Characterization of the Psychrotrophic Lactic Acid Bacterium Leuconostoc gelidum subsp. aenigmaticum LS4 Isolated from Kimchi Based on Comparative Analyses of Its Genomic and Phenotypic Properties
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation and Identification of Psychrotrophic LAB from Kimchi
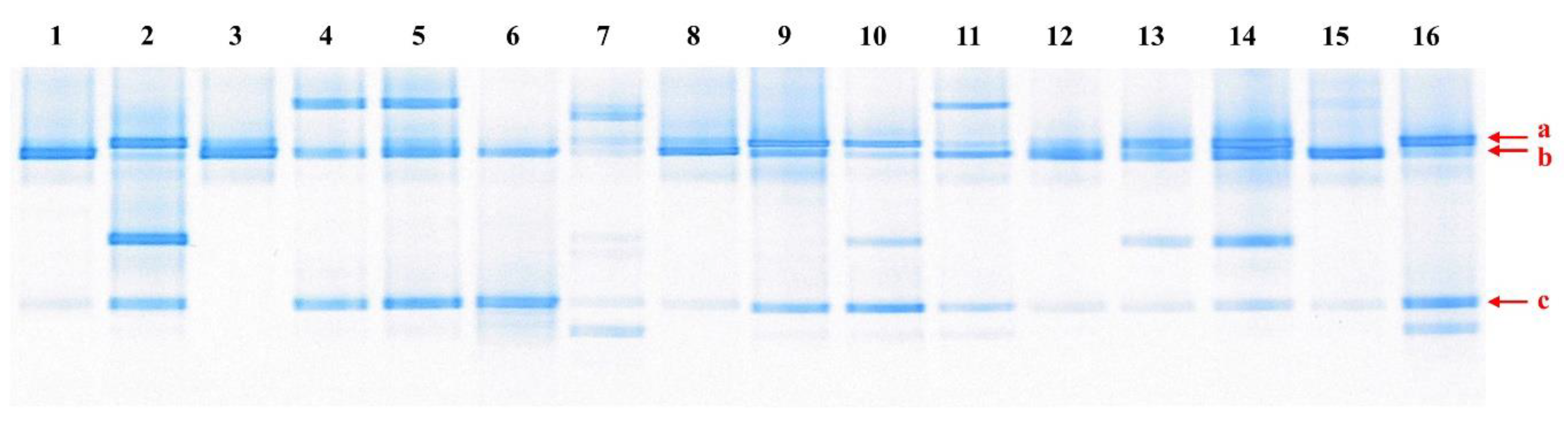
2.2. Polymerase Chain Reaction (PCR)-Denaturing Gradient Gel Electrophoresis (DGGE)
2.3. Physiological and Biochemical Properties of LAB Isolates
2.3.1. Growth at a NaCl Concentration of 6.5%
2.3.2. Heme-Stimulated Aerobic Growth
2.3.3. Carbohydrate Utilization
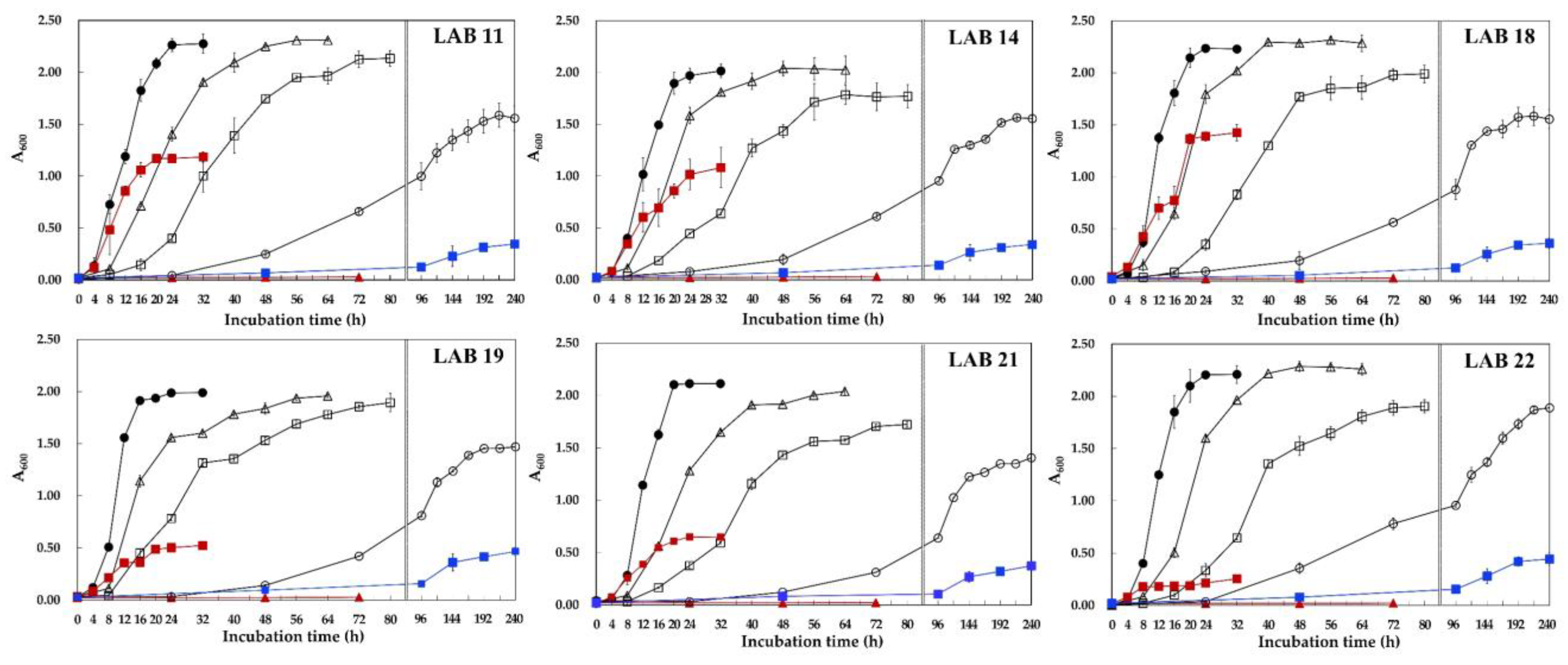
2.3.4. Growth at Low Temperature
2.4. Phenotypic Characteristics as Starter Cultures
2.4.1. Sensory Properties
2.4.2. Hemolysis and Antibiotic Resistance
2.4.3. Enzymatic Activities
2.4.4. Biogenic Amine Production
2.4.5. Stress Tolerance: Temperature, pH, and NaCl concentration
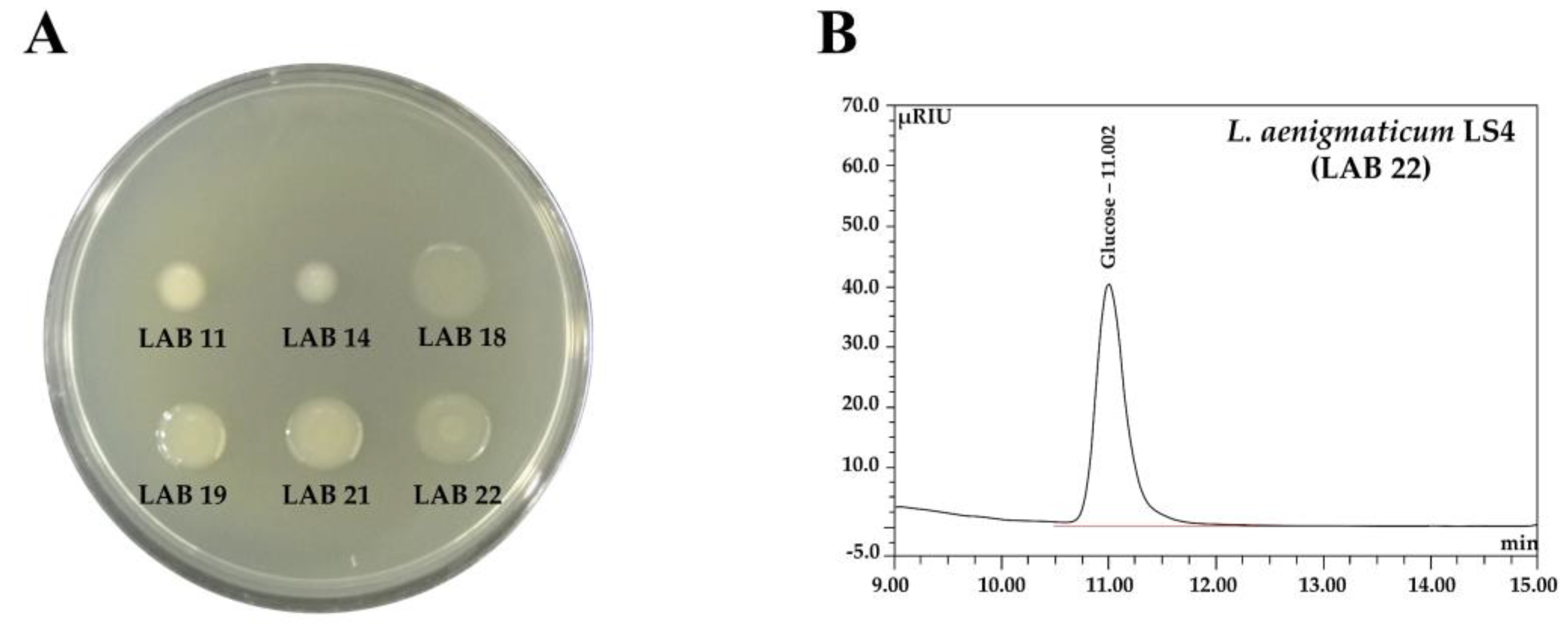
2.4.6. Exopolysaccharide (EPS) Production and Determination of EPS Monosaccharide Composition
2.4.7. Antimicrobial Activity
2.5. Genome Sequencing and Analysis
2.6. Statistical Analysis
3. Results and Discussion
3.1. Microorganisms Present in Kimchi Samples, and Isolation and Identification of Psychrotropic LAB
3.2. Growth at Low Temperature
3.3. Phenotypic Characteristics as Starter Cultures
3.3.1. Sensory Properties
3.3.2. Safety Aspects
3.3.3. Technical Aspects
3.3.4. Functional Aspects
3.4. General Genome Features and Functional Annotations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Park, K.Y.; Kim, H.Y.; Jeong, J.K. Kimchi and its health benefits. In Fermented Foods in Health and Disease Prevention; Frias, J., Martinez-Villaluenga, C., Peñas, E., Eds.; Academic Press: Cambridge, MA, USA, 2017; pp. 477–502. [Google Scholar]
- Moon, S.H.; Kim, C.R.; Chang, H.C. Heterofermentative lactic acid bacteria as a starter culture to control kimchi fermentation. LWT Food Sci. Technol. 2018, 88, 181–188. [Google Scholar] [CrossRef]
- Mun, S.Y.; Moon, S.H.; Choon, C.H. Characterization of High-Ornithine-Producing Weissella koreensis DB1 Isolated from Kimchi and Its Application in Rice Bran Fermentation as a Starter Culture. Foods 2020, 9, 1545. [Google Scholar] [CrossRef]
- Hong, Y.; Yang, H.-S.; Chang, H.; Kim, H.-Y. Comparison of Bacterial Community Changes in Fermenting Kimchi at Two Different Temperatures Using a Denaturing Gradient Gel Electrophoresis Analysis. J. Microbiol. Biotechnol. 2013, 23, 76–84. [Google Scholar] [CrossRef]
- Lee, M.; Song, J.H.; Lee, S.H.; Jung, M.Y.; Chang, J.Y. Effect of seasonal production on bacterial communities in Korean industrial kimchi fermentation. Food Control 2018, 91, 381–389. [Google Scholar] [CrossRef]
- Ahmadsah, L.S.F.; Min, S.-G.; Han, S.-K.; Hong, Y.; Kim, H.-Y. Effect of Low Salt Concentrations on Microbial Changes During Kimchi Fermentation Monitored by PCR-DGGE and Their Sensory Acceptance. J. Microbiol. Biotechnol. 2015, 25, 2049–2057. [Google Scholar] [CrossRef]
- Mun, S.Y.; Chang, H.C. Characterization of Weissella koreensis SK Isolated from Kimchi Fermented at Low Temperature (around 0 °C) Based on Complete Genome Sequence and Corresponding Phenotype. Microorganisms 2020, 8, 1147. [Google Scholar] [CrossRef]
- Jung, J.Y.; Lee, S.H.; Jeon, C.O. Kimchi microflora: History, current status, and perspectives for industrial kimchi production. Appl. Microbiol. Biotechnol. 2014, 98, 2385–2393. [Google Scholar] [CrossRef] [PubMed]
- Ryu, E.H.; Chang, H.C. In Vitro study of potentially probiotic lactic acid bacteria strains isolated from kimchi. Ann. Microbiol. 2013, 63, 1387–1395. [Google Scholar] [CrossRef]
- Chae, S.J.; Kim, E.J.; Chang, H.C. A novel NADH fluorescence-based method for identifying and monitoring lactic acid bacteria growths in kimchi. Int. J. Food Sci. Technol. 2020, 56, 2946–2960. [Google Scholar] [CrossRef]
- Cho, J.; Lee, D.; Yang, C.; Jeon, J.; Kim, J.; Han, H. Microbial population dynamics of kimchi, a fermented cabbage product. FEMS Microbiol. Lett. 2006, 257, 262–267. [Google Scholar] [CrossRef]
- Jeong, S.E.; Chun, B.H.; Kim, K.H.; Park, D.; Roh, S.W.; Lee, S.H.; Jeon, C.O. Genomic and metatranscriptomic analyses of Weissella koreensis reveal its metabolic and fermentative features during kimchi fermentation. Food Microbiol. 2018, 76, 1–10. [Google Scholar] [CrossRef]
- Park, J.-M.; Shin, J.-H.; Lee, D.-W.; Song, J.-C.; Suh, H.-J.; Chang, U.-J.; Kim, J.-M. Identification of the lactic acid bacteria in Kimchi according to initial and over-ripened fermentation using PCR and 16S rRNA gene sequence analysis. Food Sci. Biotechnol. 2010, 19, 541–546. [Google Scholar] [CrossRef]
- Rahkila, R.; de Bruyne, K.; Johansson, P.; Vandamme, P.; Bjorkroth, J. Reclassification of Leuconostoc gasicomitatum as Leuconostoc gelidum subsp. gasicomitatum comb. nov., description of Leuconostoc gelidum subsp. aenigmaticum subsp. nov., designation of Leuconostoc gelidum subsp. gelidum subsp. nov. and emended description of Leuconostoc gelidum. Int. J. Syst. Evol. Microbiol. 2014, 64, 1290–1295. [Google Scholar] [CrossRef]
- Hébraud, M.; Pituer, P. Cold shock response and low temperature adaptation in psychrotrophic bacteria. J. Mol. Microbial. Biotechnol. 1999, 1, 211–219. [Google Scholar]
- Mahon, C.R.; Lehman, D.C.; Manuselis, G., Jr. Textbook of Diagnostic Microbiology, 6th ed.; Elsvier: Amsterdam, The Netherlands, 2018; pp. 11–504. [Google Scholar]
- Watthanasakphuban, N.; Virginia, L.; Haltrich, D.; Peterbauer, C. Analysis and Reconstitution of the Menaquinone Biosynthesis Pathway in Lactiplantibacillus plantarum and Lentilactibacillus buchneri. Microorganisms 2021, 9, 1476. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.-W.; Heo, S.; Lee, J.-H. Safety assessment of Tetragenococcus halophilus isolates from doenjang, a Korean high-salt-fermented soybean paste. Food Microbiol. 2017, 62, 92–98. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA). Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J. 2012, 10, 2740. [Google Scholar]
- Bover-Cid, S.; Holzapfel, W.H. Improved screening procedure for biogenic amine production by lactic acid bacteria. Int. J. Food Microbiol. 1999, 53, 33–41. [Google Scholar] [CrossRef]
- Diez-Ozaeta, I.; Amárita, F.; Lavilla, M.; Rainieri, S. Ecology of indigenous lactic acid bacteria from Rioja Alavesa red wines, focusing on biogenic amine production ability. LWT Food Sci. Technol. 2019, 116, 108544. [Google Scholar] [CrossRef]
- Bargossi, E.; Tabanelli, G.; Montanari, C.; Gatto, V.; Chinnici, F.; Gardini, F.; Torriani, S. Growth, biogenic amine production and tyrDC transcription of Enterococcus faecalisin synthetic medium containing defined amino acid concentrations. J. Appl. Microbiol. 2017, 122, 1078–1091. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Chang, H.C. Isolation and characterization of exopolysaccharide producing lactic acid bacteria from Kimchi. Microbiol. Biotechnol. Lett. 2006, 34, 196–203. [Google Scholar]
- Bukhari, S.A.; Salman, M.; Numan, M.; Javed, M.R.; Zubair, M.; Mustafa, G. Characterization of antifungal metabolites produced by Lactobacillus plantarum and Lactobacillus coryniformis isolated from rice rinsed water. Mol. Biol. Rep. 2020, 47, 1871–1881. [Google Scholar] [CrossRef]
- Mun, S.Y.; Kim, S.K.; Woo, E.R.; Chang, H.C. Purification and characterization of an antimicrobial compound produced by Lactobacillus plantarum EM showing both antifungal and antibacterial activities. LWT Food Sci. Technol. 2019, 114, 108403. [Google Scholar] [CrossRef]
- Kim, B.J.; Lee, H.J.; Park, S.Y.; Kim, J.H.; Han, H.U. Identification and characterization of Leuconostoc gelidum, isolated from kimchi, a fermented cabbage product. J. Microbiol. 2000, 38, 132–136. [Google Scholar]
- Kim, B.; Lee, J.; Jang, J.; Kim, J.; Han, H. Leuconostoc inhae sp. nov., a lactic acid bacterium isolated from kimchi. Int. J. Syst. Evol. Microbiol. 2003, 53, 1123–1126. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.Y.; Lee, S.H.; Jeon, C.O. Complete Genome Sequence of Leuconostoc gelidum Strain JB7, Isolated from Kimchi. J. Bacteriol. 2012, 194, 6665. [Google Scholar] [CrossRef]
- Kim, D.-S.; Choi, S.-H.; Kim, D.-W.; Kim, R.N.; Nam, S.-H.; Kang, A.; Kim, A.; Park, H.-S. Genome Sequence of Leuconostoc gelidum KCTC 3527, Isolated from Kimchi. J. Bacteriol. 2011, 193, 799–800. [Google Scholar] [CrossRef]
- Pothakos, V.; Snauwaert, C.; de Vos, P.; Huys, G.; Devlieghere, F. Psychrotrophic members of Leuconostoc gasicomitatum, Leuconostoc gelidum and Lactococcus piscium dominate at the end of shelf-life in packaged and chilled-stored food products in Belgium. Food Microbiol. 2014, 39, 61–67. [Google Scholar] [CrossRef]
- Shaw, B.G.; Harding, C.D. Leuconostoc gelidum sp. nov. and Leuconostoc carnosum sp. nov. from Chill-Stored Meats. Int. J. Syst. Bacteriol. 1989, 39, 217–223. [Google Scholar] [CrossRef]
- ISO 15214:1998. Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Mesophilic Lactic Acid Bacteria—Colony-Count Technique at 30 Degrees C; International Organization for Standardization: Geneva, Switzerland, 1998. [Google Scholar]
- ISO 4833:2003. Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Microorganisms—Colony-Count Technique at 30 Degrees C; International Organization for Standardization: Geneva, Switzerland, 2003. [Google Scholar]
- Gounot, A.-M. Psychrophilic and psychrotrophic microorganisms. Cell. Mol. Life Sci. 1986, 42, 1192–1197. [Google Scholar] [CrossRef] [PubMed]
- Jääskeläinen, E.; Vesterinen, S.; Parshintsev, J.; Johansson, P.; Riekkola, M.-L.; Bjorkroth, J. Production of Buttery-Odor Compounds and Transcriptome Response in Leuconostoc gelidum subsp. gasicomitatum LMG18811T during Growth on Various Carbon Sources. Appl. Environ. Microbiol. 2015, 81, 1902–1908. [Google Scholar] [CrossRef]
- Johansson, P.; Paulin, L.; Säde, E.; Salovuori, N.; Alatalo, E.R.; Björkroth, K.J.; Auvinen, P. Genome Sequence of a Food Spoilage Lactic Acid Bacterium, Leuconostoc gasicomitatum LMG 18811T, in Association with Specific Spoilage Reactions. Appl. Environ. Microbiol. 2011, 77, 4344–4351. [Google Scholar] [CrossRef]
- Pothakos, V.; Devlieghere, F.; Villani, F.; Bjorkroth, J.; Ercolini, D. Lactic acid bacteria and their controversial role in fresh meat spoilage. Meat Sci. 2015, 109, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.W.; Shim, J.M.; Park, S.-K.; Heo, H.-J.; Kim, H.-J.; Ham, K.-S.; Kim, J.H. Isolation of lactic acid bacteria with probiotic potentials from kimchi, traditional Korean fermented vegetable. LWT Food Sci. Technol. 2016, 71, 130–137. [Google Scholar] [CrossRef]
- Benmechernene, Z.; Chentouf, H.F.; Yahia, B.; Fatima, G.; Quintela-Baluja, M.; Calo-Mata, P.; Barros-Velázquez, J. Technological Aptitude and Applications of Leuconostoc mesenteroides Bioactive Strains Isolated from Algerian Raw Camel Milk. BioMed Res. Int. 2013, 2013, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Choi, I.-K.; Jung, S.-H.; Kim, B.-J.; Park, S.-Y.; Kim, J.; Han, H.-U. Novel Leuconostoc citreum starter culture system for the fermentation of kimchi, a fermented cabbage product. Antonie Van Leeuwenhoek 2003, 84, 247–253. [Google Scholar] [CrossRef]
- Development of Technique for Microbial Succession of Kimchi to Control the Fermentation and Constant Quality: Final Report; Ministry of Agriculture, Food and Rural Affairs: Sejong, Korea, 2013.
- Lynch, K.; Zannini, E.; Coffey, A.; Arendt, E.K. Lactic Acid Bacteria Exopolysaccharides in Foods and Beverages: Isolation, Properties, Characterization, and Health Benefits. Annu. Rev. Food Sci. Technol. 2018, 9, 155–176. [Google Scholar] [CrossRef] [PubMed]
- Korcz, E.; Varga, L. Exopolysaccharides from lactic acid bacteria: Techno-functional application in the food industry. Trends Food Sci. Technol. 2021, 110, 375–384. [Google Scholar] [CrossRef]
- Zikmanis, P.; Brants, K.; Kolesovs, S.; Semjonovs, P. Extracellular polysaccharides produced by bacteria of the Leuconostoc genus. World J. Microbiol. Biotechnol. 2020, 36, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Duru, I.C.; Ylinen, A.; Belanov, S.; Pulido, A.A.; Paulin, L.; Auvinen, P. Transcriptomic time-series analysis of cold- and heat-shock response in psychrotrophic lactic acid bacteria. BMC Genom. 2021, 22, 1–16. [Google Scholar] [CrossRef]
- Capozzi, V.; Fiocco, D.; Spano, G. Responses of lactic acid bacteria to cold stress. In Stress Responses of Lactic Acid Bacteria; Tsakalidou, E., Papadimitriou, K., Eds.; Springer: Boston, MA, USA, 2011; pp. 91–110. [Google Scholar]
- Mokoena, M.P. Lactic Acid Bacteria and Their Bacteriocins: Classification, Biosynthesis and Applications against Uropathogens: A Mini-Review. Molecules 2017, 22, 1255. [Google Scholar] [CrossRef] [PubMed]
| Sampling Location | Kimchi Sample | Isolate | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Province | City/District | Kimchi | Storage Temperature | Storage Period | Strain | Length of 16S rRNA Gene (bp) | Similarity (%) | Identified as LAB 1 | Accession No. 2 |
| Seoul/ Incheon/ Gyeonggi | Seoul | Kimchi 1 | 0 °C | 1 month | LAB 1 | 1350 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 |
| LAB 2 | 1517 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |||||
| Ganghwa | Kimchi 2 | 10 °C | 3 months | LAB 3 | 1510 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |
| Yangpyeong | Kimchi 3 | 4 °C | 2 months | LAB 4 | 1522 | 99.46 | Lb. algidus | NR_028617.1 | |
| Hwaseong | Kimchi 4 | 4 °C | 3 months | LAB 5 | 1509 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |
| Gangwon | Wonju | Kimchi 5 | 3 °C ↓ | 3 months | LAB 6 | 1467 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 |
| Chungcheong | Seocheon | Kimchi 6 | 0 °C | 2~3 months | LAB 7 | 1468 | 99.59 | L. gelidum subsp. gasicomitatum | NR_074997.2 |
| LAB 8 | 1489 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |||||
| Daegu/ Busan/ Gyeongsang | Daegu | Kimchi 7 | 0 °C | 2 months | LAB 9 | 1507 | 99.93 | L. carnosum | NR_040811.1 |
| Kimchi 8 | 10 °C ↓ | 3 months | LAB 10 | 1504 | 99.66 | L. inhae | NR_025204.1 | ||
| Busan | Kimchi 9 | 4 °C | 3 months | LAB 11 | 1518 | 100.00 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |
| LAB 12 | 1509 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |||||
| LAB 13 | 1512 | 99.67 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |||||
| Andong | Kimchi 10 | 2~3 °C | 2~3 months | LAB 14 | 1501 | 99.93 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |
| LAB 15 | 1500 | 99.66 | L. inhae | NR_025204.1 | |||||
| Gimhae | Kimchi 11 | 5~10 °C | 1 month | LAB 16 | 1519 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |
| LAB 17 | 1518 | 99.33 | Lb. algidus | NR_028617.1 | |||||
| LAB 18 | 1501 | 100.00 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |||||
| Uiryeong | Kimchi 12 | 0 °C | 2 months | LAB 19 | 1479 | 100.00 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |
| Gwangju/ Jeolla | Gwangju | Kimchi 13 | 6.5 °C | 2 months | LAB 20 | 1519 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 |
| Sunchang | Kimchi 14 | 4 °C | 3 months | LAB 21 | 1519 | 99.93 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |
| Iksan | Kimchi 15 | 0 °C | 1 month | LAB 22 | 1511 | 100.00 | L. gelidum subsp. aenigmaticum | NR_133769.1 | |
| Jinan | Kimchi 16 | 0 °C | 3~4 months | LAB 23 | 1429 | 100.00 | L. gelidum subsp. gasicomitatum | NR_074997.2 | |
| Kimchi Juice | Sourness | Fresh Taste | Carbonate Taste | Pleasant Fermentative Smell | Sewerage-Like Smell | Overall Acceptability |
|---|---|---|---|---|---|---|
| Control juice | 1.00 ± 0.00 c | 2.00 ± 0.00 c | 1.00 ± 0.00 b | 1.00 ± 0.00 d | 1.22 ± 0.44 c | 3.00 ± 0.00 c |
| LAB 11 juice | 4.11 ± 0.14 a | 1.89 ± 0.60 c | 3.78 ± 0.44 a | 2.89 ± 0.33 c | 3.44 ± 1.13 ab | 2.00 ± 0.87 de |
| LAB 14 juice | 4.22 ± 0.44 a | 2.00 ± 0.71 c | 3.78 ± 0.44 a | 3.00 ± 0.00 c | 3.67 ± 0.87 ab | 2.00 ± 0.71 de |
| LAB 18 juice | 3.56 ± 0.53 b | 1.56 ± 0.73 c | 3.78 ± 0.44 a | 3.11 ± 0.33 c | 4.22 ± 0.67 a | 1.44 ± 0.73 e |
| LAB 19 juice | 3.56 ± 0.53 b | 2.11 ± 1.05 c | 3.78 ± 0.44 a | 3.00 ± 0.00 c | 3.00 ± 1.32 b | 2.22 ± 0.33 d |
| LAB 21 juice | 3.56 ± 0.53 b | 4.33 ± 0.50 a | 3.78 ± 0.44 a | 4.11 ± 0.48 a | 1.11 ± 0.22 c | 4.78 ± 0.44 a |
| LAB 22 juice | 3.44 ± 0.53 b | 3.11 ± 0.33 b | 3.78 ± 0.44 a | 3.67 ± 0.71 b | 1.89 ± 0.93 c | 4.00 ± 0.71 b |
| Antibiotics 1 | AMP | CHL | ERY | GEN | KAN | STR | TET | VAN |
|---|---|---|---|---|---|---|---|---|
| Breakpoints for Leuconostocs 2 | 2 | 4 | 1 | 16 | 16 | 64 | 8 | N.R 3 |
| L. aenigmaticum LS4 | 2 | 4 | 0.06 | 0.125 | 2 | 4 | 1 | 512 |
| Stress | Viable Cell Counts (log CFU/mL) | ||||
|---|---|---|---|---|---|
| 0 h | 24 h | 48 h | 72 h | ||
| Temperature | −2 °C | 9.40 ± 0.04 az | 9.44 ± 0.00 az | 9.43 ± 0.00 az | 9.42 ± 0.01 az |
| 0 °C | 9.40 ± 0.04 az | 9.42 ± 0.01 az | 9.37 ± 0.03 az | 9.37 ± 0.07 az | |
| 4 °C | 9.40 ± 0.04 azA | 9.46 ± 0.01 azA | 9.38 ± 0.04 azA | 9.38 ± 0.05 azA | |
| 10 °C | 9.40 ± 0.04 az | 9.46 ± 0.03 az | 9.41 ± 0.00 az | 9.36 ± 0.06 az | |
| 20 °C | 9.40 ± 0.04 az | 9.49 ± 0.06 az | 9.47 ± 0.02 az | 9.18 ± 0.03 by | |
| 25 °C (Control) | 9.40 ± 0.04 az | 9.33 ± 0.10 az | 8.77 ± 0.04 by | 5.06 ± 0.10 cx | |
| 30 °C | 9.40 ± 0.04 az | 8.82 ± 0.31 bz | 4.36 ± 0.48 cy | N.D 1 | |
| 50 °C | 9.40 ± 0.04 az | N.D | N.D | N.D | |
| 70 °C | 9.40 ± 0.04 az | N.D | N.D | N.D | |
| pH | pH 2.0 | 9.40 ± 0.04 az | N.D | N.D | N.D |
| pH 4.0 | 9.40 ± 0.04 azA | 8.77 ± 0.07 byC | 6.11 ± 0.05 cxC | 3.03 ± 0.10 cwD | |
| pH 6.0 | 9.40 ± 0.04 az | 9.34 ± 0.02 az | 8.46 ± 0.12 by | 4.94 ± 0.05 bx | |
| pH 6.5 (Control) | 9.40 ± 0.04 az | 9.33 ± 0.10 az | 8.77 ± 0.04 ay | 5.06 ± 0.10 bx | |
| pH 8.0 | 9.40 ± 0.04 az | 9.37 ± 0.09 az | 8.41 ± 0.01 by | 6.19 ± 0.27 ax | |
| pH 10.0 | 9.40 ± 0.04 az | N.D | N.D | N.D | |
| NaCl | 0 % (Control) | 9.40 ± 0.04 az | 9.33 ± 0.10 az | 8.77 ± 0.04 ay | 5.06 ± 0.10 cx |
| 1.0% | 9.40 ± 0.04 az | 9.37 ± 0.01 az | 8.14 ± 0.10 by | 5.85 ± 0.40 bx | |
| 3.0% | 9.40 ± 0.04 azA | 9.19 ± 0.09 ayB | 8.60 ± 0.07 abxB | 8.56 ± 0.04 bxC | |
| 5.0% | 9.40 ± 0.04 az | 9.12 ± 0.09 ay | 8.51 ± 0.10 abx | 8.39 ± 0.06 ax | |
| 7.0% | 9.40 ± 0.04 az | 8.60 ± 0.08 by | 6.82 ± 0.10 cx | 3.82 ± 0.31 dw | |
| 9.0% | 9.40 ± 0.04 az | 4.42 ± 0.50 cy | 2.75 ± 0.27 dx | N.D | |
| 12.0% | 9.40 ± 0.04 az | 3.92 ± 0.13 cy | 2.93 ± 0.11 dx | N.D | |
| 15.0% | 9.40 ± 0.04 az | 4.20 ± 0.05 cy | 2.96 ± 0.46 dx | N.D | |
| 4 °C, pH 4.0 | 9.40 ± 0.04 azA | 9.31 ± 0.01 azB | 9.32 ± 0.05 azA | 9.10 ± 0.06 ayB | |
| 4 °C, pH 4.0, NaCl 3% | 9.40 ± 0.04 azA | 9.31 ± 0.03 azB | 9.33 ± 0.05 azA | 9.19 ± 0.04 ayB | |
| Indicator Species | Diameter of Inhibition Zone (mm) | |||
|---|---|---|---|---|
| Culture Filtrate of LS4 1 | Protease Treated Culture Filtrate 2 | Mixture of Organic Acids 3 | ||
| Molds | Aspergillus flavus ATCC 22546™ | 12.71 ± 0.23 a | 12.44 ± 0.08 a | 12.53 ± 0.12 a |
| Aspergillus fumigatus ATCC 96918™ | 14.19 ± 0.25 a | 13.95 ± 0.15 a | 13.81 ± 0.17 a | |
| Aspergillus nidulans PF-3 | 13.31 ± 0.37 a | 13.43 ± 0.06 a | 13.18 ± 0.18 a | |
| Aspergillus ochraceus PF-2 | 11.98 ± 0.50 a | 11.98 ± 0.16 a | 11.88 ± 0.37 a | |
| Penicillium roqueforti ATCC 10110™ | 10.13 ± 0.53 a | 10.67 ± 0.23 a | 10.10 ± 0.60 a | |
| Bacteria | Bacillus cereus ATCC 14579™ | 14.47 ± 0.30 a | 12.39 ± 0.13 b | 12.77 ± 0.24 b |
| Escherichia coli O157:H7 ATCC 43895™ | 13.03 ± 0.07 a | 12.29 ± 0.29 b | 12.39 ± 0.03 b | |
| Listeria monocytogenes ATCC 19113™ | 12.28 ± 0.26 a | 11.43 ± 0.10 b | 11.33 ± 0.25 b | |
| Micrococcus luteus ATCC 4698™ | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a | |
| Pseudomonas aeruginosa KCCM 11328 | 16.10 ± 0.25 a | 14.68 ± 0.24 b | 14.46 ± 0.19 b | |
| Salmonella enterica serovar. Typhi ATCC 14028™ | 14.72 ± 0.12 a | 12.55 ± 0.10 b | 12.43 ± 0.30 b | |
| Staphylococcus aureus KCCM 40881 | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a | |
| Vibrio parahaemolyticus KCCM 11965 | 18.17 ± 0.30 a | 15.22 ± 0.21 b | 14.98 ± 0.18 b | |
| Attribute | Chromosome | Plasmid 1 | Plasmid 2 |
|---|---|---|---|
| Genome size (bp) | 1,988,425 | 19,308 | 11,283 |
| G + C content (%) | 36.8 | 36.7 | 33.3 |
| Total genes (no.) | 2033 | 23 | 13 |
| Protein coding genes (no.) | 1954 | 23 | 13 |
| rRNAs (no.) | 12 | 0 | 0 |
| tRNAs (no.) | 67 | 0 | 0 |
| GenBank Accession No. | CP071950 | CP071951 | CP071952 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mun, S.Y.; Seo, Y.J.; Chang, H.C. Characterization of the Psychrotrophic Lactic Acid Bacterium Leuconostoc gelidum subsp. aenigmaticum LS4 Isolated from Kimchi Based on Comparative Analyses of Its Genomic and Phenotypic Properties. Foods 2021, 10, 1899. https://doi.org/10.3390/foods10081899
Mun SY, Seo YJ, Chang HC. Characterization of the Psychrotrophic Lactic Acid Bacterium Leuconostoc gelidum subsp. aenigmaticum LS4 Isolated from Kimchi Based on Comparative Analyses of Its Genomic and Phenotypic Properties. Foods. 2021; 10(8):1899. https://doi.org/10.3390/foods10081899
Chicago/Turabian StyleMun, So Yeong, Ye Jin Seo, and Hae Choon Chang. 2021. "Characterization of the Psychrotrophic Lactic Acid Bacterium Leuconostoc gelidum subsp. aenigmaticum LS4 Isolated from Kimchi Based on Comparative Analyses of Its Genomic and Phenotypic Properties" Foods 10, no. 8: 1899. https://doi.org/10.3390/foods10081899
APA StyleMun, S. Y., Seo, Y. J., & Chang, H. C. (2021). Characterization of the Psychrotrophic Lactic Acid Bacterium Leuconostoc gelidum subsp. aenigmaticum LS4 Isolated from Kimchi Based on Comparative Analyses of Its Genomic and Phenotypic Properties. Foods, 10(8), 1899. https://doi.org/10.3390/foods10081899







