Non-IgE-Mediated Gastrointestinal Food Protein-Induced Allergic Disorders. Clinical Perspectives and Analytical Approaches
Abstract
:1. Digestive Manifestations of Food Allergy
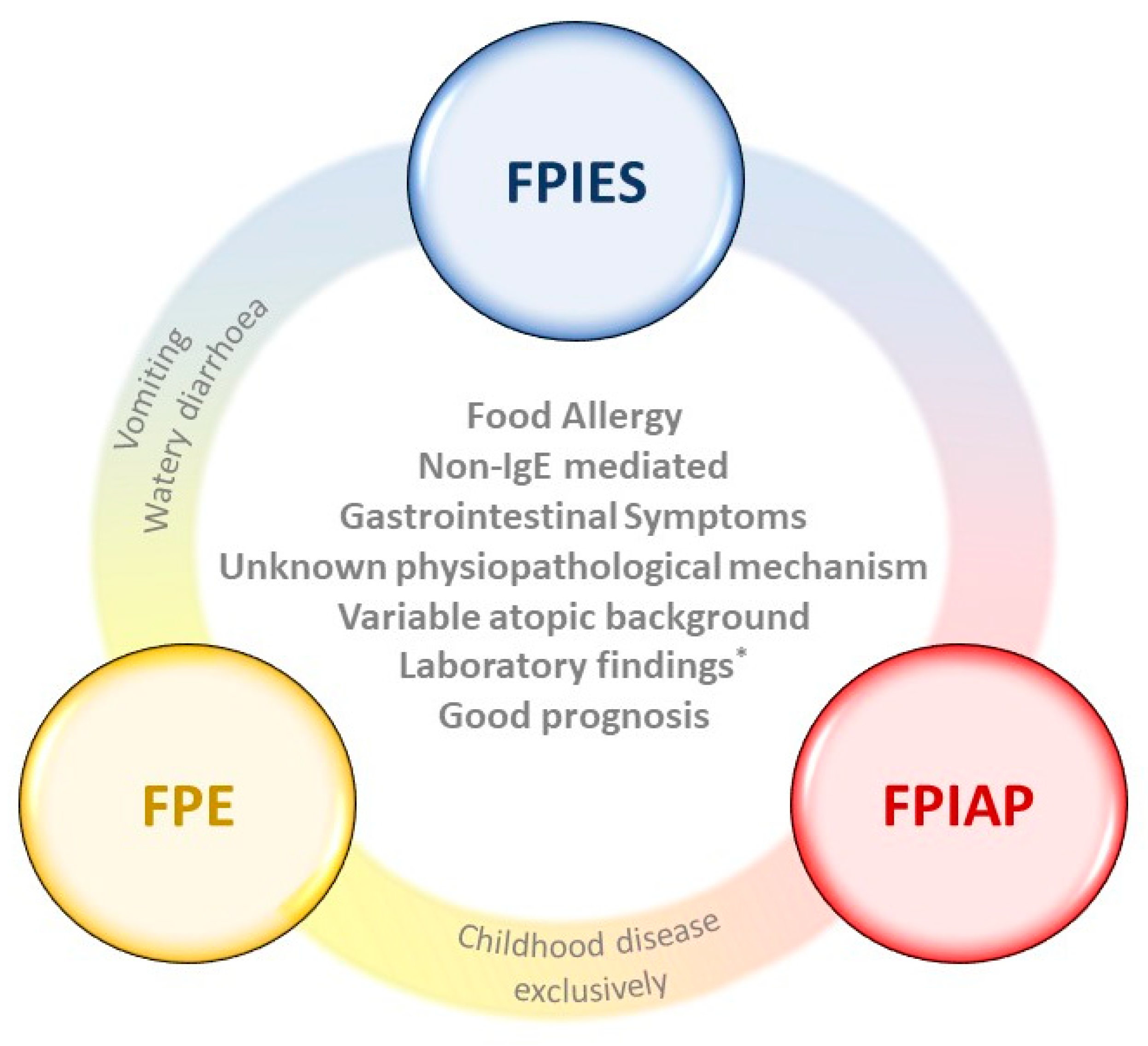
1.1. Food Protein-Induced Allergic Proctocolitis (FPIAP)
1.2. Food Protein-Induced Enteropathy (FPE)
1.3. Food Protein-Induced Enterocolitis Syndrome (FPIES)
2. Approaches for the Study of Non-IgE-GI-FA
2.1. Animal Models
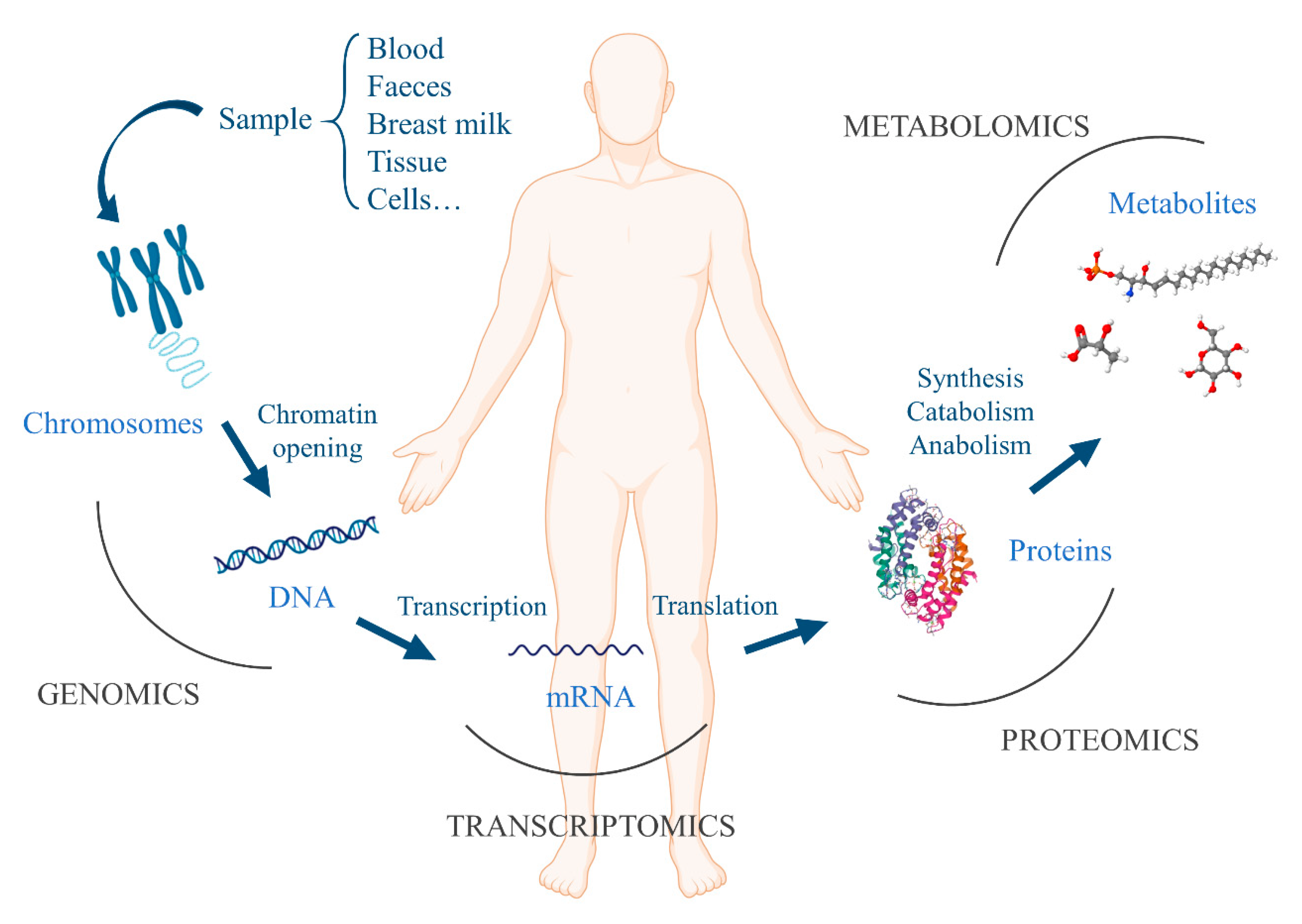
2.2. Gut Microbiota and Omics Techniques
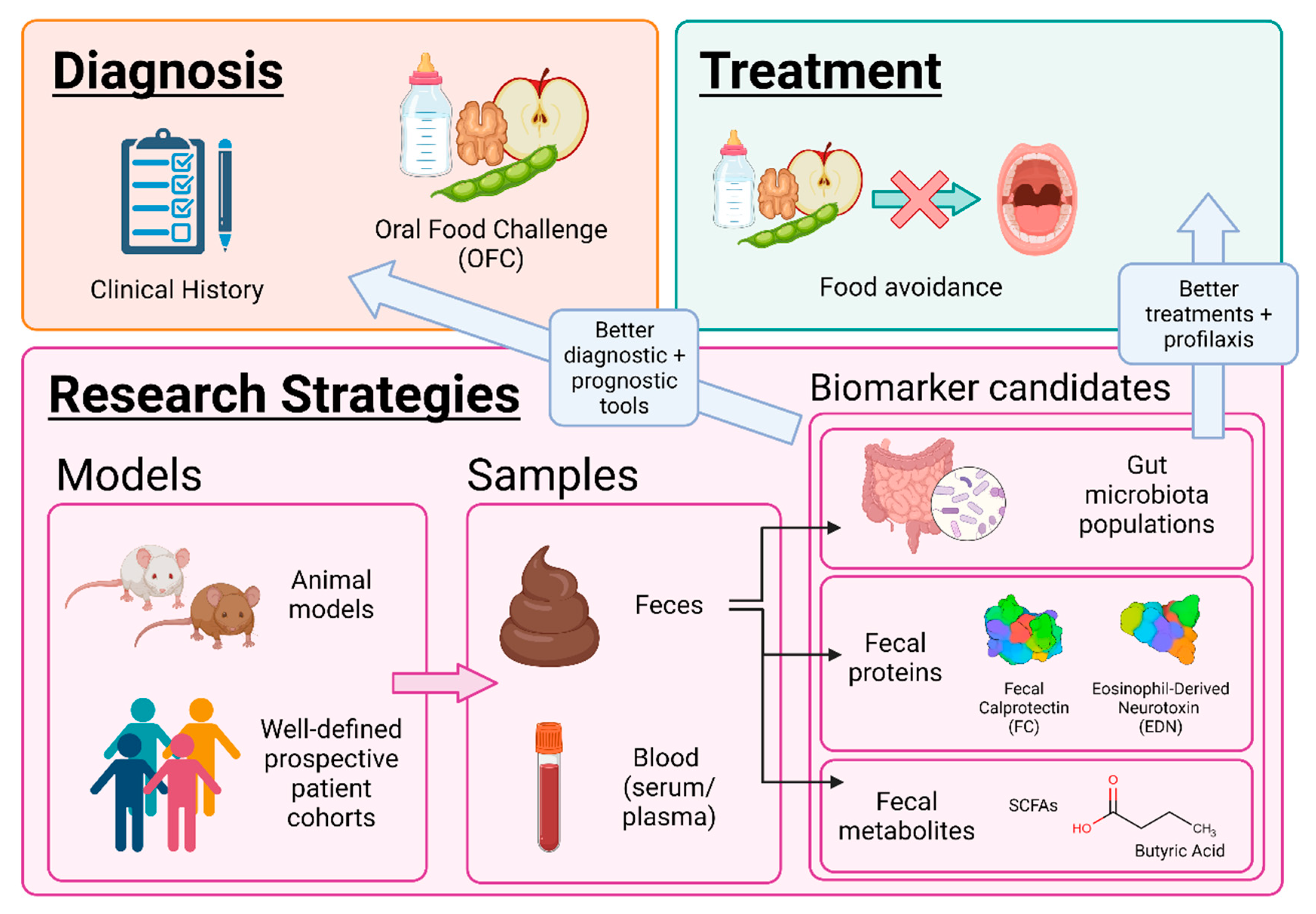
2.3. Fecal Biomarkers. Protein Analysis
3. Therapeutical Perspectives
4. Unmet Needs
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAF | amino acid-based formula |
| AE | allergic enteritis |
| BCFAs | branched chain fatty acids |
| CCR8 | CC chemokine receptor 8 |
| CM | cow’s milk |
| CMA | cow’s milk allergy |
| CMF | cow’s milk formula |
| ECP | eosinophilic cationic protein |
| EDN | eosinophil-derived neurotoxin |
| EHF | extensively hydrolyzed formula |
| EoE | eosinophilic esophagitis |
| FA | food allergy |
| FC | fecal calprotectin |
| FPE | food protein-induced enteropathy |
| FPIAP | food protein-induced allergic proctocolitis |
| FPIES | food protein-induced enterocolitis syndrome |
| GI-FA | gastrointestinal food allergy |
| HBI | healthy breastfed infants |
| HRF | hydrolyzed rice formula |
| MLNs | mesenteric lymph nodes |
| non-IgE-GI-FA | non-IgE-mediated gastrointestinal food allergy |
| OFC | oral food challenge |
| PPs | Peyer’s patches |
| SCFAs | short-chain fatty acids |
| SPBF | soy protein-based formula |
| TNF-α | tumor necrosis factor-α |
References
- Boyce, J.A.; Assa’ad, A.; Burks, A.W.; Jones, S.M.; Sampson, H.A.; Wood, R.A.; Plaut, M.; Cooper, S.F.; Fenton, M.J.; NIAID-Sponsored Expert Panel; et al. Guidelines for the Diagnosis and Management of Food Allergy in the United States: Report of the NIAID-Sponsored Expert Panel. J. Allergy Clin. Immunol. 2010, 126, S1–S58. [Google Scholar] [CrossRef] [PubMed]
- Nowak-Węgrzyn, A. Food Protein-Induced Enterocolitis Syndrome and Allergic Proctocolitis. Allergy Asthma Proc. 2015, 36, 172–184. [Google Scholar] [CrossRef] [PubMed]
- Lake, A.M. Food-Induced Eosinophilic Proctocolitis. J. Pediatr. Gastroenterol. Nutr. 2000, 30, S58–S60. [Google Scholar] [CrossRef] [PubMed]
- Lake, A.M.; Whitington, P.F.; Hamilton, S.R. Dietary Protein-Induced Colitis in Breast-Fed Infants. J. Pediatr. 1982, 101, 906–910. [Google Scholar] [CrossRef]
- Fagundes-Neto, U.; Ganc, A.J. Allergic Proctocolitis: The Clinical Evolution of a Transitory Disease with a Familial Trend. Case Reports. Einstein 2013, 11, 229–233. [Google Scholar] [CrossRef] [Green Version]
- Chung, H.L.; Hwang, J.B.; Park, J.J.; Kim, S.G. Expression of Transforming Growth Factor Beta1, Transforming Growth Factor Type I and II Receptors, and TNF-Alpha in the Mucosa of the Small Intestine in Infants with Food Protein-Induced Enterocolitis Syndrome. J. Allergy Clin. Immunol. 2002, 109, 150–154. [Google Scholar] [CrossRef]
- Sampson, H.A.; Anderson, J.A. Summary and Recommendations: Classification of Gastrointestinal Manifestations Due to Immunologic Reactions to Foods in Infants and Young Children. J. Pediatr. Gastroenterol. Nutr. 2000, 30, S87–S94. [Google Scholar] [CrossRef]
- Liacouras, C.A. Food Protein-Induced Allergic Proctocolitis of Infancy—UpToDate. Available online: https://www.uptodate.com/contents/food-protein-induced-allergic-proctocolitis-of-infancy (accessed on 8 April 2021).
- Elizur, A.; Cohen, M.; Goldberg, M.R.; Rajuan, N.; Cohen, A.; Leshno, M.; Katz, Y. Cow’s Milk Associated Rectal Bleeding: A Population Based Prospective Study. Pediatr. Allergy Immunol. 2012, 23, 766–770. [Google Scholar] [CrossRef]
- Arvola, T.; Ruuska, T.; Keränen, J.; Hyöty, H.; Salminen, S.; Isolauri, E. Rectal Bleeding in Infancy: Clinical, Allergological, and Microbiological Examination. Pediatrics 2006, 117, e760–e768. [Google Scholar] [CrossRef] [Green Version]
- Caubet, J.-C.; Szajewska, H.; Shamir, R.; Nowak-Węgrzyn, A. Non-IgE-Mediated Gastrointestinal Food Allergies in Children. Pediatr. Allergy Immunol. 2017, 28, 6–17. [Google Scholar] [CrossRef]
- Connors, L.; O’Keefe, A.; Rosenfield, L.; Kim, H. Non-IgE-Mediated Food Hypersensitivity. Allergy Asthma Clin. Immunol. 2018, 14, 56. [Google Scholar] [CrossRef]
- Odze, R.D.; Bines, J.; Leichtner, A.M.; Goldman, H.; Antonioli, D.A. Allergic Proctocolitis in Infants: A Prospective Clinicopathologic Biopsy Study. Hum. Pathol. 1993, 24, 668–674. [Google Scholar] [CrossRef]
- Hwang, J.-B.; Hong, J. Food Protein-Induced Proctocolitis: Is This Allergic Disorder a Reality or a Phantom in Neonates? Korean J. Pediatr. 2013, 56, 514–518. [Google Scholar] [CrossRef] [Green Version]
- Xanthakos, S.A.; Schwimmer, J.B.; Melin-Aldana, H.; Rothenberg, M.E.; Witte, D.P.; Cohen, M.B. Prevalence and Outcome of Allergic Colitis in Healthy Infants with Rectal Bleeding: A Prospective Cohort Study. J. Pediatr. Gastroenterol. Nutr. 2005, 41, 16–22. [Google Scholar] [CrossRef]
- Calvani, M.; Anania, C.; Cuomo, B.; D’Auria, E.; Decimo, F.; Indirli, G.C.; Marseglia, G.; Mastrorilli, V.; Sartorio, M.U.A.; Santoro, A.; et al. Non–IgE- or Mixed IgE/Non–IgE-Mediated Gastrointestinal Food Allergies in the First Years of Life: Old and New Tools for Diagnosis. Nutrients 2021, 13, 226. [Google Scholar] [CrossRef]
- Su, K.-W.; Shreffler, W.G.; Yuan, Q. Gastrointestinal Immunopathology of Food Protein–Induced Enterocolitis Syndrome and Other Non-Immunoglobulin E–Mediated Food Allergic Diseases. Ann. Allergy Asthma Immunol. 2021, 126, 516–523. [Google Scholar] [CrossRef]
- Cianferoni, A. Food Protein-Induced Enterocolitis Syndrome Epidemiology. Ann. Allergy Asthma Immunol. 2021, 126, 469–477. [Google Scholar] [CrossRef]
- Hoffmann, N.V.; Ahmed, A.; Fortunato, J.E. Food Protein-Induced Enterocolitis Syndrome (FPIES): Dynamic Relationship among Gastrointestinal Symptoms, Immune Response and the Autonomic Nervous System. Ann. Allergy Asthma Immunol. 2021, 126, 498–505. [Google Scholar] [CrossRef]
- Nowak-Węgrzyn, A.; Chehade, M.; Groetch, M.E.; Spergel, J.M.; Wood, R.A.; Allen, K.; Atkins, D.; Bahna, S.; Barad, A.V.; Berin, C.; et al. International Consensus Guidelines for the Diagnosis and Management of Food Protein–Induced Enterocolitis Syndrome: Executive Summary—Workgroup Report of the Adverse Reactions to Foods Committee, American Academy of Allergy, Asthma & Immunology. J. Allergy Clin. Immunol. 2017, 139, 1111–1126.e4. [Google Scholar] [CrossRef] [Green Version]
- Caubet, J.C.; Ford, L.S.; Sickles, L.; Järvinen, K.M.; Sicherer, S.H.; Sampson, H.A.; Nowak-Węgrzyn, A. Clinical Features and Resolution of Food Protein–Induced Enterocolitis Syndrome: 10-Year Experience. J. Allergy Clin. Immunol. 2014, 134, 382–389.e4. [Google Scholar] [CrossRef]
- Agyemang, A.; Nowak-Wegrzyn, A. Food Protein-Induced Enterocolitis Syndrome: A Comprehensive Review. Clin. Rev. Allerg Immunol. 2019, 57, 261–271. [Google Scholar] [CrossRef]
- Ishige, T.; Yagi, H.; Tatsuki, M.; Hatori, R.; Nishida, Y.; Takizawa, T.; Arakawa, H. Endoscopic Findings in the Acute Phase of Food Protein-Induced Enterocolitis Syndromae. Pediatr. Allergy Immunol. 2015, 26, 90–91. [Google Scholar] [CrossRef]
- Hwang, J.-B.; Sohn, S.M.; Kim, A.S. Prospective Follow-up Oral Food Challenge in Food Protein-Induced Enterocolitis Syndrome. Arch. Dis. Child. 2009, 94, 425–428. [Google Scholar] [CrossRef]
- Katz, Y.; Goldberg, M.R.; Rajuan, N.; Cohen, A.; Leshno, M. The Prevalence and Natural Course of Food Protein–Induced Enterocolitis Syndrome to Cow’s Milk: A Large-Scale, Prospective Population-Based Study. J. Allergy Clin. Immunol. 2011, 127, 647–653.e3. [Google Scholar] [CrossRef]
- Ruffner, M.A.; Ruymann, K.; Barni, S.; Cianferoni, A.; Brown-Whitehorn, T.; Spergel, J.M. Food Protein-Induced Enterocolitis Syndrome: Insights from Review of a Large Referral Population. J. Allergy Clin. Immunol. Pract. 2013, 1, 343–349. [Google Scholar] [CrossRef]
- Infante, S.; Cabrera-Freitag, P.; Morales-Cabeza, C.; Alvarez-Perea, A. Geographical Variations in Food Protein-Induced Enterocolitis Syndrome. Curr. Treat. Options Allergy 2019, 6, 309–321. [Google Scholar] [CrossRef]
- Bird, J.A.; Barni, S.; Brown-Whitehorn, T.F.; du Toit, G.; Infante, S.; Nowak-Wegrzyn, A. FPIES Oral Food Challenge: Time for a Change? Ann. Allergy Asthma Immunol. 2021, 126, 506–515. [Google Scholar] [CrossRef]
- Infante, S.; Marco-Martín, G.; Sánchez-Domínguez, M.; Rodríguez-Fernández, A.; Fuentes-Aparicio, V.; Alvarez-Perea, A.; Cabrera-Freitag, P.; Morales-Cabeza, C.; Zubeldia, J.M.; Zapatero, L. Food Protein-Induced Enterocolitis Syndrome by Fish: Not Necessarily a Restricted Diet. Allergy 2018, 73, 728–732. [Google Scholar] [CrossRef]
- Infante, S.; Marco-Martín, G.; Zubeldia, J.M.; Fuentes-Aparicio, V.; Alvarez-Perea, A.; Cabrera-Freitag, P.; Morales-Cabeza, C.; Zapatero, L. Oral Food Challenge in Food Protein-Induced Enterocolitis Syndrome by Fish: Is There Any Room for Improvement? Int. Arch. Allergy Immunol. 2019, 179, 215–220. [Google Scholar] [CrossRef]
- Mehr, S.; Campbell, D.E. Food Protein-induced Enterocolitis Syndrome: Guidelines Summary and Practice Recommendations. Med. J. Aust. 2019, 210, 94–99. [Google Scholar] [CrossRef]
- Barni, S.; Vazquez-Ortiz, M.; Giovannini, M.; Liccioli, G.; Sarti, L.; Cianferoni, A.; Mori, F. Diagnosing Food Protein-induced Enterocolitis Syndrome. Clin. Exp. Allergy 2021, 51, 14–28. [Google Scholar] [CrossRef] [PubMed]
- Zubeldia-Varela, E.; Barber, D.; Barbas, C.; Perez-Gordo, M.; Rojo, D. Sample Pre-Treatment Procedures for the Omics Analysis of Human Gut Microbiota: Turning Points, Tips and Tricks for Gene Sequencing and Metabolomics. J. Pharm. Biomed. Anal. 2020, 191, 113592. [Google Scholar] [CrossRef] [PubMed]
- Eguiluz-Gracia, I.; Tay, T.R.; Hew, M.; Escribese, M.M.; Barber, D.; O’Hehir, R.E.; Torres, M.J. Recent Developments and Highlights in Biomarkers in Allergic Diseases and Asthma. Allergy 2018, 73, 2290–2305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fritsché, R. Animal Models in Food Allergy: Assessment of Allergenicity and Preventive Activity of Infant Formulas. Toxicol. Lett. 2003, 140, 303–309. [Google Scholar] [CrossRef]
- Schülke, S.; Albrecht, M. Mouse Models for Food Allergies: Where Do We Stand? Cells 2019, 8, 546. [Google Scholar] [CrossRef] [Green Version]
- Burggraf, M.; Nakajima-Adachi, H.; Hachimura, S.; Ilchmann, A.; Pemberton, A.D.; Kiyono, H.; Vieths, S.; Toda, M. Oral Tolerance Induction Does Not Resolve Gastrointestinal Inflammation in a Mouse Model of Food Allergy. Mol. Nutr. Food Res. 2011, 55, 1475–1483. [Google Scholar] [CrossRef]
- Mishra, A.; Hogan, S.P.; Brandt, E.B.; Rothenberg, M.E. An Etiological Role for Aeroallergens and Eosinophils in Experimental Esophagitis. J. Clin. Investig. 2001, 107, 83–90. [Google Scholar] [CrossRef] [Green Version]
- Mishra, A.; Hogan, S.P.; Brandt, E.B.; Rothenberg, M.E. IL-5 Promotes Eosinophil Trafficking to the Esophagus. J. Immunol. 2002, 168, 2464–2469. [Google Scholar] [CrossRef] [Green Version]
- Hogan, S.P.; Mishra, A.; Brandt, E.B.; Royalty, M.P.; Pope, S.M.; Zimmermann, N.; Foster, P.S.; Rothenberg, M.E. A Pathological Function for Eotaxin and Eosinophils in Eosinophilic Gastrointestinal Inflammation. Nat. Immunol. 2001, 2, 353–360. [Google Scholar] [CrossRef]
- Gonipeta, B.; Kim, E.; Gangur, V. Mouse Models of Food Allergy: How Well Do They Simulate the Human Disorder? Crit. Rev. Food Sci. Nutr. 2015, 55, 437–452. [Google Scholar] [CrossRef]
- Helm, R.M. Food Allergy Animal Models: An Overview. Ann. N. Y. Acad. Sci. 2002, 964, 139–150. [Google Scholar] [CrossRef]
- Kanagaratham, C.; Sallis, B.F.; Fiebiger, E. Experimental Models for Studying Food Allergy. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 356–369.e1. [Google Scholar] [CrossRef]
- Liu, T.; Navarro, S.; Lopata, A.L. Current Advances of Murine Models for Food Allergy. Mol. Immunol. 2016, 70, 104–117. [Google Scholar] [CrossRef]
- Huang, J.; Liu, C.; Wang, Y.; Wang, C.; Xie, M.; Qian, Y.; Fu, L. Application of in Vitro and in Vivo Models in the Study of Food Allergy. Food Sci. Hum. Wellness 2018, 7, 235–243. [Google Scholar] [CrossRef]
- Blanco-Pérez, F.; Kato, Y.; Gonzalez-Menendez, I.; Laiño, J.; Ohbayashi, M.; Burggraf, M.; Krause, M.; Kirberg, J.; Iwakura, Y.; Martella, M.; et al. CCR8 Leads to Eosinophil Migration and Regulates Neutrophil Migration in Murine Allergic Enteritis. Sci. Rep. 2019, 9, 9608. [Google Scholar] [CrossRef]
- Nakajima-Adachi, H.; Kikuchi, A.; Fujimura, Y.; Shibahara, K.; Makino, T.; Goseki-Sone, M.; Kihara-Fujioka, M.; Nochi, T.; Kurashima, Y.; Igarashi, O.; et al. Peyer’s Patches and Mesenteric Lymph Nodes Cooperatively Promote Enteropathy in a Mouse Model of Food Allergy. PLoS ONE 2014, 9, e107492. [Google Scholar] [CrossRef]
- Chehade, M.; Aceves, S.S. Food Allergy and Eosinophilic Esophagitis. Curr. Opin. Allergy Clin. Immunol. 2010, 10, 231–237. [Google Scholar] [CrossRef]
- Mishra, A.; Rothenberg, M.E. Intratracheal IL-13 Induces Eosinophilic Esophagitis by an IL-5, Eotaxin-1, and STAT6-Dependent Mechanism1. Gastroenterology 2003, 125, 1419–1427. [Google Scholar] [CrossRef]
- Mishra, A.; Schlotman, J.; Wang, M.; Rothenberg, M.E. Critical Role for Adaptive T Cell Immunity in Experimental Eosinophilic Esophagitis in Mice. J. Leukoc. Biol. 2007, 81, 916–924. [Google Scholar] [CrossRef]
- Niranjan, R.; Mavi, P.; Rayapudi, M.; Dynda, S.; Mishra, A. Pathogenic Role of Mast Cells in Experimental Eosinophilic Esophagitis. Am. J. Physiol.-Gastrointest. Liver Physiol. 2013, 304, G1087–G1094. [Google Scholar] [CrossRef]
- iHMP Consortium. The Integrative HMP (iHMP) Research Network Consortium The Integrative Human Microbiome Project. Nature 2019, 569, 641–648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Falony, G.; Joossens, M.; Vieira-Silva, S.; Wang, J.; Darzi, Y.; Faust, K.; Kurilshikov, A.; Bonder, M.J.; Valles-Colomer, M.; Vandeputte, D.; et al. Population-Level Analysis of Gut Microbiome Variation. Science 2016, 352, 560–564. [Google Scholar] [CrossRef] [PubMed]
- Goodacre, R. Metabolomics of a Superorganism. J. Nutr. 2007, 137, 259S–266S. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human Gut Microbiome Viewed across Age and Geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef]
- Rojo, D.; Méndez-García, C.; Raczkowska, B.A.; Bargiela, R.; Moya, A.; Ferrer, M.; Barbas, C. Exploring the Human Microbiome from Multiple Perspectives: Factors Altering Its Composition and Function. FEMS Microbiol. Rev. 2017, 41, 453–478. [Google Scholar] [CrossRef] [Green Version]
- Ruiz-Ruiz, S.; Sanchez-Carrillo, S.; Ciordia, S.; Mena, M.C.; Méndez-García, C.; Rojo, D.; Bargiela, R.; Zubeldia-Varela, E.; Martínez-Martínez, M.; Barbas, C.; et al. Functional Microbiome Deficits Associated with Ageing: Chronological Age Threshold. Aging Cell 2020, 19, e13063. [Google Scholar] [CrossRef]
- Cseh, A.; Molnár, K.; Pintér, P.; Szalay, B.; Szebeni, B.; Treszl, A.; Arató, A.; Vásárhelyi, B.; Veres, G. Regulatory T Cells and T Helper Subsets in Breast-Fed Infants with Hematochezia Caused by Allergic Colitis. J. Pediatr. Gastroenterol. Nutr. 2010, 51, 675–677. [Google Scholar] [CrossRef]
- Tan, J.; McKenzie, C.; Vuillermin, P.J.; Goverse, G.; Vinuesa, C.G.; Mebius, R.E.; Macia, L.; Mackay, C.R. Dietary Fiber and Bacterial SCFA Enhance Oral Tolerance and Protect against Food Allergy through Diverse Cellular Pathways. Cell Rep. 2016, 15, 2809–2824. [Google Scholar] [CrossRef] [Green Version]
- Pascal, M.; Perez-Gordo, M.; Caballero, T.; Escribese, M.M.; Lopez Longo, M.N.; Luengo, O.; Manso, L.; Matheu, V.; Seoane, E.; Zamorano, M.; et al. Microbiome and Allergic Diseases. Front. Immunol. 2018, 9, 1584. [Google Scholar] [CrossRef]
- Bunyavanich, S.; Shen, N.; Grishin, A.; Wood, R.; Burks, W.; Dawson, P.; Jones, S.M.; Leung, D.Y.M.; Sampson, H.; Sicherer, S.; et al. Early-Life Gut Microbiome Composition and Milk Allergy Resolution. J. Allergy Clin. Immunol. 2016, 138, 1122–1130. [Google Scholar] [CrossRef] [Green Version]
- Fazlollahi, M.; Chun, Y.; Grishin, A.; Wood, R.A.; Burks, A.W.; Dawson, P.; Jones, S.M.; Leung, D.Y.M.; Sampson, H.A.; Sicherer, S.H.; et al. Early-Life Gut Microbiome and Egg Allergy. Allergy 2018, 73, 1515–1524. [Google Scholar] [CrossRef]
- Berni Canani, R.; De Filippis, F.; Nocerino, R.; Paparo, L.; Di Scala, C.; Cosenza, L.; Della Gatta, G.; Calignano, A.; De Caro, C.; Laiola, M.; et al. Gut Microbiota Composition and Butyrate Production in Children Affected by Non-IgE-Mediated Cow’s Milk Allergy. Sci. Rep. 2018, 8, 12500. [Google Scholar] [CrossRef] [Green Version]
- Pannaraj, P.S.; Li, F.; Cerini, C.; Bender, J.M.; Yang, S.; Rollie, A.; Adisetiyo, H.; Zabih, S.; Lincez, P.J.; Bittinger, K.; et al. Association Between Breast Milk Bacterial Communities and Establishment and Development of the Infant Gut Microbiome. JAMA Pediatr. 2017, 171, 647–654. [Google Scholar] [CrossRef]
- Azad, M.B.; Konya, T.; Guttman, D.S.; Field, C.J.; Sears, M.R.; HayGlass, K.T.; Mandhane, P.J.; Turvey, S.E.; Subbarao, P.; Becker, A.B.; et al. Infant Gut Microbiota and Food Sensitization: Associations in the First Year of Life. Clin. Exp. Allergy 2015, 45, 632–643. [Google Scholar] [CrossRef]
- Trompette, A.; Gollwitzer, E.S.; Yadava, K.; Sichelstiel, A.K.; Sprenger, N.; Ngom-Bru, C.; Blanchard, C.; Junt, T.; Nicod, L.P.; Harris, N.L.; et al. Gut Microbiota Metabolism of Dietary Fiber Influences Allergic Airway Disease and Hematopoiesis. Nat. Med. 2014, 20, 159–166. [Google Scholar] [CrossRef]
- Obeso, D.; Mera-Berriatua, L.; Rodríguez-Coira, J.; Rosace, D.; Fernández, P.; Martín-Antoniano, I.A.; Santaolalla, M.; Marco Martín, G.; Chivato, T.; Fernández-Rivas, M.; et al. Multi-Omics Analysis Points to Altered Platelet Functions in Severe Food-Associated Respiratory Allergy. Allergy 2018, 73, 2137–2149. [Google Scholar] [CrossRef]
- Wörheide, M.A.; Krumsiek, J.; Kastenmüller, G.; Arnold, M. Multi-Omics Integration in Biomedical Research—A Metabolomics-Centric Review. Anal. Chim. Acta 2021, 1141, 144–162. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Snyder, M.P. Integrative Omics for Health and Disease. Nat. Rev. Genet. 2018, 19, 299–310. [Google Scholar] [CrossRef]
- Segal, J.P.; Mullish, B.H.; Quraishi, M.N.; Acharjee, A.; Williams, H.R.T.; Iqbal, T.; Hart, A.L.; Marchesi, J.R. The Application of Omics Techniques to Understand the Role of the Gut Microbiota in Inflammatory Bowel Disease. Ther. Adv. Gastroenterol. 2019, 12, 175628481882225. [Google Scholar] [CrossRef] [Green Version]
- Zubeldia-Varela, E.; Raczkowska, B.A.; Ferrer, M.; Perez-Gordo, M.; Rojo, D. Chapter 4—Techniques for Phenotyping the Gut Microbiota Metabolome. In Microbiome and Metabolome in Diagnosis, Therapy, and Other Strategic Applications; Faintuch, J., Faintuch, S., Eds.; Academic Press: Cambridge, MA, USA, 2019; pp. 33–41. ISBN 978-0-12-815249-2. [Google Scholar]
- Díaz, M.; Guadamuro, L.; Espinosa-Martos, I.; Mancabelli, L.; Jiménez, S.; Molinos-Norniella, C.; Pérez-Solis, D.; Milani, C.; Rodríguez, J.M.; Ventura, M.; et al. Microbiota and Derived Parameters in Fecal Samples of Infants with Non-IgE Cow’s Milk Protein Allergy under a Restricted Diet. Nutrients 2018, 10, 1481. [Google Scholar] [CrossRef] [Green Version]
- Candy, D.C.A.; Van Ampting, M.T.J.; Nijhuis, M.M.O.; Wopereis, H.; Butt, A.M.; Peroni, D.G.; Vandenplas, Y.; Fox, A.T.; Shah, N.; West, C.E.; et al. A Synbiotic-Containing Amino-Acid-Based Formula Improves Gut Microbiota in Non-IgE-Mediated Allergic Infants. Pediatr. Res. 2018, 83, 677–686. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fox, A.T.; Wopereis, H.; Van Ampting, M.T.J.; Nijhuis, M.M.O.; Butt, A.M.; Peroni, D.G.; Vandenplas, Y.; Candy, D.C.A.; Shah, N.; West, C.E.; et al. A Specific Synbiotic-Containing Amino Acid-Based Formula in Dietary Management of Cow’s Milk Allergy: A Randomized Controlled Trial. Clin. Transl. Allergy 2019, 9, 5. [Google Scholar] [CrossRef] [PubMed]
- Wopereis, H.; van Ampting, M.T.J.; Cetinyurek-Yavuz, A.; Slump, R.; Candy, D.C.A.; Butt, A.M.; Peroni, D.G.; Vandenplas, Y.; Fox, A.T.; Shah, N.; et al. A Specific Synbiotic-Containing Amino Acid-Based Formula Restores Gut Microbiota in Non-IgE Mediated Cow’s Milk Allergic Infants: A Randomized Controlled Trial. Clin. Transl. Allergy 2019, 9, 27. [Google Scholar] [CrossRef] [PubMed]
- Guadamuro, L.; Diaz, M.; Jimenez, S.; Molinos-Norniella, C.; Perez-Solis, D.; Miguel Rodriguez, J.; Bousono, C.; Gueimonde, M.; Margolies, A.; Delgado, S.; et al. Fecal Changes Following Introduction of Milk in Infants With Outgrowing Non-IgE Cow’s Milk Protein Allergy Are Influenced by Previous Consumption of the Probiotic LGG. Front. Immunol. 2019, 10, 1819. [Google Scholar] [CrossRef] [Green Version]
- Aparicio, M.; Alba, C.; Rodríguez, J.M.; Fernández, L.; Proctocolitis Study Group of CAM Public Health Area 6. Microbiological and Immunological Markers in Milk and Infant Feces for Common Gastrointestinal Disorders: A Pilot Study. Nutrients 2020, 12, 634. [Google Scholar] [CrossRef] [Green Version]
- Xiong, L.-J.; Xie, X.-L.; Li, Y.; Deng, X.-Z. Current Status of Fecal Calprotectin as a Diagnostic or Monitoring Biomarker for Cow’s Milk Protein Allergy in Children: A Scoping Review. World J. Pediatr. 2021, 17, 63–70. [Google Scholar] [CrossRef]
- Stríz, I.; Trebichavský, I. Calprotectin—A Pleiotropic Molecule in Acute and Chronic Inflammation. Physiol. Res. 2004, 53, 245–253. [Google Scholar]
- Berni Canani, R.; Rapacciuolo, L.; Romano, M.T.; de Horatio, L.T.; Terrin, G.; Manguso, F.; Cirillo, P.; Paparo, F.; Troncone, R. Diagnostic Value of Faecal Calprotectin in Paediatric Gastroenterology Clinical Practice. Dig. Liver Dis. 2004, 36, 467–470. [Google Scholar] [CrossRef]
- Bunn, S.K.; Bisset, W.M.; Main, M.J.C.; Golden, B.E. Fecal Calprotectin as a Measure of Disease Activity in Childhood Inflammatory Bowel Disease. J. Pediatric Gastroenterol. Nutr. 2001, 32, 171–177. [Google Scholar] [CrossRef]
- Chang, M.-H.; Chou, J.-W.; Chen, S.-M.; Tsai, M.-C.; Sun, Y.-S.; Lin, C.-C.; Lin, C.-P. Faecal Calprotectin as a Novel Biomarker for Differentiating between Inflammatory Bowel Disease and Irritable Bowel Syndrome. Mol. Med. Rep. 2014, 10, 522–526. [Google Scholar] [CrossRef]
- Beşer, Ö.F.; Sancak, S.; Erkan, T.; Kutlu, T.; Çokuğraş, H.; Çokuğraş, F.Ç. Can Fecal Calprotectin Level Be Used as a Markers of Inflammation in the Diagnosis and Follow-Up of Cow’s Milk Protein Allergy? Allergy Asthma Immunol. Res. 2014, 6, 33–38. [Google Scholar] [CrossRef] [Green Version]
- Ataee, P.; Zoghali, M.; Nikkhoo, B.; Ghaderi, E.; Mansouri, M.; Nasiri, R.; Eftekhari, K. Diagnostic Value of Fecal Calprotectin in Response to Mother’s Diet in Breast-Fed Infants with Cow’s Milk Allergy Colitis. Iran. J. Pediatr. 2018, 28, e66172. [Google Scholar] [CrossRef]
- Carrell, R.W.; Jeppsson, J.-O.; Laurell, C.-B.; Brennan, S.O.; Owen, M.C.; Vaughan, L.; Boswell, D.R. Structure and Variation of Human α 1-Antitrypsin. Nature 1982, 298, 329–334. [Google Scholar] [CrossRef]
- Elli, L.; Topa, M.; Rimondi, A. Protein-Losing Enteropathy. Curr. Opin. Gastroenterol. 2020, 36, 238–244. [Google Scholar] [CrossRef]
- Singer, A.A.M. Food Protein–Induced Enterocolitis and Alpha-1-Antitrypsin Deficiency. J. Allergy Clin. Immunol. 2020, 145, 444. [Google Scholar] [CrossRef]
- Kalach, N.; Kapel, N.; Waligora-Dupriet, A.-J.; Castelain, M.-C.; Cousin, M.O.; Sauvage, C.; Ba, F.; Nicolis, I.; Campeotto, F.; Butel, M.J.; et al. Intestinal Permeability and Fecal Eosinophil-Derived Neurotoxin Are the Best Diagnosis Tools for Digestive Non-IgE-Mediated Cow’s Milk Allergy in Toddlers. Clin. Chem. Lab. Med. 2013, 51, 351–361. [Google Scholar] [CrossRef]
- Malik, A.; Batra, J.K. Antimicrobial Activity of Human Eosinophil Granule Proteins: Involvement in Host Defence against Pathogens. Crit. Rev. Microbiol. 2012, 38, 168–181. [Google Scholar] [CrossRef]
- Yamada, Y. Unique Features of Non-IgE-Mediated Gastrointestinal Food Allergy during Infancy in Japan. Curr. Opin. Allergy Clin. Immunol. 2020, 20, 299–304. [Google Scholar] [CrossRef]
- Roca, M.; Rodriguez Varela, A.; Donat, E.; Cano, F.; Hervas, D.; Armisen, A.; Vaya, M.J.; Sjölander, A.; Ribes-Koninckx, C. Fecal Calprotectin and Eosinophil-Derived Neurotoxin in Healthy Children Between 0 and 12 Years. J. Pediatr. Gastroenterol. Nutr. 2017, 65, 394–398. [Google Scholar] [CrossRef]
- Rycyk, A.; Cudowska, B.; Lebensztejn, D.M. Eosinophil-Derived Neurotoxin, Tumor Necrosis Factor Alpha, and Calprotectin as Non-Invasive Biomarkers of Food Protein-Induced Allergic Proctocolitis in Infants. J. Clin. Med. 2020, 9, 3147. [Google Scholar] [CrossRef]
- Wada, H.; Horisawa, T.; Inoue, M.; Yoshida, T.; Toma, T.; Yachie, A. Sequential Measurement of Fecal Parameters in a Case of Non-Immunoglobulin E-Mediated Milk Allergy. Pediatr. Int. 2007, 49, 109–111. [Google Scholar] [CrossRef] [PubMed]
- Wada, T.; Toma, T.; Muraoka, M.; Matsuda, Y.; Yachie, A. Elevation of Fecal Eosinophil-Derived Neurotoxin in Infants with Food Protein-Induced Enterocolitis Syndrome. Pediatr. Allergy Immunol. 2014, 25, 617–619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mennini, M.; Fiocchi, A.G.; Cafarotti, A.; Montesano, M.; Mauro, A.; Villa, M.P.; Di Nardo, G. Food Protein-Induced Allergic Proctocolitis in Infants: Literature Review and Proposal of a Management Protocol. World Allergy Organ. J. 2020, 13, 100471. [Google Scholar] [CrossRef] [PubMed]
- Miceli Sopo, S.; Monaco, S.; Bersani, G.; Romano, A.; Fantacci, C. Proposal for Management of the Infant with Suspected Food Protein-Induced Allergic Proctocolitis. Pediatr. Allergy Immunol. 2018, 29, 215–218. [Google Scholar] [CrossRef]
- Joshi, S.R.; Nicolaides, R.E.; Bird, J.A. Acute Food Protein-Induced Enterocolitis Syndrome. In Food Protein Induced Enterocolitis (FPIES): Diagnosis and Management; Brown-Whitehorn, T.F., Cianferoni, A., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 31–67. ISBN 978-3-030-21229-2. [Google Scholar]
- Jarocka-Cyrta, E.; Valverde-Monge, M.; Nowak-Węgrzyn, A. Management of Food Protein-Induced Enterocolitis Syndrome (FPIES): Current Approach and Future Needs. Curr. Treat. Options Allergy 2017, 4, 383–394. [Google Scholar] [CrossRef]
- Wood, R.A. Advances in Food Allergy in 2015. J. Allergy Clin. Immunol. 2016, 138, 1541–1547. [Google Scholar] [CrossRef] [Green Version]
- Potaczek, D.P.; Alashkar Alhamwe, B.; Miethe, S.; Garn, H. Epigenetic Mechanisms in Allergy Development and Prevention. In Handbook of Experimental Pharmacology; Springer: Berlin/Heidelberg, Germany, 2021. [Google Scholar] [CrossRef]
- Berin, C. Advances in Understanding Immune Mechanisms of Food Protein Induced Enterocolitis Syndrome. Ann. Allergy Asthma Immunol. 2021, 126, 478–481. [Google Scholar] [CrossRef]
| Disease | Description | Diet of Allergic Infants | Microbial Communities (Genomics) | Metabolites (Metabolomics) | Reference |
|---|---|---|---|---|---|
| Non-IgE-mediated CMA | Allergic vs. healthy children | No dietary restriction (collection at diagnosis) | ↑ Phylum Bacteroidetes ↑ Genera Bacteroides, Alistipes, and Sarcina | ↓ Fecal butyrate | Berni Canani et al. [63] |
| Non-IgE-mediated CMA | Allergic vs. healthy infants (unrestricted conventional diet) | Milk exclusion diet for six months:
| EHF, SPBF & HRF: ↑ Families Lachnospiraceae and Coriobacteriaceae ↓ Genus Bacteroides HRF: ↓ Genus Bifidobacteria | ↑ Isobutyric and isovaleric acids = Acetic, propionic, and butyric acids | Díaz et al. [72] |
| Non-IgE-mediated CMA | Allergic with test formula vs. allergic with control formula vs. HBI at week 8 | Amino acid-based formula for 8 weeks:
| Bifidobacteria: HBI > test formula > control formula Eubacterium rectale/Clostridium coccoides: HBI < test formula < control formula | - | Candy et al. [73] |
| Non-IgE-mediated CMA | Allergic with test formula vs. allergic with control formula at week 26 | Formula for 26 weeks:
| Bifidobacteria: test formula > control formula Eubacterium rectale/Clostridium coccoides: test formula < control formula | - | Fox et al. [74] |
| Non-IgE-mediated CMA | Allergic with test formula vs. allergic with control formula vs. HBI | Amino acid-based formula:
| Test vs. control: ↑ Species of Bifidobacterium and Veillonella families ↓ Species of Lachnospiraceae family and Ruminococcus and Alistipes genera HBI vs. test formula: ≈ Bifidobacterium spp. and Lachnospiraceae spp. | Test vs. control: ↑ L-lactate ↓ Valeric and isobutyric acids = Acetate, propionate, butyrate, iso-valerate, and D-lactate | Wopereis et al. [75] |
| Non-IgE-mediated CMA | Allergic children before and after milk challenges | Hypoallergenic hydrolyzed formulas | ↑ Lactic acid bacteria (genus Lactobacillus) ↓ Bacterial groups of Lachnospiraceae and Ruminococcaceae families | ↑ Skatole ↓ p-cresol and BCFAs = Acetic, propionic, and butyric acids | Guadamuro et al. [76] |
| Gastrointestinal disorders | Colic vs. non-IgE-mediated CMA vs. proctocolitis vs. healthy controls | Exclusive breastfeeding | Fecal microbiota in non-IgE-mediated CMA: ↑ Families Eggerthellaceae, Lachnospiraceae, and Peptostreptococcaceae ↑ Genera Intestinibacter (more increased in proctocolitis) and Rothia↓ Genera Erysipelatoclostridium (higher in controls) and Bifidobacterium Breast milk: ↑ Family Eggerthellaceae in non-IgE-mediated CMA ↑ Family Prolixibacteraceae in proctocolitis | - | Aparicio et al. [77] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zubeldia-Varela, E.; Barker-Tejeda, T.C.; Blanco-Pérez, F.; Infante, S.; Zubeldia, J.M.; Pérez-Gordo, M. Non-IgE-Mediated Gastrointestinal Food Protein-Induced Allergic Disorders. Clinical Perspectives and Analytical Approaches. Foods 2021, 10, 2662. https://doi.org/10.3390/foods10112662
Zubeldia-Varela E, Barker-Tejeda TC, Blanco-Pérez F, Infante S, Zubeldia JM, Pérez-Gordo M. Non-IgE-Mediated Gastrointestinal Food Protein-Induced Allergic Disorders. Clinical Perspectives and Analytical Approaches. Foods. 2021; 10(11):2662. https://doi.org/10.3390/foods10112662
Chicago/Turabian StyleZubeldia-Varela, Elisa, Tomás Clive Barker-Tejeda, Frank Blanco-Pérez, Sonsoles Infante, José M. Zubeldia, and Marina Pérez-Gordo. 2021. "Non-IgE-Mediated Gastrointestinal Food Protein-Induced Allergic Disorders. Clinical Perspectives and Analytical Approaches" Foods 10, no. 11: 2662. https://doi.org/10.3390/foods10112662
APA StyleZubeldia-Varela, E., Barker-Tejeda, T. C., Blanco-Pérez, F., Infante, S., Zubeldia, J. M., & Pérez-Gordo, M. (2021). Non-IgE-Mediated Gastrointestinal Food Protein-Induced Allergic Disorders. Clinical Perspectives and Analytical Approaches. Foods, 10(11), 2662. https://doi.org/10.3390/foods10112662








