Sampling Dynamics for Volatile Organic Compounds Using Headspace Solid-Phase Microextraction Arrow for Microbiological Samples
Abstract
1. Introduction
2. Materials and Methods
2.1. Volatiles Reference Mix (VRM)
2.2. Gas Chromatography–Mass Spectrometry (GC–MS) Analysis and Data Processing
2.3. Optimization of SPME Arrow Method
2.4. Application to Bacillus subtilis Headspace
2.5. Statistical Analysis
3. Results and Discussion
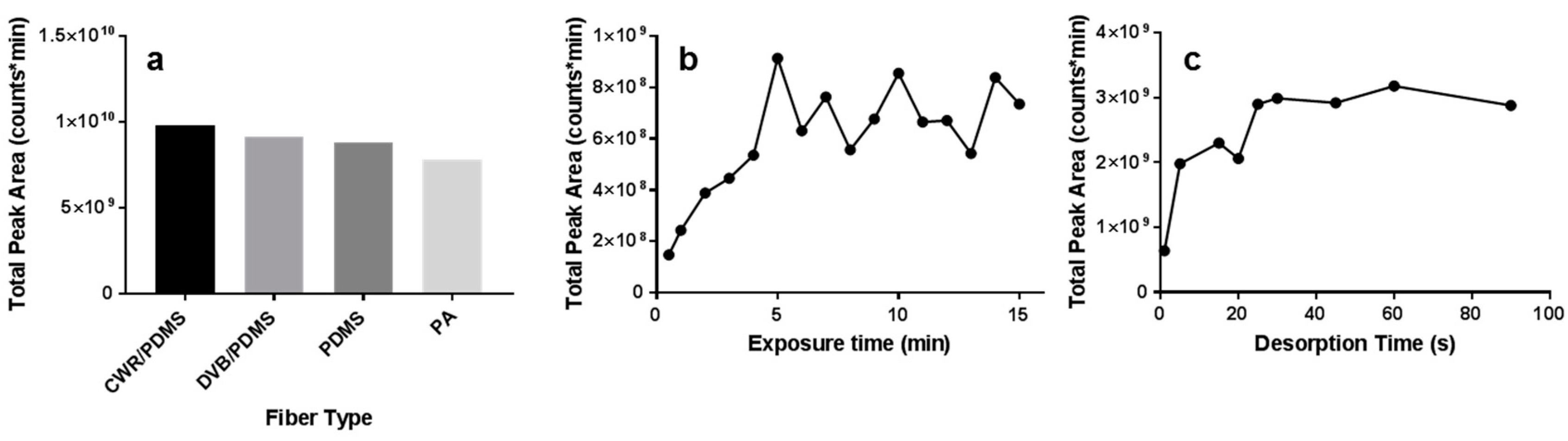
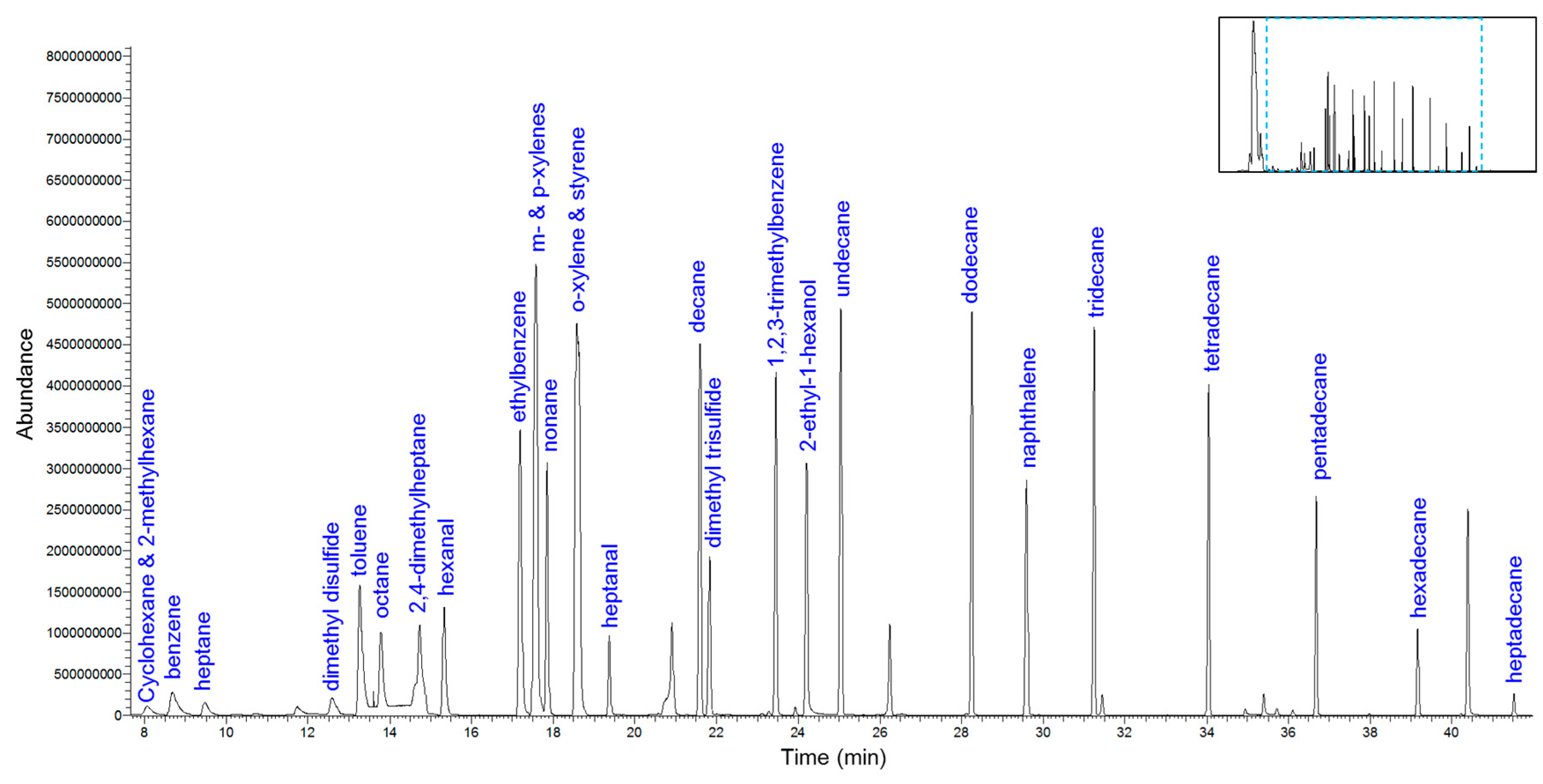
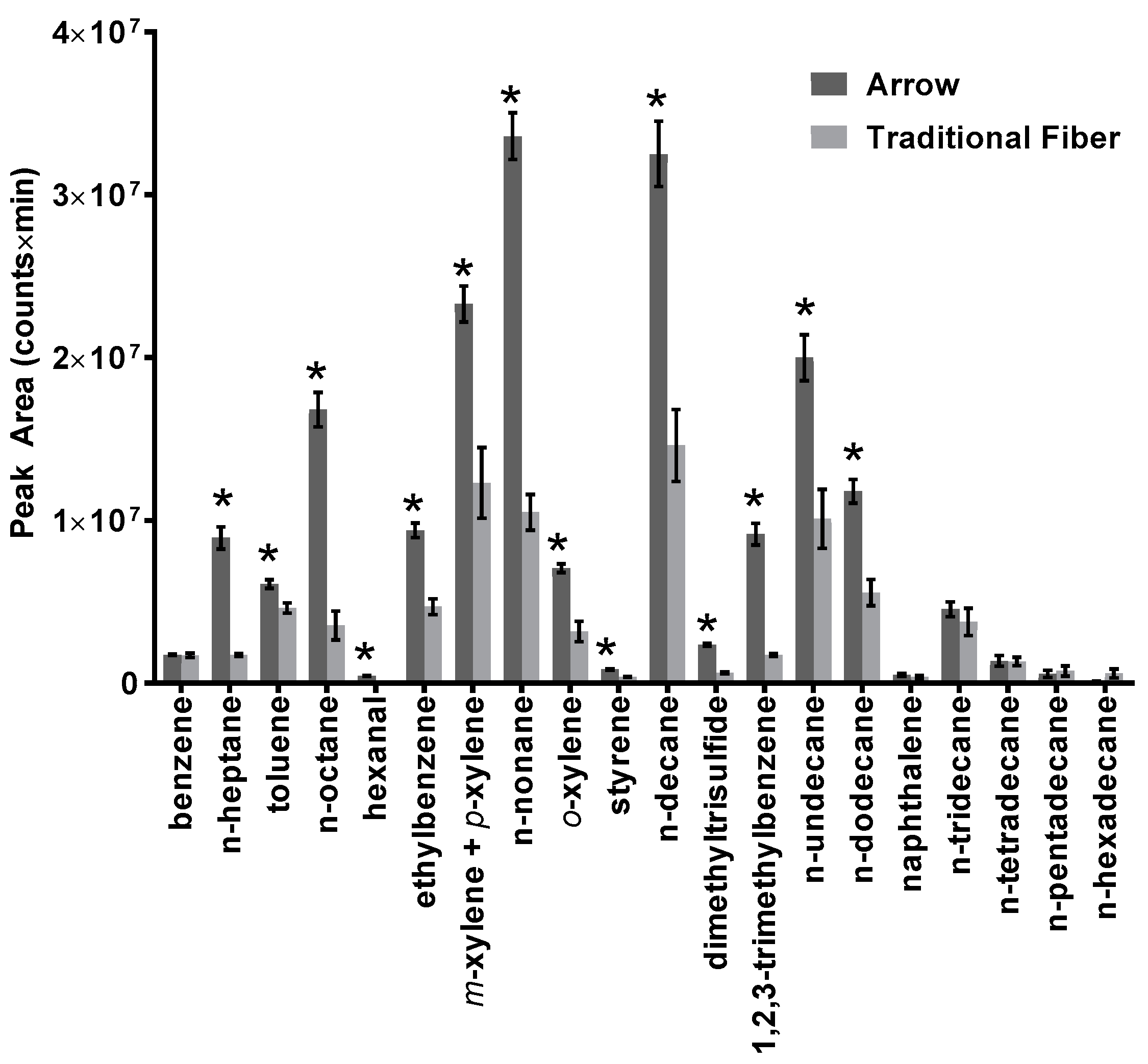
3.1. Arrow Method Optimization
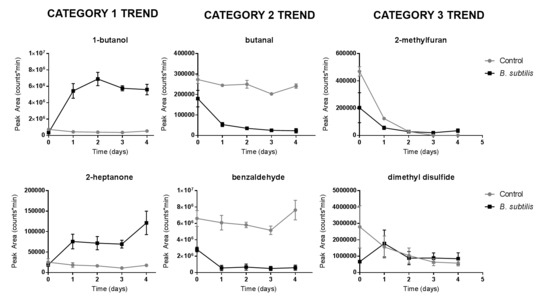
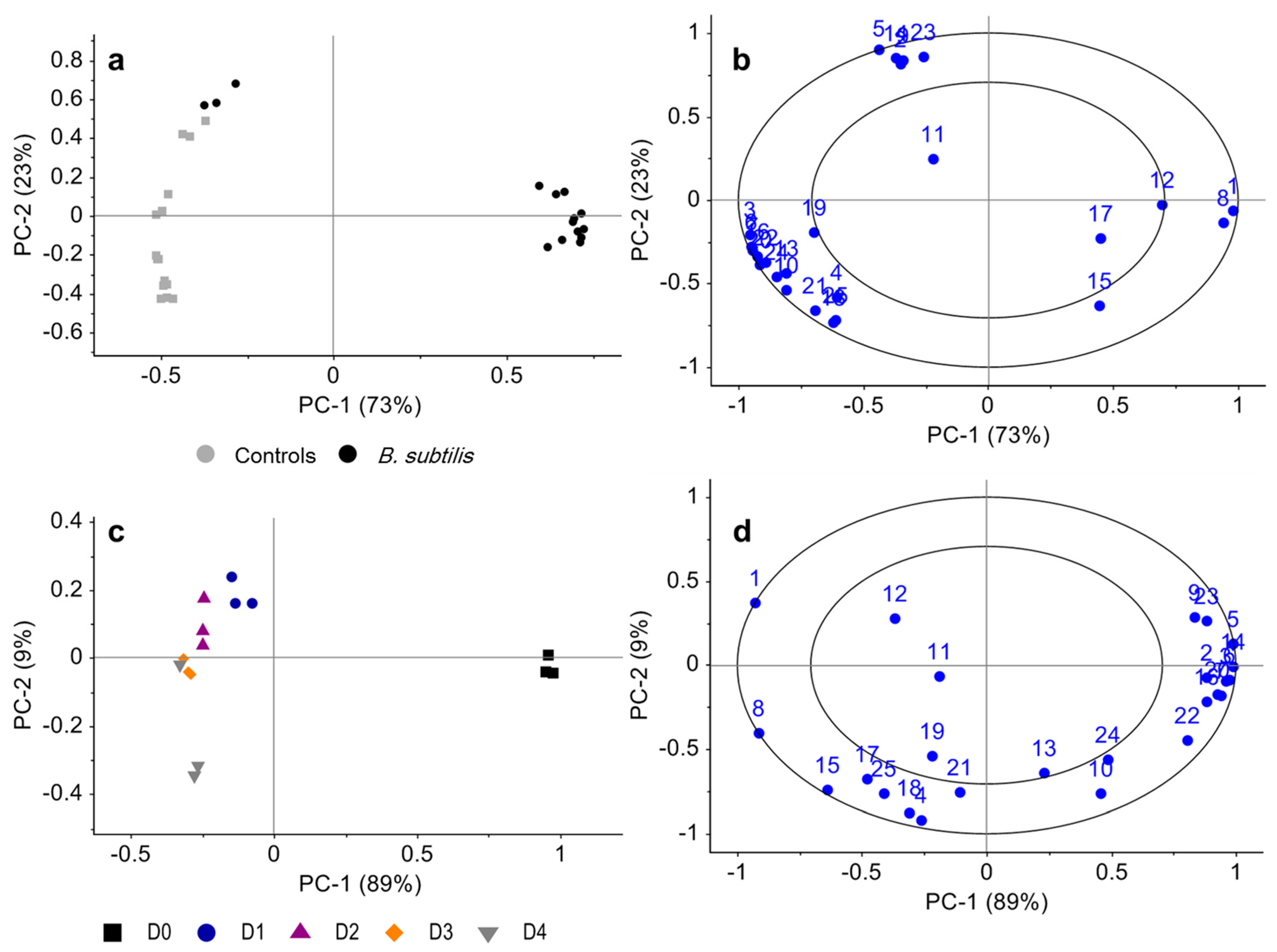
3.2. Bacillus subtilis Profiling
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tranchida, P.Q.; Dugo, P.; Dugo, G.; Mondello, L. Comprehensive two-dimensional chromatography in food analysis. J. Chromatogr. A 2004, 1054, 3–16. [Google Scholar] [CrossRef]
- Tranchida, P.Q.; Donato, P.; Cacciola, F.; Beccaria, M.; Dugo, P.; Mondello, L. Potential of comprehensive chromatography in food analysis. Trends Anal. Chem. 2013, 52, 186–205. [Google Scholar] [CrossRef]
- Serrano, A.; Gallego, M. Sorption study of 25 volatile organic compounds in several Mediterranean soils using headspace-gas chromatography–mass spectrometry. J. Chromatogr. A 2006, 1118, 261–270. [Google Scholar] [CrossRef] [PubMed]
- Hewitt, A.D. Comparison of sample preparation methods for the analysis of volatile organic compounds in soil samples: Solvent extraction vs. vapor partitioning. Environ. Sci. Technol. 1998, 32, 143–149. [Google Scholar] [CrossRef]
- Kurán, P.; Sojak, L. Environmental analysis of volatile organic compounds in water and sediment by gas chromatography. J. Chromatogr. A 1996, 733, 119–141. [Google Scholar] [CrossRef]
- Voice, T.C.; Kolb, B. Static and dynamic headspace analysis of volatile organic compounds in soils. Environ. Sci. Technol. 1993, 27, 709–713. [Google Scholar] [CrossRef]
- Kleeberg, K.K.; Liu, Y.; Jans, M.; Schlegelmilch, M.; Streese, J.; Stegmann, R. Development of a simple and sensitive method for the characterization of odorous waste gas emissions by means of solid-phase microextraction (SPME) and GC–MS/olfactometry. Waste Manag. 2005, 25, 872–879. [Google Scholar] [CrossRef] [PubMed]
- Tholl, D.; Boland, W.; Hansel, A.; Loreto, F.; Röse, U.S.R.; Schnitzler, J.-P. Practical approaches to plant volatile analysis. Plant J. 2006, 45, 540–560. [Google Scholar] [CrossRef] [PubMed]
- Heath, R.R.; Manukian, A. An automated system for use in collecting volatile chemicals released from plants. J. Chem. Ecol. 1994, 20, 593–608. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, M.A.; Nizio, K.D.; Ueland, M.; Forbes, S.L. Forensic decomposition odour profiling: A review of experimental designs and analytical techniques. Trends Anal. Chem. 2017, 91, 112–124. [Google Scholar] [CrossRef]
- Verheggen, F.; Perrault, K.A.; Caparros Megido, R.; Dubois, L.M.; Francis, F.; Haubruge, E.; Forbes, S.L.; Focant, J.-F.; Stefanuto, P.-H. The odour of death: An overview of current knowledge on characterization and applications. Bioscience 2017, 67, 600–613. [Google Scholar] [CrossRef]
- Cao, W.; Duan, Y. Breath analysis: Potential for clinical diagnosis and exposure assessment. Clin. Chem. 2006, 52, 800–811. [Google Scholar] [CrossRef] [PubMed]
- Bijland, L.R.; Bomers, M.K.; Smulders, Y.M. Smelling the diagnosis A review on the use of scent in diagnosing disease. Neth. J. Med. 2013, 71, 300–307. [Google Scholar] [PubMed]
- Hamm, S.; Bleton, J.; Connan, J.; Tchapla, A. A chemical investigation by headspace SPME and GC–MS of volatile and semi-volatile terpenes in various olibanum samples. Phytochemistry 2005, 66, 1499–1514. [Google Scholar] [CrossRef] [PubMed]
- Cnuts, D.; Perrault, K.A.; Stefanuto, P.-H.; Dubois, L.M.; Focant, J.-F.; Rots, V. Fingerprinting glues using HS-SPME GC×GC-HRTOFMS: A new powerful method allows tracking glues back in time. Archaeometry 2018, 1–16. [Google Scholar] [CrossRef]
- Perrault, K.A.; Stefanuto, P.; Dubois, L.; Cnuts, D.; Rots, V.; Focant, J.-F. A new approach for the characterization of organic residues from stone tools using GC×GC-TOFMS. Separations 2016, 3, 1–16. [Google Scholar] [CrossRef]
- Jerković, I.; Marijanović, Z.; Gugić, M.; Roje, M. Chemical profile of the organic residue from ancient amphora found in the Adriatic Sea determined by direct GC and GC–MS analysis. Molecules 2011, 16, 7936–7948. [Google Scholar] [CrossRef] [PubMed]
- Kremser, A.; Jochmann, M.A.; Schmidt, T.C. Systematic comparison of static and dynamic headspace sampling techniques for gas chromatography. Anal. Bioanal. Chem. 2016, 408, 6567–6579. [Google Scholar] [CrossRef] [PubMed]
- Harper, M. Sorbent trapping of volatile organic compounds from air. J. Chromatogr. A 2000, 885, 129–151. [Google Scholar] [CrossRef]
- Arthur, C.L.; Pawliszyn, J. Solid phase microextraction with thermal desorption using fused silica optical fibers. Anal. Chem. 1990, 62, 2145–2148. [Google Scholar] [CrossRef]
- Zhang, Z.; Pawliszyn, J. Headspace solid-phase microextraction. Anal. Chem. 1993, 65, 1843–1852. [Google Scholar] [CrossRef]
- Pawliszyn, J. Theory of solid-phase microextraction. J. Chromatogr. Sci. 2012, 38, 13–59. [Google Scholar] [CrossRef]
- Bagheri, H.; Piri-Moghadam, H.; Naderi, M. Towards greater mechanical, thermal and chemical stability in solid-phase microextraction. Trends Anal. Chem. 2012, 34, 126–139. [Google Scholar] [CrossRef]
- Spietelun, A.; Kloskowski, A.; Chrzanowski, W.; Namieśnik, J. Understanding solid-phase microextraction: Key factors influencing the extraction process and trends in improving the technique. Chem. Rev. 2013, 113, 1667–1685. [Google Scholar] [CrossRef] [PubMed]
- Baltussen, E.; Sandra, P.; David, F.; Cramers, C. Stir bar sorptive extraction (SBSE), a novel extraction technique for aqueous samples: Theory and principles. J. Microcolumn Sep. 1999, 11, 737–747. [Google Scholar] [CrossRef]
- Tienpont, B.; David, F.; Bicchi, C.; Sandra, P. High capacity headspace sorptive extraction. J. Microcolumn Sep. 2000, 12, 577–584. [Google Scholar] [CrossRef]
- Helin, A.; Rönkkö, T.; Parshintsev, J.; Hartonen, K.; Schilling, B.; Läubli, T.; Riekkola, M.L. Solid phase microextraction arrow for the sampling of volatile amines in wastewater and atmosphere. J. Chromatogr. A 2015, 1426, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Kremser, A.; Jochmann, M.A.; Schmidt, T.C. PAL SPME Arrow—Evaluation of a novel solid-phase microextraction device for freely dissolved PAHs in water. Anal. Bioanal. Chem. 2016, 408, 943–952. [Google Scholar] [CrossRef] [PubMed]
- Forbes, S.L.; Troobnikoff, A.N.; Ueland, M.; Nizio, K.D.; Perrault, K.A. Profiling the decomposition odour at the grave surface before and after probing. Forensic Sci. Int. 2016, 259, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, P.; Nizio, K.D.; Perrault, K.A.; Forbes, S.L. Establishing the volatile profile of pig carcasses as analogues for human decomposition during the early postmortem period. Heliyon 2016, 2, e00070. [Google Scholar] [CrossRef] [PubMed]
- Perrault, K.A.; Stefanuto, P.-H.; Stuart, B.H.; Rai, T.; Focant, J.-F.; Forbes, S.L. Detection of decomposition volatile organic compounds in soil following removal of remains from a surface deposition site. Forensic Sci. Med. Pathol. 2015, 11, 376–387. [Google Scholar] [CrossRef] [PubMed]
- Carter, D.O.; Tomberlin, J.K.; Benbow, M.E.; Metcalf, J.L. (Eds.) Forensic Microbiology, 1st ed.; John Wiley & Sons Ltd.: Chichester, UK, 2017; ISBN 978-1-119-06255-4. [Google Scholar]
- Bean, H.D.; Dimandja, J.-M.D.; Hill, J.E. Bacterial volatile discovery using solid phase microextraction and comprehensive two-dimensional gas chromatography–time-of-flight mass spectrometry. J. Chromatogr. B 2012, 901, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Bean, H.D.; Rees, C.A.; Hill, J.E. Comparative analysis of the volatile metabolomes of Pseudomonas aeruginosa clinical isolates. J. Breath Res. 2016, 10, 47102. [Google Scholar] [CrossRef] [PubMed]
- Nizio, K.D.; Perrault, K.A.; Troobnikoff, A.N.; Ueland, M.; Shoma, S.; Iredell, J.R.; Middleton, P.G.; Forbes, S.L. In vitro volatile organic compound profiling using GC×GC-TOFMS to differentiate bacteria associated with lung infections: A proof-of-concept study. J. Breath Res. 2016, 10, 26008. [Google Scholar] [CrossRef] [PubMed]
- Filipiak, W.; Sponring, A.; Filipiak, A.; Ager, C.; Schubert, J.; Miekisch, W.; Amann, A.; Troppmair, J. TD-GC–MS analysis of volatile metabolites of human lung cancer and normal cells in vitro. Cancer Epidemiol. Biomark. Prev. 2010, 19, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Perrault, K.A.; Nizio, K.D.; Forbes, S.L. A comparison of one-dimensional and comprehensive two-dimensional gas chromatography for decomposition odour profiling using inter-year replicate field trials. Chromatographia 2015, 78, 1057–1070. [Google Scholar] [CrossRef]
- Logan, N.A.; De Vos, P. Genus I: Bacillus. In Bergey’s Manual of Systematic Bacteriology; De Vos, P., Garrity, G.M., Jones, D., Krieg, N.R., Ludwig, W., Rainey, F.A., Schleifer, K.-H., Whitman, W.B., Eds.; Springer: New York, NY, USA, 2019; pp. 21–128. [Google Scholar]
- Dyer, B.D. A Field Guide to Bacteria; Cornell University Press: Ithaca, NY, USA, 2003. [Google Scholar]
- Chun, L.P.; Miguel, M.J.; Junkins, E.N.; Forbes, S.L.; Carter, D.O. An initial investigation into the ecology of culturable aerobic postmortem bacteria. Sci. Justice 2015, 55, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Carter, D.O.; Yellowlees, D.; Tibbett, M. Cadaver decomposition in terrestrial ecosystems. Naturwissenschaften 2007, 94, 12–24. [Google Scholar] [CrossRef] [PubMed]
- Logan, N.A. Bacillus species of medical and veterinary importance. J. Med. Microbiol. 1988, 25, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Barreira, L.M.F.; Parshintsev, J.; Kärkkäinen, N.; Hartonen, K.; Jussila, M.; Kajos, M.; Kulmala, M.; Riekkola, M.L. Field measurements of biogenic volatile organic compounds in the atmosphere by dynamic solid-phase microextraction and portable gas chromatography-mass spectrometry. Atmos. Environ. 2015, 115, 214–222. [Google Scholar] [CrossRef]
- Smith, D.; Španěl, P. On the importance of accurate quantification of individual volatile metabolites in exhaled breath. J. Breath Res. 2017, 11, 4. [Google Scholar] [CrossRef] [PubMed]
- Stefanuto, P.-H.; Perrault, K.A.; Lloyd, R.M.; Stuart, B.H.; Rai, T.; Forbes, S.L.; Focant, J.-F. Exploring new dimensions in cadaveric decomposition odour analysis. Anal. Methods 2015, 7, 2287–2294. [Google Scholar] [CrossRef]
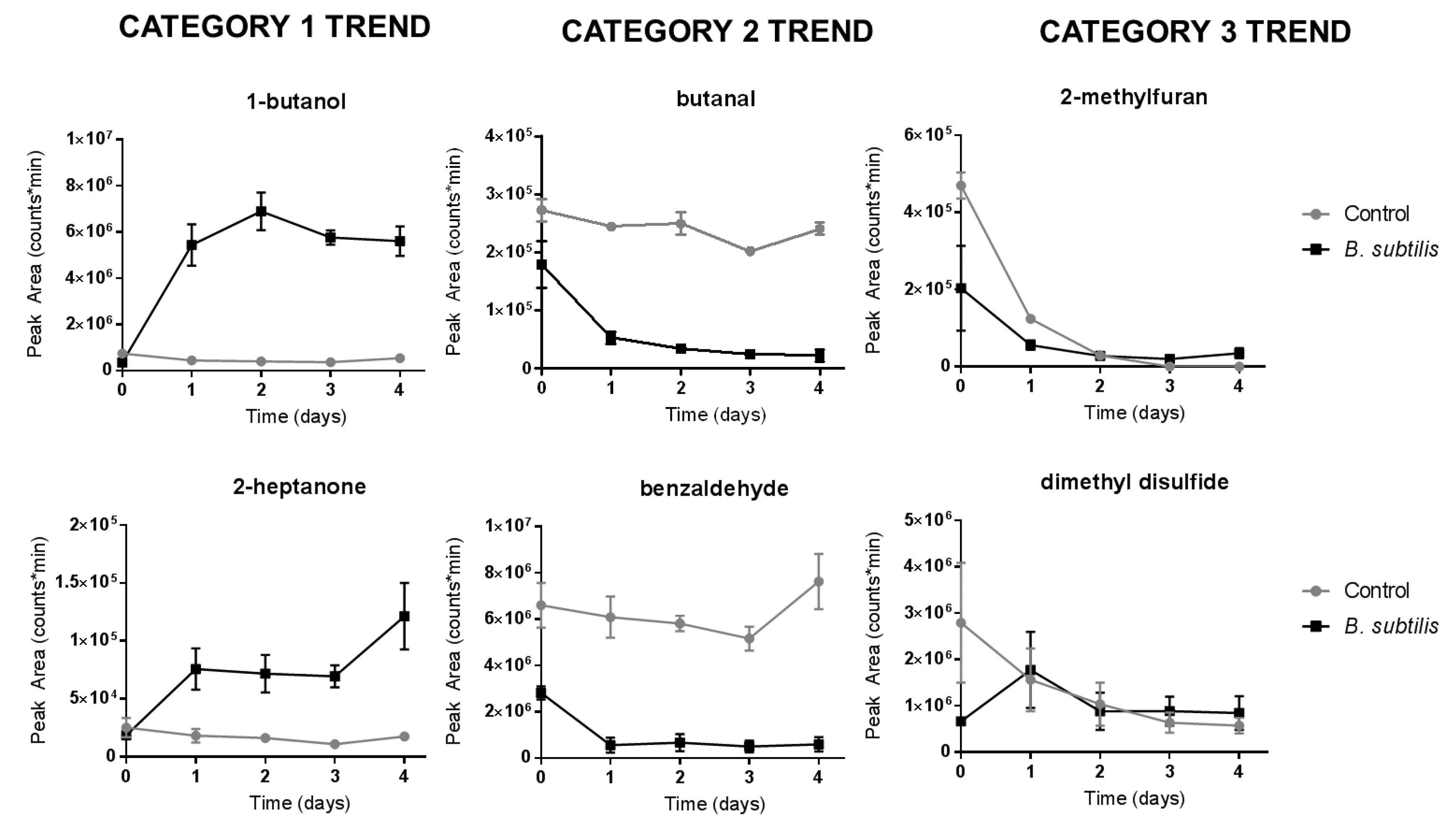
| Label | Compound | CAS | Retention Time (min) | Retention Index | Trend Category |
|---|---|---|---|---|---|
| 1 | 1,3-pentadiene | 504-60-9 | 3.282 ± 0.015 (n = 6) | 591.2 ± 0.2 (n = 6) | 1 * |
| 2 | 2-methylfuran | 534-22-5 | 6.049 ± 0.017 (n = 9) | 639.5 ± 0.3 (n = 9) | 3 |
| 3 | butanal | 123-72-8 | 6.452 ± 0.016 (n = 12) | 647.0 ± 0.3 (n = 12) | 2 * |
| 4 | 2-butanone | 78-93-3 | 6.774 ± 0.027 (n = 14) | 653.0 ± 0.5 (n = 14) | 2 * |
| 5 | trichloromethane | 67-66-3 | 7.495 ± 0.008 (n = 27) | 666.7 ± 0.2 (n = 27) | 3 |
| 6 | 3-methylbutanal | 590-86-3 | 8.986 ± 0.005 (n = 18) | 696.3 ± 0.1 (n = 18) | 2 * |
| 7 | 2-methylbutanal | 96-17-3 | 9.298 ± 0.008 (n = 18) | 702.6 ± 0.2 (n = 18) | 2 * |
| 8 | 1-butanol | 71-36-3 | 10.052 ± 0.008 (n = 12) | 718.2 ± 0.2 (n = 12) | 1 * |
| 9 | bromodichloromethane | 75-27-4 | 11.294 ± 0.009 (n = 27) | 744.8 ± 0.2 (n = 27) | 3 |
| 10 | 1,3-diazine | 289-95-2 | 12.157 ± 0.007 (n = 27) | 763.8 ± 0.2 (n = 27) | 2 * |
| 11 | dimethyl disulfide | 624-92-0 | 12.387 ± 0.005 (n = 30) | 769.0 ± 0.1 (n = 30) | 3 |
| 12 | 1-methylcyclohexene | 591-49-1 | 12.632 ± 0.002 (n = 7) | 774.5 ± 0.1 (n = 7) | 1 * |
| 13 | 2-methylbutanenitrile | 18936-17-9 | 12.794 ± 0.004 (n = 10) | 778.2 ± 0.1 (n = 10) | 2 * |
| 14 | toluene | 108-88-3 | 13.105 ± 0.006 (n = 29) | 785.3 ± 0.1 (n = 29) | 3 |
| 15 | 3-methyl-2-pentanone | 209.282-1 | 13.277 ± 0.005 (n = 8) | 789.2 ± 0.1 (n = 8) | 1 |
| 16 | furfural | 98-01-1 | 17.408 ± 0.005 (n = 17) | 890.1 ± 0.1 (n = 17) | 2 * |
| 17 | 2-heptanone | 110-43-0 | 18.913 (n = 1) | 929.5 (n = 1) | 1 * |
| 18 | 2,5-dimethylpyrazine | 123-32-0 | 19.281 ± 0.003 (n = 30) | 939.4 ± 0.1 (n = 30) | 2 * |
| 19 | methoxy-phenyl-oxime | 21.691 ± 0.006 (n = 6) | 1006.3 ± 0.2 (n = 6) | 2 * | |
| 20 | benzaldehyde | 100-52-7 | 22.087 ± 0.003 (n = 18) | 1017.6 ± 0.1 (n = 18) | 2 * |
| 21 | 2-ethyl-5-methylpyrazine | 13360-64-0 | 22.535 ± 0.004 (n = 25) | 1030.6 ± 0.1 (n = 25) | 2 * |
| 22 | limonene | 22.969 ± 0.004 (n = 15) | 1043.3 ± 0.1 (n = 15) | 2 * | |
| 23 | dodecane | 112-40-3 | 28.078 ± 0.003 (n = 29) | 1201.7 ± 0.1 (n = 29) | 3 |
| 24 | decanal | 112-31-2 | 29.572 ± 0.003 (n = 15) | 1251.2 ± 0.1 (n = 15) | 2 * |
| 25 | butylated hydroxytoluene | 128-37-0 | 37.804 ± 0.004 (n = 12) | 1550.2 ± 0.2 (n = 12) | 2 * |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eckert, K.E.; Carter, D.O.; Perrault, K.A. Sampling Dynamics for Volatile Organic Compounds Using Headspace Solid-Phase Microextraction Arrow for Microbiological Samples. Separations 2018, 5, 45. https://doi.org/10.3390/separations5030045
Eckert KE, Carter DO, Perrault KA. Sampling Dynamics for Volatile Organic Compounds Using Headspace Solid-Phase Microextraction Arrow for Microbiological Samples. Separations. 2018; 5(3):45. https://doi.org/10.3390/separations5030045
Chicago/Turabian StyleEckert, Kevin E., David O. Carter, and Katelynn A. Perrault. 2018. "Sampling Dynamics for Volatile Organic Compounds Using Headspace Solid-Phase Microextraction Arrow for Microbiological Samples" Separations 5, no. 3: 45. https://doi.org/10.3390/separations5030045
APA StyleEckert, K. E., Carter, D. O., & Perrault, K. A. (2018). Sampling Dynamics for Volatile Organic Compounds Using Headspace Solid-Phase Microextraction Arrow for Microbiological Samples. Separations, 5(3), 45. https://doi.org/10.3390/separations5030045