Characterization and Action Mechanism Analysis of VvmiR156b/c/d-VvSPL9 Module Responding to Multiple-Hormone Signals in the Modulation of Grape Berry Color Formation
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Hormone Treatments
2.2. Ultra-Performance Liquid Chromatography (UPLC) Analysis
2.3. Gas Chromatography-Mass Spectrometer (GC-MS) Analysis
2.4. Total RNA and DNA Isolation, cDNA Synthesis
2.5. Cis-Element Analysis of The Promoters from VvMIR156b/c/d
2.6. Construction of the VvMIR156b/c/d Promoter Vector and Agrobacterium-Mediated Tobacco Transient Transformation
2.7. β-Glucuronidase (GUS) Staining and Activity Detection
2.8. Construction of vvmiR156b/c/d and VvSPL9 Overexpression Vectors and Instantaneous Injection of Tomato
2.9. Gene Expression Analysis
2.10. Statistical Analysis
3. Results
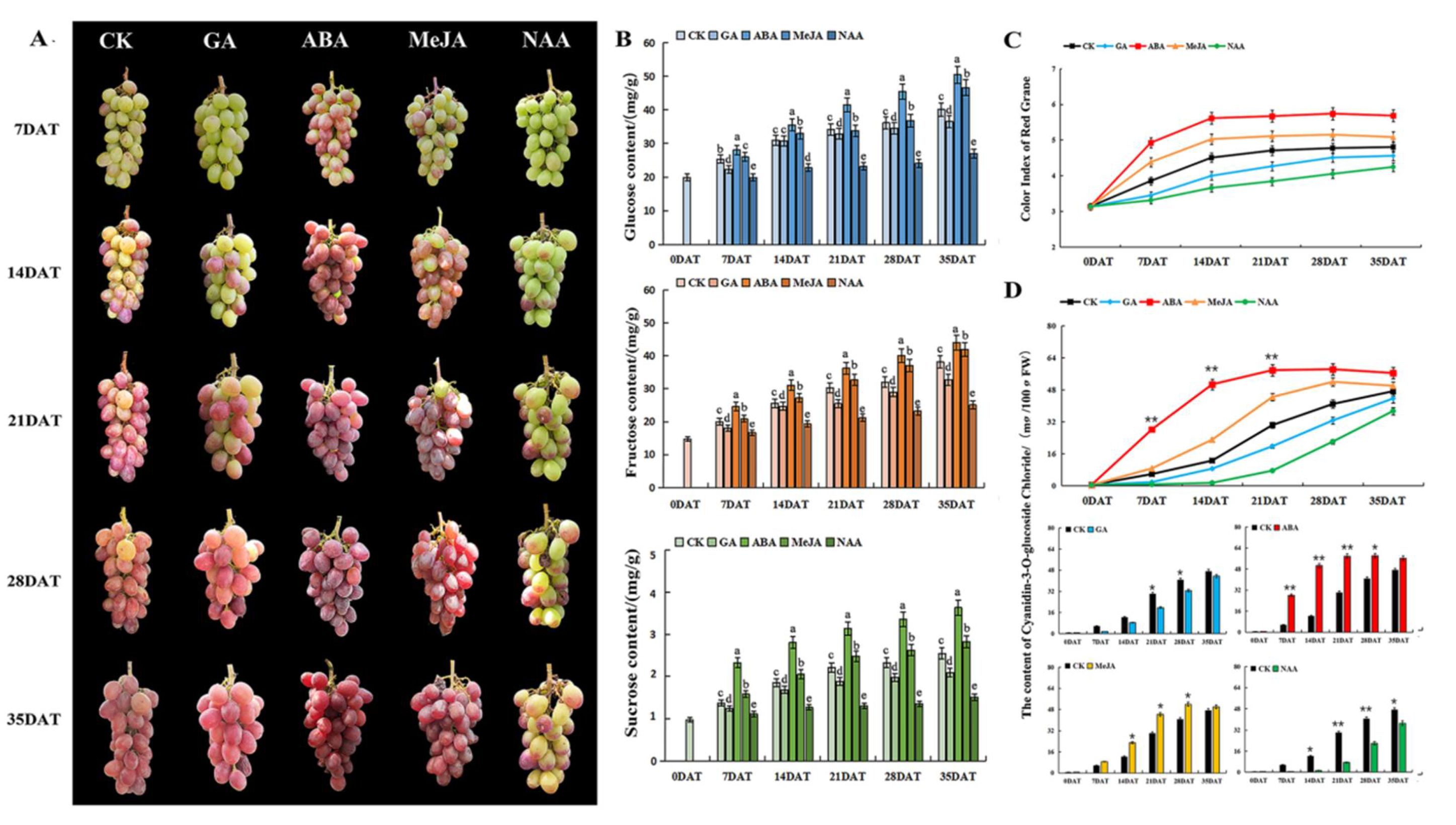
3.1. The Effect of Different Hormones on the Color and Quality Traits of Grape Berry
3.1.1. Effects of Different Hormones on Grape Phenotypic Characteristics
3.1.2. Effects of Different Hormones on Grape Sugar Content
3.1.3. Effects of Different Hormones on Grape Coloring
3.1.4. Effects of Different Hormones on Grape Volatile Compound
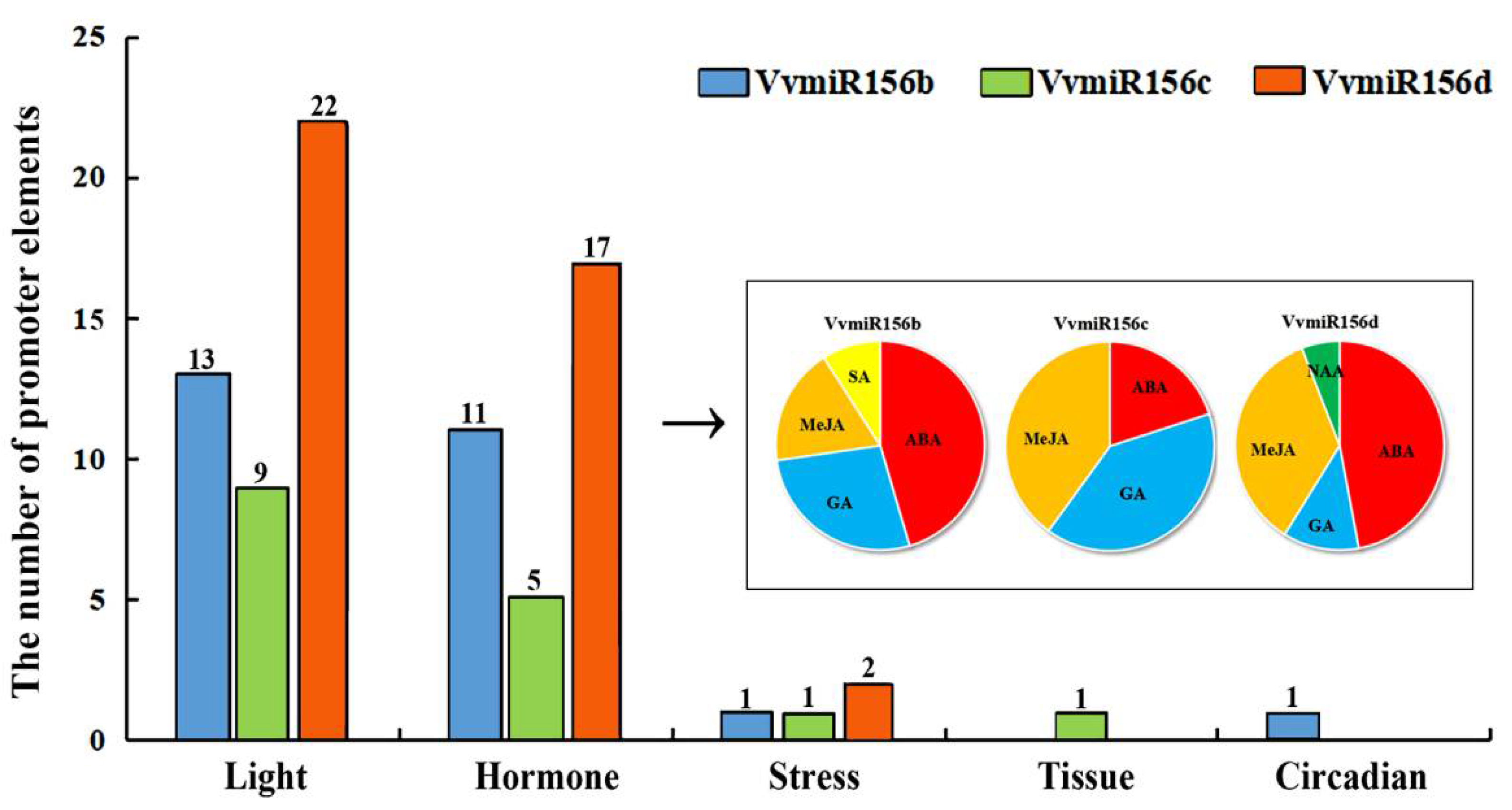
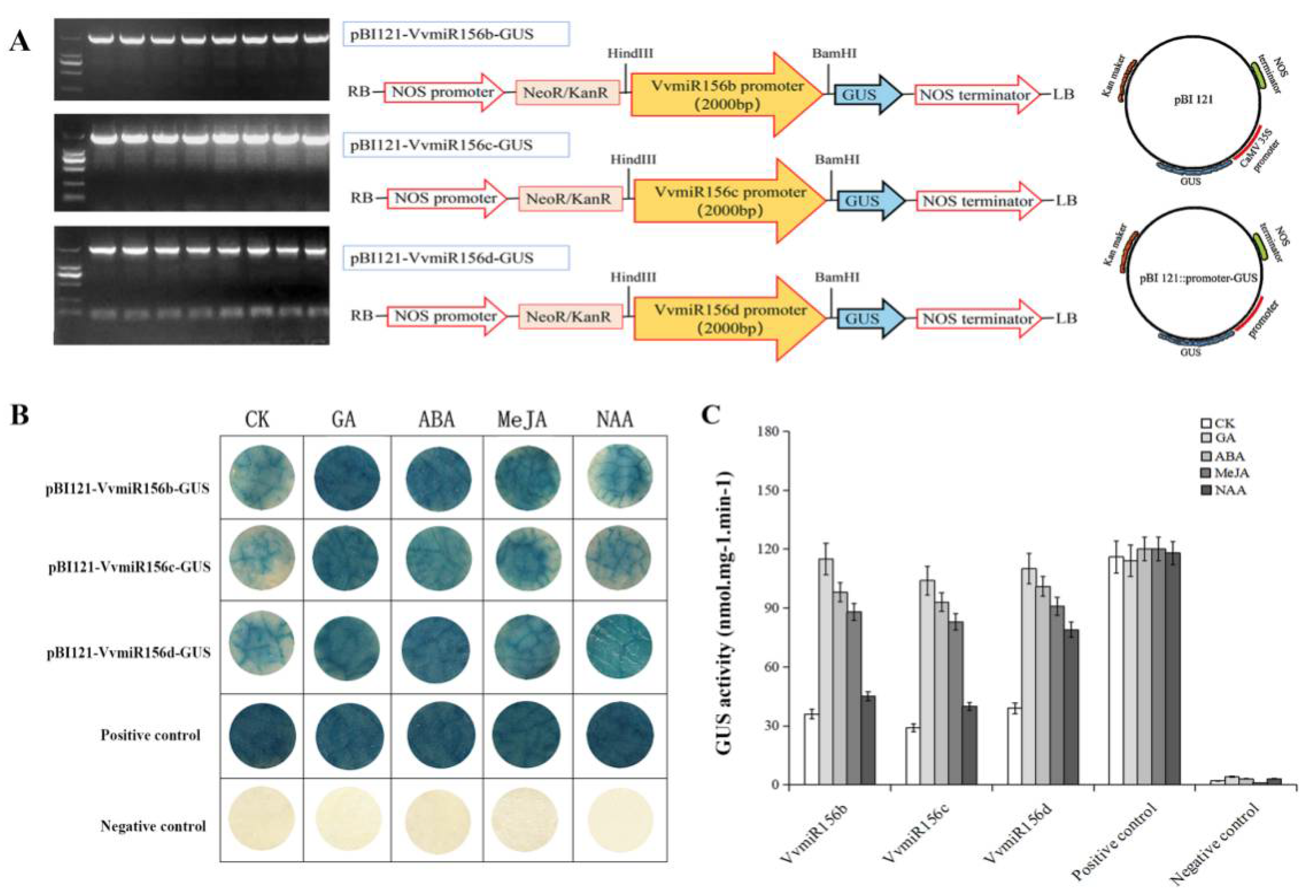
3.2. Cis-element Analysis and GUS Activity Detection of VvMIR156b/c/d Promoters in Response to Exogenous Hormones
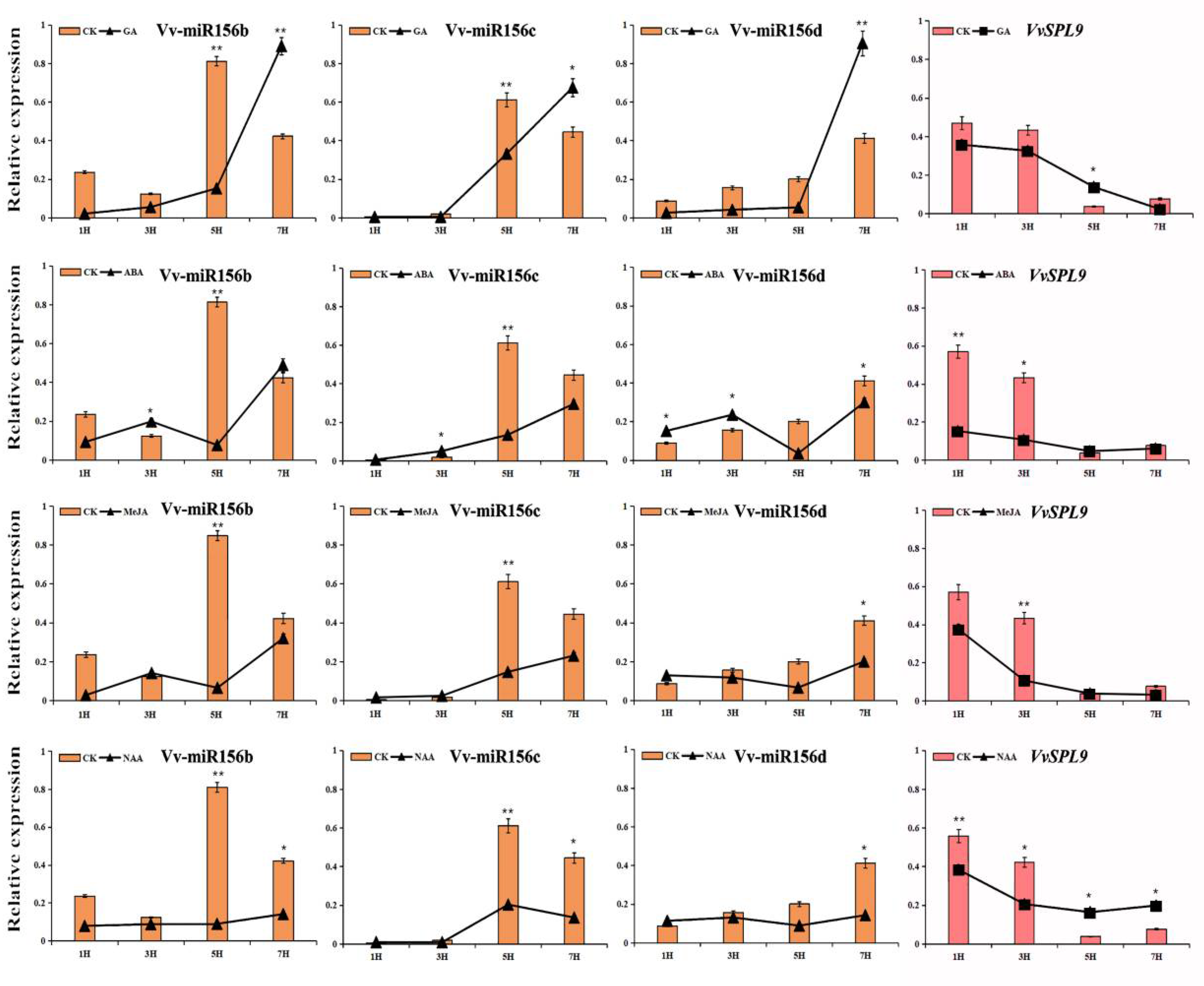
3.3. The Expression Characteristics of VvmiR156b/c/d and VvSPL9 Responsive to Various Hormones in the Modulation of Grape Color Formation
3.3.1. Short-Term Response of VvmiR156b/c/d and VvSPL9 to Exogenous Hormones in ‘Wink’ Leaves
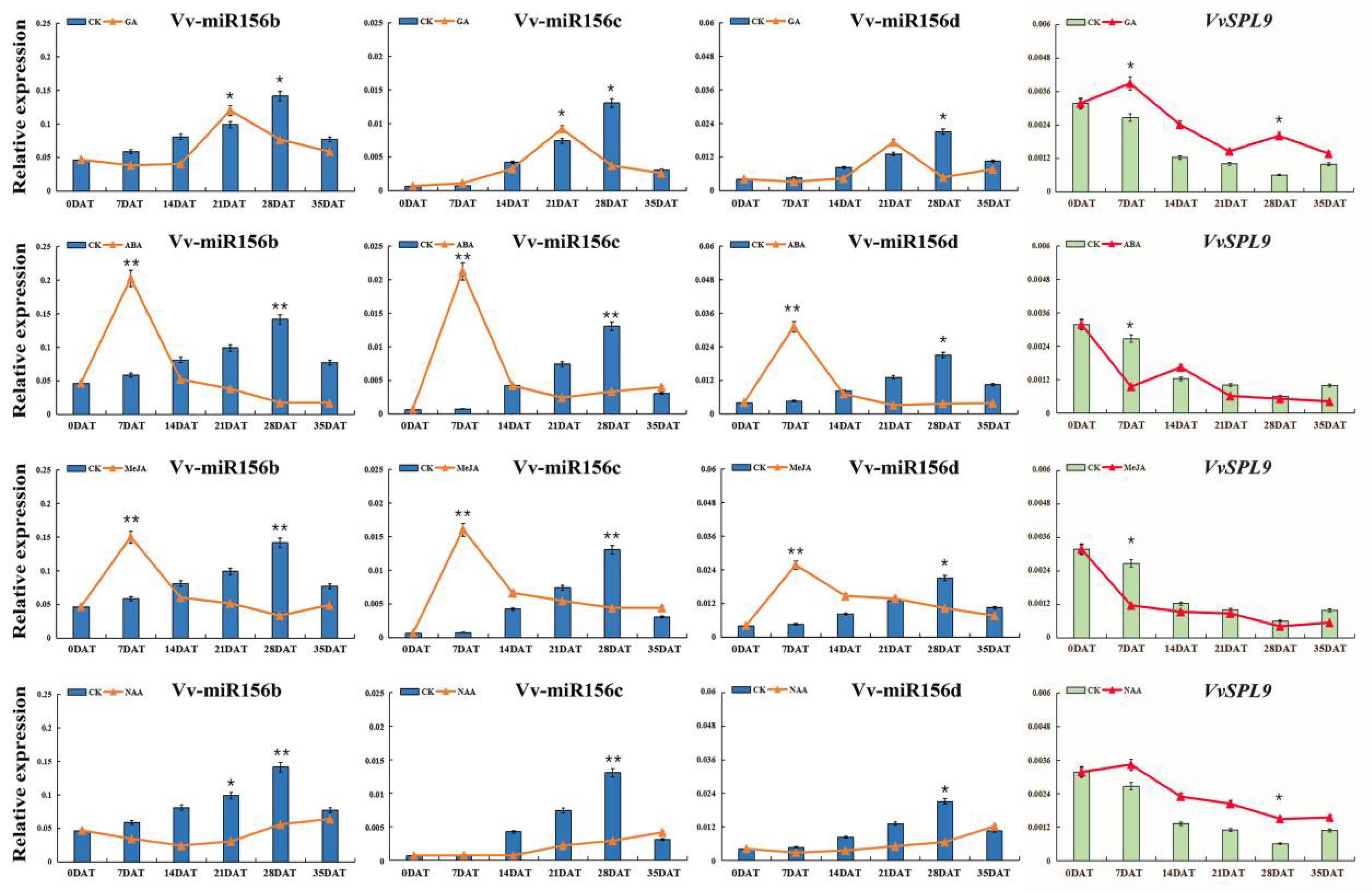
3.3.2. Long-Term Response of VvmiR156b/c/d and VvSPL9 to Exogenous Hormones in ‘Wink’ Berry Skin
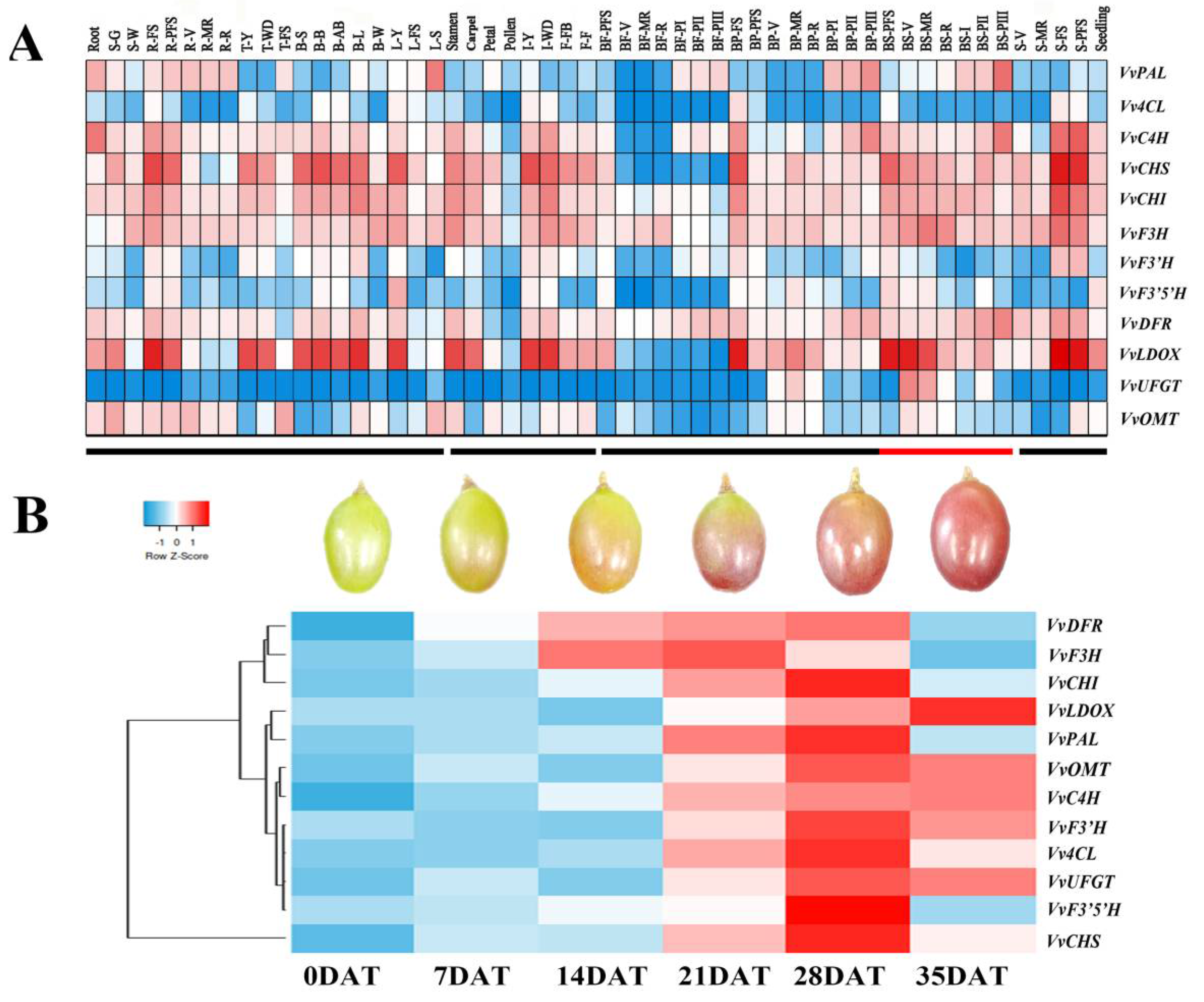
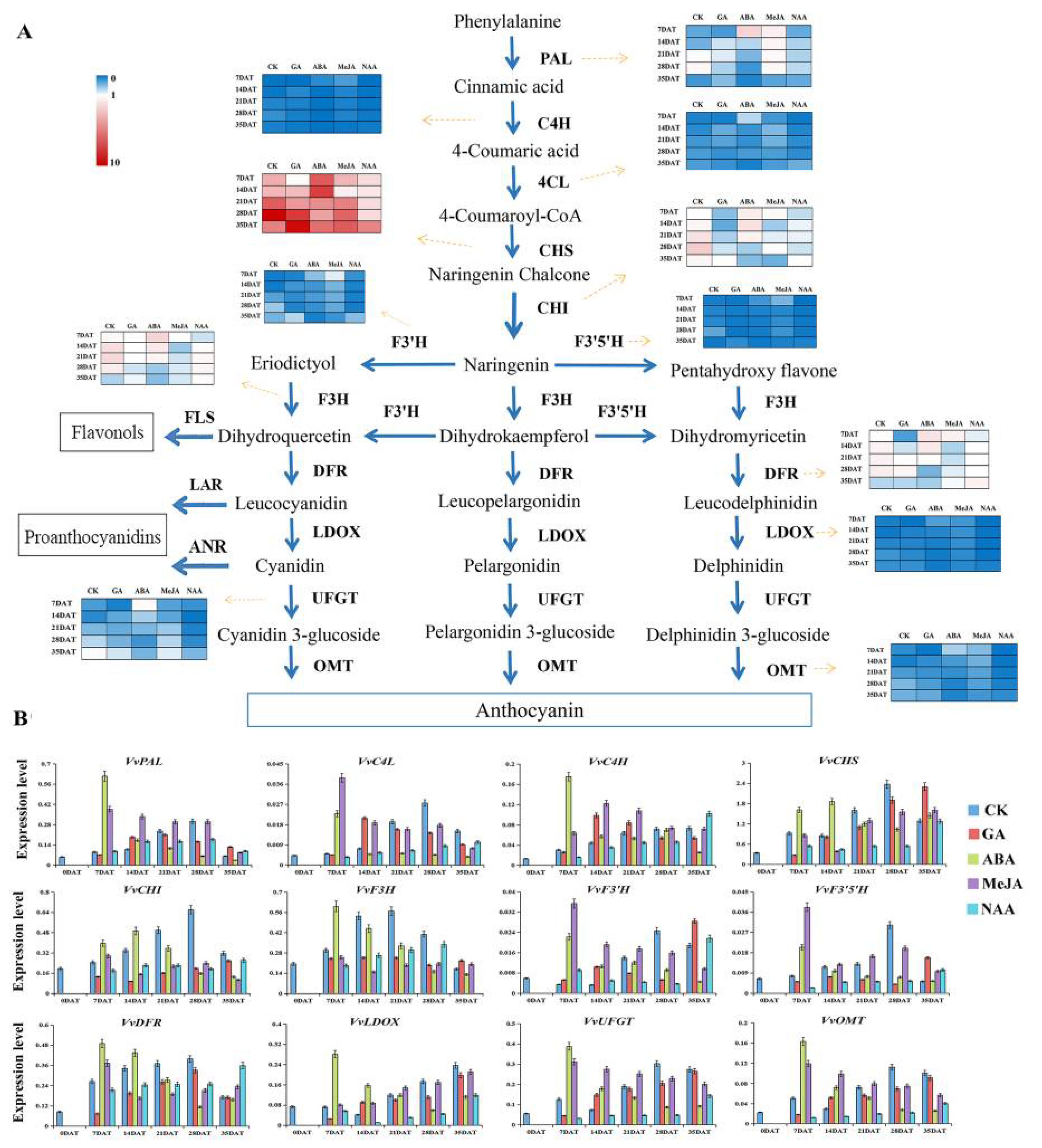
3.4. Expression Modes of Genes Related to Anthocyanin Synthesis by Different Hormone Treatments
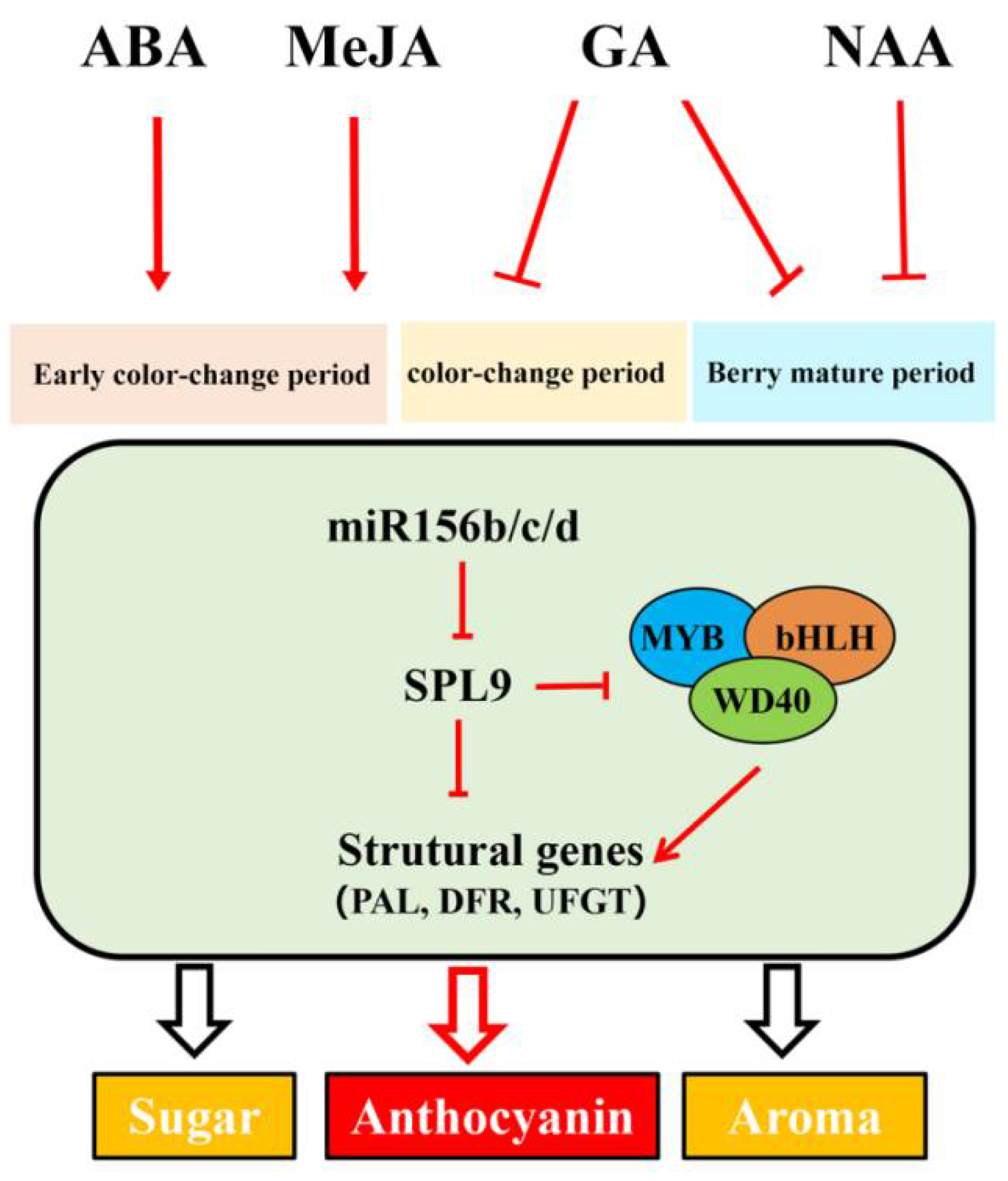
3.5. Interaction Mode Variation of VvmiR156b/c/d-VvSPL9 Modules in Response to Hormone Signal in the Regulation of the Grape Color Formation
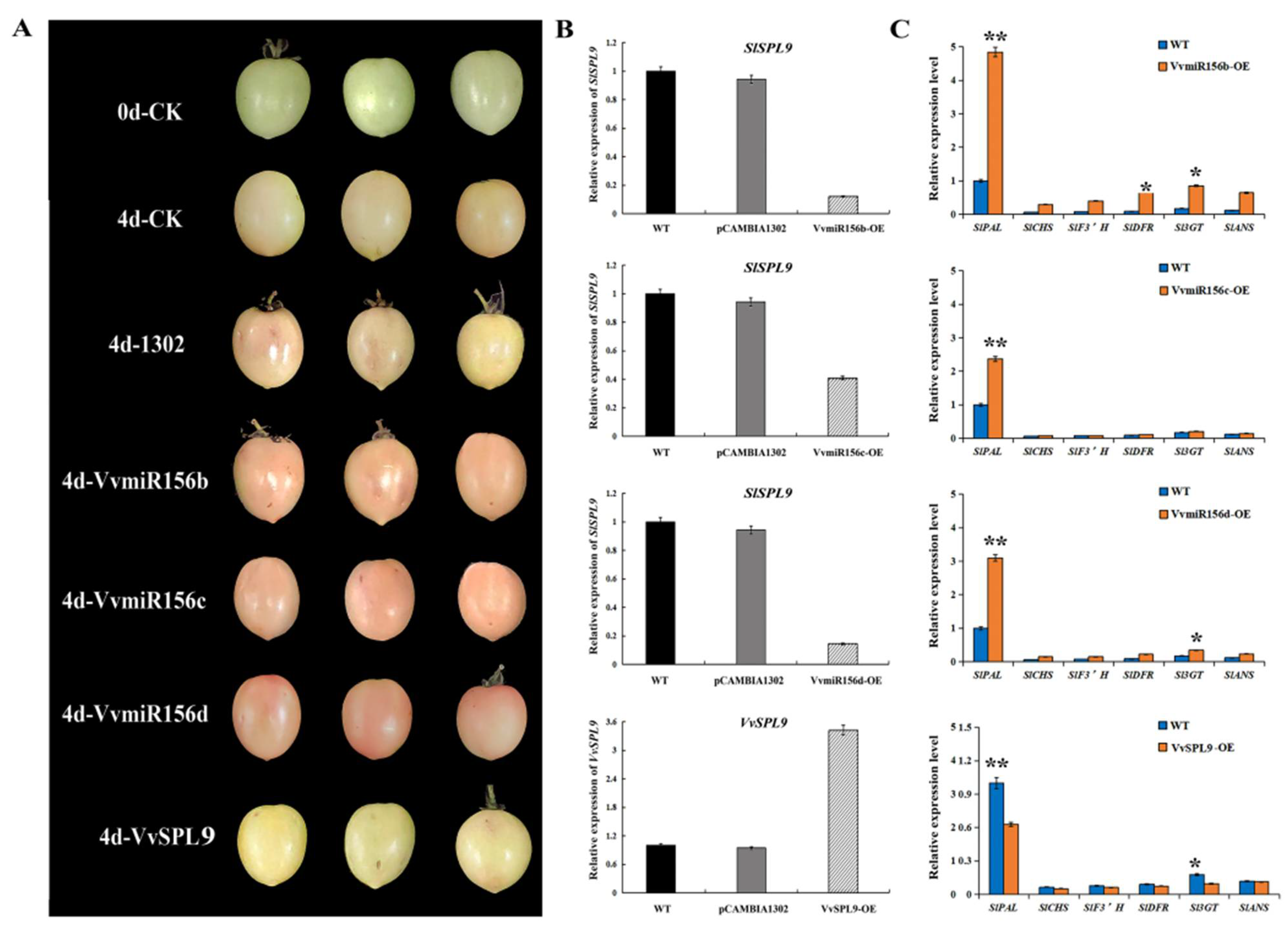
3.6. Functional Verification of Overexpression of VvMIR156b/c/d and VvSPL9 in Tomatoes
4. Discussion
4.1. Regulatory Genes for Anthocyanin Biosynthesis
4.2. Regulatory Mechanism of Hormones on Anthocyanin Synthesis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, G.; Zhao, J.; Qin, B.; Yin, Y.; An, W.; Mu, Z.X.; Cao, Y.L. ABA mediates development-dependent anthocyanin biosynthesis and fruit coloration in Lycium plants. BMC Plant Biol. 2019, 19. [Google Scholar] [CrossRef] [PubMed]
- Ma, G.; Zhang, L.; Kudaka, R.; Inaba, H.; Kato, M.; Furuya, T.; Kitamura, M.; Kitaya, Y.; Yamamoto, R.; Yahata, M.; et al. Exogenous Application of ABA and NAA Alleviates the Delayed Coloring Caused by Puffing Inhibitor in Citrus Fruit. Cells 2021, 10, 308. [Google Scholar] [CrossRef] [PubMed]
- Crupi, P.; Alba, V.; Masi, G.; Caputo, A.R.; Tarricone, L. Effect of two exogenous plant growth regulators on the color and quality parameters of seedless table grape berries. Food Res. Int. 2019, 126, 108667. [Google Scholar] [CrossRef] [PubMed]
- Dong, T.; Zheng, T.; Fu, W.; Guan, L.; Fang, J.; Jia, H. The Effect of Ethylene on the Color Change and Resistance to Botrytis cinerea Infection in ‘Kyoho’ Grape Fruits. Foods 2020, 9, 892. [Google Scholar] [CrossRef]
- Sarker, R.; Bandyopadhyay, S.; Maulik, U. An Overview of Computational Approaches for Prediction of miRNA Genes and their Targets. Curr. Bioinform. 2011, 6, 129–143. [Google Scholar] [CrossRef]
- Runmao, L.; Liye, H.; Jiayu, H.; Peigang, Q.; Yanran, W.; Qiming, D.; Xiaoting, Y.; Shuangcheng, L.; Shiquan, W.; Wenming, W. Comprehensive analysis of microRNA-Seq and target mRNAs of rice sheath blight pathogen provides new insights into pathogenic regulatory mechanisms. DNA Res. 2016, 23, 415–425. [Google Scholar]
- Yang, F.; Cai, J.; Yang, Y.; Liu, Z. Overexpression of microRNA828 reduces anthocyanin accumulation in Arabidopsis. Plant Cell Tissue Organ Cult. 2013, 115, 159–167. [Google Scholar] [CrossRef]
- Borevitz, J.O.; Xia, Y.J.; Blount, J.; Dixon, R.A.; Lamb, C. Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 2000, 12, 2383–2393. [Google Scholar] [CrossRef]
- Rajagopalan, R.; Vaucheret, H.; Trejo, J.; Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 2006, 20. [Google Scholar] [CrossRef]
- Hsieh, L.C.; Lin, S.I.; Shih, A.C.; Chen, J.W.; Lin, W.Y.; Tseng, C.Y.; Li, W.H.; Chiou, T.J. Uncovering small RNA-mediated responses to phosphate deficiency in Arabidopsis by deep sequencing. Plant Physiol. 2009, 151, 2120–2132. [Google Scholar] [CrossRef]
- Tirumalai, V.; Swetha, C.; Nair, A.; Pandit, A.; Shivaprasad, P.V. miR828 and miR858 regulate VvMYB114 to promote anthocyanin and flavonol accumulation in grapes. J. Exp. Bot. 2019, 70, 4775–4792. [Google Scholar] [CrossRef] [PubMed]
- Xia, R.; Zhu, H.; An, Y.Q.; Beers, E.P.; Liu, Z. Apple miRNAs and tasiRNAs with novel regulatory networks. Genome Biol. 2012, 13, R47. [Google Scholar] [CrossRef] [PubMed]
- Vaucheret, H.; Vazquez, F.; Crété, P.; Bartel, D.P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev. 2004, 18, 1187–1197. [Google Scholar] [CrossRef] [PubMed]
- Chiou, T.J. The role of microRNAs in sensing nutrient stress. Plant Cell Environ. 2010, 30, 323–332. [Google Scholar] [CrossRef]
- Pant, B.D.; Musialak-Lange, M.; Nuc, P.; May, P.; Buhtz, A.; Kehr, J.; Walther, D.; Scheible, W.R. Identification of nutrient-responsive Arabidopsis and rapeseed microRNAs by comprehensive real-time polymerase chain reaction profiling and small RNA sequencing. Plant Physiol. 2009, 150, 1541–1555. [Google Scholar] [CrossRef]
- Frank, M.; Harald, K.; Pawel, B.; Bernd, W. The Arabidopsis Transcription Factor MYB12 Is a Flavonol-Specific Regulator of Phenylpropanoid Biosynthesis. Plant Physiol. 2005, 138, 1083–1096. [Google Scholar]
- Jaillon, O.; Aury, J.M.; Noel, B.; Policriti, A.; Clepet, C.; Casagrande, A.; Choisne, N.; Aubourg, S.; Vitulo, N.; Jubin, C.; et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 2007, 449, 463–467. [Google Scholar]
- Velasco, R.; Zharkikh, A.; Troggio, M.; Cartwright, D.A.; Cestaro, A.; Pruss, D.; Pindo, M.; Fitzgerald, L.M.; Vezzulli, S.; Reid, J.; et al. A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2007, 2, e1326. [Google Scholar] [CrossRef]
- Miao, C.; Wang, Z.; Zhang, L.; Yao, J.; Zhu, J.K.; Hua, K.; Liu, X.; Shi, H. The grain yield modulator miR156 regulates seed dormancy through the gibberellin pathway in rice. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef]
- Xu, M.; Leichty, A.; Hu, T.; Poethig, R. H2A.Z promotes the transcription of MIR156A and MIR156C in Arabidopsis by facilitating the deposition of H3K4me3. Development 2018, 145, dev152868. [Google Scholar] [CrossRef]
- Long, J.M.; Liu, C.Y.; Feng, M.Q.; Yun, L.; Wu, X.M.; Guo, W.W. miR156-SPL modules regulate induction of somatic embryogenesis in citrus callus. J. Exp. Bot. 2018, 12, 2979–2993. [Google Scholar] [CrossRef]
- Gou, J.Y.; Felippes, F.F.; Liu, C.J.; Weigel, D.; Wang, J.W. Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell 2011, 23, 1512–1522. [Google Scholar] [CrossRef]
- Cui, L.G.; Shan, J.X.; Shi, M.; Gao, J.P.; Lin, H.X. The miR156-SPL9-DFR pathway coordinates the relationship between development and abiotic stress tolerance in plants. Plant J. Cell Mol. Biol. 2014, 80, 1108–1117. [Google Scholar] [CrossRef]
- Liu, R.; Lai, B.; Hu, B.; Qin, Y.; Hu, G.; Zhao, J. Identification of MicroRNAs and Their Target Genes Related to the Accumulation of Anthocyanins in Litchi chinensis by High-Throughput Sequencing and Degradome Analysis. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Fang, J.; Wang, C.; Yin, Y.; Sun, X.; Leng, X.; Song, C. Grapevine microRNAs responsive to exogenous gibberellin. BMC Genom. 2014, 15, 111. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Wang, J.; Wang, C.; Shen, W.; Jia, H.; Zhu, X.; Li, X. Study on Expression Modes and Cleavage Role of miR156b/c/d and its Target Gene Vv-SPL9 During the Whole Growth Stage of Grapevine. J. Hered. 2016, 107, 626–634. [Google Scholar] [CrossRef] [PubMed]
- Cui, M.; Chen, W.; Zhang, W.; Tariq, P.; Salman, H.M.; Wei, T.; Fang, J. Characterization of Vv-miR156: Vv-SPL pairs involved in the modulation of grape berry development and ripening. Mol. Genet. Genom. 2018, 293, 1333–1354. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Zhang, Z.; Zhang, S.; Fu, W.; Su, L.; Fang, J.; Jia, H. Effect of methylation level on grape fruit development process. J. Agric. Food Chem. 2020, 68, 2099–2115. [Google Scholar] [CrossRef]
- Tian, Y.; Zhao, Y.; Huang, J.; Zeng, H.; Zheng, B. Effects of different drying methods on the product quality and volatile compounds of whole shiitake mushrooms. Food Chem. 2016, 2016, 714–722. [Google Scholar] [CrossRef]
- Zhang, W.; Abdelrahman, M.; Jiu, S.; Guan, L.; Han, J.; Zheng, T.; Jia, H.; Song, C.; Fang, J.; Wang, C. VvmiR160s/VvARFs interaction and their spatio-temporal expression/cleavage products during GA-induced grape parthenocarpy. BMC Plant Biol. 2019, 19. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Jiu, S.; Zhang, C.; Wang, C.; Tariq, P.; Liu, Z.; Wang, B.; Cui, L.; Fang, J. Abscisic acid and sucrose regulate tomato and strawberry fruit ripening through the abscisic acid-stress-ripening transcription factor. Plant Biotechnol. J. 2016, 14, 2045–2065. [Google Scholar] [CrossRef]
- Fung, R.W.M.; Gonzalo, M.; Fekete, C.; Kovacs, L.G.; He, Y.; Marsh, E.; Mcintyre, L.M.; Qiu, S.W.; Schachtman, D.P. Powdery Mildew Induces Defense-Oriented Reprogramming of the Transcriptome in a Susceptible but Not in a Resistant Grapevine. Plant Physiol. 2008, 146, 236–249. [Google Scholar] [CrossRef]
- Koes, R.; Verweij, W.; Quattrocchio, F. Flavonoids: A colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci. 2005, 10, 236–242. [Google Scholar] [CrossRef]
- Saito, K.; Yonekura-Sakakibara, K.; Nakabayashi, R.; Higashi, Y.; Yamazaki, M.; Tohge, T.; Fernie, A.R. The flavonoid biosynthetic pathway in Arabidopsis: Structural and genetic diversity. Plant Physiol. Biochem. 2013, 72, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Wang, M.; Li, X.; Jiu, S.; Wang, C.; Fang, J. Genome-Wide Analysis of the Sucrose Synthase Gene Family in Grape (Vitis vinifera): Structure, Evolution, and Expression Profiles. Genes 2017, 8, 111. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhao, X.; Li, J.; Cai, H.; Li, L. MicroRNA408 Is Critical for the HY5-SPL7 Gene Network That Mediates the Coordinated Response to Light and Copper. Plant Cell 2014, 26, 4933. [Google Scholar] [CrossRef] [PubMed]
- Santino, A.; Taurino, M.; De Domenico, S.; Bonsegna, S.; Poltronieri, P.; Pastor, V.; Flors, V. Jasmonate signaling in plant development and defense response to multiple (a)biotic stresses. Plant Cell Rep. 2013, 32, 1085–1098. [Google Scholar] [CrossRef]
- Das, P.K.; Shin, D.H.; Choi, S.B.; Park, Y.I. Sugar-hormone cross-talk in anthocyanin biosynthesis. Mol. Cells 2012, 34, 501–507. [Google Scholar] [CrossRef]
- Leng, P.; Yuan, B.; Guo, Y. The role of abscisic acid in fruit ripening and responses to abiotic stress. J. Exp. Bot. 2014, 65, 4577–4588. [Google Scholar] [CrossRef]
- Flores, G.; Blanch, G.P.; Ruiz del Castillo, M.L. Postharvest treatment with (−) and (+)-methyl jasmonate stimulates anthocyanin accumulation in grapes. LWT Food Sci. Technol. 2015, 62, 807–812. [Google Scholar] [CrossRef]
- Shan, X.; Zhang, Y.; Peng, W.; Wang, Z.; Xie, D. Molecular mechanism for jasmonate-induction of anthocyanin accumulation in Arabidopsis. J. Exp. Bot. 2009, 60, 3849–3860. [Google Scholar] [CrossRef]
- Xie, Z.; Su, Z.; Wang, W.; Guan, L.; Wang, C.; Bai, Y.; Zhu, X.; Wang, X.; Jia, H.; Fang, J. Characterization of VvSPL18 and Its Expression in Response to Exogenous Hormones during Grape Berry Development and Ripening. Cytogenet. Genome Res. 2019, 159. [Google Scholar] [CrossRef] [PubMed]
- Jeong, S.T.; Goto-Yamamoto, N.; Kobayashi, S.; Esaka, M. Effects of plant hormones and shading on the accumulation of anthocyanins and the expression of anthocyanin biosynthetic genes in grape berry skins. Plant Sci. 2004, 167, 247–252. [Google Scholar] [CrossRef]
- Chen, J.; Mao, L.; Mi, H.; Lu, W.; Ying, T.; Luo, Z. Involvement of abscisic acid in postharvest water-deficit stress associated with the accumulation of anthocyanins in strawberry fruit. Postharvest Biol. Technol. 2016, 111, 99–105. [Google Scholar] [CrossRef]
- Shen, X.; Zhao, K.; Liu, L.; Zhang, K.; Li, T.; Yuan, H.; Liao, X.; Wang, Q.; Guo, X.; Li, F. A Role for PacMYBA in ABA-Regulated Anthocyanin Biosynthesis in Red-Colored Sweet Cherry cv. Hong Deng (Prunus avium L.). Plant Cell Physiol. 2014, 55. [Google Scholar] [CrossRef] [PubMed]
- Gagné, S.; Cluzet, S.; Mérillon, J.-M.; Gény, L. ABA Initiates Anthocyanin Production in Grape Cell Cultures. J. Plant Growth Regul. 2011, 30, 1–10. [Google Scholar] [CrossRef]
- El-Kereamy, A.; Chervin, C.; Roustan, J.P.; Cheynier, V.; Souquet, J.M.; Moutounet, M.; Raynal, J.; Ford, C.; Latché, A.; Pech, J.C.; et al. Exogenous ethylene stimulates the long-term expression of genes related to anthocyanin biosynthesis in grape berries. Physiol. Plant. 2003, 119, 175–182. [Google Scholar] [CrossRef]
- Loreti, E.; Povero, G.; Novi, G.; Solfanelli, C.; Alpi, A.; Perata, P. Gibberellins, jasmonate and abscisic acid modulate the sucrose-induced expression of anthocyanin biosynthetic genes in Arabidopsis. New Phytol. 2008, 179, 1004–1016. [Google Scholar] [CrossRef]
- Qi, T.; Song, S.; Ren, Q.; Wu, D.; Huang, H.; Chen, Y.; Fan, M.; Peng, W.; Ren, C.; Xie, D. The jasmonate-ZIM-domain proteins interact with the WD-repeat/bHLH/MYB complexes to regulate jasmonate-mediated anthocyanin accumulation and trichome initiation in Arabidopsis thaliana. Plant Cell 2011, 23, 1795–1814. [Google Scholar] [CrossRef]
- Li, Z.; Jun, W. Effects of NAA and ABA treatments on the expression of anthocyanin biosynthetic genes in ‘Jing You’ grape berry skin. J. China Agric. Univ. 2011, 16, 30–37. [Google Scholar]
- Wu, Y.F.; Reed, G.W.; Tian, C.Q. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Wang, L.J.; Mao, Y.B.; Cai, W.J.; Xue, H.W.; Chen, X.Y. Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 2005, 17, 2204–2216. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Galvão, V.C.; Zhang, Y.C.; Horrer, D.; Zhang, T.Q.; Hao, Y.H.; Feng, Y.Q.; Wang, S.; Schmid, M.; Wang, J.W. Gibberellin regulates the Arabidopsis floral transition through miR156-targeted SQUAMOSA promoter binding-like transcription factors. Plant Cell 2012, 24, 3320–3332. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Herr, A.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar] [CrossRef]
- Reyes, J.L.; Chua, N.H. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J. 2007, 49, 592–606. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Zhang, Y.C.; Wang, C.Y.; Luo, Y.C.; Huang, Q.J.; Chen, S.Y.; Zhou, H.; Qu, L.H.; Chen, Y.Q. Expression analysis of phytohormone-regulated microRNAs in rice, implying their regulation roles in plant hormone signaling. FEBS Lett. 2009, 583, 723–728. [Google Scholar] [CrossRef] [PubMed]
| Physiological Indexes | Different Treatment Periods | |||||
|---|---|---|---|---|---|---|
| Treatment | 7DAT | 14DAT | 21DAT | 28DAT | 35DAT | |
| transverse diameter (mm) | CK | 20.53 ± 0.59 b | 21.35 ± 0.61 bc | 20.57 ± 0.58 b | 21.75 ± 0.73 b | 21.54 ± 0.79 b |
| GA | 19.87 ± 0.63 c | 21.19 ± 0.74 c | 20.34 ± 0.36 b | 20.98 ± 0.63 c | 21.31 ± 0.82 b | |
| ABA | 21.01 ± 0.34 a | 21.96 ± 0.58 a | 21.54 ± 0.41 a | 22.12 ± 0.25 b | 22.27 ± 0.29 a | |
| MeJA | 20.94 ± 0.64 a | 21.73 ± 0.62 ab | 21.23 ± 0.76 a | 22.67 ± 0.59 a | 22.16 ± 0.31 a | |
| NAA | 19.28 ± 0.58 d | 19.43 ± 0.54 d | 19.31 ± 0.49 c | 19.84 ± 0.62 d | 20.06 ± 0.76 c | |
| longitudinal diameter (mm) | CK | 28.12 ± 0.62 b | 28.91 ± 0.77 b | 29.14 ± 0.59 ab | 29.11 ± 0.99 ab | 29.42 ± 0.89 bc |
| GA | 27.36 ± 0.86 c | 28.20 ± 0.94 c | 28.85 ± 0.63 b | 28.82 ± 0.83 b | 29.17 ± 0.68 c | |
| ABA | 28.58 ± 0.74 a | 29.31 ± 0.96 a | 29.45 ± 0.74 a | 29.50 ± 0.39 a | 29.90 ± 0.43 a | |
| MeJA | 28.49 ± 0.57 ab | 29.15 ± 0.62 ab | 29.32 ± 0.92 a | 29.31 ± 0.65 a | 29.61 ± 1.03 ab | |
| NAA | 26.82 ± 0.94 d | 27.31 ± 0.54 d | 27.89 ± 0.91 c | 28.33 ± 0.78 c | 28.62 ± 0.73 d | |
| berry weigh (g) | CK | 6.13 ± 0.16 a | 6.81 ± 0.14 bc | 8.06 ± 0.09 ab | 8.31 ± 0.15 a | 8.53 ± 0.14 b |
| GA | 6.05 ± 0.16 a | 6.56 ± 0.22 c | 7.78 ± 0.35 b | 8.23 ± 0.11 ab | 8.38 ± 0.19 b | |
| ABA | 6.23 ± 0.19 a | 7.72 ± 0.21 a | 8.39 ± 0.38 a | 8.50 ± 0.22 a | 8.92 ± 0.29 a | |
| MeJA | 6.19 ± 0.12 a | 6.97 ± 0.23 b | 8.18 ± 0.23 a | 8.41 ± 0.27 a | 8.61 ± 0.26 ab | |
| NAA | 6.10 ± 0.18 a | 6.57 ± 0.15 c | 7.32 ± 0.17 c | 7.84 ± 0.14 b | 7.97 ± 0.13 c | |
| soluble solid (%) | CK | 12.57 ± 0.22 c | 15.61 ± 0.21 b | 17.05 ± 0.39 c | 18.12 ± 0.47 c | 21.21 ± 0.73 c |
| GA | 12.25 ± 0.37 c | 12.57 ± 0.22 c | 15.41 ± 0.21 d | 17.58 ± 0.39 d | 19.09 ± 0.61 d | |
| ABA | 14.63 ± 0.41 a | 16.46 ± 0.52 a | 18.81 ± 0.52 a | 20.13 ± 0.52 a | 22.71 ± 0.67 a | |
| MeJA | 13.84 ± 0.52 b | 15.89 ± 0.34 b | 17.93 ± 0.62 b | 19.44 ± 0.69 b | 21.82 ± 0.59 b | |
| NAA | 11.39 ± 0.23 d | 12.46 ± 0.31 c | 14.82 ± 0.39 e | 16.37 ± 0.43 e | 17.54 ± 0.41 e | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, Z.; Wang, X.; Xuan, X.; Sheng, Z.; Jia, H.; Emal, N.; Liu, Z.; Zheng, T.; Wang, C.; Fang, J. Characterization and Action Mechanism Analysis of VvmiR156b/c/d-VvSPL9 Module Responding to Multiple-Hormone Signals in the Modulation of Grape Berry Color Formation. Foods 2021, 10, 896. https://doi.org/10.3390/foods10040896
Su Z, Wang X, Xuan X, Sheng Z, Jia H, Emal N, Liu Z, Zheng T, Wang C, Fang J. Characterization and Action Mechanism Analysis of VvmiR156b/c/d-VvSPL9 Module Responding to Multiple-Hormone Signals in the Modulation of Grape Berry Color Formation. Foods. 2021; 10(4):896. https://doi.org/10.3390/foods10040896
Chicago/Turabian StyleSu, Ziwen, Xicheng Wang, Xuxian Xuan, Zilu Sheng, Haoran Jia, Naseri Emal, Zhongjie Liu, Ting Zheng, Chen Wang, and Jinggui Fang. 2021. "Characterization and Action Mechanism Analysis of VvmiR156b/c/d-VvSPL9 Module Responding to Multiple-Hormone Signals in the Modulation of Grape Berry Color Formation" Foods 10, no. 4: 896. https://doi.org/10.3390/foods10040896
APA StyleSu, Z., Wang, X., Xuan, X., Sheng, Z., Jia, H., Emal, N., Liu, Z., Zheng, T., Wang, C., & Fang, J. (2021). Characterization and Action Mechanism Analysis of VvmiR156b/c/d-VvSPL9 Module Responding to Multiple-Hormone Signals in the Modulation of Grape Berry Color Formation. Foods, 10(4), 896. https://doi.org/10.3390/foods10040896






