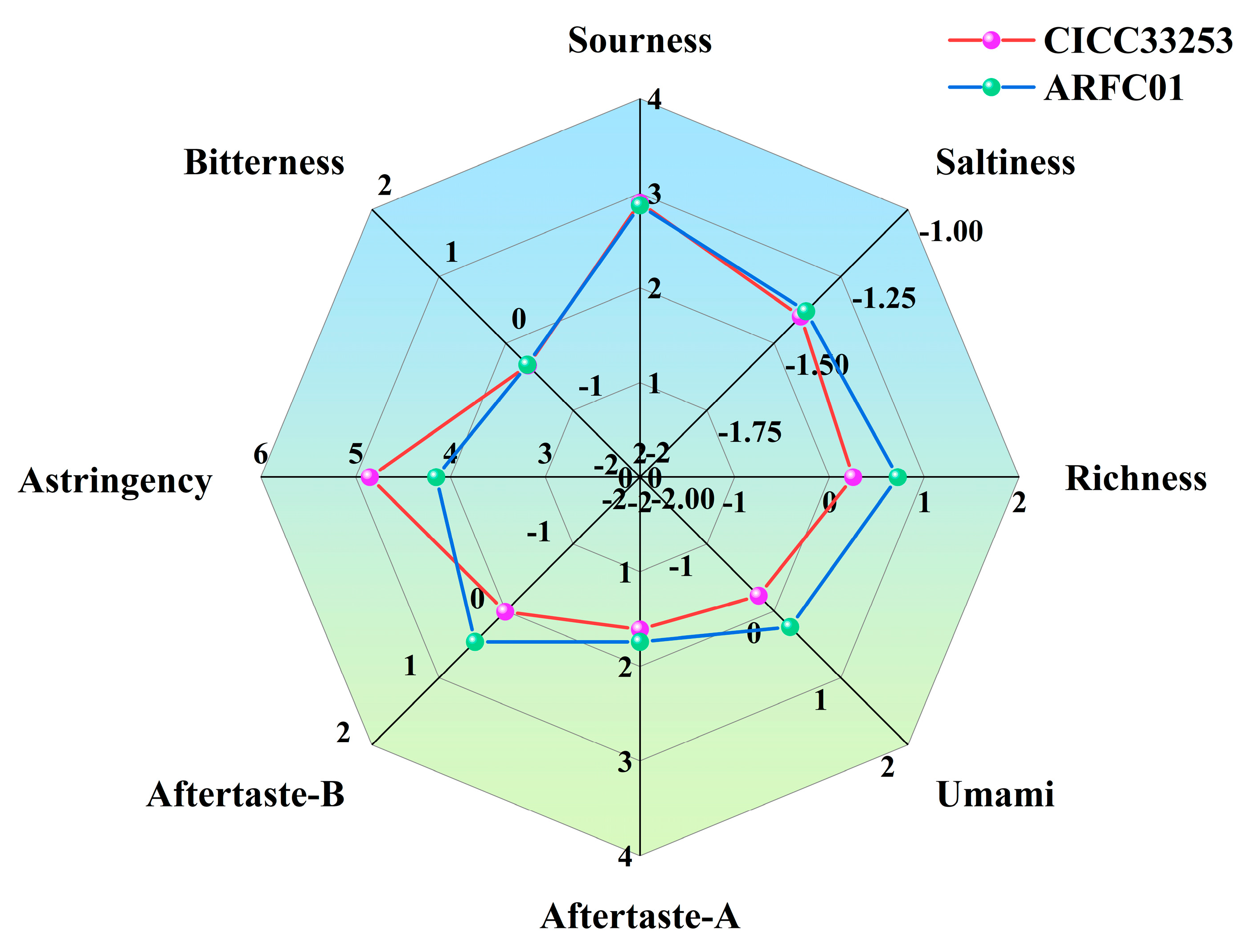
3.3. Sensory Analysis of Strawberry Wine
3.4. qRT-PCR of Analysis
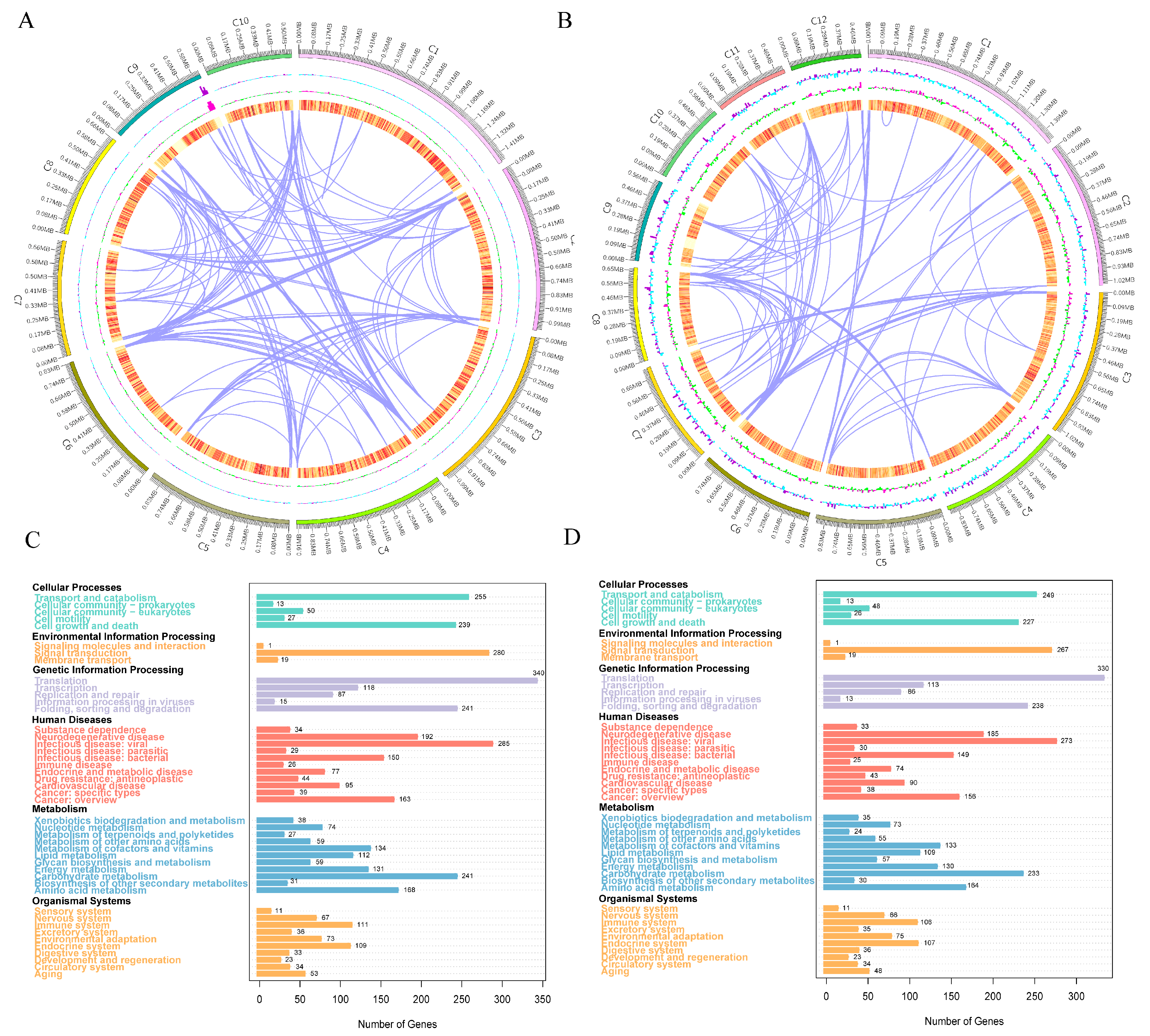
3.5. Genome Assembly and Functional Annotation Analysis
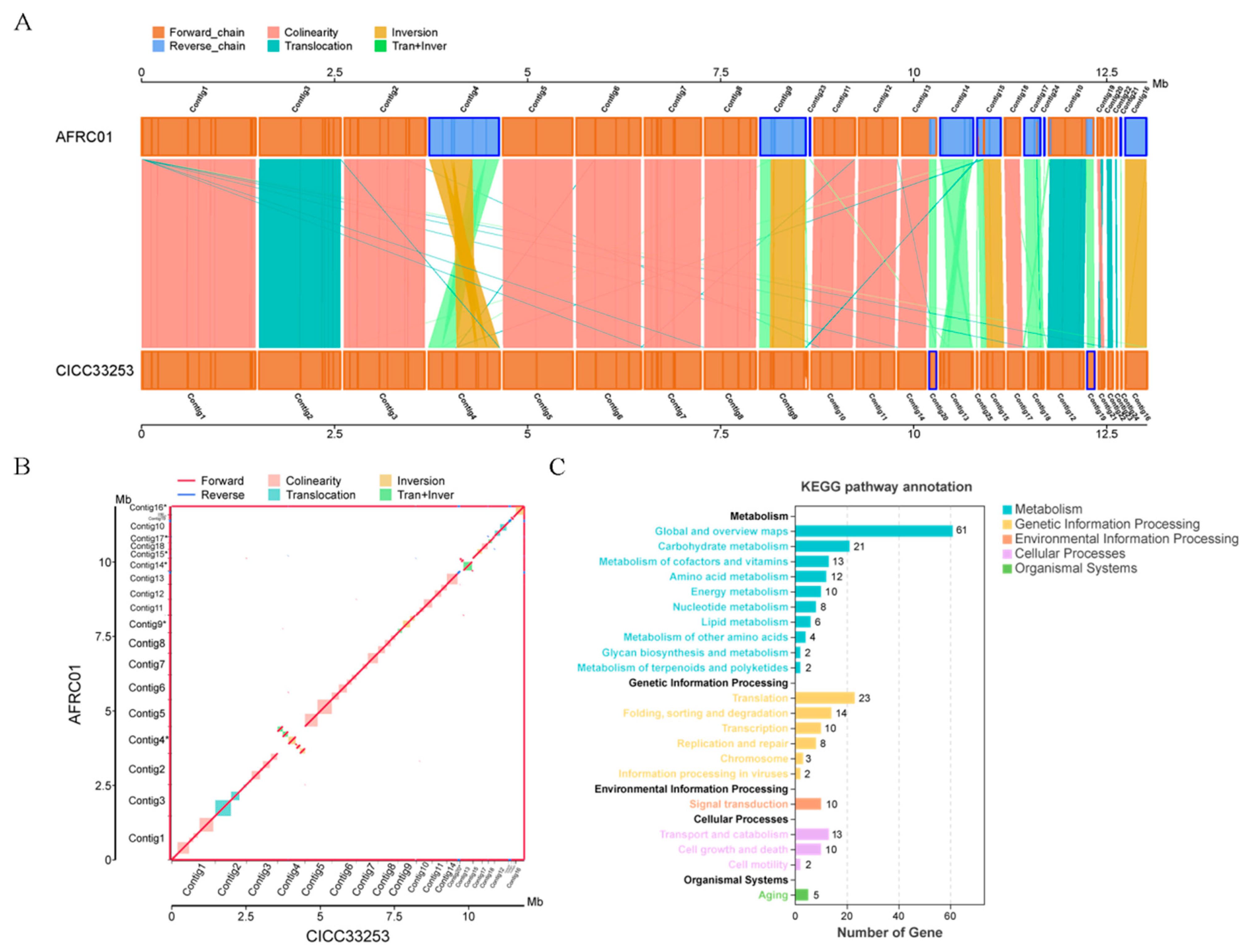
3.6. Comparative Genomics and SNP Analysis
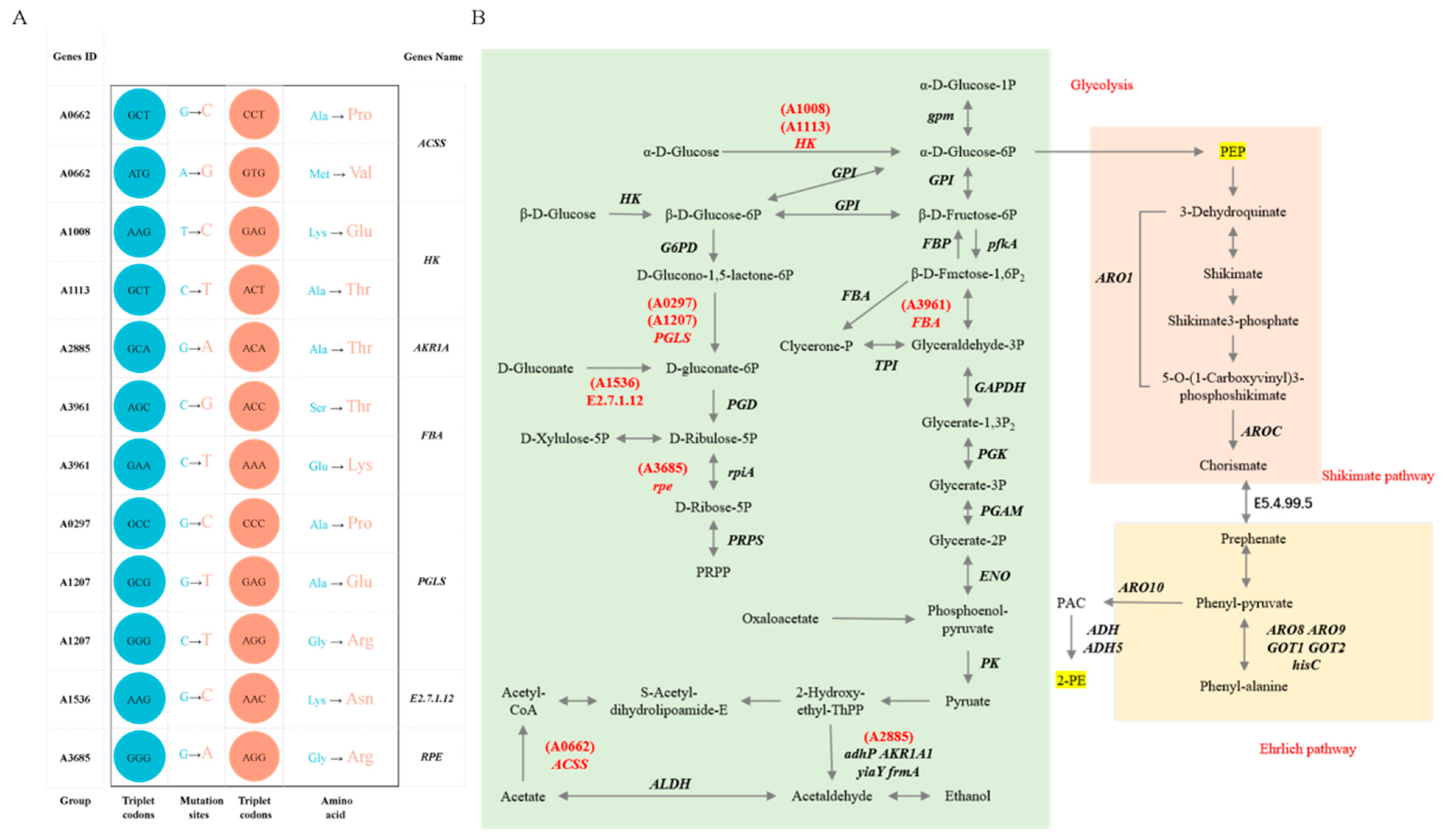
3.7. Analysis of Mutation Loci Associated with 2-Phenylethanol Metabolic Pathways
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Šic, Ž.J.; Bogdanović, S.; Voća, S.; Skendrović, B.M. Biological potential of fruit and leaves of strawberry tree (Arbutus unedo L.) from Croatia. Molecules 2020, 25, 5102. [Google Scholar] [CrossRef] [PubMed]
- Miller, K.; Feucht, W.; Schmid, M. Bioactive compounds of strawberry and blueberry and their potential health effects based on human intervention studies: A brief overview. Nutrients 2019, 11, 1510. [Google Scholar] [CrossRef] [PubMed]
- Dai, H.P.; Xue, L.; Tan, C.H.; Lei, J.J. Strawberry cultivation in Chinese solar greenhouse with three thermal walls in northern China. Acta Hortic. 2016, 1156, 569–572. [Google Scholar] [CrossRef]
- Salazar-Orbea, G.L.; García-Villalba, R.; Bernal, M.J.; Hernández, A.; Tomás-Barberán, F.A.; Sánchez-Siles, L.M. Stability of phenolic compounds in apple and strawberry: Effect of different processing techniques in industrial set up. Food Chem. 2023, 1, 134099. [Google Scholar] [CrossRef]
- Modise, D.M. Does freezing and thawing affect the volatile profile of strawberry fruit (Fragaria × ananassa Duch.). Postharvest Biol. Technol. 2008, 50, 25–30. [Google Scholar] [CrossRef]
- Giampieri, F.; Tulipani, S.; Alvarez-Suarez, J.M.; Quiles, J.L.; Mezzetti, B.; Battino, M. The strawberry: Composition, nutritional quality, and impact on human health. Nutrition 2012, 28, 9–19. [Google Scholar] [CrossRef]
- Song, Y.Y.; Zhang, Y.J.; Liu, N.; Ye, D.Q.; Gong, X.; Qin, Y.; Liu, Y.L. Volatile compounds in wild strawberry and their odorants of wild strawberry wines: Effects of different stages of fermentation. Int. J. Food Prop. 2017, 20, S399–S415. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhang, J.; Sun, Y.F.; Li, Y.Z.; Chen, R.; Jiang, C. Effects of different aging periods on physicochemical indexes and volatile components of strawberry wine. Food Ferment. Ind. 2025, 51, 158–166. (In Chinese) [Google Scholar]
- Li, N.; Wang, L.Y.; Yin, J.; Ma, N.; Tao, Y.S. Adjustment of impact odorants in Hutai-8 rose wine by co-fermentation of Pichia fermentans and Saccharomyces cerevisiae. Food Res. Int. 2022, 153, 110959. [Google Scholar] [CrossRef]
- Wilson, A.; Johnson, J.B.; Naiker, M. Biotechnological Modification of Cider Brewing Processes for the En-hanced Production of 2-Phenylethanol. Beverages 2022, 8, 64. [Google Scholar] [CrossRef]
- Celińska, E.; Kubiak, P.; Biaas, W.; Dziadas, M.; Grajek, W. Yarrowia lipolytica: The novel and promising 2-phenylethanol producer. J. Ind. Microbiol. Biotechnol. 2013, 40, 389–392. [Google Scholar] [CrossRef]
- Bagheri, B.; Bauer, F.F.; Cardinali, G.; Setati, M.E. Ecological interactions are a primary driver of population dynamics in wine yeast microbiota during fermentation. Sci. Rep. 2020, 10, 4911. [Google Scholar] [CrossRef]
- Liu, P.; Ivanova-Petropulos, V.; Duan, C.; Yan, G. Effect of unsaturated fatty acids on intra-metsabolites and aroma compounds of Saccharomyces cerevisiae in wine fermentation. Foods 2021, 10, 277. [Google Scholar] [CrossRef]
- Lambrechts, M.G.; Pretorius, I.S. Yeast and its Importance to Wine Aroma—A Review. S. Afr. J. Enol. Vitic. 2019, 21, 97–129. [Google Scholar] [CrossRef]
- Drężek, K.; Antunovics, Z.; Grabiec, A.K. Novel Saccharomyces uvarum x Saccharomyces kudriavzevii synthetic hybrid with enhanced 2-phenylethanol production. Microb. Cell Factories 2024, 23, 203. [Google Scholar] [CrossRef]
- Mei, J.; Min, H.; Lü, Z. Enhanced biotransformation of l-phenylalanine to 2-phenylethanol using an in situ product adsorption technique. Process Biochem. 2009, 44, 886–890. [Google Scholar] [CrossRef]
- Chantasuban, T.; Santomauro, F.; Gore-Lloyd, D.; Parsons, S.; Henk, D.; Scott, R.J.; Chuck, C. Elevated production of the aromatic fragrance molecule, 2-phenylethanol, using Metschnikowia pulcherrima through both de novo and ex novo conversion in batch and continuous modes. J. Chem. Technol. Biotechnol. 2018, 93, 2118–2130. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Wang, Y.; Ye, D.; Duan, L.; Duan, C.; Yan, G. Effect of the addition of branched-chain amino acids to non-limited nitrogen synthetic grape must on volatile compounds and global gene expression during alcoholic fermentation. Aust. J. Grape Wine Res. 2018, 24, 197–205. [Google Scholar] [CrossRef]
- Wilson, A.; Johnson, J.B.; Batley, R.; Lal, P.; Wakeling, L.; Naiker, M. Authentication Using Volatile Composition: A Proof-of-Concept Study on the Volatile Profiles of Fourteen Queensland Ciders. Beverages 2021, 7, 28. [Google Scholar] [CrossRef]
- Tian, S.; Liang, X.L.; Chen, J.; Zeng, W.Z.; Zhou, J.W.; Du, G.C. Enhancement of 2-phenylethanol production by a wild-type Wickerhamomyces anomalus strain isolated from rice wine. Bioresour. Technol. 2020, 318, 124257. [Google Scholar] [CrossRef]
- Wang, X.Z.; Wang, Z.G.; Feng, T. Screening of Yeast in Various Vineyard Soil and Study on Its Fla-vor Compounds from Brewing Grape Wine. Molecules 2022, 27, 512. [Google Scholar] [CrossRef]
- Ntuli, R.G.; Saltman, Y.; Ponangi, R.; Jeffery, D.W.; Bindon, K.; Wilkinson, K.L. Impact of fermentation temperature and grape solids content on the chemical composition and sensory profiles of Caber-net Sauvignon wines made from flash détente treated must fermented off-skins. Food Chem. 2022, 369, 130861. [Google Scholar] [CrossRef]
- Tian, T.T.; Yang, H.; Yang, F.; Li, B.W.; Sun, J.Y.; Wu, D.H.; Lu, J. Optimization of fermentation conditions and comparison of flavor compounds for three fermented greengage wines. LWT-Food Sci. Technol. 2018, 89, 542–550. [Google Scholar] [CrossRef]
- Debra, R.; Tormod, N.; Florian, B. Linking gene regulation and the exo-metabolome: A comparative transcriptomics approach to identify genes that impact on the production of volatile aroma com-pounds in yeast. BMC Genom. 2008, 9, 530. [Google Scholar]
- De-la-Fuente-Blanco, A.; Sáenz-Navajas, M.; Ferreira, V. On the effects of higher alcohols on red wine aroma. Food Chem. 2016, 210, 107–114. [Google Scholar] [CrossRef]
- Wang, S.; Wang, Y.; Yuan, Q.; Liu, Y.; Zhao, F.G.; Lin, Y.; Han, S.Y. Development of high methanol-tolerance Pichia pastoris based on iterative adaptive laboratory evolution. Green Chem. 2023, 25, 8845–8857. [Google Scholar] [CrossRef]
- Martino, M.E.; Bayjanov, J.R.; Caffrey, B.E.; Wels, M.; Joncour, P.; Hughes, S.; Gillet, B.; Kleerebezem, M.; van Hijum, S.A.F.; Leulier, F. Nomadic lifestyle of Lactobacillus plantarum revealed by comparative genomics of 54 strains isolated from different habitats. Environ. Microbiol. 2016, 18, 4974–4989. [Google Scholar] [CrossRef]
- Liu, S.P.; Bai, M.; Zhou, J.B.; Jin, Z.M.; Xu, Y.Z.; Yang, Q.L.; Zhou, J.D.; Zhang, S.J.; Mao, J. Analysis of genes from Saccharomyces cerevisiae HJ01 participating in aromatic alcohols biosynthesis during huangjiu fermentation. LWT-Food Sci. Technol. 2024, 154, 112705. [Google Scholar] [CrossRef]
- Kalinowski, J.; Bathe, B.; Bartels, D.; Bischoff, N.; Bott, M.; Burkovski, A.; Dusch, N.; Eggeling, L.; Eikmanns, B.J.; Gaigalat, L.; et al. The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins. J. Biotechnol. 2003, 104, 5–25. [Google Scholar] [CrossRef]
- Dong, Y.; Chen, R.; Graham, E.B.; Yu, B.; Bao, Y.Y.; Li, X.; You, X.W.; Feng, Y.Z. Eco-evolutionary strategies for relieving carbon limitation under salt stress differ across microbial clades. Nat. Commun. 2024, 15, 6013. [Google Scholar] [CrossRef] [PubMed]
- Fang, G.; Chai, L.; Zhong, X.; Lu, Z.M.; Zhang, X.J.; Wu, L.H.; Wang, S.T.; Shen, C.H.; Shi, J.S.; Xu, Z.H. Comparative Genomics Unveils the Habitat Adaptation and Metabolic Profiles of Clostridium in an Artificial Ecosystem for Liquor Production. MSystems 2022, 7, e0029722. [Google Scholar] [CrossRef]
- Li, Y.Z.; Gao, J.Y.; Wang, F.H.; Liao, Y.H. Advances in synthesis of 2-phenylethanol. Chin. J. Biotechnol. 2024, 40, 1694–1710. (In Chinese) [Google Scholar]
- Liu, L.; Zeng, W.Z.; Yu, S.Q.; Li, J.H.; Zhou, J.W. Rapid Enabling of Gluconobacter oxydans Resistance to High D-Sorbitol Concentration and High Temperature by Microdroplet-Aided Adaptive Evolution. Front. Bioeng. Biotechnol. 2021, 3, 731247. [Google Scholar] [CrossRef]
- Lan, W.; Zhang, M.; Xie, X.; Li, R.; Cheng, W.; Ma, T.T.; Zhou, Y.B. Effects of Cultivar Factors on Fermentation Characteristics and Volatile Organic Components of Strawberry Wine. Foods 2024, 13, 2874. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Hu, Z.; Hu, X.; Li, D.; Tian, S. Electronic tongue and electronic nose for food quality and safety. Food Res. Int. 2022, 162 Pt B, 112214. [Google Scholar] [CrossRef] [PubMed]
- Imam, M.; Nagpal, K. The electronic tongue: An advanced taste-sensing multichannel sensory tool with global selectivity for application in the pharmaceutical and food industry. Pharm. Dev. Technol. 2023, 28, 318–332. [Google Scholar] [CrossRef] [PubMed]
- Jolly, N.P.; Varela, C.; Pretorius, I.S. Not your ordinary yeast: Non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Res. 2014, 14, 215–237. [Google Scholar] [CrossRef]
- Wang, J.; Jian, X.; Xing, X.H.; Zhang, C.; Fei, Q. Empowering a Methanol-Dependent Escherichia coli via Adaptive Evolution Using a High-Throughput Microbial Microdroplet Culture System. Front. Bioeng. Biotechnol. 2020, 8, 570. [Google Scholar] [CrossRef]
- Trick, A.Y.; Chen, F.E.; Schares, J.A.; Freml, B.E.; Lor, P.; Yun, Y.; Wang, T.H. High resolution estimates of relative gene abundance with quantitative ratiometric regression PCR (qRR-PCR). Analyst 2021, 146, 6463–6469. [Google Scholar] [CrossRef]
- Zhang, C.; Fan, T.; Wang, Z.; Yu, J.; Guo, X.; Jiang, W.; Miao, L.; Yang, H. Screening of High-Yield 2-Phenylethanol Producing Strain from Wild-Type Saccharomyces cerevisiae and Optimization of Fermentation Parameters. Foods 2025, 14, 2444. [Google Scholar] [CrossRef]
- Bezerra, M.; Ribeiro, M.; Cosme, F.; Nunes, F.M. Overview of the Distinctive Characteristics of Strawberry, Raspberry, and Blueberry in Berries, Berry Wines, and Berry Spirits. Compr. Rev. Food Sci. Food Saf. 2024, 23, e13354. [Google Scholar] [CrossRef]
- Qaderi, R.; Mezzetti, B.; Capocasa, F.; Mazzoni, L. Stability of Strawberry Fruit (Fragaria x ananassa Duch.) Nutritional Quality at Different Storage Conditions. Appl. Sci. 2023, 13, 313. [Google Scholar] [CrossRef]
- Ostrowska-Ligęza, E.; Szulc, K.; Jakubczyk, E.; Dolatowska-Żebrowska, K.; Wirkowska-Wojdyła, M.; Bryś, J.; Górska, A. The Influence of a Chocolate Coating on the State Diagrams and Thermal Behaviour of Freeze-Dried Strawberries. Appl. Sci. 2022, 12, 1342. [Google Scholar] [CrossRef]
- Cakar, U.; Petrovic, A.; Colovic, M.; Krstic, D.; Dordevic, B. Strawberry Wine Antioxidant Properties in The Protection Against Free Radicals. Free Radic. Biol. Med. 2021, 8, 157. [Google Scholar]
- Kumar, S.; Kumar, J.; Tripathi, J.; Gupta, S.; Gautam, S. Secondary product from strawberry (Fragaria ananassa) fruit for extended preservation and value addition. J. Food Sci. Technol. 2022, 59, 1598–1609. [Google Scholar] [CrossRef]
- Merlino, V.M.; Fracassetti, D.; Di Canito, A.; Pizzi, S.; Borra, D.; Giuggioli, N.R.; Vigentini, I. Is the consumer ready for innovative fruit wines? Perception and acceptability of young consumers. Foods 2021, 10, 1545. [Google Scholar] [CrossRef]
- Thompson, B.; Machas, M.; Nielsen, D.R. Creating pathways towards aromatic building blocks and fine chemicals. Curr. Opin. Biotechnol. 2015, 36, 1–7. [Google Scholar] [CrossRef]
- Huang, Y.H.; Yang, S.T.; Zhou, Z.Q.; Xiong, J.; Chen, H.; Liu, Y.H.; Jiang, X.T. Study on synthesis of 2-phenylethanol by microbes and plants. J. Food Sci. Technol. 2014, 32, 41–44. (In Chinese) [Google Scholar]
- Peng, B.Z.; Li, F.L.; Cui, L.; Guo, Y.D. Effects of Fermentation Temperature on Key Aroma Compounds and Sensory Properties of Apple Wine. J. Food Sci. 2015, 80, S2937–S2943. [Google Scholar] [CrossRef] [PubMed]
- Yao, F.; Wu, P.; Liu, P.; Wang, L.X. Optimization of fermentation conditions and antioxidant activity analysis of Aronia melanocarpa wine. China Brew 2023, 42, 206–211. (In Chinese) [Google Scholar]
- Aldrete-tapia, J.A.; Escalante-Minakata, P.; Mirandacastilleja, D.E.; Hernández-Iturriaga, M. Fermentation conditions for yeast selection and effect of yeast-bacterial interaction in developing a starter culture for tequila fermentation. J. Food Sci. 2022, 87, 5089–5098. [Google Scholar] [CrossRef]
- Liu, Z.X.; Wang, W.X.; Shangguan, L.L.; Chen, X.; Dai, J. Advances in the 2-phenylethanol tolerance of yeast. Acta Microbiol. Sin. 2024, 64, 981–998. (In Chinese) [Google Scholar]
- Wu, F.L.; Cao, P.; Song, G.T.; Chen, W.J.; Wang, Q.H. Expanding the repertoire of aromatic chemicals by microbial production. J. Chem. Technol. Biot. 2018, 93, 2804–2816. [Google Scholar] [CrossRef]
- Xu, H.; Li, Z.; Li, L.Y.; Xie, X.M.; Cai, D.B.; Wang, Z.; Zhan, Y.Y.; Chen, S.W. Sustainable production of 2-phenylethanol from agro-industrial wastes by metabolically engineered Bacillus licheniformis. LWT 2023, 173, 114414. [Google Scholar] [CrossRef]
- Wang, G.L.; Wang, M.Y.; Yang, J.C.; Li, Q.; Zhu, N.Q.; Liu, L.X.; Hu, X.M.; Yang, X.P. De novo synthesis of 2-phenylethanol from glucose by metabolically engineered Escherichia coli. J. Ind. Microbiol. Biotechnol. 2023, 49, kuac026. [Google Scholar] [CrossRef]
- Hassing, E.J.; de Groot, P.A.; Marquenie, V.R.; Pronk, J.T.; Daran, J.G. Connecting central carbon andaromatic amino acid metabolisms to improve de novo 2-phenylethanol production in Saccharomyces cerevisiae. Metab. Eng. 2019, 56, 165–180. [Google Scholar] [CrossRef]
- Fukuda, N.; Honda, S.; Fujiwara, M.; Yoshimura, Y.; Nakamura, T. Polyploid engineering by increasing mutant gene dosage in yeasts. Microb. Biotechnol. 2021, 14, 979–992. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, D.; Yang, Z.; Wang, X.; Xu, M.; Zhang, X.; Wu, T.; Wu, W.; Zheng, S.; Li, Y.; He, F.; et al. Fermentation Process Optimization of Strawberry Wine Using the 2-Phenylethanol Tolerant Saccharomyces cerevisiae AFRC01 and Comparative Genomic Analysis. Foods 2025, 14, 3043. https://doi.org/10.3390/foods14173043
Liu D, Yang Z, Wang X, Xu M, Zhang X, Wu T, Wu W, Zheng S, Li Y, He F, et al. Fermentation Process Optimization of Strawberry Wine Using the 2-Phenylethanol Tolerant Saccharomyces cerevisiae AFRC01 and Comparative Genomic Analysis. Foods. 2025; 14(17):3043. https://doi.org/10.3390/foods14173043
Chicago/Turabian StyleLiu, Dan, Zelin Yang, Xue Wang, Meng Xu, Xinqi Zhang, Tong Wu, Wenhui Wu, Shu Zheng, Yan Li, Fumeng He, and et al. 2025. "Fermentation Process Optimization of Strawberry Wine Using the 2-Phenylethanol Tolerant Saccharomyces cerevisiae AFRC01 and Comparative Genomic Analysis" Foods 14, no. 17: 3043. https://doi.org/10.3390/foods14173043
APA StyleLiu, D., Yang, Z., Wang, X., Xu, M., Zhang, X., Wu, T., Wu, W., Zheng, S., Li, Y., He, F., Xu, Y., Li, F., & Lan, W. (2025). Fermentation Process Optimization of Strawberry Wine Using the 2-Phenylethanol Tolerant Saccharomyces cerevisiae AFRC01 and Comparative Genomic Analysis. Foods, 14(17), 3043. https://doi.org/10.3390/foods14173043




