Microbial and Qualitative Traits of Quinoa and Amaranth Seeds from Experimental Fields in Southern Italy
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials and Reagents
2.2. Experimental Field and Samples
2.3. Biomass, Yield and Seed Chemical Composition Analysis
2.4. Color Measurement
2.5. Microbiological Analysis
2.6. Bacillus spp. Isolation and Phenotypic Characteristics
2.7. Molecular Identification
2.8. DGGE Analysis and Sequencing
2.9. Statistical Analysis
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Devi, R.; Kaur, T.; Kour, D.; Yadav, A.N. Microbial consortium of mineral solubilizing and nitrogen fixing bacteria for plant growth promotion of amaranth (Amaranthus hypochondrius L.). Biocatal. Agric. Biotechnol. 2022, 43, 102404. [Google Scholar] [CrossRef]
- Schmidt, D.; Verruma-Bernardi, M.R.; Forti, V.A.; Borges, M.T.M.R. Quinoa and amaranth as functional foods: A review. Food Rev. Int. 2021, 39, 2277–2296. [Google Scholar] [CrossRef]
- Di Renzo, T.; Reale, A.; Boscaino, F.; Messia, M.C. Flavoring production in Kamut®, quinoa and wheat doughs fermented by Lactobacillus paracasei, Lactobacillus plantarum and Lactobacillus brevis: A SPME-GC/MS study. Front. Microbiol. 2018, 9, 429. [Google Scholar] [CrossRef] [PubMed]
- Alandia, G.; Rodriguez, J.P.; Jacobsen, S.E.; Bazile, D.; Condori, B. Global expansion of quinoa and challenges for the Andean region. Glob. Food Sec. 2020, 26, 100429. [Google Scholar] [CrossRef]
- Sellami, M.H.; Pulvento, C.; Lavini, A. Agronomic practices and performances of quinoa under field conditions: A systematic review. Plants 2020, 10, 72. [Google Scholar] [CrossRef] [PubMed]
- Asher, A.; Galili, S.; Whitney, T.; Rubinovich, L. The potential of quinoa (Chenopodium quinoa) cultivation in Israel as a dual-purpose crop for grain production and livestock feed. Sci. Hortic. 2020, 272, 109534. [Google Scholar] [CrossRef]
- Sellami, M.H.; Pulvento, C.; Aria, M.; Stellacci, A.M.; Lavini, A. A Systematic Review of Field Trials to Synthesize Existing Knowledge and Agronomic Practices on Protein Crops in Europe. Agronomy 2019, 9, 292. [Google Scholar] [CrossRef]
- De Santis, G.; Ronga, D.; Caradonia, F.; Ambrosio, T.D.; Troisi, J.; Rascio, A.; Fragasso, M.; Pecchioni, N.; Rinaldi, M. Evaluation of two groups of quinoa (Chenopodium quinoa Willd.) accessions with different seed colours for adaptation to the Mediterranean environment. Crop Pasture Sci. 2018, 69, 1264–1275. [Google Scholar] [CrossRef]
- El Gendy, A.N.G.; Tavarini, S.; Conte, G.; Pistelli, L.; Hendawy, S.F.; Omer, E.A.; Angelini, L.G. Yield and qualitative characterisation of seeds of Amaranthus hypochondriacus L. and Amaranthus cruentus L. grown in central Italy. Ital. J. Agron. 2018, 13, 63–73. [Google Scholar] [CrossRef]
- Pulvento, C.; Lavini, A.; Riccardi, M.; d’Andria, R.; Ragab, R. Assessing amaranth adaptability in a Mediterranean area of south Italy under different climatic scenarios. Irrig. Drain. 2013, 64, 50–58. [Google Scholar] [CrossRef]
- Lavini, A.; Pulvento, C.; d’Andria, R.; Riccardi, M.; Choukr-Allah, R.; Belhabib, O.; Yazar, A.; İncekaya, Ç.; Metin Sezen, S.; Qadir, M.; et al. Quinoa’s potential in the Mediterranean region. J. Agron. Crop Sci. 2014, 200, 344–360. [Google Scholar] [CrossRef]
- Yazar, A.; Incekaya, Ç.; Sezen, S.M.; Jacobsen, S.E. Saline water irrigation of quinoa (Chenopodium quinoa) under Mediterranean conditions. Crop Past. Sci. 2015, 66, 993–1002. [Google Scholar] [CrossRef]
- Park, S.J.; Sharma, A.; Lee, H.J. A review of recent studies on the antioxidant activities of a third-millennium food: Amaranthus spp. Antioxidants 2020, 9, 1236. [Google Scholar] [CrossRef]
- Gebreil, S.Y.; Ali, M.I.K.; Mousa, E.A.M. Utilization of amaranth flour in preparation of high nutritional value bakery products. Food Nutr. Sci. 2020, 11, 336–354. [Google Scholar] [CrossRef]
- Whang, S.; Zhu, F. Formulation and quality attributes of quinoa food products. Food Bioproc. Technol. 2016, 9, 49–68. [Google Scholar] [CrossRef]
- Brito, I.L.; de Souza, E.L.; Felex, S.S.S.; Madruga, M.S.; Yamashita, F.; Magnani, M. Nutritional and sensory characteristics of gluten-free quinoa (Chenopodium quinoa Willd)-based cookies development using an experimental mixture design. J. Food Sci. Technol. 2015, 52, 5866–5873. [Google Scholar] [CrossRef]
- Azizi, S.; Azizi, M.H.; Moogouei, R.; Rajaei, P. The effect of Quinoa flour and enzymes on the quality of gluten-free bread. Food Sci. Nutr. 2020, 8, 2373–2382. [Google Scholar] [CrossRef]
- Ballester-Sánchez, J.; Millán-Linares, M.C.; Fernández-Espinar, M.T.; Haros, C.M. Development of healthy, nutritious bakery products by incorporation of quinoa. Foods 2019, 8, 379. [Google Scholar] [CrossRef]
- Kurek, M.A.; Sokolova, N. Optimization of bread quality with quinoa flour of different particle size and degree of wheat flour replacement. Food Sci. Technol. 2020, 40, 307–314. [Google Scholar] [CrossRef]
- Franco, W.; Evert, K.; Van Nieuwenhove, C. Quinoa flour, the germinated grain flour, and sourdough as alternative sources for gluten-free bread formulation: Impact on chemical, textural and sensorial characteristics. Fermentation 2021, 7, 115. [Google Scholar] [CrossRef]
- Pereira, A.P.M.; Stradiotto, G.C.; Freire, L.; Alvarenga, V.O.; Crucello, A.; Morassi, L.P.L.; Silva, F.P.; Sant’Ana, A.S. Occurrence and enumeration of rope-producing spore forming bacteria in flour and their spoilage potential in different bread formulations. LWT 2020, 133, 110108. [Google Scholar] [CrossRef]
- Valerio, F.; De Bellis, P.; Di Biase, M.; Lonigro, S.L.; Giussani, B.; Visconti, A.; Lavermicocca, P.; Sisto, A. Diversity of spore-forming bacteria and identification of Bacillus amyloliquefaciens as a species frequently associated with the ropy spoilage of bread. Int. J. Food Microbiol. 2012, 153, 278–285. [Google Scholar] [CrossRef] [PubMed]
- De Bellis, P.; Minervini, F.; Di Biase, M.; Valerio, F.; Lavermicocca, P.; Sisto, A. Toxigenic potential and heat survival of spore-forming bacteria isolated from bread and ingredients. Int. J. Food. Microbiol. 2015, 197, 30–39. [Google Scholar] [CrossRef] [PubMed]
- ICC. Standard Methods of the International Association for Cereal Science and Technology; ICC: Vienna, Austria, 1995. [Google Scholar]
- AACC. Approved Methods of the American Association of Cereal Chemists, Volume 1; American Association of Cereal Chemists Inc.: St. Paul, MN, USA, 2000. [Google Scholar]
- Santagata, G.; Di Renzo, T.; Mallardo, S.; Reale, A.; Cascone, G.; Boscaino, F.; Volpe, M.G. Innovative technologies optimizing the production process of “Castagne del Prete”: Impact on microstructure and volatile compounds. LWT 2022, 168, 113881. [Google Scholar] [CrossRef]
- Garbeva, P.; vanVeen, J.; van Elsas, J. Predominant Bacillus spp. In agricultural soil under different management regimes detected via PCR-DGGE. Microb. Ecol. 2003, 45, 302–316. [Google Scholar] [CrossRef] [PubMed]
- Heuer, H.; Krsek, M.; Baker, P.; Smalla, K.; Wellington, E.M.H. Analysis of actinomycete communities by specific amplification of genes encoding 16S rDNA and gel-electrophoretic separation in denaturing gradients. Appl. Environ. Microbiol. 1997, 63, 3233–3241. [Google Scholar] [CrossRef]
- Araújo, W.L.; Marcon, J.; Maccheroni, W.; van Elsas, J.D.; van Vuurde, J.W.L.; Azevedo, J.L. Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plants. Appl. Environ. Microbiol. 2002, 68, 4906–4914. [Google Scholar] [CrossRef] [PubMed]
- Succi, M.; Sorrentino, E.; Di Renzo, T.; Tremonte, P.; Reale, A.; Tipaldi, L.; Pannella, G.; Russo, A.; Coppola, R. Lactic acid bacteria in pharmaceutical formulations: Presence and viability of “healthy microorganisms”. J. Pharm. Nutr. Sci. 2014, 4, 66–75. [Google Scholar] [CrossRef]
- Pulvento, C.; Sellami, M.H.; Lavini, A. Yield and quality of Amaranthus hypochondriacus grain amaranth under drought and salinity at various phenological stages in southern Italy. J. Sci. Food Agric. 2022, 102, 5022–5033. [Google Scholar] [CrossRef]
- Lavini, A.; Pulvento, C.; d’Andria, R.; Riccardi, M.; Jacobsen, S.E. Effects of saline irrigation on yield and qualitative characterization of seed of an amaranth accession grown under Mediterranean conditions. J. Agric. Sci. 2016, 154, 858–869. [Google Scholar] [CrossRef]
- Thiam, E.; Allaoui, A.; Benlhabib, O. Quinoa productivity and stability evaluation through varietal and environmental interaction. Plants 2021, 10, 714. [Google Scholar] [CrossRef]
- Repo-Carrasco, R.; Espinoza, C.; Jacobsen, S.E. Nutritional value and use of the andean crops quinoa (Chenopodium quinoa) and kaniwa (Chenopodium pallidicaule). Food Rev. Int. 2003, 19, 179–189. [Google Scholar] [CrossRef]
- Vega-Galvez., A.; Miranda, M.; Vergara, J.; Uribe, E.; Puente, L.; Martinez, E.A. Nutrition facts and functional potential of quinoa (Chenopodium quinoa Willd.) an ancient andean grain: Areview. J. Sci. Food Agric. 2010, 90, 2541–2547. [Google Scholar] [CrossRef]
- Smirnova, T.B.; Temereva, I.V.; Tolstoguzova, T.T. The use of whole-ground amaranth flour in the production of bread. IOPConf. Series: Earth Environ. Sci. 2022, 1052, 012027. [Google Scholar] [CrossRef]
- Hager, A.S.; Wolter, A.; Jacob, F.; Zannini, E.; Arendt, E.K. Nutritional properties and ultra-structure of commercial gluten free flours from different botanical sources compared to wheat flours. J. Cereal Sci. 2012, 56, 239–247. [Google Scholar] [CrossRef]
- Taylor, J.R.N.; Parker, M.L. Quinoa. In Pseudocereals and Less Common Cereals; Belton, P.S., Taylors, J.R.N., Eds.; Springer: Berlin, Germany, 2002; pp. 93–122. [Google Scholar]
- Wright, K.H.; Pike, O.A.; Fairbanks, D.J.; Huber, S.C. Composition of Atriplex hortensis, sweet and bitter Chenopodium quinoa seeds. Food Chem. Toxicol. 2002, 67, 1383–1385. [Google Scholar] [CrossRef]
- De Bock, P.; Daelemans, L.; Selis, L.; Raes, K.; Vermeir, P.; Eeckhout, M.; Van Bockstaele, F. Comparison of the chemical and technological characteristics of wholemeal flours obtained from amaranth (Amaranthus sp.), quinoa (Chenopodium quinoa) and buckwheat (Fagopyrum sp.) seeds. Foods 2021, 10, 651. [Google Scholar] [CrossRef] [PubMed]
- Juárez Arana, C.D.; Martínez Peniche, R.A.; Martínez, M.G.; Iturriaga, M.H. Microbiological profile, incidence, and behaviour of Salmonella on seeds traded in mexican markets. J. Food Prot. 2021, 84, 1, 99–105. [Google Scholar] [CrossRef]
- Hernández, E.H.M.; Coronado, A.C.M.; Ardila, W.L.R. Seed quality of 22 quinoa materials (Chenopodium quinoa Willd.) from the department of Boyacá. Rev. Ceres 2020, 67, 306–314. [Google Scholar] [CrossRef]
- Kanbar, A.; Beisel, J.; Wetters, S.; Gutierrez, M.T.; Graeff-Hönninger, S.; Nick, P. A rapid, simple, an reliable assay to authenticate Peruvian kiwicha (A. caudatus) for food applications. Eur. Food Res. Technol. 2022, 248, 2779–2797. [Google Scholar] [CrossRef]
- Granado-Rodríguez, S.; Aparicio, N.; Matías, J.; Pérez-Romero, L.F.; Maestro, I.; Garcés, I.; Pedroche, J.J.; Haros, C.M.; Fernández-Garcia, N.; Navarro del Hierro, J.; et al. Studying the impact of different field environmental conditions on seed quality of quinoa: The case of three different years changing seed nutritional traits in southern Europe. Front. Plant Sci. 2021, 12, 649132. [Google Scholar] [CrossRef]
- Johnston-Monje, D.; Lundberg, D.S.; Lazarovits, G.; Reis, V.M.; Raizada, M.N. Bacterial populations in juvenile maize rhizospheres originate from both seed and soil. Plant Soil 2016, 405, 337–355. [Google Scholar] [CrossRef]
- Paz, P.C.; Janny, R.J.; Håkansson, Å. Safeguarding of quinoa beverage production by fermentation with Lactobacillus plantarum DSM9843. Int. J. Food Microbiol. 2020, 324, 108630. [Google Scholar] [CrossRef]
- Noelting, M.C.; Sisterna, M.N.; Lori, G.; Sandoval, M.C.; Molina, M.C.; Mónaco, C.I. First report of Alternaria alternate causingdiscoloration on amaranthus seeds in Argentina. Australas. Plant Dis. Notes 2011, 6, 1–2. [Google Scholar] [CrossRef]
- Hariram, U.; Labbè, R. Spore prevalence and toxigenicity of Bacillus cereus and Bacillus thuringiensis isolates from U.S. retail spices. J. Food Prot. 2015, 78, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Berghofer, L.K.; Hocking, A.D.; Miskelly, D.; Jansson, E. Microbiology of wheat and flour milling in Australia. Int. J. Food Microbiol. 2002, 85, 137–149. [Google Scholar] [CrossRef]
- Viedma, P.M.; Abriouel, H.; Omar, N.B.; López, R.L.; Gálvez, A. Inhibition of spoilage and toxigenic Bacillus species in dough from wheat flour by the cyclic peptide enterocin AS-48. Food Control 2011, 22, 756–761. [Google Scholar] [CrossRef]
- Postollec, F.; Mathot, A.-G.; Bernard, M.; Divanac’h, M.-L.; Pavan, S.; Sohier, D. Tracking spore-forming bacteria in food: From natural biodiversity to selection by processes. Int. J. Food Microbiol. 2012, 158, 1–8. [Google Scholar] [CrossRef]
- Cook, F.K.; Johnson, B.L. Microbiological spoilage of cereal products. In Compendium of the Microbiological Spoilage of Foods and Beverages; Springer: Berlin/Heidelberg, Germany, 2009; pp. 223–244. [Google Scholar]
- Pitzschke, A. Developmental peculiarities and seed-borne endophytes in quinoa: Omnipresent, robust bacilli contribute to plant fitness. Front. Microbiol. 2016, 7, 2. [Google Scholar] [CrossRef]
- Castillo, J.A.; Conde, G.; Claros, M.; Ortuño, N. Diversity of cultivable microorganisms associated with Quinoa (Chenopodium quinoa) and their potential for plant growth-promotion. Rev. Bionatura 2022, 7, 61. [Google Scholar] [CrossRef]
- Iurlina, M.O.; Saiz, A.I.; Fuselli, S.R.; Fritz, R. Prevalence of Bacillus spp. in different food products collected in Argentina. LWT 2006, 39, 105–110. [Google Scholar] [CrossRef]
- Fangio, M.F.; Roura, S.I.; Fritz, R. Isolation and identification of Bacillus spp. and related genera from different starchy foods. J. Food Sci. 2010, 75, M218–M221. [Google Scholar] [CrossRef] [PubMed]
- Rodrigo, D.; Rosell, C.M.; Martinez, A. Risk of Bacillus cereus in relation to rice and derivatives. Foods 2021, 10, 302. [Google Scholar] [CrossRef] [PubMed]
- Yong-Bae, P.; Jung-Beom, K.; Sang-Woon, S.; Jong-Chan, K.; Seung-Hak, C.; Bok-Kwon, L.; Juhee, A.; Jae-Myung, K.; Deog-Hwan, O. Prevalence, genetic diversity, and antibiotic susceptibility of Bacillus cereus strains isolated from rice and cereals collected in Korea. J. Food Prot. 2009, 72, 612–617. [Google Scholar] [CrossRef]
- Földes, T.; Bánhegyi, I.; Herpai, Z.; Varga, L.; Szigeti, J. Isolation of Bacillus strains from the rhizosphere of cereals and in vitro screening for antagonism against phytopathogenic, food-borne pathogenic and spoilage micro-organisms. J. Appl. Microbiol. 2000, 89, 840–846. [Google Scholar] [CrossRef] [PubMed]
- Reale, A.; Di Renzo, T.; Succi, M.; Tremonte, P.; Coppola, R.; Sorrentino, E. Microbiological and fermentative properties of baker’s yeast starter used in breadmaking. J. Food Sci. 2013, 78, M1224–M1231. [Google Scholar] [CrossRef]
- Pacher, N.; Burtscher, J.; Johler, S.; Etter, D.; Bender, D.; Fieseler, L.; Domig, K.J. Ropiness in bread—A re-emerging spoilage phenomenon. Foods 2022, 11, 3021. [Google Scholar] [CrossRef]
- TeGiffel, M.; Beumer, R.; Leijendekkers, S.; Rombouts, F. Incidence of Bacillus cereus and Bacillus subtilis in foods in The Netherlands. Food Microbiol. 1996, 13, 53–58. [Google Scholar] [CrossRef]
- Satomi, M.; Duc, M.T.L.; Venkateswaran, K. Bacillus safensis sp. nov., isolated from spacecraft and assembly-facility surfaces. Int. J. Syst. Evol. Microbiol. 2006, 56, 1735–1740. [Google Scholar] [CrossRef]
- Rong, S.; Xu, H.; Li, L.; Chen, R.; Gao, X.; Xu, Z. Antifungal activity of endophytic Bacillus safensis B21 and its potential application as a biopesticide to control rice blast. Pestic. Biochem. Phys. 2020, 162, 69–77. [Google Scholar] [CrossRef]
- Abril, A.G.; Rama, J.L.R.; Feijoo-Siota, L.; Calo-Mata, P.; Salazar, S.; Peix, A.; Velázquez, E.; Villa, T.G. Bacillus safensis subsp. Osmophilus subsp. nov., isolated from condensed milk, and description of Bacillus safensis subsp. Safensis subsp. Nov. Int. J. Syst. Evol. Microbiol. 2019, 69, 189–195. [Google Scholar] [CrossRef]
- Ishag, A.E.S.A.; Abdelbagi, A.O.; Hammad, A.M.A.; Elsheikh, E.A.E.; Elsaid, O.E.; Hur, J.-H.; Laing, M.D. Biodegradation of chlorpyrifos, malathion, and dimethoate by three strains of bacteria isolated from pesticide-polluted soils in Sudan. J. Agric. Food Chem. 2016, 64, 8491–8498. [Google Scholar] [CrossRef] [PubMed]
- Abdelli, F.; Jardak, M.; Elloumi, J.; Stien, D.; Cherif, S.; Mnif, S.; Aifa, S. Antibacterial, anti-adherent and cytotoxic activities of surfactin(s) from a lipolytic strain Bacillus safensis F4. Biodegrad. 2019, 30, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Kronstad, J.W. Disarming fungal pathogens: Bacillus safensis inhibits virulence factor production and biofilm formation by Cryptococcus neoformans and Candida albicans. MBio 2017, 8, e01537-17. [Google Scholar] [CrossRef] [PubMed]
- Lateef, A.; Adelere, I.A.; Gueguim-Kana, E.B. The biology and potential biotechnological applications of Bacillus safensis. Biologia 2015, 70, 411–419. [Google Scholar] [CrossRef]
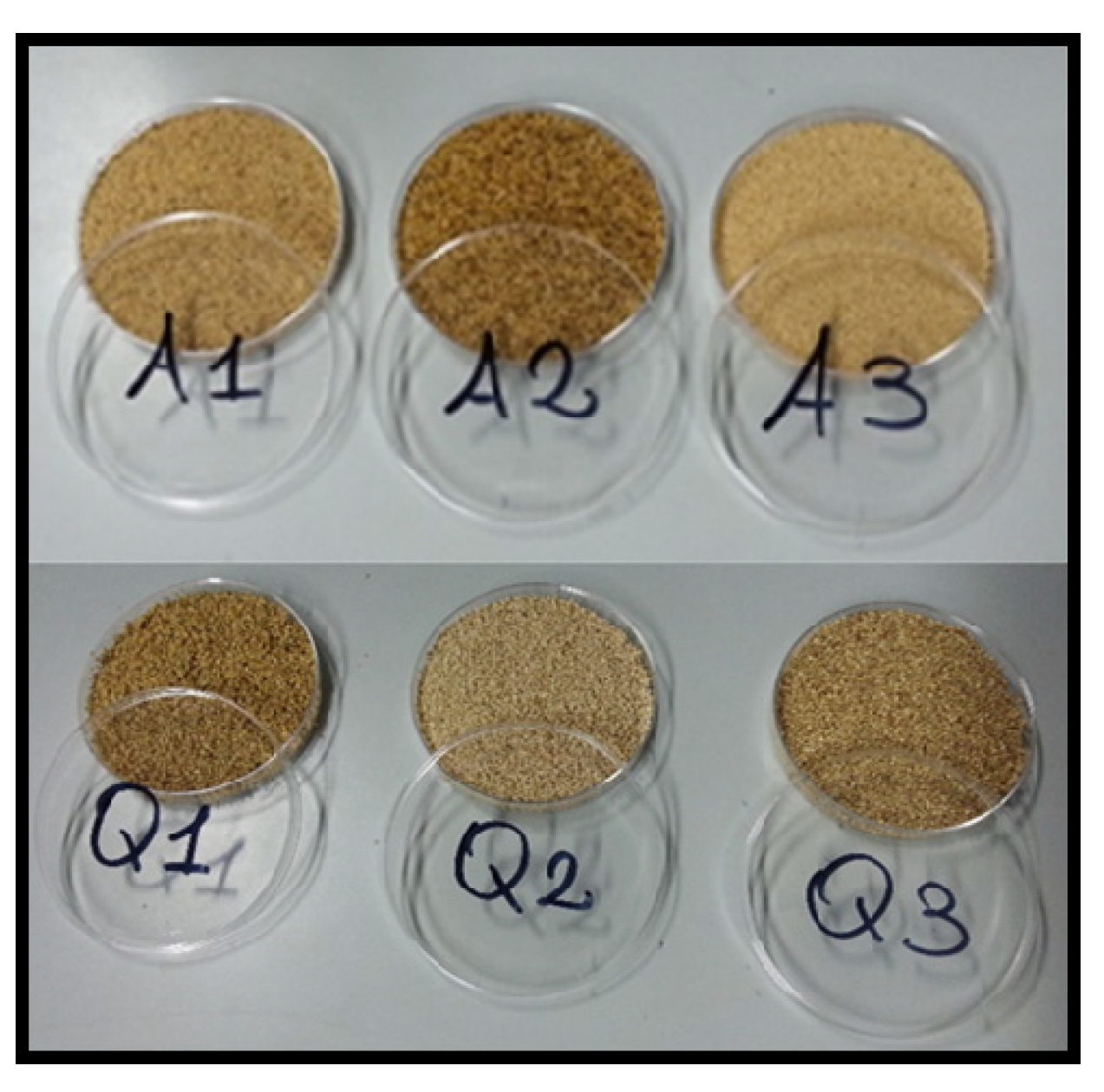

| Sample | Type of Pseudocereal | Location | Geographic Position | Altitude (m.a.s.l.) | Soil Type | Average Annual Rainfall (mm) | |
|---|---|---|---|---|---|---|---|
| AMARANTH | A1 | Amaranth accession 5 | Vitulazio (CE) | 14°50′ E, 40°07′ N | 25 | clay loam | 805 |
| A2 | Amaranth accession 12 | Vitulazio (CE) | 14°50′ E, 40°07′ N | 25 | clay loam | 805 | |
| A3 | Amaranth accession 14 | Vitulazio (CE) | 14°50′ E, 40°07′ N | 25 | clay loam | 805 | |
| QUINOA | Q1 | Quinoa var. Titicaca | Ercolano (NA) | 14°21′ E, 40°50′ N | 175 | sandy | 1080 |
| Q2 | Quinoa var. Titicaca | Vitulazio (CE) | 14°50′ E, 40°07′ N | 25 | clay loam | 805 | |
| Q3 | Quinoa var. Titicaca | Montecalvo Irpino (AV) | 15°3′ E, 41°15′ N | 518 | clay loam | 1068 | |
| Sample | Seed Yield | Biomass | Harvest Index | 1000 Seeds Weight |
|---|---|---|---|---|
| t/ha | % | g | ||
| A1 | 1.34 b | 23.70 b | 6 | 0.95 |
| A2 | 2.20 a | 28.90 a | 8 | 0.68 |
| A3 | 1.96 a | 28.4 a | 7 | 0.95 |
| p-value | <0.05 | <0.05 | n.s | n.s |
| Q1 | 2.3 a | 6.7 a | 34 a | 2.45 |
| Q2 | 1.33 b | 5.85 ab | 23 b | 3.00 |
| Q3 | 0.19 c | 2.10 b | 9 c | 2.97 |
| p-value | <0.05 | <0.05 | <0.05 | n.s |
| Sample | Protein | Lipid | Ash | Total Starch | Dietary Fiber | Water Activity (aw) | pH | |
|---|---|---|---|---|---|---|---|---|
| AMARANTH | A1 | 13.7 ± 0.02 a | 4.0 ± 0.05 b | 2.0 ± 0.11 a | 54.3 ± 0.55 b | 15.1 ± 1.00 a | 0.468 ± 0.022 a | 5.42 ± 0.11 a |
| A2 | 14.4 ± 0.15 b | 3.2 ± 0.03 a | 3.2 ± 0.13 c | 48.9 ± 0.61 a | 19.1 ± 1.23 b | 0.496 ± 0.020 a | 5.43 ± 0.09 a | |
| A3 | 13.7 ± 0.01 a | 3.3 ± 0.09 a | 2.4 ± 0.20 b | 56.0 ± 0.43 c | 13.5 ± 0.98 a | 0.514 ± 0.024 a | 5.36 ± 0.20 a | |
| QUINOA | Q1 | 16.5 ± 0.09 c | 5.9 ± 0.06 a | 6.0 ± 0.11 b | 54.9 ± 0.69 a | 10.8 ± 0.94 a | 0.436 ± 0.014 a | 5.39 ± 0.15 a |
| Q2 | 15.9 ± 0.12 a | 6.5 ± 0.09 b | 5.3 ± 0.23 a | 58.7 ± 0.75 c | 10.2 ± 0.85 a | 0.445 ± 0.012 a | 5.47 ± 0.08 a | |
| Q3 | 14.3 ± 0.04 b | 6.6 ± 0.10 b | 5.6 ± 0.24 a | 56.8 ± 0.73 b | 10.6 ± 0.70 a | 0.427 ± 0.024 a | 5.62 ± 0.17 a | |
| Sample | Colorimetric Indices | |||
|---|---|---|---|---|
| L* | a* | b* | ||
| AMARANTH | A1 | 48.02 ± 0.39 b | 5.99 ± 0.07 a | 20.71 ± 0.32 b |
| A2 | 38.93 ± 0.58 a | 5.97 ± 0.05 a | 19.55 ± 0.40 a | |
| A3 | 50.04 ± 0.26 c | 5.66 ± 0.03 b | 21.13 ± 0.14 b | |
| QUINOA | Q1 | 44.14 ± 0.16 a | 4.92 ± 0.02 c | 18.46 ± 0.07 b |
| Q2 | 46.74 ± 0.17 b | 4.24 ± 0.05 a | 18.87 ± 0.13 c | |
| Q3 | 44.74 ± 0.63 a | 4.58 ± 0.14 b | 17.39 ± 0.25 a | |
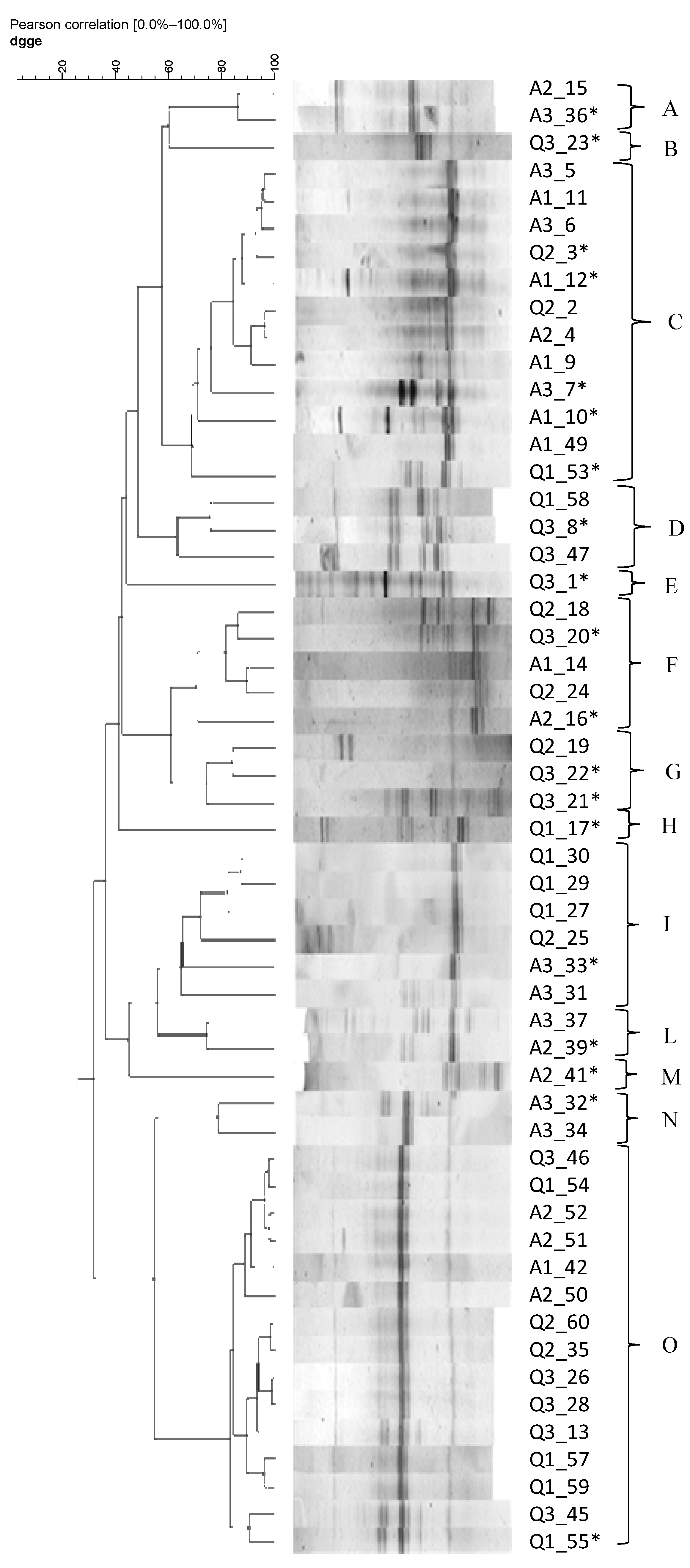
| Cluster | Strain | Size (bp) | Closest Relative | % Identity | Source a |
|---|---|---|---|---|---|
| A | A3_36 | 1363 | Uncultured bacterium clone | 100 | KF066707 |
| B | Q3_23 | 1372 | Uncultured bacterium clone | 97 | GQ017951 |
| C | Q2_3 | 1462 | B. subtilis | 99 | JN942155 |
| C | A1_12 | 1414 | B. subtilis | 99 | KF830999 |
| C | A3_7 | 878 | B. subtilis | 99 | KP699115 |
| C | A1_10 | 1417 | B. subtilis | 99 | KT720106 |
| C | Q1_53 | 1416 | B. subtilis | 99 | KP347686 |
| D | Q3_8 | 919 | Uncultured bacterium clone | 99 | JX283599 |
| E | Q3_1 | 1417 | B. subtilis | 99 | KT720106 |
| F | Q3_20 | 940 | B. safensis | 99 | HM583998 |
| F | A2_16 | 878 | B. safensis | 99 | HM583998 |
| G | Q3_21 | 1002 | B. licheniformis | 99 | KR782291 |
| G | Q3_22 | 1395 | B. licheniformis | 97 | KT318805 |
| H | Q1_17 | 1464 | Uncultured bacterium clone | 99 | JQ940780 |
| I | A3_33 | 1417 | B. subtilis | 99 | KT720106 |
| L | A2_39 | 1401 | B. subtilis | 99 | KM458977 |
| M | A2_41 | 1395 | B. licheniformis | 99 | KT318805 |
| N | A3_32 | 1504 | B. cereus | 99 | GU056810 |
| O | Q1_55 | 734 | B. cereus | 99 | DQ339658 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reale, A.; Messia, M.C.; Pulvento, C.; Lavini, A.; Nazzaro, S.; Di Renzo, T. Microbial and Qualitative Traits of Quinoa and Amaranth Seeds from Experimental Fields in Southern Italy. Foods 2023, 12, 1866. https://doi.org/10.3390/foods12091866
Reale A, Messia MC, Pulvento C, Lavini A, Nazzaro S, Di Renzo T. Microbial and Qualitative Traits of Quinoa and Amaranth Seeds from Experimental Fields in Southern Italy. Foods. 2023; 12(9):1866. https://doi.org/10.3390/foods12091866
Chicago/Turabian StyleReale, Anna, Maria Cristina Messia, Cataldo Pulvento, Antonella Lavini, Stefania Nazzaro, and Tiziana Di Renzo. 2023. "Microbial and Qualitative Traits of Quinoa and Amaranth Seeds from Experimental Fields in Southern Italy" Foods 12, no. 9: 1866. https://doi.org/10.3390/foods12091866
APA StyleReale, A., Messia, M. C., Pulvento, C., Lavini, A., Nazzaro, S., & Di Renzo, T. (2023). Microbial and Qualitative Traits of Quinoa and Amaranth Seeds from Experimental Fields in Southern Italy. Foods, 12(9), 1866. https://doi.org/10.3390/foods12091866










