Allergens in Food: Analytical LC-MS/MS Method for the Qualitative Detection of Pistacia vera
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Reagents and Chemicals
2.3. Sample Preparation
2.4. Instrumental Conditions for LC-MS/MS Analysis
2.5. LC-MS/MS Method Optimization and Validation
2.6. PCR and ELISA Analysis
3. Results
3.1. LC-MS/MS Validation
3.2. Applicability of the Method for Routine Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| LC-MS/MS | Liquid Chromatography–Tandem Mass Spectrometry |
| CRMs | Certified Reference Materials |
| SRM | Single Reaction Monitoring |
| TFA | Trifluoroacetic acid |
| ACN | Acetonitrile |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| PCR | Polymerase Chain Reaction |
| TOF | Time-Of-Flight |
| QqQ | Triple Quadrupole |
| tR | Retention Time |
| PT | Proficiency Testing |
| CV% | Coefficient of Variation |
| SD | Standard Deviation |
| SDL | Screening Detection Limit |
| S/N | Signal-to-Noise |
| HESI | Heated Electrospray Ionization |
| HPLC | High-Performance Liquid Chromatography |
| PAL | Precautionary Allergen Labelling |
| MRM3 | Multiple Reaction Monitoring Cubed |
| IgE | Immunoglobulin E |
References
- Mendes, C.; Costa, J.; Vicente, A.A.; Oliveira, M.B.P.P.; Mafra, I. Cashew Nut Allergy: Clinical Relevance and Allergen Characterisation. Clin. Rev. Allergy Immunol. 2019, 57, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Costa, J.; Silva, I.; Vicente, A.A.; Oliveira, M.B.P.P.; Mafra, I. Pistachio Nut Allergy: An Updated Overview. Crit. Rev. Food Sci. Nutr. 2019, 59, 546–562. [Google Scholar] [CrossRef] [PubMed]
- USDA-ARS. Germplasm Resources Information Network (GRIN). Available online: https://www.ars-grin.gov/ (accessed on 11 June 2025).
- Ak, B.E.; Karadag, S.; Sakar, E. Pistachio Production and Industry in Turkey: Current Status and Future Perspective. Options Méditerranéennes Ser. A Mediterr. Semin. 2016, 118, 115–118. [Google Scholar]
- Zhao, Y.; Sun, X.; Marquis, C.P.; Lee, N.A. Development of a Sensitive Sandwich ELISA Specific to 2S Albumin (Ana o 3) as a Stable Protein Marker for Cashew Nut Residue Detection in Pre-Packaged Food Products. Food Control 2019, 96, 432–440. [Google Scholar] [CrossRef]
- Costa, J.; Bavaro, S.L.; Benedé, S.; Diaz-Perales, A.; Bueno-Diaz, C.; Gelencser, E.; Klueber, J.; Larré, C.; Lozano-Ojalvo, D.; Lupi, R.; et al. Are Physicochemical Properties Shaping the Allergenic Potency of Plant Allergens? Clin. Rev. Allergy Immunol. 2020, 62, 37–63. [Google Scholar] [CrossRef]
- Gaskin, F.E.; Taylor, S.L. Sandwich Enzyme-Linked Immunosorbent Assay (ELISA) for Detection of Cashew Nut in Foods. J. Food Sci. 2011, 76, T218–T226. [Google Scholar] [CrossRef]
- Linacero, R.; Sanchiz, A.; Ballesteros, I.; Cuadrado, C. Application of Real-Time PCR for Tree Nut Allergen Detection in Processed Foods. Crit. Rev. Food Sci. Nutr. 2020, 60, 1077–1093. [Google Scholar] [CrossRef] [PubMed]
- Elizur, A.; Appel, M.Y.; Nachshon, L.; Levy, M.B.; Epstein-Rigbi, N.; Golobov, K.; Goldberg, M.R. NUT Co Reactivity–ACquiring Knowledge for Elimination Recommendations (NUT CRACKER) Study. Allergy Eur. J. Allergy Clin. Immunol. 2018, 73, 593–601. [Google Scholar] [CrossRef]
- Sicherer, S.H.; Noone, S.A.; Muñoz-Furlong, A. The Impact of Childhood Food Allergy on Quality of Life. Ann. Allergy Asthma Immunol. 2001, 87, 461–464. [Google Scholar] [CrossRef]
- Relazione Annuale 2004 | EFSA. Available online: https://www.efsa.europa.eu/it/corporate/pub/ar04 (accessed on 14 April 2025).
- Luyt, D.K.; Vaughan, D.; Oyewole, E.; Stiefel, G. Ethnic Differences in Prevalence of Cashew Nut, Pistachio Nut and Almond Allergy. Pediatr. Allergy Immunol. 2016, 27, 651–654. [Google Scholar] [CrossRef]
- Kuźmiński, A.; Przybyszewski, M.; Przybyszewska, J.; Ukleja-Sokołowska, N.; Pałgan, K.; Bartuzi, Z. Tree Nut Allergy. Postep. Dermatol. Alergol. 2021, 38, 544–549. [Google Scholar] [CrossRef]
- Regulation–178/2002–EN–EUR-Lex. Available online: https://eur-lex.europa.eu/eli/reg/2002/178/oj/eng (accessed on 16 July 2025).
- Regulation–1169/2011–EN–Food Information to Consumers Regulation–EUR-Lex. Available online: https://eur-lex.europa.eu/eli/reg/2011/1169/oj/eng (accessed on 16 July 2025).
- Soon, J.M.; Manning, L. “May Contain” Allergen Statements: Facilitating or Frustrating Consumers? J. Consum. Policy 2017, 40, 447–472. [Google Scholar] [CrossRef]
- Scharf, A.; Kasel, U.; Wichmann, G.; Besler, M. Performance of ELISA and PCR Methods for the Determination of Allergens in Food: An Evaluation of Six Years of Proficiency Testing for Soy (Glycine max L.) and Wheat Gluten (Triticum aestivum L.). J. Agric. Food Chem. 2013, 61, 10261–10272. [Google Scholar] [CrossRef]
- Stephan, O.; Vieths, S. Development of a Real-Time PCR and a Sandwich ELISA for Detection of Potentially Allergenic Trace Amounts of Peanut (Arachis Hypogaea) in Processed Foods. J. Agric. Food Chem. 2004, 52, 3754–3760. [Google Scholar] [CrossRef] [PubMed]
- Scaravelli, E.; Brohée, M.; Marchelli, R.; Van Hengel, A.J. The Effect of Heat Treatment on the Detection of Peanut Allergens as Determined by ELISA and Real-Time PCR. Anal. Bioanal. Chem. 2009, 395, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Mari, A.; Rasi, C.; Palazzo, P.; Scala, E. Allergen Databases: Current Status and Perspectives. Curr. Allergy Asthma Rep. 2009, 9, 376–383. [Google Scholar] [CrossRef]
- Poms, R.E.; Klein, C.L.; Anklam, E. Methods for Allergen Analysis in Food: A Review. Food Addit. Contam. 2004, 21, 1–31. [Google Scholar] [CrossRef]
- Midun, E.; Radulovic, S.; Brough, H.; Caubet, J.C. Recent Advances in the Management of Nut Allergy. World Allergy Organ. J. 2021, 14, 100491. [Google Scholar] [CrossRef] [PubMed]
- Van Der Valk, J.P.M.; Dubois, A.E.J.; Gerth Van Wijk, R.; Wichers, H.J.; De Jong, N.W. Systematic Review on Cashew Nut Allergy. Allergy Eur. J. Allergy Clin. Immunol. 2014, 69, 692–698. [Google Scholar] [CrossRef]
- Wang, F.; Robotham, J.M.; Teuber, S.S.; Sathe, S.K.; Roux, K.H. Ana o 2, a Major Cashew (Anacardium occidentale L.) Nut Allergen of the Legumin Family. Int. Arch. Allergy Immunol. 2003, 132, 27–39. [Google Scholar] [CrossRef]
- Robotham, J.M.; Wang, F.; Seamon, V.; Teuber, S.S.; Sathe, S.K.; Sampson, H.A.; Beyer, K.; Seavy, M.; Roux, K.H. Ana o 3, an Important Cashew Nut (Anacardium occidentale L.) Allergen of the 2S Albumin Family. J. Allergy Clin. Immunol. 2005, 115, 1284–1290. [Google Scholar] [CrossRef] [PubMed]
- Careri, M.; Costa, A.; Elviri, L.; Lagos, J.B.; Mangia, A.; Terenghi, M.; Cereti, A.; Garoffo, L.P. Use of Specific Peptide Biomarkers for Quantitative Confirmation of Hidden Allergenic Peanut Proteins Ara h 2 and Ara h 3/4 for Food Control by Liquid Chromatography-Tandem Mass Spectrometry. Anal. Bioanal. Chem. 2007, 389, 1901–1907. [Google Scholar] [CrossRef]
- Posada-Ayala, M.; Alvarez-Llamas, G.; Maroto, A.S.; Maes, X.; Muñoz-Garcia, E.; Villalba, M.; Rodríguez, R.; Perez-Gordo, M.; Vivanco, F.; Pastor-Vargas, C.; et al. Novel Liquid Chromatography–Mass Spectrometry Method for Sensitive Determination of the Mustard Allergen Sin a 1 in Food. Food Chem. 2015, 183, 58–63. [Google Scholar] [CrossRef]
- Bianco, M.; Ventura, G.; Calvano, C.D.; Losito, I.; Cataldi, T.R.I. Food Allergen Detection by Mass Spectrometry: From Common to Novel Protein Ingredients. Proteomics 2023, 23, e2200427. [Google Scholar] [CrossRef]
- Spolidoro, G.C.I.; Lisik, D.; Nyassi, S.; Ioannidou, A.; Ali, M.M.; Amera, Y.T.; Rovner, G.; Khaleva, E.; Venter, C.; van Ree, R.; et al. Prevalence of Tree Nut Allergy in Europe: A Systematic Review and Meta-Analysis. Allergy Eur. J. Allergy Clin. Immunol. 2024, 79, 302–323. [Google Scholar] [CrossRef] [PubMed]
- Van Hengel, A.J. Food Allergen Detection Methods and the Challenge to Protect Food-Allergic Consumers. Anal. Bioanal. Chem. 2007, 389, 111–118. [Google Scholar] [CrossRef]
- Andjelkovic, U.; Martinovic, T.; Josic, D. Foodomic Investigations of Food Allergies. Curr. Opin. Food Sci. 2015, 4, 92–98. [Google Scholar] [CrossRef]
- Picariello, G.; Mamone, G.; Addeo, F.; Ferranti, P. The Frontiers of Mass Spectrometry-Based Techniques in Food Allergenomics. J. Chromatogr. A 2011, 1218, 7386–7398. [Google Scholar] [CrossRef] [PubMed]
- Cho, C.Y.; Nowatzke, W.; Oliver, K.; Garber, E.A.E. Multiplex Detection of Food Allergens and Gluten. Anal. Bioanal. Chem. 2015, 407, 4195–4206. [Google Scholar] [CrossRef]
- Ben Rejeb, S.; Abbott, M.; Davies, D.; Cléroux, C.; Delahaut, P. Multi-Allergen Screening Immunoassay for the Detection of Protein Markers of Peanut and Four Tree Nuts in Chocolate. Food Addit. Contam. 2005, 22, 709–715. [Google Scholar] [CrossRef]
- Ladenburger, E.M.; Dehmer, M.; Grünberg, R.; Waiblinger, H.U.; Stoll, D.; Bergemann, J. Highly Sensitive Matrix-Independent Quantification of Major Food Allergens Peanut and Soy by Competitive Real-Time PCR Targeting Mitochondrial DNA. J. AOAC Int. 2018, 101, 170–184. [Google Scholar] [CrossRef] [PubMed]
- The Probability of Obtaining: A Correct and Representative Result in Allergen Analysis. Available online: https://www.researchgate.net/publication/285150237_The_probability_of_obtaining_A_correct_and_representative_result_in_allergen_analysis (accessed on 21 July 2025).
- Brun, V.; Masselon, C.; Garin, J.; Dupuis, A. Isotope Dilution Strategies for Absolute Quantitative Proteomics. J. Proteom. 2009, 72, 740–749. [Google Scholar] [CrossRef]
- Ahsan, N.; Rao, R.S.P.; Gruppuso, P.A.; Ramratnam, B.; Salomon, A.R. Targeted Proteomics: Current Status and Future Perspectives for Quantification of Food Allergens. J. Proteom. 2016, 143, 15. [Google Scholar] [CrossRef]
- Planque, M.; Arnould, T.; Gillard, N.; Planque, M.; Arnould, T.; Gillard, N. Food Allergen Analysis: Detection, Quantification and Validation by Mass Spectrometry. Allergen 2017, 2, 1–38. [Google Scholar] [CrossRef]
- Planque, M.; Arnould, T.; Renard, P.; Delahaut, P.; Dieu, M.; Gillard, N. Highlight on Bottlenecks in Food Allergen Analysis: Detection and Quantification by Mass Spectrometry. J. AOAC Int. 2017, 100, 1126–1130. [Google Scholar] [CrossRef] [PubMed][Green Version]
- James, C.A.; Welham, S.; Rose, P. Evaluation of Food Allergen Information, Labelling and Unintended Food Allergen Presence in Imported Prepacked Foods and Drinks Purchased Online in the UK. Food Control 2024, 162, 110462. [Google Scholar] [CrossRef]
- Sealey-Voyksner, J.; Zweigenbaum, J.; Voyksner, R. Discovery of Highly Conserved Unique Peanut and Tree Nut Peptides by LC-MS/MS for Multi-Allergen Detection. Food Chem. 2016, 194, 201–211. [Google Scholar] [CrossRef]
- Sun New, L.; Schreiber, A.; Stahl-Zeng, J.; Liu, H.F. Simultaneous Analysis of Multiple Allergens in Food Products by LC-MS/MS. J. AOAC Int. 2018, 101, 132–145. [Google Scholar] [CrossRef] [PubMed]
- Luparelli, A.; Losito, I.; De Angelis, E.; Pilolli, R.; Monaci, L. Multi-Target Detection of Nuts and Peanuts as Hidden Allergens in Bakery Products through Bottom-Up Proteomics and High-Resolution Mass Spectrometry. Foods 2023, 12, 726. [Google Scholar] [CrossRef]
- Mattarozzi, M.; Bignardi, C.; Elviri, L.; Careri, M. Rapid Shotgun Proteomic Liquid Chromatography-Electrospray Ionization-Tandem Mass Spectrometry-Based Method for the Lupin (Lupinus albus L.) Multi-Allergen Determination in Foods. J. Agric. Food Chem. 2012, 60, 5841–5846. [Google Scholar] [CrossRef]
- EURL Guidance Document on the Quality Control During Routine Analysis (Ongoing Method Performance Verification). Version 1.2 (3 April 2023). Available online: https://eurl-residues.eu/wp-content/uploads/2023/04/Guidance_On-going-performance-verification_v1_2_20230403.pdf (accessed on 1 August 2025).
- European EURL Guidance Document on Screening Method Validation. Version 1.1, 21 September 2023. Available online: https://sitesv2.anses.fr/en/system/files/Guidance_screening_20230921_v1_0.pdf (accessed on 1 August 2025).
- EUR-Lex–02021R0808-20250217–EN–EUR-Lex. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX%3A02021R0808-20250217 (accessed on 31 July 2025).
- Karageorgou, E.; Samanidou, V. Youden Test Application in Robustness Assays during Method Validation. J. Chromatogr. A 2014, 1353, 131–139. [Google Scholar] [CrossRef]
- Andorf, S.; Borres, M.P.; Block, W.; Tupa, D.; Bollyky, J.B.; Sampath, V.; Elizur, A.; Lidholm, J.; Jones, J.E.; Galli, S.J.; et al. Association of Clinical Reactivity with Sensitization to Allergen Components in Multifood-Allergic Children. J. Allergy Clin. Immunol. Pr. 2017, 5, 1325–1334.e4. [Google Scholar] [CrossRef] [PubMed]
- Maloney, J.M.; Rudengren, M.; Ahlstedt, S.; Bock, S.A.; Sampson, H.A. The Use of Serum-Specific IgE Measurements for the Diagnosis of Peanut, Tree Nut, and Seed Allergy. J. Allergy Clin. Immunol. 2008, 122, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Noorbakhsh, R.; Mortazavi, S.A.; Sankian, M.; Shahidi, F.; Tehrani, M.; Azad, F.J.; Behmanesh, F.; Varasteh, A.R. Pistachio Allergy-Prevalence and In Vitro Cross-Reactivity with Other Nuts. Allergol. Int. 2011, 60, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Bastiaan-Net, S.; Reitsma, M.; Cordewener, J.H.G.; Van Der Valk, J.P.M.; America, T.A.H.P.; Dubois, A.E.J.; Gerth Van Wijk, R.; Savelkoul, H.F.J.; De Jong, N.W.; Wichers, H.J. IgE Cross-Reactivity of Cashew Nut Allergens. Int. Arch. Allergy Immunol. 2019, 178, 19–32. [Google Scholar] [CrossRef]
- Korte, R.; Brockmeyer, J. MRM3-Based LC-MS Multi-Method for the Detection and Quantification of Nut Allergens. Anal. Bioanal. Chem. 2016, 408, 7845–7855. [Google Scholar] [CrossRef]
- Toda, M.; Hellwig, M.; Henle, T.; Vieths, S. Influence of the Maillard Reaction on the Allergenicity of Food Proteins and the Development of Allergic Inflammation. Curr. Allergy Asthma Rep. 2019, 19, 4. [Google Scholar] [CrossRef]
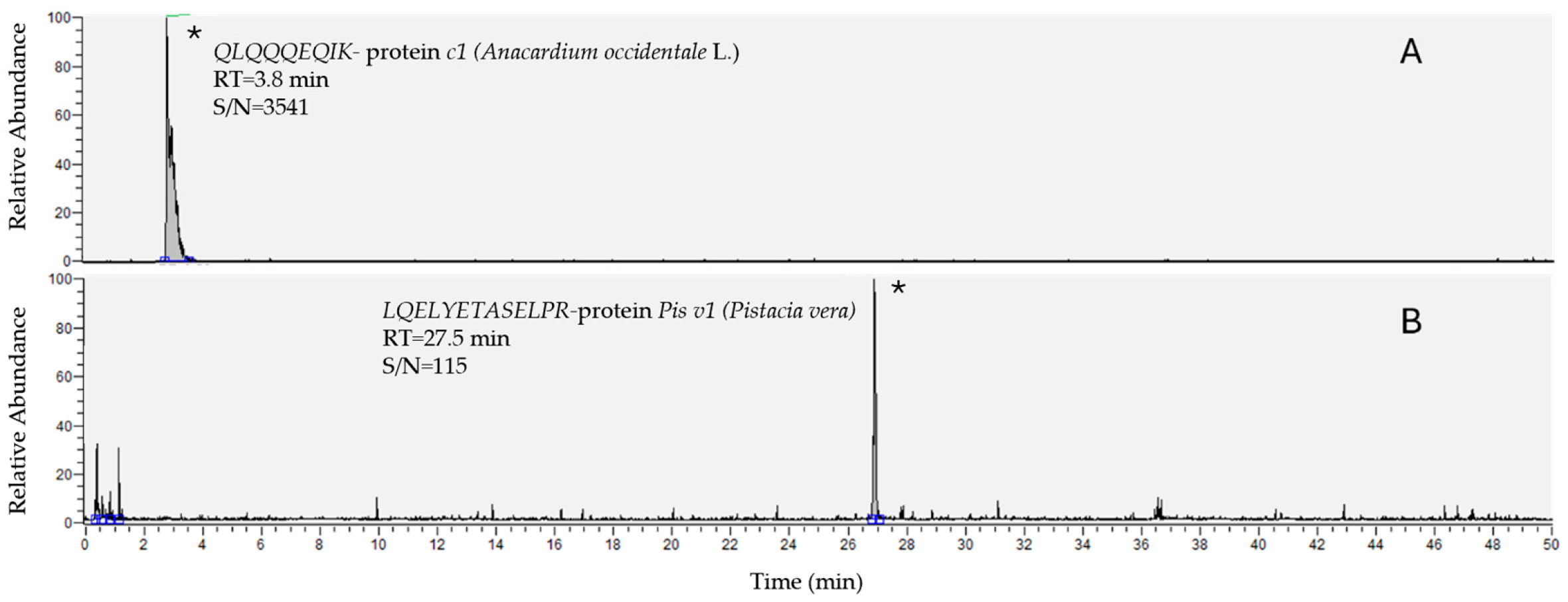
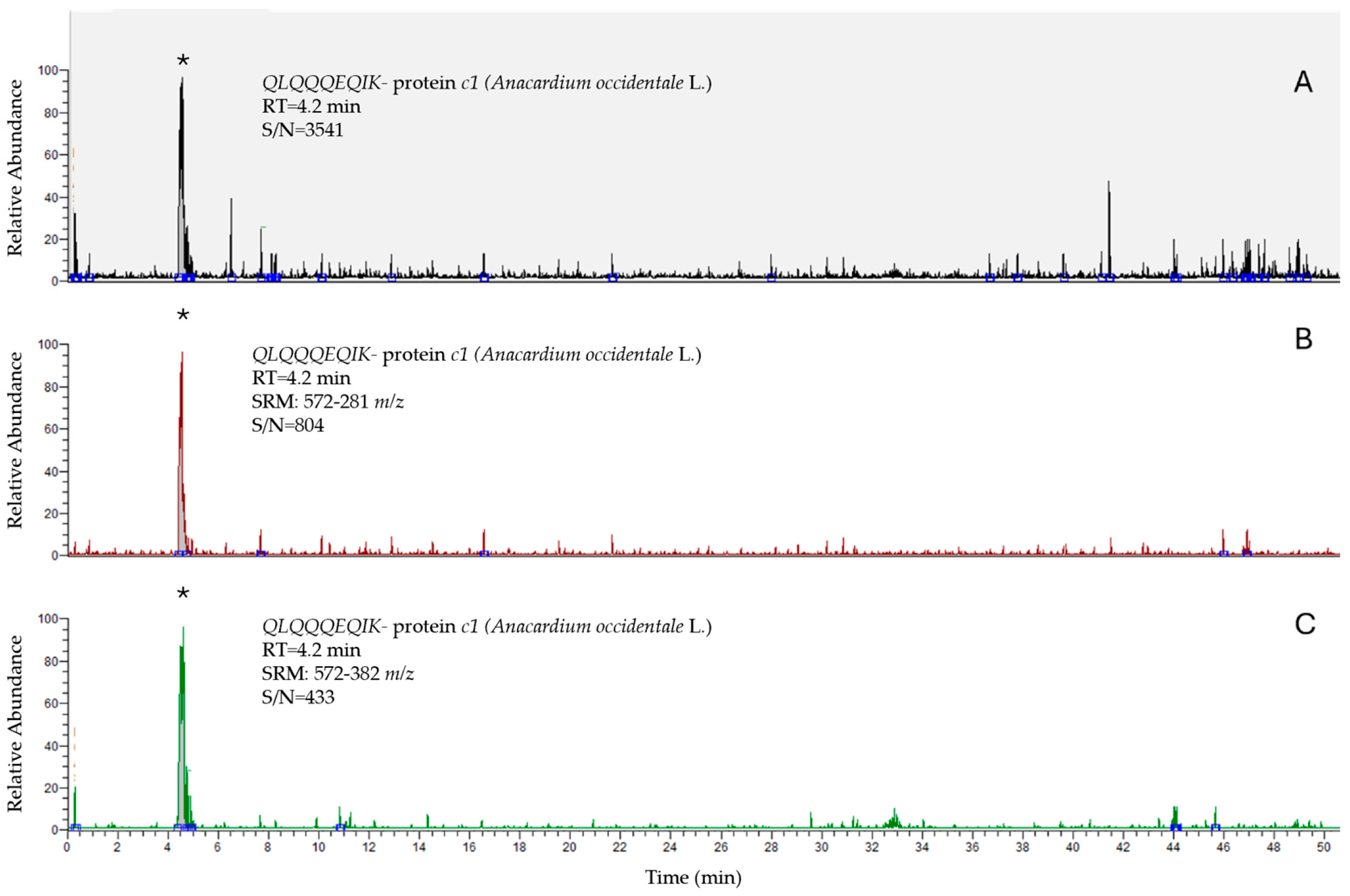
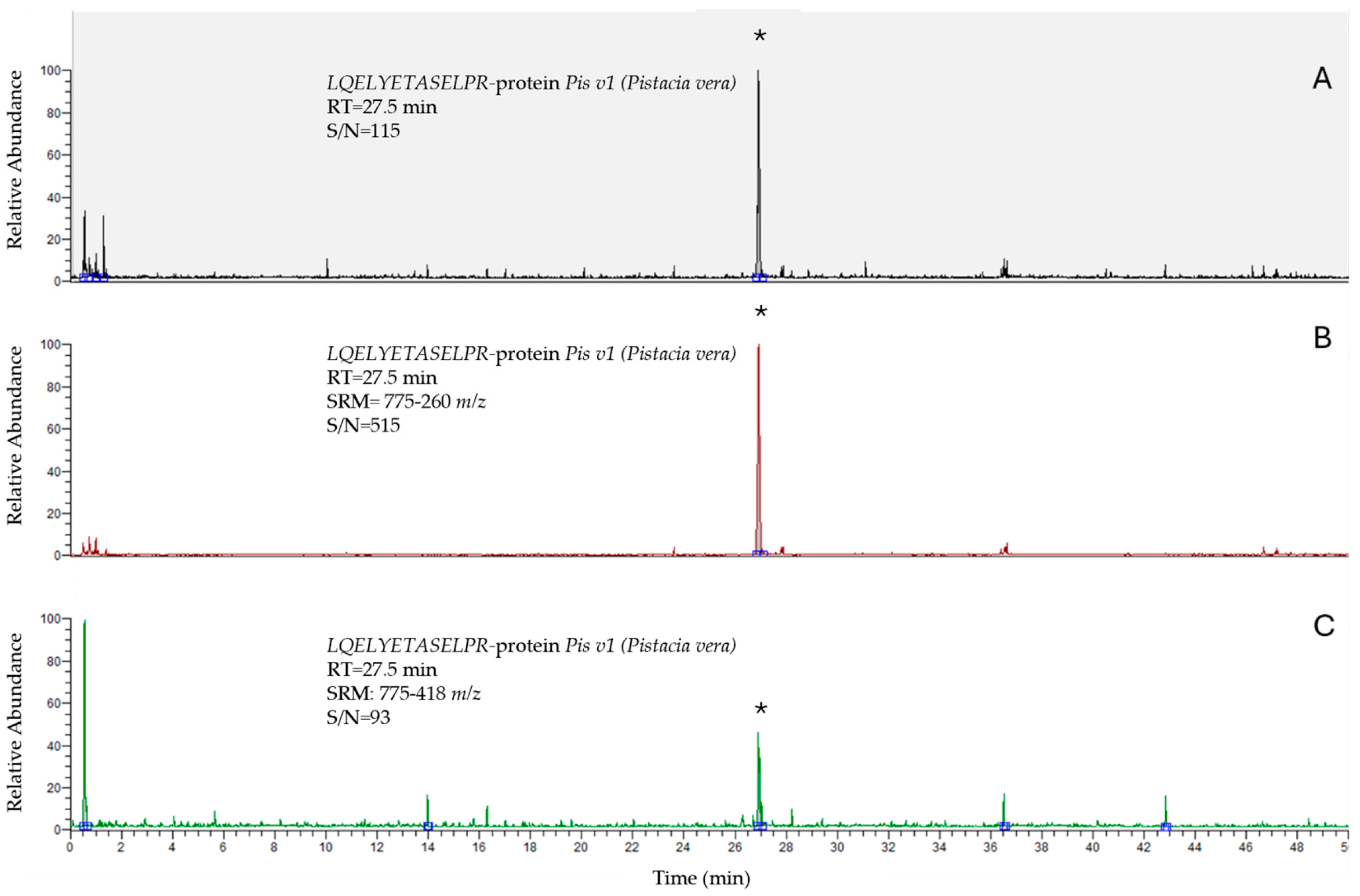
| Peptide | Polarity | Precursor Ion (m/z) | Product Ion (m/z) | Collision Energy (V) | S-Lens (V) |
|---|---|---|---|---|---|
| QLQQQEQIK | + | 572 | 281 | 53 | 125 |
| 572 | 382 | 34 | 125 | ||
| 572 | 563 | 20 | 136 | ||
| LQELYETASELPR | + | 775 | 260 | 50 | 120 |
| 775 | 418 | 47 | 120 | ||
| 775 | 542 | 17 | 120 |
| Parameters | Values | Value Ranges |
|---|---|---|
| Centrifugation force | 7871 g | 6375–9524 g |
| Centrifuge time | 10 min | 9–11 min |
| Digestion temperature | 38 °C | 34.2–41.8 °C |
| Digestion time | 120 min | 108–132 min |
| Trypsin volume | 25 µL | 22.5–27.5 µL |
| TRIS HCl volume | 1000 µL | 900–1100 µL |
| Acetonitrile volume | 200 µL | 180–220 µL |
| Pistacia vera | Anacardium occidentale L. | |||
|---|---|---|---|---|
| Matrix | tR (min) | Sample (Area) | tR (min) | Sample (Area) |
| Bakery product | 26.8 | 967 | / | not detected |
| Cocoa | / | not detected | 3.8 | 1452 |
| Olive Sauce | / | not detected | / | not detected |
| PT cashew | / | not detected | 3.9 | 1013 |
| PT pistachio | 27.1 | 927 | / | not detected |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giugliano, R.; Morello, S.; Lupi, S.; Vivaldi, B.; Bianchi, D.M.; Razzuoli, E. Allergens in Food: Analytical LC-MS/MS Method for the Qualitative Detection of Pistacia vera. Foods 2025, 14, 3031. https://doi.org/10.3390/foods14173031
Giugliano R, Morello S, Lupi S, Vivaldi B, Bianchi DM, Razzuoli E. Allergens in Food: Analytical LC-MS/MS Method for the Qualitative Detection of Pistacia vera. Foods. 2025; 14(17):3031. https://doi.org/10.3390/foods14173031
Chicago/Turabian StyleGiugliano, Roberta, Sara Morello, Samantha Lupi, Barbara Vivaldi, Daniela Manila Bianchi, and Elisabetta Razzuoli. 2025. "Allergens in Food: Analytical LC-MS/MS Method for the Qualitative Detection of Pistacia vera" Foods 14, no. 17: 3031. https://doi.org/10.3390/foods14173031
APA StyleGiugliano, R., Morello, S., Lupi, S., Vivaldi, B., Bianchi, D. M., & Razzuoli, E. (2025). Allergens in Food: Analytical LC-MS/MS Method for the Qualitative Detection of Pistacia vera. Foods, 14(17), 3031. https://doi.org/10.3390/foods14173031





