Optical Imaging in Brainsmatics
Abstract
1. Neurosciences Call for Brainsmatics
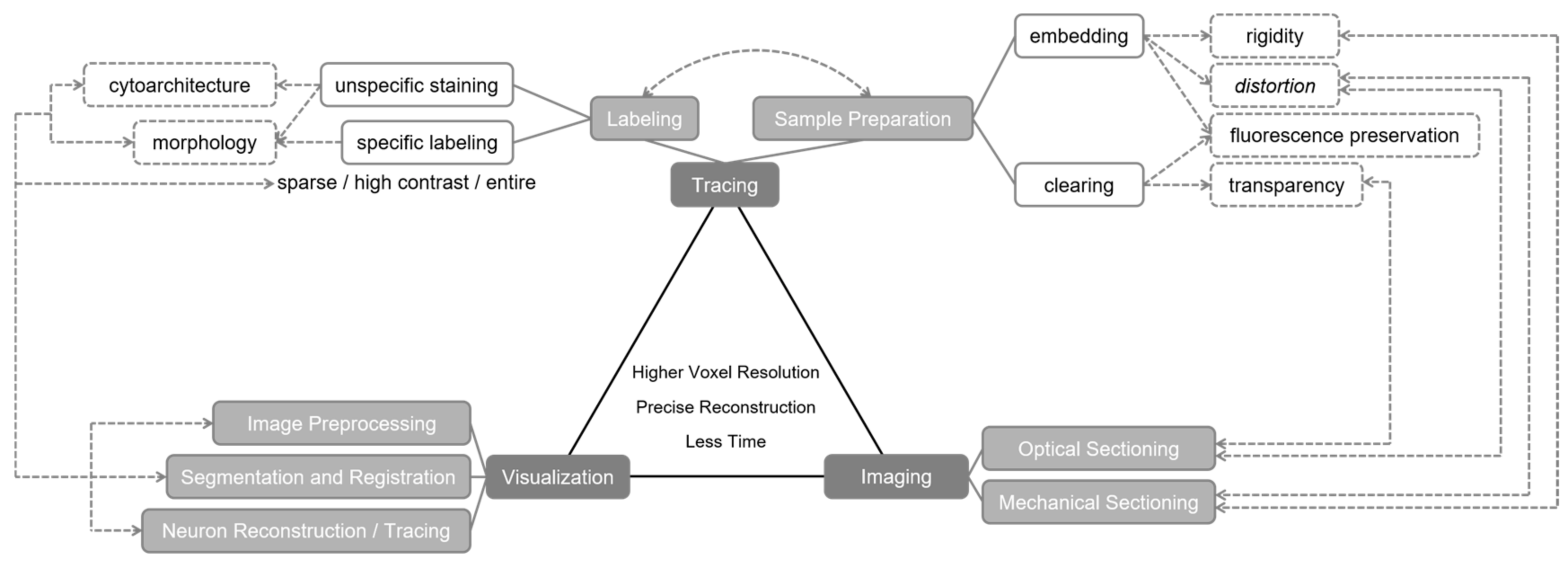
2. Roles of Optical Imaging in Brainsmatics
2.1. Tracing Approaches Prior to Optical Imaging
2.1.1. Specific Labeling
2.1.2. Sample Preparation
2.2. Optical Imaging in Brain Surveying
2.2.1. Mechanical Sectioning Strategy
2.2.2. Optical Sectioning Strategy
2.3. Data Processing after Optical Imaging
3. Ideal Optical Imaging in Brainsmatics
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sudhof, T.C. Molecular Neuroscience in the 21st Century: A Personal Perspective. Neuron 2017, 96, 536–541. [Google Scholar] [CrossRef] [PubMed]
- Jorgenson, L.A.; Newsome, W.T.; Anderson, D.J.; Bargmann, C.I.; Brown, E.N.; Deisseroth, K.; Donoghue, J.P.; Hudson, K.L.; Ling, G.S.; MacLeish, P.R.; et al. The BRAIN Initiative: Developing technology to catalyse neuroscience discovery. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140164:1–20140164:12. [Google Scholar] [CrossRef] [PubMed]
- Bassett, D.S.; Sporns, O. Network neuroscience. Nat. Neurosci. 2017, 20, 353–364. [Google Scholar] [CrossRef] [PubMed]
- Lichtman, J.W.; Sanes, J.R. Ome sweet ome: What can the genome tell us about the connectome? Curr. Opin. Neurobiol. 2008, 18, 346–353. [Google Scholar] [CrossRef] [PubMed]
- White, J.G.; Southgate, E.; Thomson, J.N.; Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R. Soc. B Biol. Sci. 1986, 314, 1–340. [Google Scholar] [CrossRef] [PubMed]
- Chiang, A.; Lin, C.; Chuang, C.; Chang, H.; Hsieh, C.; Yeh, C.; Shih, C.; Wu, J.; Wang, G.; Chen, Y.; et al. Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution. Curr. Biol. 2011, 21, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Hildebrand, D.G.C.; Cicconet, M.; Torres, R.M.; Choi, W.; Quan, T.M.; Moon, J.; Wetzel, A.W.; Scott Champion, A.; Graham, B.J.; Randlett, O.; et al. Whole-brain serial-section electron microscopy in larval zebrafish. Nature 2017, 545, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.W.; Harris, J.A.; Ng, L.; Winslow, B.; Cain, N.; Mihalas, S.; Wang, Q.; Lau, C.; Kuan, L.; Henry, A.M.; et al. A mesoscale connectome of the mouse brain. Nature 2014, 508, 207–214. [Google Scholar] [CrossRef]
- Craddock, R.C.; James, G.A.; Holtzheimer, P.E.; Hu, X.P.; Mayberg, H.S. A whole brain fMRI atlas generated via spatially constrained spectral clustering. Hum. Brain Mapp. 2012, 33, 1914–1928. [Google Scholar] [CrossRef]
- Amunts, K.; Lepage, C.; Borgeat, L.; Mohlberg, H.; Dickscheid, T.; Rousseau, M.E.; Bludau, S.; Bazin, P.L.; Lewis, L.B.; Oros-Peusquens, A.M.; et al. BigBrain: An ultrahigh-resolution 3D human brain model. Science 2013, 340, 1472–1475. [Google Scholar] [CrossRef]
- Fan, L.; Li, H.; Zhuo, J.; Zhang, Y.; Wang, J.; Chen, L.; Yang, Z.; Chu, C.; Xie, S.; Laird, A.R.; et al. The Human Brainnetome Atlas: A New Brain Atlas Based on Connectional Architecture. Cereb. Cortex 2016, 26, 3508–3526. [Google Scholar] [CrossRef] [PubMed]
- Lein, E.S.; Hawrylycz, M.J.; Ao, N.; Ayres, M.; Bensinger, A.; Bernard, A.; Boe, A.F.; Boguski, M.S.; Brockway, K.S.; Byrnes, E.J.; et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 2007, 445, 168–176. [Google Scholar] [CrossRef] [PubMed]
- Bakken, T.E.; Miller, J.A.; Ding, S.; Sunkin, S.M.; Smith, K.A.; Ng, L.; Szafer, A.; Dalley, R.A.; Royall, J.J.; Lemon, T.; et al. A comprehensive transcriptional map of primate brain development. Nature 2016, 535, 367–375. [Google Scholar] [CrossRef]
- Miller, J.A.; Ding, S.; Sunkin, S.M.; Smith, K.A.; Ng, L.; Szafer, A.; Ebbert, A.; Riley, Z.L.; Royall, J.J.; Aiona, K.; et al. Transcriptional landscape of the prenatal human brain. Nature 2014, 508, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, B.; Sun, Q.; Zhang, Y.; Ren, M.; Zhang, X.; Li, A.; Yuan, J.; Madisen, L.; Luo, Q.; et al. Generation of a whole-brain atlas for the cholinergic system and mesoscopic projectome analysis of basal forebrain cholinergic neurons. Proc. Natl. Acad. Sci. USA 2018, 115, 415–420. [Google Scholar] [CrossRef]
- Beliveau, V.; Ganz, M.; Feng, L.; Ozenne, B.; Højgaard, L.; Fisher, P.M.; Svarer, C.; Greve, D.N.; Knudsen, G.M. A High-Resolution in vivo Atlas of the Human Brain’s Serotonin System. J. Neurosci. 2017, 37, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Lei, V.L.C.; Li, D.; Hu, Z.; Xiang, Y.; Yuan, Z. Mapping the small-world properties of brain networks in Chinese to English simultaneous interpreting by using functional near-infrared spectroscopy. J. Innov. Opt. Heal. Sci. 2018, 11, 1840001:1–1840001:12. [Google Scholar] [CrossRef]
- Lamprecht, R.; LeDoux, J. Structural plasticity and memory. Nat. Rev. Neurosci. 2004, 5, 45–54. [Google Scholar] [CrossRef]
- Burke, S.N.; Barnes, C.A. Neural plasticity in the ageing brain. Nat. Rev. Neurosci. 2006, 7, 30–40. [Google Scholar] [CrossRef] [PubMed]
- Young, M.P.; Scannell, J.W. Brain Structure-Function Relationships: Advances from Neuroinformatics. Philos. Trans. R. Soc. B Biol. Sci. 2000, 355, 3–6. [Google Scholar] [CrossRef][Green Version]
- Nayak, L.; Dasgupta, A.; Das, R.; Ghosh, K.; De Rajat, K. Computational neuroscience and neuroinformatics: Recent progress and resources. J. Biosci. 2018, 43, 1037–1054. [Google Scholar] [CrossRef] [PubMed]
- Frackowiak, R.; Markram, H. The future of human cerebral cartography: A novel approach. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140171:1–20140171:13. [Google Scholar] [CrossRef] [PubMed]
- Bjerke, I.E.; Øvsthus, M.; Papp, E.A.; Yates, S.C.; Silvestri, L.; Fiorilli, J.; Pennartz, C.M.A.; Pavone, F.S.; Puchades, M.A.; Leergaard, T.B.; et al. Data integration through brain atlasing: Human Brain Project tools and strategies. Eur. Psychiat. 2018, 50, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q. Brainsmatics—bridging the brain science and brain-inspired artificial intelligence. Sci. Sin. Vitae 2017, 47, 1015–1024. (In Chinese) [Google Scholar] [CrossRef]
- Reardon, S. Giant neuron encircles entire brain of a mouse. Nature 2017, 543, 14–15. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H. Mesoscale connectomics. Curr. Opin. Neurobiol. 2018, 50, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, L.; Mascaro, A.A.; Lotti, J.; Sacconi, L.; Pavone, F.S. Advanced optical techniques to explore brain structure and function. J. Innov. Opt. Heal. Sci. 2013, 6, 1230002:1–1230002:15. [Google Scholar] [CrossRef]
- Golgi, C. Sulla struttura della sostanza grigia del cervello. Gazz. Med. Ital. 1873, 33, 244–246. [Google Scholar]
- Ecker, J.R.; Geschwind, D.H.; Kriegstein, A.R.; Ngai, J.; Osten, P.; Polioudakis, D.; Regev, A.; Sestan, N.; Wickersham, I.R.; Zeng, H. The BRAIN Initiative Cell Census Consortium: Lessons Learned toward Generating a Comprehensive Brain Cell Atlas. Neuron 2017, 96, 542–557. [Google Scholar] [CrossRef]
- Kuzmenkov, A.I.; Vassilevski, A.A. Labelled animal toxins as selective molecular markers of ion channels: Applications in neurobiology and beyond. Neurosci. Lett. 2018, 679, 15–23. [Google Scholar] [CrossRef]
- Micheva, K.D.; Smith, S.J. Array tomography: A new tool for imaging the molecular architecture and ultrastructure of neural circuits. Neuron 2007, 55, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Zingg, B.; Hintiryan, H.; Gou, L.; Song, M.Y.; Bay, M.; Bienkowski, M.S.; Foster, N.N.; Yamashita, S.; Bowman, I.; Toga, A.W.; et al. Neural Networks of the Mouse Neocortex. Cell 2014, 156, 1096–1111. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Mellor, R.H.; Bernstein, M.; Keller-Peck, C.; Nguyen, Q.T.; Wallace, M.; Nerbonne, J.M.; Lichtman, J.W.; Sanes, J.R. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 2000, 28, 41–51. [Google Scholar] [CrossRef]
- Madisen, L.; Zwingman, T.A.; Sunkin, S.M.; Oh, S.W.; Zariwala, H.A.; Gu, H.; Ng, L.L.; Palmiter, R.D.; Hawrylycz, M.J.; Jones, A.R.; et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 2010, 13, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Livet, J.; Weissman, T.A.; Kang, H.; Draft, R.W.; Lu, J.; Bennis, R.A.; Sanes, J.R.; Lichtman, J.W. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 2007, 450, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Zhao, T.; Petralia, R.S.; Yu, Y.; Peng, H.; Myers, E.; Magee, J.C. mGRASP enables mapping mammalian synaptic connectivity with light microscopy. Nat. Methods 2012, 9, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Chamberlin, N.L.; Du, B.; de Lacalle, S.; Saper, C.B. Recombinant adeno-associated virus vector: Use for transgene expression and anterograde tract tracing in the CNS. Brain Res. 1998, 793, 169–175. [Google Scholar] [CrossRef]
- Zingg, B.; Chou, X.; Zhang, Z.; Mesik, L.; Liang, F.; Tao, H.W.; Zhang, L.I. AAV-Mediated Anterograde Transsynaptic Tagging: Mapping Corticocollicular Input-Defined Neural Pathways for Defense Behaviors. Neuron 2017, 93, 33–47. [Google Scholar] [CrossRef]
- Luo, L.; Callaway, E.M.; Svoboda, K. Genetic Dissection of Neural Circuits. Neuron 2008, 57, 634–660. [Google Scholar] [CrossRef]
- Huang, Z.J.; Zeng, H. Genetic approaches to neural circuits in the mouse. Annu. Rev. Neurosci. 2013, 36, 183–215. [Google Scholar] [CrossRef]
- Lin, R.; Wang, R.; Yuan, J.; Feng, Q.; Zhou, Y.; Zeng, S.; Ren, M.; Jiang, S.; Ni, H.; Zhou, C.; et al. Cell-type-specific and projection-specific brain-wide reconstruction of single neurons. Nat. Methods 2018, 15, 1033–1036. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Hu, B.; Zhang, Y.; Luo, Q.; Gong, H. Development of a Plastic Embedding Method for Large-Volume and Fluorescent-Protein-Expressing Tissues. PLoS ONE 2013, 8, e60877:1–e60877:5. [Google Scholar] [CrossRef] [PubMed]
- Ragan, T.; Kadiri, L.R.; Venkataraju, K.U.; Bahlmann, K.; Sutin, J.; Taranda, J.; Arganda-Carreras, I.; Kim, Y.; Seung, H.S.; Osten, P. Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat. Methods 2012, 9, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Dodt, H.; Leischner, U.; Schierloh, A.; Jährling, N.; Mauch, C.P.; Deininger, K.; Deussing, J.M.; Eder, M.; Zieglgänsberger, W.; Becker, K. Ultramicroscopy: Three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 2007, 4, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Ariel, P. A beginner’s guide to tissue clearing. Int. J. Biochem. Cell Biol. 2017, 84, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Erturk, A.; Becker, K.; Jahrling, N.; Mauch, C.P.; Hojer, C.D.; Egen, J.G.; Hellal, F.; Bradke, F.; Sheng, M.; Dodt, H.U. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 2012, 7, 1983–1995. [Google Scholar] [CrossRef] [PubMed]
- Renier, N.; Wu, Z.; Simon, D.J.; Yang, J.; Ariel, P.; Tessier-Lavigne, M. iDISCO: A Simple, Rapid Method to Immunolabel Large Tissue Samples for Volume Imaging. Cell 2014, 159, 896–910. [Google Scholar] [CrossRef] [PubMed]
- Pan, C.; Cai, R.; Quacquarelli, F.P.; Ghasemigharagoz, A.; Lourbopoulos, A.; Matryba, P.; Plesnila, N.; Dichgans, M.; Hellal, F.; Ertürk, A. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat. Methods 2016, 13, 859–867. [Google Scholar] [CrossRef]
- Cai, R.; Pan, C.; Ghasemigharagoz, A.; Todorov, M.I.; Förstera, B.; Zhao, S.; Bhatia, H.S.; Parra-Damas, A.; Mrowka, L.; Theodorou, D.; et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull–meninges connections. Nat. Neurosci. 2019, 22, 317–327. [Google Scholar] [CrossRef]
- Qi, Y.; Yu, T.; Xu, J.; Wan, P.; Ma, Y.; Zhu, J.; Li, Y.; Gong, H.; Luo, Q.; Zhu, D. FDISCO: Advanced solvent-based clearing method for imaging whole organs. Sci. Adv. 2019, 5, eaau8355:1–eaau8355:13. [Google Scholar] [CrossRef]
- Kuwajima, T.; Sitko, A.A.; Bhansali, P.; Jurgens, C.; Guido, W.; Mason, C. ClearT: A detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 2013, 140, 1364–1368. [Google Scholar] [CrossRef] [PubMed]
- Ke, M.; Fujimoto, S.; Imai, T. SeeDB: A simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 2013, 16, 1154–1161. [Google Scholar] [CrossRef] [PubMed]
- Costantini, I.; Ghobril, J.; Di Giovanna, A.P.; Mascaro, A.L.A.; Silvestri, L.; Müllenbroich, M.C.; Onofri, L.; Conti, V.; Vanzi, F.; Sacconi, L.; et al. A versatile clearing agent for multi-modal brain imaging. Sci. Rep. 2015, 5, 9808:1–9808:9. [Google Scholar] [CrossRef] [PubMed]
- Hama, H.; Kurokawa, H.; Kawano, H.; Ando, R.; Shimogori, T.; Noda, H.; Fukami, K.; Sakaue-Sawano, A.; Miyawaki, A. Scale: A chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 2011, 14, 1481–1488. [Google Scholar] [CrossRef] [PubMed]
- Susaki, E.A.; Tainaka, K.; Perrin, D.; Kishino, F.; Tawara, T.; Watanabe, T.M.; Yokoyama, C.; Onoe, H.; Eguchi, M.; Yamaguchi, S.; et al. Whole-Brain Imaging with Single-Cell Resolution Using Chemical Cocktails and Computational Analysis. Cell 2014, 157, 726–739. [Google Scholar] [CrossRef] [PubMed]
- Hama, H.; Hioki, H.; Namiki, K.; Hoshida, T.; Kurokawa, H.; Ishidate, F.; Kaneko, T.; Akagi, T.; Saito, T.; Saido, T.; et al. ScaleS: An optical clearing palette for biological imaging. Nat. Neurosci. 2015, 18, 1518–1529. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.; Wallace, J.; Kim, S.; Kalyanasundaram, S.; Andalman, A.S.; Davidson, T.J.; Mirzabekov, J.J.; Zalocusky, K.A.; Mattis, J.; Denisin, A.K.; et al. Structural and molecular interrogation of intact biological systems. Nature 2013, 497, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.; Deisseroth, K. CLARITY for mapping the nervous system. Nat. Methods 2013, 10, 508–513. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Treweek, J.B.; Kulkarni, R.P.; Deverman, B.E.; Chen, C.K.; Lubeck, E.; Shah, S.; Cai, L.; Gradinaru, V. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 2014, 158, 945–958. [Google Scholar] [CrossRef]
- Richardson, D.S.; Lichtman, J.W. Clarifying Tissue Clearing. Cell 2015, 162, 246–257. [Google Scholar] [CrossRef]
- Xiong, H.; Zhou, Z.; Zhu, M.; Lv, X.; Li, A.; Li, S.; Li, L.; Yang, T.; Wang, S.; Yang, Z.; et al. Chemical reactivation of quenched fluorescent protein molecules enables resin-embedded fluorescence microimaging. Nat. Commun. 2014, 5, 3992:1–3992:9. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Yang, X.; Chen, S.; Zhang, L.; Long, B.; Tan, C.; Yuan, J.; Gong, H. On-line optical clearing method for whole-brain imaging in mice. Biomed. Opt. Express 2019, 10, 2612–2622. [Google Scholar] [CrossRef] [PubMed]
- Mayerich, D.; Abbott, L.; McCormick, B. Knife-edge scanning microscopy for imaging and reconstruction of three-dimensional anatomical structures of the mouse brain. J. Microsc. 2008, 231, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Gong, H.; Zhang, B.; Wang, Q.; Yan, C.; Wu, J.; Liu, Q.; Zeng, S.; Luo, Q. Micro-Optical Sectioning Tomography to Obtain a High-Resolution Atlas of the Mouse Brain. Science 2010, 330, 1404–1408. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zeng, S.; Yan, C.; Lv, X.; Yang, Z.; Xu, T.; Feng, Z.; Ding, W.; Qi, X.; Li, A.; et al. Continuously tracing brain-wide long-distance axonal projections in mice at a one-micron voxel resolution. Neuroimage 2013, 74, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Yang, Z.; Li, A.; Lv, X.; Zhou, Z.; Wang, X.; Qi, X.; Li, S.; Luo, Q.; Gong, H.; et al. Visualization of brain circuits using two-photon fluorescence micro-optical sectioning tomography. Opt. Express 2013, 21, 9839–9850. [Google Scholar] [CrossRef]
- Gong, H.; Xu, D.; Yuan, J.; Li, X.; Guo, C.; Peng, J.; Li, Y.; Schwarz, L.A.; Li, A.; Hu, B.; et al. High-throughput dual-colour precision imaging for brain-wide connectome with cytoarchitectonic landmarks at the cellular level. Nat. Commun. 2016, 7, 12142:1–12142:12. [Google Scholar] [CrossRef] [PubMed]
- Seiriki, K.; Kasai, A.; Hashimoto, T.; Schulze, W.; Niu, M.; Yamaguchi, S.; Nakazawa, T.; Inoue, K.; Uezono, S.; Takada, M.; et al. High-Speed and Scalable Whole-Brain Imaging in Rodents and Primates. Neuron 2017, 94, 1085–1100. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Tillberg, P.W.; Boyden, E.S. Expansion microscopy. Science 2015, 347, 543–548. [Google Scholar] [CrossRef]
- Nguyen, D.; Marchand, P.J.; Planchette, A.L.; Nilsson, J.; Sison, M.; Extermann, J.; Lopez, A.; Sylwestrzak, M.; Sordet-Dessimoz, J.; Schmidt-Christensen, A.; et al. Optical projection tomography for rapid whole mouse brain imaging. Biomed. Opt. Express 2017, 8, 5637–5650. [Google Scholar] [CrossRef]
- Economo, M.N.; Clack, N.G.; Lavis, L.D.; Gerfen, C.R.; Svoboda, K.; Myers, E.W.; Chandrashekar, J. A platform for brain-wide imaging and reconstruction of individual neurons. ELife 2016, 5, e10566:1–e10566:22. [Google Scholar] [CrossRef] [PubMed]
- Lichtman, J.W.; Denk, W. The big and the small: Challenges of imaging the brain’s circuits. Science 2011, 334, 618–623. [Google Scholar] [CrossRef] [PubMed]
- Malik, W.Q.; Schummers, J.; Sur, M.; Brown, E.N. Denoising Two-Photon Calcium Imaging Data. PLoS ONE 2011, 6, e20490:1–e20490:11. [Google Scholar] [CrossRef] [PubMed]
- Santi, P.A.; Johnson, S.B.; Hillenbrand, M.; Grandpre, P.Z.; Glass, T.J.; Leger, J.R. Thin-sheet laser imaging microscopy for optical sectioning of thick tissues. Biotechniques 2009, 46, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zeng, T.; Peng, H.; Ji, S. Deep Learning Segmentation of Optical Microscopy Images Improves 3-D Neuron Reconstruction. IEEE Trans. Med. Imaging 2017, 36, 1533–1541. [Google Scholar] [CrossRef] [PubMed]
- Haker, S.; Tannenbaum, A.; Kikinis, R. Mass Preserving Mappings and Image Registration; Niessen, W.J., Viergever, M.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2001; pp. 120–127. [Google Scholar]
- Willig, K.I.; Rizzoli, S.O.; Westphal, V.; Jahn, R.; Hell, S.W. STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis. Nature 2006, 440, 935–939. [Google Scholar] [CrossRef] [PubMed]
- Zhong, H. Applying superresolution localization-based microscopy to neurons. Synapse 2015, 69, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Nehme, E.; Weiss, L.E.; Michaeli, T.; Shechtman, Y. Deep-STORM: Super-resolution single-molecule microscopy by deep learning. Optica 2018, 5, 458–464. [Google Scholar] [CrossRef]
- Wang, H.; Rivenson, Y.; Jin, Y.; Wei, Z.; Gao, R.; Günaydın, H.; Bentolila, L.A.; Kural, C.; Ozcan, A. Deep learning enables cross-modality super-resolution in fluorescence microscopy. Nat. Methods 2019, 16, 103–110. [Google Scholar] [CrossRef]
- Kim, K.H.; Buehler, C.; Bahlmann, K.; Ragan, T.; Lee, W.A.; Nedivi, E.; Heffer, E.L.; Fantini, S.; So, P.T.C. Multifocal multiphoton microscopy based on multianode photomultiplier tubes. Opt. Express 2007, 15, 11658–11678. [Google Scholar] [CrossRef]
- Zhang, X.; Chen, Y.; Ning, K.; Zhou, C.; Han, Y.; Gong, H.; Yuan, J. Deep learning optical-sectioning method. Opt. Express 2018, 26, 30762:1–30762:11. [Google Scholar] [CrossRef] [PubMed]
- Cyranoski, D. China launches brain-imaging factory. Nature 2017, 548, 268–269. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Procedures | Methods/Properties | Strategies | ||
|---|---|---|---|---|
| Brain-Wide Positioning System (BPS) [67] | Serial Two-Photon Tomography (STPT) [71] | Block-Face Serial Microscopy Tomography (FAST) [68] | ||
| Tracing Approach | Staining | Cell nuclei stained with propidium iodide (PI) in real time | NuclearID-Red | Cell nuclei stained with Hoechst 33,258 dye; Vascular structures stained with fluorescein-labeled tomato lectin |
| Specific Labeling | AAV-CAG injected into the Cg of a C57BL/6J mouse; AAV-CAGFLEX injected into the posterior region of the anterior hypothalamic area of a SOM-Cre mouse; CAV-Cre and AAV-FLEx(loxP)-TVA-GFP injected into the olfactory bulb and the ipsilateral locus coeruleus of a wild-type mouse | AAV 2/1 Syn-iCre and AAV 2/1 CAG-Flex-eGFP injected into the motor cortex of C57/BL6 mice | Mature oligodendrocytes in proteolipid protein (PLP)::ChR2-EYFP mice; Vasoactive intestinal peptide (VIP)-expressing interneurons and long projecting neurons in VIP: tdTomato mice; Long-range axonal projections addressed by an adeno-associated virus (AAV) serotype 2-CMV-tdTomato vector injected into the anterior cingulate cortex | |
| Embedded in | Resin | Gelatin | Agarose gel | |
| Clearing | / | CUBIC-1 | / | |
| Sample Preparation Time | 5 days | 9–13 days | 8 days | |
| Imaging | Tomography | Structured Illumination Microscopy | Two-Photon Tomography | Spinning Disk-Based Confocal Microscopy |
| Voxel Resolution/μm3 | 0.32 × 0.32 × 2.0 | 0.3 × 0.3 × 1.0 | 0.7 × 0.7 × 5.0 | |
| Acquisition Time | 3.2 days | 1 week | 2.4 h | |
| Data Size (num. of coronal sections) | 10.9 TB (4834 × 2) | 30 TB | 1 TB (2400) | |
| Demonstration | Positioning Info | Yes | No | No |
| Knowledge | Co-localization of axonal long-range projections with cellular landmarks and complete 3D morphology of barrel cortex layer V/VI pyramidal neurons | Axonal morphology (length range: 23.6–121.2 mm) of five neurons innervated 28 distinct brain regions in the motor cortex | Whole-brain cytoarchitecture, vasculature, mature oligodendrocytes, interneurons, and long projecting neurons | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, H.; Guan, Y.; Chen, J.; Luo, Q. Optical Imaging in Brainsmatics. Photonics 2019, 6, 98. https://doi.org/10.3390/photonics6030098
Shi H, Guan Y, Chen J, Luo Q. Optical Imaging in Brainsmatics. Photonics. 2019; 6(3):98. https://doi.org/10.3390/photonics6030098
Chicago/Turabian StyleShi, Hua, Yue Guan, Jianwei Chen, and Qingming Luo. 2019. "Optical Imaging in Brainsmatics" Photonics 6, no. 3: 98. https://doi.org/10.3390/photonics6030098
APA StyleShi, H., Guan, Y., Chen, J., & Luo, Q. (2019). Optical Imaging in Brainsmatics. Photonics, 6(3), 98. https://doi.org/10.3390/photonics6030098





