1. Introduction
Adulteration and mislabeling of food has been known since biblical times and probably goes back as far as when food started to be traded [
1]. Unsurprisingly, laws and regulations to combat these practices can also be found throughout history: ancient food regulations are referred to in Egyptian, Chinese, Hindu, Greek, and Roman texts. Today, the EU General Food Law—next to protecting public health—aims to ‘prevent fraudulent or deceptive practices; the adulteration of food; and any other practices which may mislead the consumer’ (Article 8 of Regulation (EC) No 178/2002 ). A commonly accepted definition is that ‘Food fraud includes adulteration, deliberate and intentional substitution, dilution, simulation, tampering, counterfeiting, or misrepresentation of food, food ingredients, or food packaging; or false or misleading statements made about a product for economic gain’ [
2]. Although food fraud is not explicitly defined in EU legislation, the Food Information to Consumers Regulation (Regulation (EU) No 1169/2011) concretises the relevant aspects of the General Food Law in Article 7 (Fair information practices) by stipulating that food information shall not be misleading, particularly as to the characteristics of the food and, in particular, as to its nature, identity, properties, composition, quantity, durability, country of origin or place of provenance, and method of manufacture or production.
Herbs and spices are among the group of ingredients most vulnerable to fraudulent manipulation, mostly due to their high economic value, which makes them an attractive target. Furthermore, international trade and highly complex supply chains, involving various actors and processing steps, contribute to the presence of adulterated herbs and spices on the market [
3,
4]. Macro- and microscopic examination of morphological features, physico-chemical methods (e.g., ash, volatile oil content), and determination of aromatic principles typical for the product (e.g., members of the terpenoid or vanillylamide families) are traditionally used for quality grading and authentication of herbs and spices.
More advanced analytical methods, often in combination with machine learning algorithms, use proteins, metabolites, or DNA, either in a targeted or untargeted manner, to authenticate plant material [
5]. Particularly, DNA based methods have equipped forensic analysts with highly specific, sensitive, and cost-effective tools because the genetic makeup of their targets is not influenced by environmental or physiological factors. The application of DNA technology for food authentication [
6,
7,
8] and specifically for plant material [
9,
10] has been extensively reviewed. They make use of DNA polymorphism between species and most of them include a polymerase chain reaction (PCR) to amplify DNA. Species-specific PCRs targeting a product specific nucleotide sequence are very attractive because of their sensitivity, specificity, reproducibility, and ability to detect low target amounts. They are particularly useful to verify the claimed identity of the product or to detect the presence of another non-declared species. This approach is less appropriate if the identity of the adulterant(s) is not known, although multiplexed assays can provide a solution as long as the number of potential adulterants is small. Non-targeted methods, on the other hand, are able to find a wider range of biological contaminants. In most such methods a ‘fingerprint’ is generated by targeting variable length sequences which is then used to confirm purity compared to a database to find contaminants [
11,
12].
DNA barcoding has gained popularity for identifying animal as well as plant species [
13,
14]. Again, the technique works well if applied to single species; if the sample contains more than one species, the barcoding regions from all of them can be amplified resulting in difficult-to-interpret sequences in the Sanger sequencing step. Meta-barcoding using Next Generation Sequencing (NGS), a high-throughput, parallel, sequencing technology, offers a solution to this problem. Although the technique is becoming more widely available, it requires access to dedicated instrumentation, bioinformatics pipelines, and experienced operators.
We have developed a method called ‘DNA accounting’ for screening the purity of single-species food ingredients using droplet digital PCR (ddPCR). The screen should be able to flag suspicious samples for further analysis by meta-barcoding, thereby reducing the number of samples and, eventually, the efforts and costs inherent to a meta-barcoding workflow. Similar approaches have been used to estimate the proportion of meat from different species in meat products [
15,
16,
17,
18], fish [
19], and products of plant origin [
20,
21]. However, these publications also rely on the correlation between DNA yield and sample intake, whereas the proposed approach does not. Therefore, by excluding a source of variation, the resulting assay should be more robust.
The core idea of the proposed DNA accounting is that, if a single species product is ‘pure’ and if the species is well characterized (in terms of genome size, ploidy, etc.), the number of target copies measured by quantitative PCR should be identical to the ‘expected’ number of target copies calculated from its fluorometrically measured DNA concentration, within the bounds of measurements uncertainty. As a corollary, deviations from the expected copy number are an indication that the product is not ‘pure’ and may be adulterated (e.g., bulking agents may have been added, leading to fewer targets as part of the DNA will come from the bulking agent).
The assumptions of the method are: (I) the target sequence is highly specific and is not present in any adulterant (or at least not in the same amount of copies), (II) all bulking agents used yield DNA with approximately the same efficiency as the product under investigation. Droplet digital PCR was chosen as a method to ‘count’ the number of target sequences since it allows direct absolute quantification unlike qPCR which relies on standard curves of a reference material to obtain absolute measurements. Moreover, it is less affected by the presence of co-extracted inhibitors due to the dilution effect.
We demonstrated the applicability of the ‘DNA accounting’ approach by analyzing 141 commercial saffron samples taken from the markets of 20 EU Member States. Saffron is derived from the stigma and styles of the saffron crocus (
Crocus sativus). Being one of the most expensive food ingredients, it is a formidable target for economically motivated adulteration [
22]. The proposed screening approach could help to systematically survey the authenticity of the product placed on the market. It is not suitable for grading its quality as other parts of
Crocus sativus such as petals and stamens, which may be used as bulking agents, cannot be recognised.
4. Discussion
DNA based methods have become a widely used tool for detecting fraud in the agri-food chain, particularly for species identification and quantification of certain food ingredient [
6,
7,
8,
9]. Species-specific PCR is the method of choice to target a particular adulterant, e.g., horse meat in beef patties, which, by multiplexing, can be extended to target several, known adulterants. Another popular approach, Sanger sequencing of barcoding regions, while efficient for identifying species, does not allow targeting several non-declared species (adulterants) at the same time.
Authentication of culinary herbs and spices goes beyond the question whether the named species, e.g.,
Crocus sativus, is present but tries to answer how much of the named species makes up the sample, in other words how ‘pure’ the sample is. The challenge in using quantitative PCR is the transformation of DNA amount (mass or number of copies) into mass (fraction) of the biological material. In its simplest format, binary mixtures (weight/weight) of the adulterant/named species are prepared, DNA is extracted, and after real-time PCR the cycle threshold (Cq) values are plotted against the log-transformed mixture percentage. Including a ubiquitous reference gene makes the assay more robust and compensates to a certain degree the effect of processing on DNA extractability and integrity [
44]. However, most published PCR assays aim at quantifying an adulterant using primers specific for the non-declared species, whereas for assessing ‘purity’ the named species itself has to be quantified.
By assuming that the DNA content is proportional to the mass of biomaterials, Rong Chen et al. [
45] developed a real-time PCR assay for estimating directly the mass of saffron in saffron containing herbal products; furthermore, they showed that the amount of DNA was fairly constant for several different batches of saffron.
We have taken this idea one step further and suggest a broadly applicable technique for estimating the ‘purity’ of biological matter using digital PCR. This technique is less prone to interference by amplification inhibitors and allows the estimation of the number of DNA copies present in a sample without external references. By measuring the number of genome copies in a sample and comparing it to the number of copies calculated from the known amount of DNA used in the reaction, ‘purity’ of a biomaterial can be estimated, provided the species of interest is well characterized (genome size, ploidy, target copy number). The number of target copies measured should be identical to the ‘expected’ number of target copies as calculated from its DNA concentration (measured flourometrically) within the bounds of measurements uncertainty.
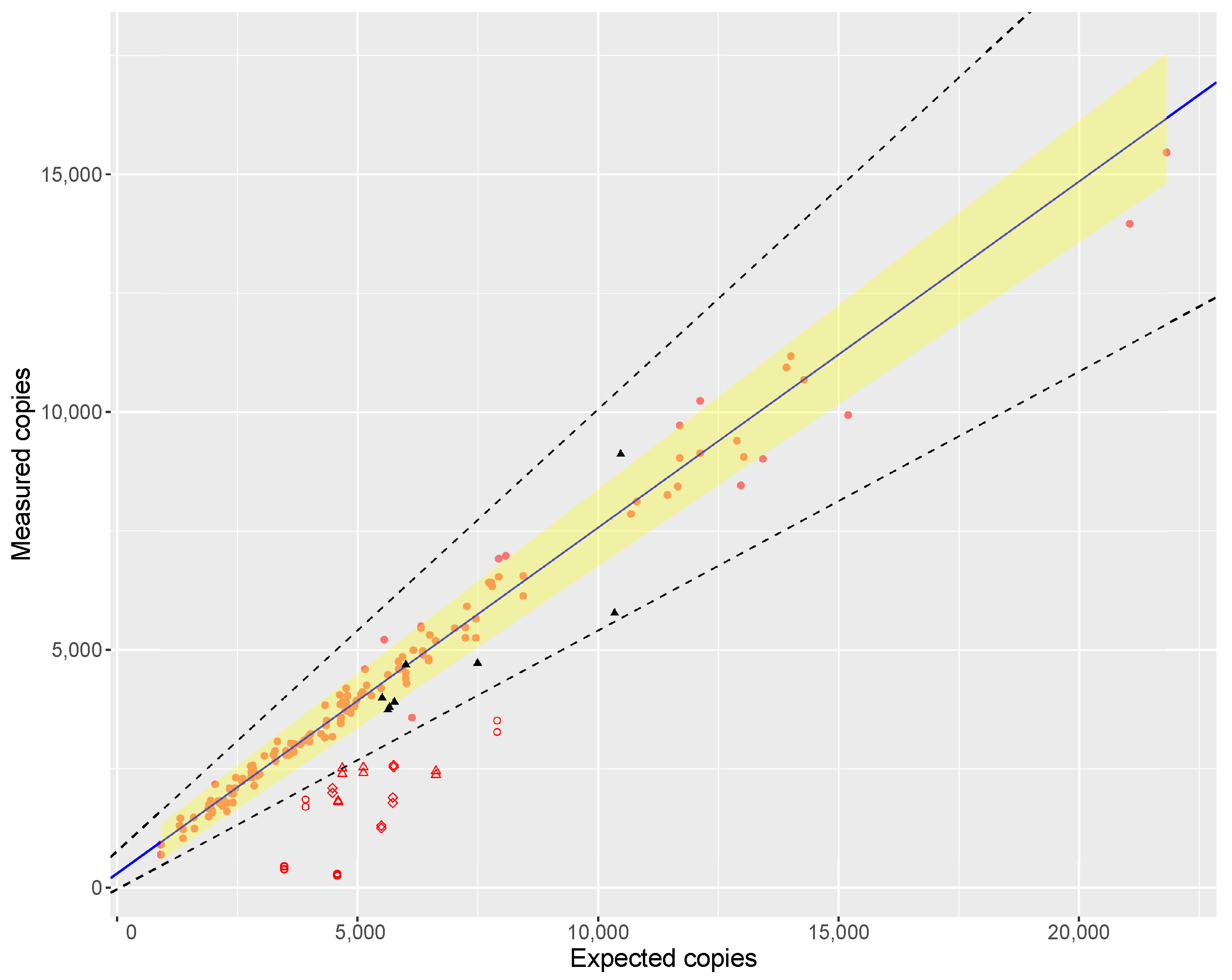
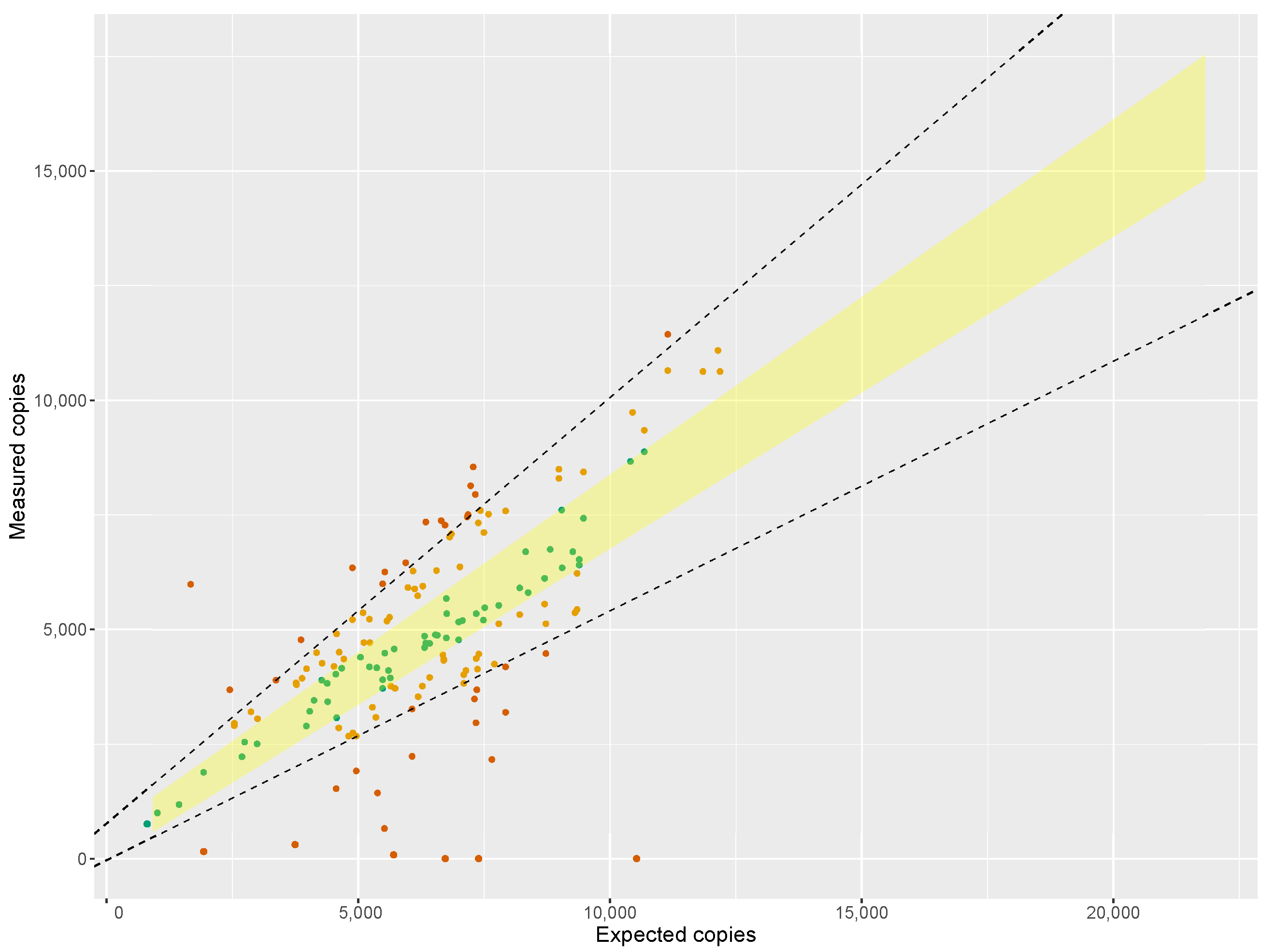
When regressing the expected against the measured copy numbers over a wide range of saffron samples, the slope of the regression line was 1.138 (
Figure 1), proving the assumption that ‘purity’ of a biomaterial can be assessed by dPCR. However, the estimated slope was not fully in line with the ideal value of 1.00. The reason for this could be inherent to the instrument used, as Low et al. [
46] also found deviations from the ideal value when comparing measured copy numbers to dilutions of a certified reference material (ERM-AD623), which were attributed to the particular brand of ddPCR instrument. Furthermore, inaccuracies in the analytical chain are inevitable: fluorometric quantification of the DNA in the extract, the 1C values used, DNA damage during extraction, PCR efficiency, etc. which may all contribute to the deviation of the slope from unity.
We have applied the technique to the analysis of 141 commercial samples of saffron taken in 20 EU Member States. It has to be stressed that the ‘purity’ estimate obtained by ddPCR is intended for screening purposes and not to draw firm conclusion whether non-declared botanicals are indeed present. For confirming this, samples flagged as suspicious were analysed using NGS metabarcoding as a means of compositional analysis. If NGS indicated species known to be potential adulterants, their presence was confirmed by qPCR of a species-specific target.
ddPCR screening indicated more samples as suspicious than could be confirmed by NGS. This was partially expected as the DNA used for the construction of the reference curve for DNA accounting was obtained from
Crocus sativus bulbs, and although precautions have been taken to minimize difference in DNA-quality between reference and samples (i.e., the bulbs were cut, dried, and milled), it stands to reason that the variability of DNA quality, and therefore of testing results, is higher amongst samples. Differences in processing, drying, milling, storage, microbial decontamination (e.g., gamma irradiation [
47]), etc. may adversely affect both DNA quantification and its specific target amplifiability.
DNA accounting has proven to be a valuable analytical approach in the food fraud detection workflow. The ability to rapidly screen large numbers of samples for their purity is a much needed capacity in today’s food chain with its rapid turn-over. The suggested approach has significant value as a screening tool and has also a potential for a confirmatory assay to estimate the purity of botanicals. For the latter approach, the inclusion of a suitable and ubiquitously present reference gene in the assay would be necessary to compensate for variations in DNA extractability and amplifiability.