Relationship between Volatile Organic Compounds and Microorganisms Isolated from Raw Sheep Milk Cheeses Determined by Sanger Sequencing and GC–IMS
Abstract
1. Introduction
2. Materials and Methods
2.1. Origin, Isolation and Preparation of Samples and Bacterial Cultures
2.2. Bacterial Identification by Sanger Sequencing
2.3. GC–IMS Analysis Conditions
2.4. Identification of VOCs
2.5. Analysis of Samples
2.6. Data Analysis
3. Results and Discussion
3.1. Sequencing of LAB Isolated from Raw Sheep Milk Cheeses
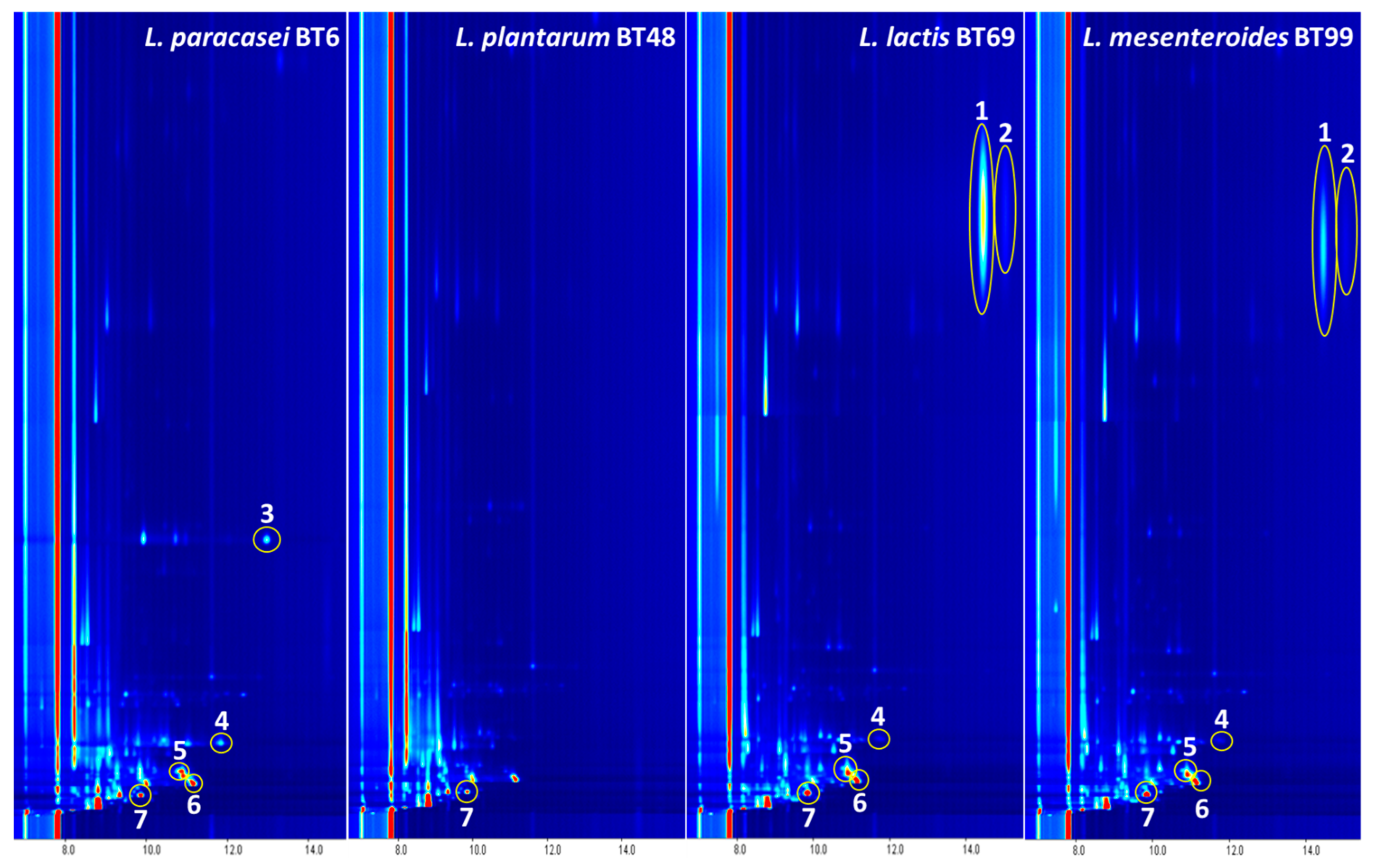
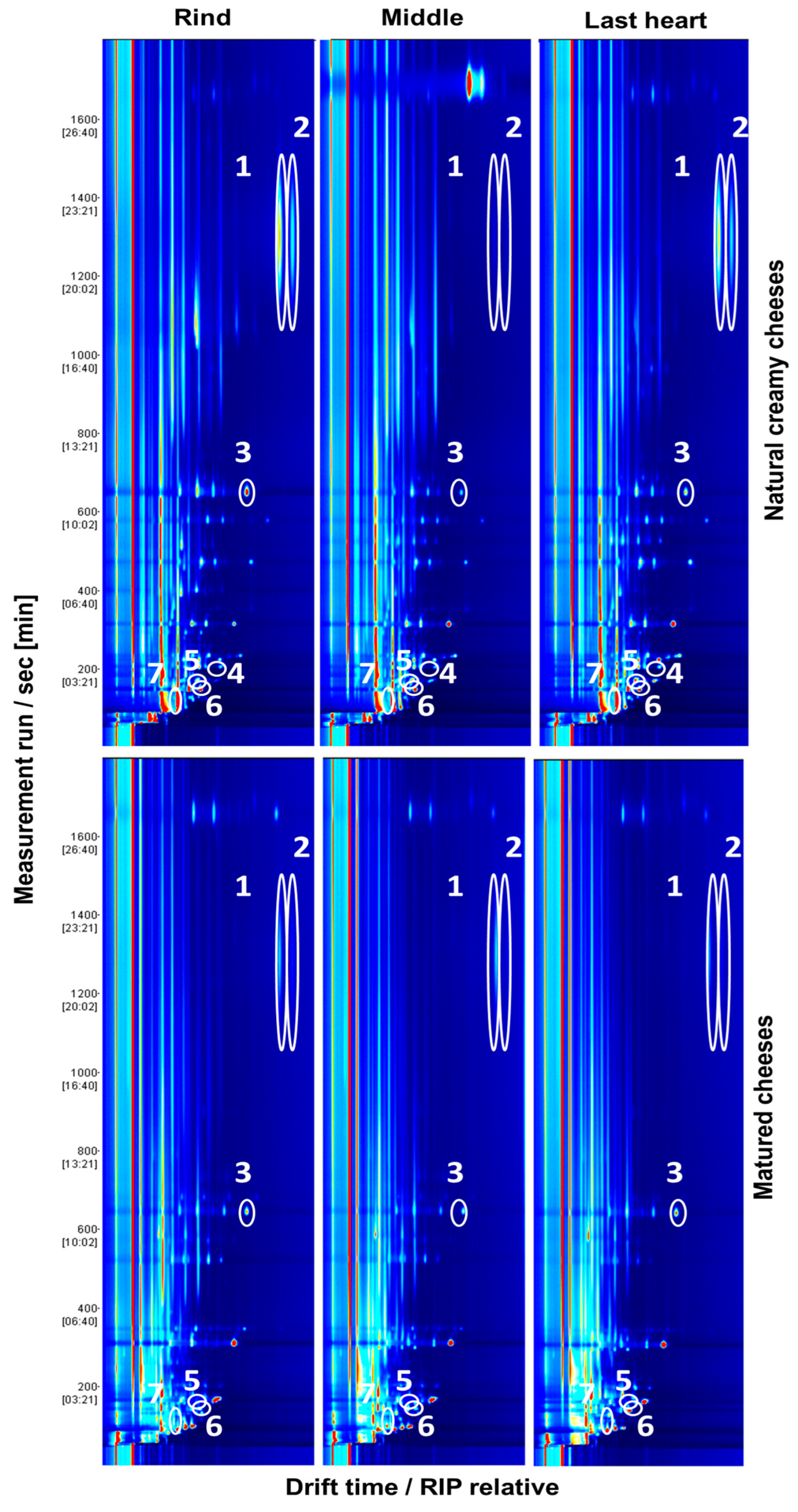
3.2. Identification of VOCs by GC–IMS
3.3. Discussion Based on Groups of VOCs
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Colonna, A.; Durham, C.; Meunier-Goddik, L. Factors affecting consumers’ preferences for and purchasing decisions regarding pasteurized and raw milk specialty cheeses. J. Dairy Sci. 2011, 94, 5217–5226. [Google Scholar] [CrossRef]
- AMRC (Ag Marketing Resource Center). Cheese Industry Profile. USDA Rural Development–Iowa State University. 2022. Available online: https://www.agmrc.org/commodities-products/livestock/dairy/cheese-industry-profile (accessed on 20 October 2022).
- Mohapatra, A.; Shinde, A.K.; Singh, R. Sheep milk: A pertinent functional food. Small Rumin. Res. 2019, 181, 6–11. [Google Scholar] [CrossRef]
- Gallegos, J.; Arce, C.; Jordano, R.; Arce, L.; Medina, L.M. Target identification of volatile metabolites to allow the differentiation of lactic acid bacteria by gas chromatography-ion mobility spectrometry. Food Chem. 2017, 220, 362–370. [Google Scholar] [CrossRef] [PubMed]
- Soltani, M.; Sahingil, D.; Gokce, Y.; Hayaloglu, A.A. Changes in volatile composition and sensory properties of Iranian ultrafiltered white cheese as affected by blends of Rhizomucor miehei protease or camel chymosin. J. Dairy Sci. 2016, 9, 7744–7754. [Google Scholar] [CrossRef]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.M.; Harris, H.M.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020, 70, 2782–2858. [Google Scholar] [CrossRef]
- Gaglio, R.; Franciosi, E.; Todaro, A.; Guarcello, R.; Alfeo, V.; Randazzo, C.L.; Settanni, L.; Todaro, M. Addition of selected starter/non-starter lactic acid bacterial inoculums to stabilise PDO Pecorino Siciliano cheese production. Food Res. Int. 2020, 136, 109335. [Google Scholar] [CrossRef]
- Galimberti, A.; Casiraghi, M.; Bruni, I.; Guzzetti, L.; Cortis, P.; Berterame, N.M.; Labra, M. From DNA barcoding to personalized nutrition: The evolution of food traceability. Curr. Res. Food Sci. 2019, 28, 41–48. [Google Scholar] [CrossRef]
- Contreras, M.M.; Jurado-Campos, N.; Arce, L.; Arroyo-Manzanares, N. A robustness study of calibration models for olive oil classification: Targeted and non-targeted fingerprint approaches based on GC-IMS. Food Chem. 2019, 288, 315–324. [Google Scholar] [CrossRef]
- Arce, L.; Gallegos, J.; Garrido-Delgado, R.; Medina, L.M.; Sielemann, S.; Wortelmann, T. Ion mobility spectrometry a versatile analytical tool for metabolomics applications in food science. Curr. Metab. 2014, 2, 264–271. [Google Scholar] [CrossRef]
- Gallegos, J.; Garrido-Delgado, R.; Arce, L.; Medina, L.M. Volatile metabolites of goat cheeses determined by Ion Mobility Spectrometry. Potential applications in quality control. Food Anal. Methods 2015, 8, 1699–1709. [Google Scholar] [CrossRef]
- Bertuzzi, A.S.; McSweeney, P.L.H.; Rea, M.C.; Kilcawley, K.N. Detection of volatile compounds of cheese and their contribution to the flavor profile of surface-ripened cheese. Compr Rev. Food Sci. Food Saf. 2018, 17, 371–390. [Google Scholar] [CrossRef] [PubMed]
- Kilcawley, K.N.; Faulkner, H.; Clarke, H.J.; O’Sullivan, M.G.; Kerry, J.P. Factors influencing the flavour of bovine milk and cheese from grass based versus non-grass based milk production systems. Foods 2018, 7, 37. [Google Scholar] [CrossRef]
- Ruiz-Moyano, S.; Goncalves dos Santos, M.T.; Galván, A.I.; Merchán, A.V.; González, E.; Córdoba, M.G.; Benito, M.J. Screening of autochthonous lactic acid bacteria strains from artisanal soft cheese: Probiotic characteristics and prebiotic metabolism. LWT. 2019, 114, 108388. [Google Scholar] [CrossRef]
- Beck, T.F.; Mullikin, J.C.; Biesecker, L.G.; Progra, N.C.S. Systematic evaluation of Sanger validation of next-generation sequencing variants. Clin. Chem. 2016, 62, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Cozzolino, R.; Martignetti, A.; De Giulio, B.; Malorni, L.; Addeo, F.; Picariello, G. SPME GC-MS monitoring of volatile organic compounds to assess typicity of Pecorino di Carmasciano ewe-milk cheese. Int. J. Dairy Technol. 2021, 74, 383–392. [Google Scholar] [CrossRef]
- Guarrasi, V.; Sannino, C.; Moschetti, M.; Bonanno, A.; Di Grigoli, A.; Settanni, L. The individual contribution of starter and non-starter lactic acid bacteria to the volatile organic compound composition of Caciocavallo Palermitano cheese. Int. J. Food Microbiol. 2017, 259, 35–42. [Google Scholar] [CrossRef]
- Marilley, L.; Casey, M.G. Flavours of cheese products: Metabolic pathways, analytical tools and identification of producing strains. Int. J. Food Microbiol. 2004, 90, 139–159. [Google Scholar] [CrossRef]
- Luz, C.; D’Opazo, V.; Quiles, J.M.; Romano, R.; Mañes, J.; Meca, G. Biopreservation of tomatoes using fermented media by lactic acid bacteria. LWT. 2020, 130, 109618. [Google Scholar] [CrossRef]
- Štefániková, J.; Árvay, J.; Miškeje, M.; Kačániová, M. Determination of volatile organic compounds in Slovak Brynda cheese by the electronic nose and the headspace solid-phase microextraction gas chromatography-mass spectrometry. Slovak J. Food Sci. 2020, 14, 767–773. [Google Scholar]
- Delgado-Martínez, F.; Carrapiso, A.I.; Contador, R.; Rosario Ramírez, M. Volatile compounds and sensory changes after high pressure processing of mature “Torta del Casar” (raw ewe’s milk cheese) during refrigerated storage. Innov. Food Sci. Emerg. Technol. 2019, 52, 34–41. [Google Scholar] [CrossRef]
- Bozoudi, D.; Kondyli, E.; Claps, S.; Hatzikamari, M.; Michaelidou, A.; Biliaderis, C.G.; Litopoulou-Tzanetaki, E. Compositional characteristics and volatile organic compounds of traditional PDO Feta cheese made in two different mountainous areas of Greece. Int. J. Dairy Technol. 2018, 71, 673–682. [Google Scholar] [CrossRef]
- Reale, A.; Di Renzo, T.; Boscaino, F.; Nazzaro, F.; Fratianni, F.; Aponte, M. Lactic acid bacteria biota and aroma profile of Italian traditional sourdoughs from the irpinian area in Italy. Front. Microbiol. 2019, 10, 1621. [Google Scholar] [CrossRef] [PubMed]
- Delgado, F.; Rodríguez-Pinilla, J.; Márquez, G.; Roa, I.; Ramírez, R. Physicochemical, proteolysis and texture changes during the storage of a mature soft cheese treated by high-pressure hydrostatic. Eur. Food Res. Technol. 2015, 240, 1167–1176. [Google Scholar] [CrossRef]
- Kirmaci, H.A.; Ozer, B.H.; Akcelik, M.; Akcelik, N. Identification and characterisation of lactic acid bacteria isolated from traditional Urfa cheese. Int. J. Dairy Technol. 2016, 69, 301–307. [Google Scholar] [CrossRef]
- Ramírez-López, C.; Vélez-Ruiz, J.F. Isolation, characterization and selection of autochthonous lactic acid bacteria from goat milk and fresh artisanal-goat cheese. Inf. Tecnol. 2016, 27, 6. [Google Scholar] [CrossRef]
- Zoumpopoulou, G.; Papadimitriou, K.; Alexandraki, V.; Mavrogonatou, E.; Alexopoulou, K.; Anastasiou, R.; Georgalaki, M.; Kletsas, D.; Tsakalidou, E.; Giaouris, E. The microbiota of Kalathaki and Melichloro Greek artisanal cheeses comprises functional lactic acid bacteria. LWT 2020, 130, 109570. [Google Scholar] [CrossRef]
- Peralta, G.H.; Wolf, I.V.; Bergamini, C.V.; Perotti, M.C.; Hynes, E.R. Evaluation of volatile compounds produced by Lactobacillus paracasei I90 in a hard-cooked cheese model using solid-phase microextraction. Dairy Sci. Technol. 2014, 94, 73–81. [Google Scholar] [CrossRef]
- Poveda, J.M.; Nieto-Arribas, P.; Seseña, S.; Chicón, R.; Castro, L.; Palop, L.; Cabezas, L. Volatile composition and improvement of the aroma of industrial Manchego cheese by using Lactobacillus paracasei subsp. paracasei as adjunct and other autochthonous strains as starters. Eur. Food Res. Technol. 2014, 238, 485–494. [Google Scholar]
- Stefanovic, E.; Kilcawley, K.N.; Roces, C.; Rea, M.C.; O’Sullivan, M.; Sheehan, J.J.; McAuliffe, O. Evaluation of the potential of Lactobacillus paracasei adjuncts for flavor compounds development and diversification in short-aged Cheddar cheese. Front. Microbiol. 2018, 9, 1506. [Google Scholar] [CrossRef]
- Terpou, A.; Bosnea, L.; Kanellaki, M.; Plessas, S.; Bekatorou, A.; Bezirtzoglou, E.; Koutinas, S.A. Growth capacity of a novel potential probiotic Lactobacillus paracasei K5 strain incorporated in industrial white brined cheese as an adjunct culture. J. Food Sci. 2018, 83, 723–731. [Google Scholar] [CrossRef]
- Shehata, M.G.; Abd El-Aziz, N.M.; Darwish, A.G.; El-Sohaimy, S.A. Lacticaseibacillus paracasei KC39 immobilized on prebiotic wheat bran to manufacture functional soft white cheese. Fermentation 2022, 8, 496. [Google Scholar] [CrossRef]
- Fernández, E.; Alegría, A.; Delgado, S.; Martín, M.C.; Mayo, B. Comparative phenotypic and molecular genetic profiling of wild Lactococcus lactis subsp. lactis strains of the L. lactis subsp. lactis and L. lactis subsp. cremoris genotypes, isolated from starter-free cheeses made of raw milk. Appl. Environ. Microbiol. 2011, 77, 5324–5335. [Google Scholar]
- Cavanagh, D.; Casey, A.; Altermann, E.; Cotter, P.D.; Fitzgerald, G.F.; McAuliffe, O. Evaluation of Lactococcus lactis Isolates from Nondairy Sources with Potential Dairy Applications Reveals Extensive Phenotype-Genotype Disparity and Implications for a Revised Species. Appl. Environ. Microbiol. 2015, 81, 3961–3972. [Google Scholar] [CrossRef] [PubMed]
- Kelleher, P.; Bottacini, F.; Mahony, J.; Kilcawley, K.N.; Van Sinderen, D. Comparative and functional genomics of the Lactococcus lactis taxon; insights into evolution and niche adaptation. BMC Genom. 2017, 18, 267. [Google Scholar] [CrossRef] [PubMed]
- Laroute, V.; Tormo, H.; Couderc, C.; Mercier-Bonin, M.; Le Bourgeois, P.; Cocaign-Bousquet, M.; Daveran-Mingot, M.L. From genome to phenotype: An integrative approach to evaluate the biodiversity of Lactococcus lactis. Microorganisms 2017, 5, 27. [Google Scholar] [CrossRef]
- McAuliffe, O. Symposium review: Lactococcus lactis from non dairy sources: Their genetic and metabolic diversity and potential applications in cheese. J. Dairy Sci. 2018, 101, 3597–3610. [Google Scholar] [CrossRef]
- Gómez-Ruiz, J.A.; Cabezas, L.; Martínez-Castro, I.; González-Viñas, M.A.; Poveda, J.M. Influence of a defined-strain starter and Lactobacillus plantarum as adjunct culture on volatile compounds and sensory characteristics of Manchego cheese. Eur. Food Res. Technol. 2008, 227, 181–190. [Google Scholar] [CrossRef]
- Duan, C.; Li, S.; Zhao, Z.; Wang, C.; Zhao, Y.; Yang, G.; Niu, C.; Gao, L.; Liu, X.; Zhao, L. Proteolytic activity of Lactobacillus plantarum strains in Cheddar cheese as adjunct cultures. J. Food Prot. 2019, 82, 2108–2118. [Google Scholar] [CrossRef]
- Jia, R.; Zhang, F.; Song, Y.; Lou, Y.; Zhao, A.; Liu, Y.; Peng, H.; Hui, Y.; Ren, R.; Wang, B. Physicochemical and textural characteristics and volatile compounds of semihard goat cheese as affected by starter cultures. J. Dairy Sci. 2021, 104, 270–280. [Google Scholar] [CrossRef]
- Pogačić, T.; Maillard, M.B.; Leclerc, A.; Hervé, C.; Chuat, V.; Valence, F.; Thierry, A. Lactobacillus and Leuconostoc volatilomes in cheese conditions. Appl. Microbiol. Biotechnol. 2016, 100, 2335–2346. [Google Scholar] [CrossRef]
- Cardinali, F.; Ferrocino, I.; Milanović, V.; Belleggia, L.; Rita Corvaglia, M.; Garofalo, C.; Foligni, R.; Mannozzi, C.; Mozzon, M.; Cocolin, L.; et al. Microbial communities and volatile profile of Queijo de Azeitão PDO cheese, a typical Mediterranean thistle curdled cheese from Portugal. Food Res. Int. 2021, 147, 110537. [Google Scholar] [CrossRef] [PubMed]
- Van Mastrigt, O.; Egas, R.A.; Abee, T.; Smid, E.J. Aroma formation in retentostat co-cultures of Lactococcus lactis and Leuconostoc mesenteroides. Food Microbiol. 2019, 82, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Majcher, M.A.; Goderska, K.; Pikul, J.; Jelen, H.H. Changes in volatile, sensory and microbial profiles during preparation of smoked ewe cheese. J. Sci. Food Agric. 2011, 91, 1416–1423. [Google Scholar] [CrossRef] [PubMed]
- Coda, R.; Brechany, E.; De Angelis, M.; De Candia, S.; Di Cagno, R.; Gobbetti, M. Comparison of the compositional, microbiological, biochemical, and volatile profile characteristics of nine Italian ewes’ milk cheeses. J. Dairy Sci. 2006, 89, 4126–4143. [Google Scholar] [CrossRef]
- The Good Scents Company Information System. Providing Information for the Flavor, Fragrance, Food and Cosmetic Industries. Available online: http://www.thegoodscentscompany.com/index.html (accessed on 20 June 2021).
- Curioni, P.M.; Bosset, J.O. Key odorants in various cheese types as determined by gas chromatography-olfactometry. Int. Dairy J. 2002, 12, 959–984. [Google Scholar] [CrossRef]

| Chemical Group | VOC | RI | tR | tD | tD (Database) | Reference * |
|---|---|---|---|---|---|---|
| Ketones | 2-Heptanone | 892.2 | 647.6; 647.0 | 1.3; 1.6 | 1.3; 1.6 | [16,17,18,19] |
| 2-Hexanone | 784.2 | 270.0; 292.0 | 1.2; 1.5 | 1.2; 1.5 | [16,18] | |
| 2-Pentanone | 688.6 | 140.5; 148.7 | 1.1; 1.4 | 1.1; 1.4 | [16,18,20] | |
| 2-Butanone | 589.4 | 96.2; 89.3 | 1.1; 1.3 | 1.1; 1.2 | [7,17,20,21,22] | |
| 3-Hydroxybutan-2-one | 715.12 | 1384.2 | 1.8 | NA | [7,23] | |
| Esters | Ethyl butanoate | 794.9 | 302.3; 300.6 | 1.2; 1.6 | 1.2; 1.6 | [16,17,18,20,22,23] |
| Ethyl hexanoate | 999.9 | 1598.8; 1603.4 | 1.3; 1.8 | 1.3; 1.8 | [7,16,17,18,22,23] | |
| Ethyl propanoate | 707.7 | 161.0; 162.6 | 1.2; 1.5 | 1.1; 1.4 | [16] | |
| Propyl butanoate | 895.9 | 660.1; 663.7 | 1.3; 1.7 | NA | [18,22] | |
| Ethyl acetate | 668.8; 673.2 | 138.1; 140.6 | 1.2; 1.4 | 1.1; 1.3 | [16,17,18,20] | |
| Aldehydes | 2-Methylbutanal | 694.6 | 119.5; 120.9 | 1.2; 1.4 | NA | [17,18,19,24] |
| Benzaldehyde | 960.9 | 1083.9; 1087.7 | 1.1; 1.5 | 1.1; 1.5 | [7,16,17,19,20,22,23] | |
| Butanal | 602.6; 602.6 | 85.5; 85.1 | 1.1; 1.3 | 1.1; 1.3 | [17,20] | |
| Hexanal | 800.2; 800.2 | 288.9; 291.5 | 1.3; 1.6 | 1.2; 1.6 | [17,18,19] | |
| 2-Methylpropanal | 551.8; 551.8 | 75.9; 75.9 | 1.1; 1.3 | 1.1; 1.3 | [18,20] | |
| 3-Methylthiopropanal | 907.7; 907.7 | 719.0; 721.9 | 1.1; 1.4 | 1.1; 1.4 | [19] | |
| Octanal | 128.2 | 1001.9; 1001.9 | 1.4; 1.8 | 1.4; 1.8 | [4,18,23] | |
| Acetaldehyde | 784.7; 785.5 | 293.4; 296.1 | 1.2; 1.4 | NA | [18,20,23] | |
| Alcohols | 2-Butanol | 602.7; 602.7 | 131.7; 133.0 | 1.2; 1.4 | NA | [16,17,18,22] |
| 2-Methyl-1-propanol | 629.0; 629.0 | 102.9; 104.4 | 1.2; 1.4 | NA | [18,20,22,23] | |
| 2-Heptanol | 970.5; 970.5 | 1329.0; 1335.3 | 1.4; 1.8 | NA | [16,17,18,22,23] | |
| n-Hexanol | 867.4; 867.4 | 561.8; 566.3 | 1.3; 1.6 | 1.9; 1.6 | [7,16,17,18,20,23] | |
| n-Butanol | 660.4; 660.4 | 129.2; 125.5 | 1.2; 1.4 | 1.2; 1.4 | [16,17,21,22] | |
| 1-Pentanol | 765.9; 765.9 | 250.9; 252.5 | 1.2; 1.5 | 1.2; 1.5 | [17,23] | |
| 3-Methyl-1-butanol | 724.0; 723.7 | 133.8; 135.2 | 1.2; 1.4 | NA | [16,17,18,19,21,22] | |
| Acids | Acetic acid | 621.3684 | 93.05; - | 1.0; - | 1.0; 1.5 | [16,19,20,21,23] |
| Chemical Group | VOC | MRS Media | Milk at 37 °C | Milk at 12 °C | Creamy Cheese | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CT | Lpp | Lms | CT | Lpp | Lms | CT | Lpp | Lms | Rind | Inner | Center | ||
| Ketones | 2-Heptanone | - | - | - | + | + | + | + | + | + | + | + | + |
| 2-Hexanone | + | + | + | - | + | - | - | - | - | + | + | + | |
| 2-Pentanone | - | - | + | + | + | + | + | + | + | + | + | + | |
| 2-Butanone | - | + | + | + | + | + | + | + | + | + | + | + | |
| 3-Hydroxybutan-2-one | - | - | + | + | + | + | + | + | + | + | + | + | |
| Esters | Ethyl butanoate | + | + | + | - | + | - | - | - | - | + | + | + |
| Ethyl hexanoate | - | - | - | - | + | - | - | - | - | + | + | + | |
| Ethyl propanoate | - | - | - | - | + | + | - | - | - | + | + | + | |
| Propyl butanoate | - | - | - | - | + | - | - | - | - | + | + | + | |
| Ethyl acetate | - | - | - | - | - | - | - | - | - | + | + | + | |
| Aldehydes | 2-Methylbutanal | - | - | - | - | + | + | - | + | + | + | + | + |
| Benzaldehyde | + | + | + | - | - | - | - | - | - | + | + | + | |
| Butanal | - | - | - | + | + | - | - | + | - | - | + | + | |
| Hexanal | - | - | - | - | - | - | - | - | + | + | + | + | |
| 2-Methylpropanal | - | - | - | - | + | + | - | + | + | + | + | + | |
| 3-Methylthiopropanal | + | + | + | + | + | + | + | + | + | + | + | + | |
| Octanal | - | - | - | - | - | - | - | - | - | - | - | + | |
| Acetaldehyde | - | - | - | - | + | + | - | + | + | + | + | + | |
| Alcohols | 2-Butanol | - | - | + | - | - | + | - | - | + | + | + | + |
| 2-Methyl-1-propanol | - | - | - | + | + | + | + | + | + | + | + | + | |
| 2-Heptanol | - | - | + | - | - | + | - | - | + | + | + | + | |
| n-Hexanol | + | + | + | - | + | - | + | + | + | + | + | + | |
| 1-Butanol | + | + | + | + | + | + | + | + | + | + | + | + | |
| Pentan-1-ol | - | - | - | - | + | + | - | + | - | + | + | + | |
| 3-Methyl-1-butanol | - | - | + | + | + | + | + | + | + | + | + | + | |
| Acids | Acetic acid | - | - | - | + | + | + | + | + | + | + | + | + |
| Chemical Group | VOC | MRS Media | Milk at 37 °C | Milk at 12 °C | Matured Cheese | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CT | Lps | Lcl | CT | Lps | Lcl | CT | Lps | Lcl | Rind | Inner | Center | ||
| Ketones | 2-Heptanone | - | + | - | + | + | + | + | + | + | + | + | + |
| 2-Hexanone | + | + | + | - | - | - | - | - | - | + | + | + | |
| 2-Pentanone | - | + | + | + | + | + | + | + | + | + | + | - | |
| 2-Butanone | - | + | + | + | + | + | + | + | + | + | + | + | |
| 3-Hydroxybutan-2-one | - | - | + | + | + | + | + | + | + | + | + | + | |
| Esters | Ethyl butanoate | + | + | + | - | - | - | - | - | - | + | + | + |
| Ethyl hexanoate | - | - | - | - | - | - | - | - | - | + | + | + | |
| Ethyl propanoate | - | - | - | - | + | + | - | - | - | - | - | - | |
| Propyl butanoate | - | - | - | - | - | - | - | - | - | - | - | - | |
| Ethyl acetate | - | - | - | - | - | - | - | - | - | - | - | - | |
| Aldehydes | 2-Methylbutanal | - | - | - | - | + | + | - | + | + | + | + | + |
| Benzaldehyde | + | + | + | - | - | - | - | - | - | + | + | + | |
| Butanal | - | - | - | + | + | - | - | + | + | + | + | + | |
| Hexanal | - | - | - | - | - | + | - | - | - | + | + | + | |
| 2-Methylpropanal | - | - | - | - | + | + | - | + | + | + | + | + | |
| 3-Methylthiopropanal | + | + | + | + | + | + | + | + | + | + | + | + | |
| Octanal | - | - | - | - | - | - | - | - | - | + | + | + | |
| Acetaldehyde | - | - | - | - | + | + | - | + | + | + | + | + | |
| Alcohols | 2-Butanol | - | + | + | - | + | + | - | + | + | + | + | + |
| 2-Methyl-1-propanol | - | - | - | + | + | + | + | + | + | + | + | + | |
| 2-Heptanol | - | - | + | - | - | + | - | - | + | + | + | + | |
| n-Hexanol | + | - | + | - | + | - | + | + | + | + | + | + | |
| 1-Butanol | + | + | + | + | + | + | + | + | + | + | + | + | |
| Pentan-1-ol | - | - | - | - | + | - | - | - | + | - | - | - | |
| 3-Methyl-1-butanol | - | + | + | + | + | + | + | + | + | - | - | - | |
| Acids | Acetic acid | - | - | - | + | + | + | + | + | + | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz, M.J.; Salatti-Dorado, J.A.; Cardador, M.J.; Frizzo, L.; Jordano, R.; Arce, L.; Medina, L.M. Relationship between Volatile Organic Compounds and Microorganisms Isolated from Raw Sheep Milk Cheeses Determined by Sanger Sequencing and GC–IMS. Foods 2023, 12, 372. https://doi.org/10.3390/foods12020372
Ruiz MJ, Salatti-Dorado JA, Cardador MJ, Frizzo L, Jordano R, Arce L, Medina LM. Relationship between Volatile Organic Compounds and Microorganisms Isolated from Raw Sheep Milk Cheeses Determined by Sanger Sequencing and GC–IMS. Foods. 2023; 12(2):372. https://doi.org/10.3390/foods12020372
Chicago/Turabian StyleRuiz, María J., José A. Salatti-Dorado, María J. Cardador, Laureano Frizzo, Rafael Jordano, Lourdes Arce, and Luis M. Medina. 2023. "Relationship between Volatile Organic Compounds and Microorganisms Isolated from Raw Sheep Milk Cheeses Determined by Sanger Sequencing and GC–IMS" Foods 12, no. 2: 372. https://doi.org/10.3390/foods12020372
APA StyleRuiz, M. J., Salatti-Dorado, J. A., Cardador, M. J., Frizzo, L., Jordano, R., Arce, L., & Medina, L. M. (2023). Relationship between Volatile Organic Compounds and Microorganisms Isolated from Raw Sheep Milk Cheeses Determined by Sanger Sequencing and GC–IMS. Foods, 12(2), 372. https://doi.org/10.3390/foods12020372








