Evaluation of Rate of Adhesion of Lactobacillus namurensis Strain GYP-74 to Porous Fine Ceramics
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation of LAB
2.2. DNA Extraction of Strain GYP-74 and Amplification of 16S rRNA Gene by Polymerase Chain Reaction (PCR)
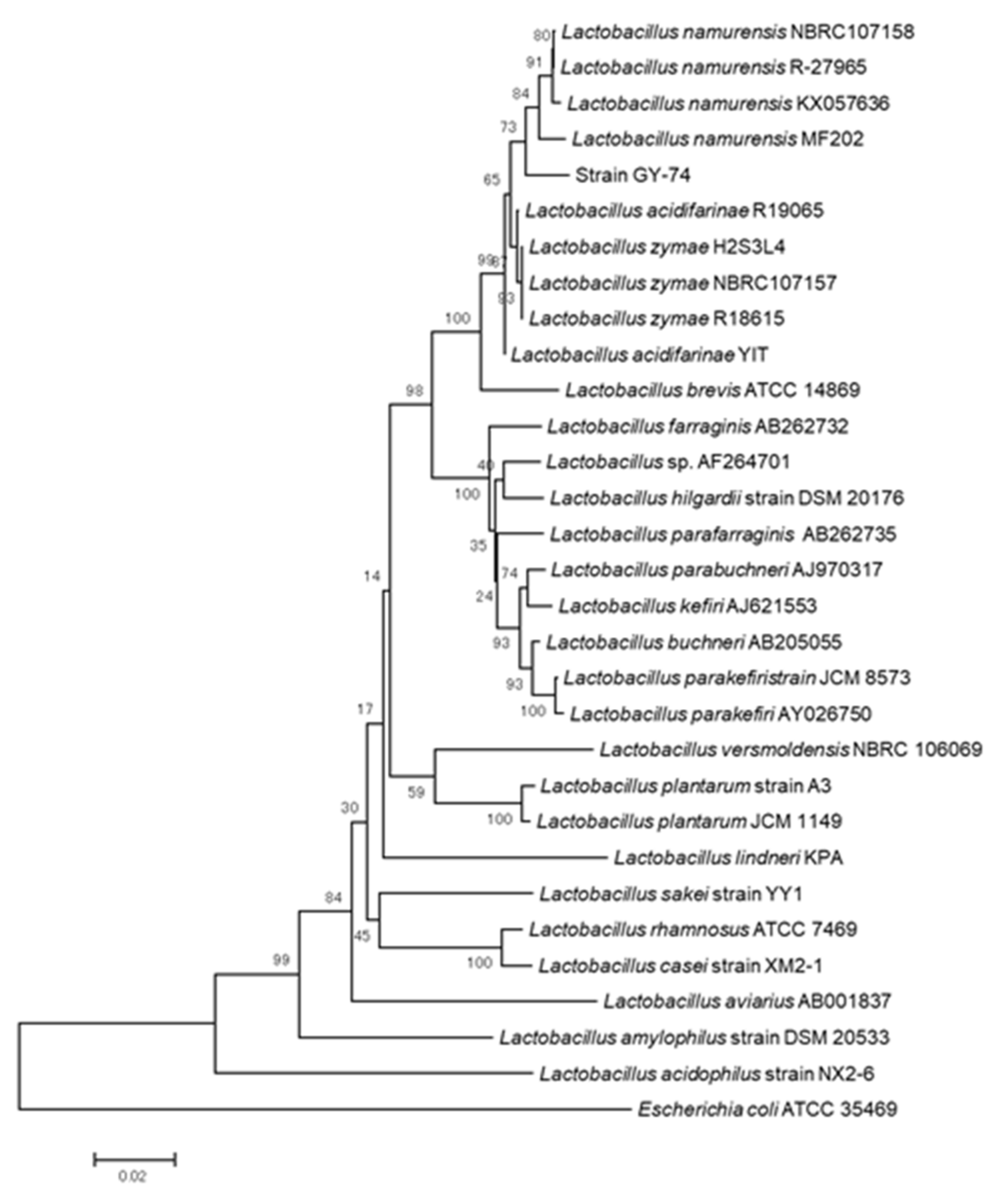
2.3. Basic Local Alignment Search Tool (BLAST) Analysis and Construction of Phylogenetic Tree
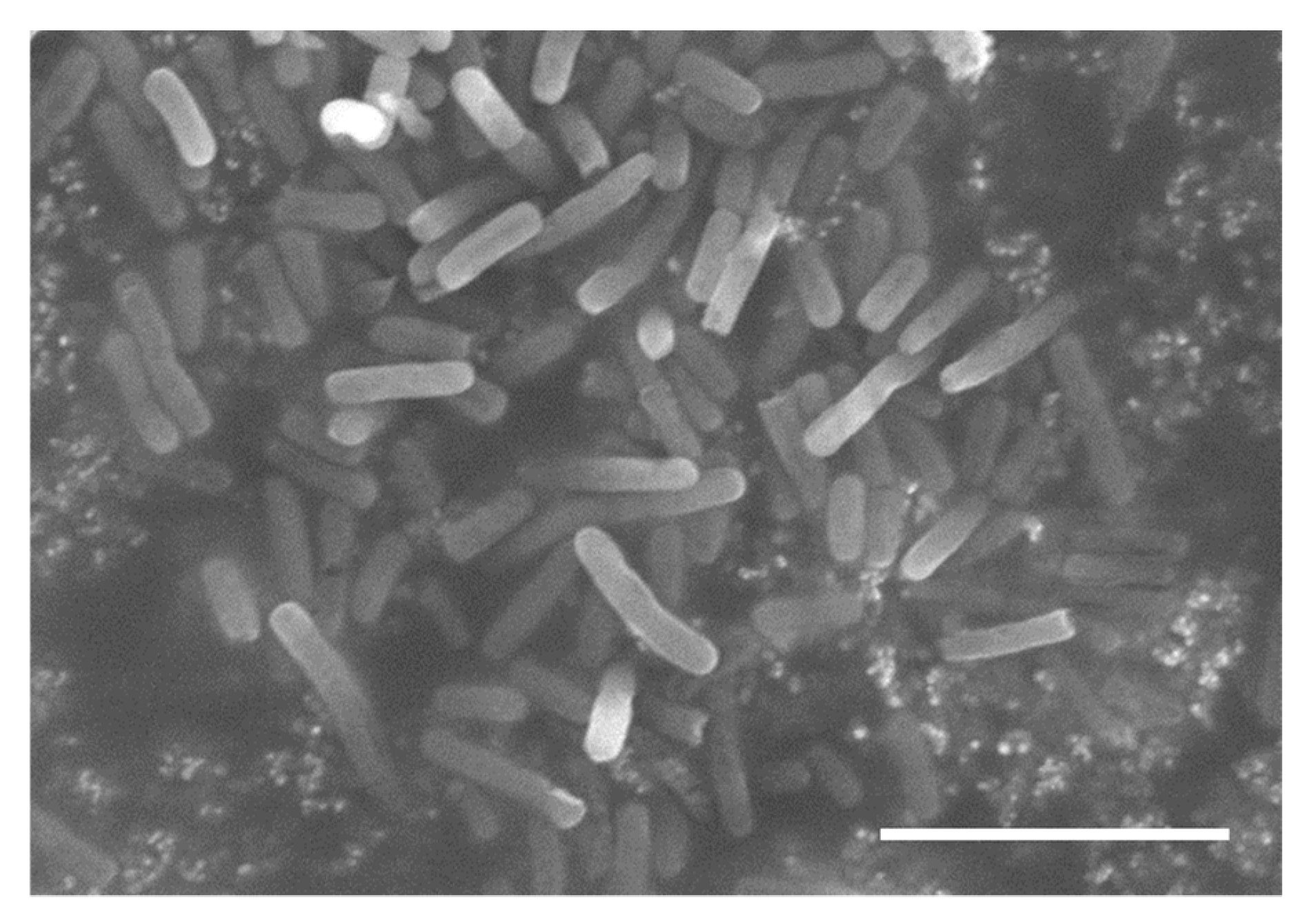
2.4. Adherent Test of Strain GYP-74 to PS and Morphological Observation with a Scanning Electron Microscope (SEM)
2.5. Analysis of the Pore Distribution Using a Mercury Porosimeter
2.6. X-ray Diffraction Analysis of the PS
2.7. Statistical Analysis
3. Results and Discussion
3.1. Isolation of LAB and Identification Based on Morphological Observation, BLAST Research and Phylogenetic Analysis
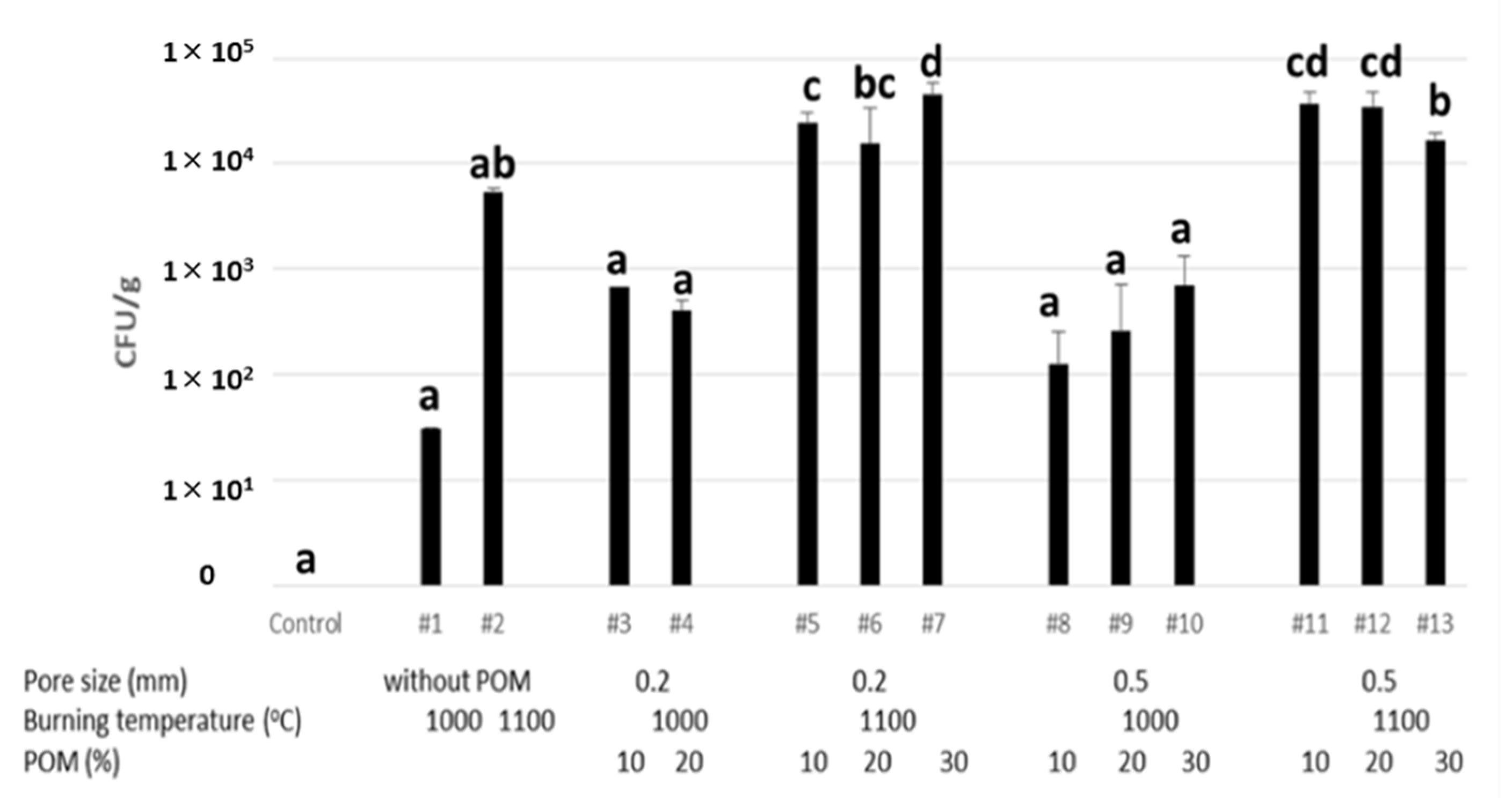
3.2. Adherent Test of Strain GYP-74 to PS
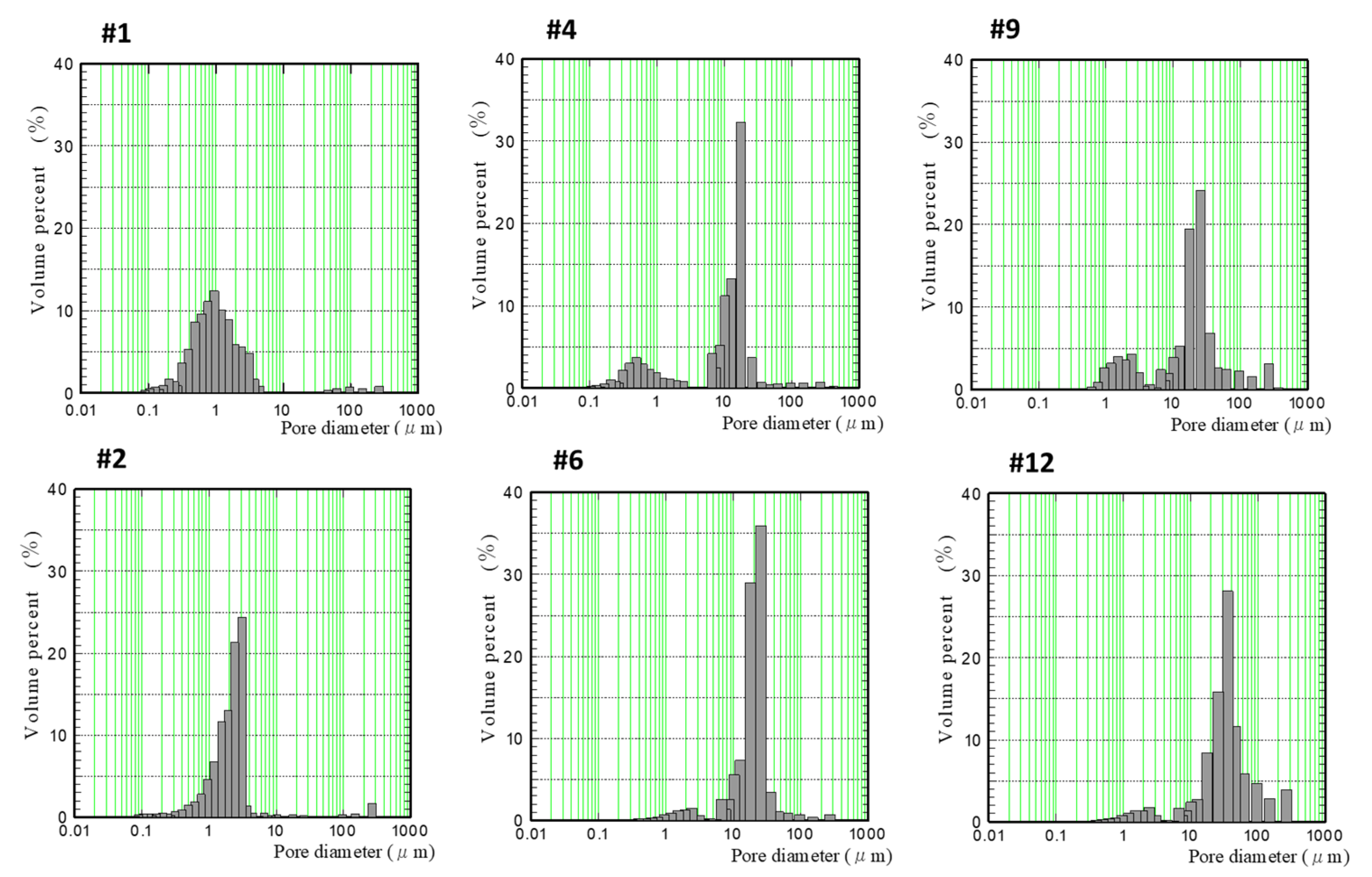
3.3. Pore Distribution Analysis
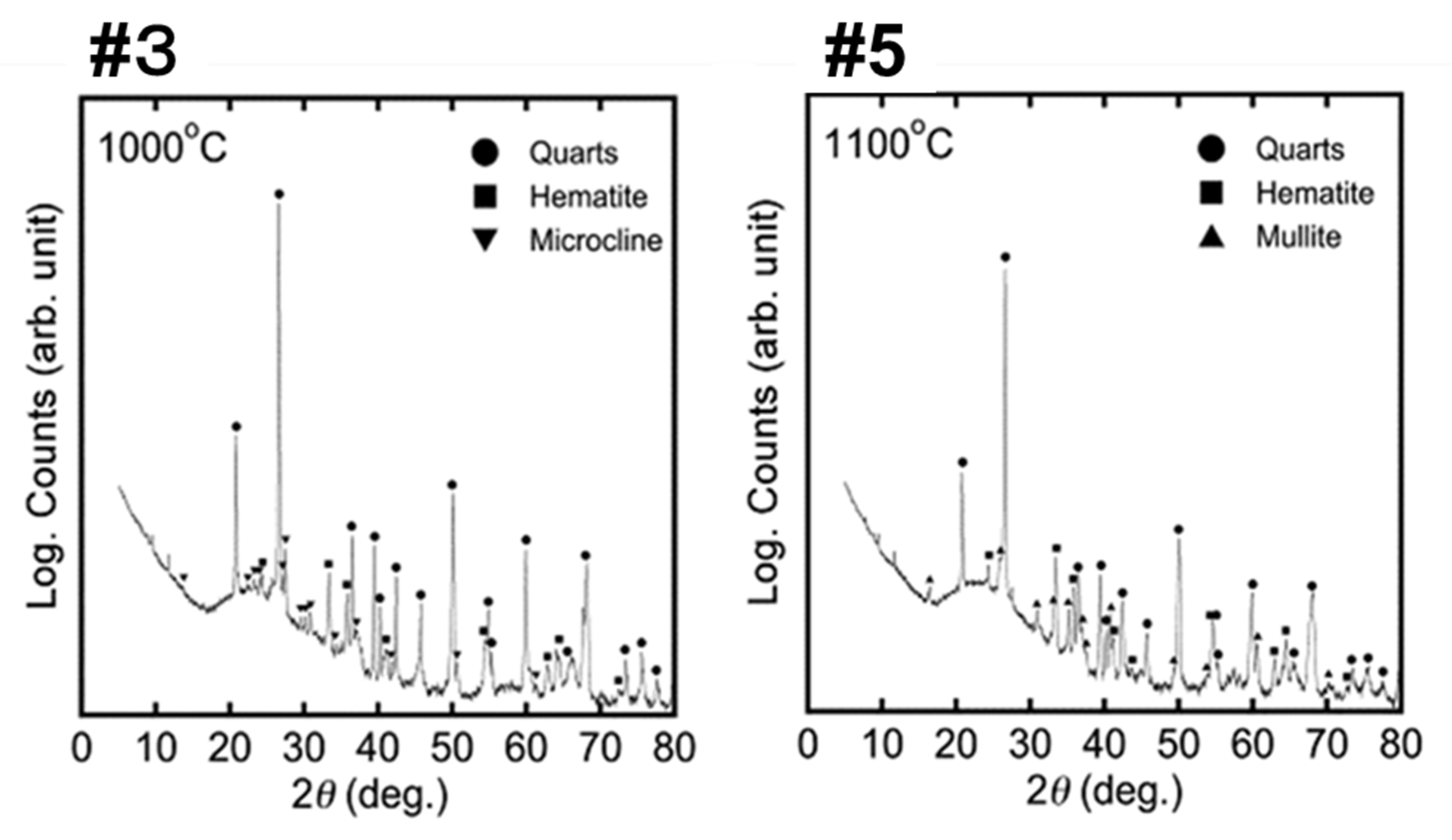
3.4. X-ray Diffraction Analysis
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sakamoto, N.; Tanaka, S.; Sonomoto, K.; Nakayama, J. 16S rRNA pyrosequencing-based investigation of the bacterial community in nukadoko, a pickling bed of fermented rice bran. Int. J. Food Microbiol. 2011, 144, 352–359. [Google Scholar] [CrossRef]
- Nakayama, J.; Hoshiko, H.; Fukuda, M.; Tanaka, H.; Sakamoto, N.; Tanaka, S.; Ohue, K.; Sakai, K.; Sonomoto, K. Molecular monitoring of bacterial community structure in long-aged nukadoko: Pickling bed of fermented rice bran dominated by slow-growing lactobacilli. J. Biosci. Bioeng. 2007, 104, 481–489. [Google Scholar] [CrossRef]
- Guan, Q.; Xiong, T.; Xie, M. Influence of Probiotic Fermented Fruit and Vegetables on Human Health and the Related Industrial Development Trend. Engineering 2021, 7, 212–218. [Google Scholar] [CrossRef]
- Yang, Y.; Park, J.H.; Kim, I.H. Effects of probiotics containing (Lactobacillus planetarium) and chlortetracycline on growth performance, nutrient digestibility, fecal microflora, diarrhea score and fecal gas emission in weanling pigs. Livest. Sci. 2020, 241, 104186. [Google Scholar] [CrossRef]
- Kuebutornye, F.K.A.; Abarike, E.D.; Lu, Y. A review on the application of Bacillus as probiotics in aquaculture. Fish Shellfish Immunol. 2019, 87, 820–828. [Google Scholar] [CrossRef]
- Fuller, R. Probiotics in man and animals. J. Appl. Bacteriol. 1989, 66, 365–378. [Google Scholar] [PubMed]
- Salminen, S.; Isolauri, E.; Salminen, E. Clinical uses of probiotics for stabilizing the gut mucosal barrier: Successful strains and future challenge. Antonie Van Leeuuwenboek 1997, 70, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Shu, Q.; Qu, F.; Gill, H.S. Probiotic treatment using Bifidobacterium lactis HNO19 reduces weanling diarrhea associated lymphoid tissue in neonatal swine. Immunol. Methods 2001, 241, 185–199. [Google Scholar]
- Sakai, M. Current research status of fish immunostimulants. Aquaculture 1998, 72, 63–92. [Google Scholar] [CrossRef]
- Anukam, K.C.; Osazuwa, E.; Osemene, G.I.; Ehigiagbe, F.; Bruce, A.W.; Reid, G. Clinical study comparing probiotic Lactobacillus GR-1 and RC-14 with metronidazole vaginal gel to treat symptomatic bacterial vaginosis. Microbes Infect. 2006, 8, 1–5. [Google Scholar] [CrossRef]
- Gionchetti, P.; Rizzello, F.; Venturi, A.; Brigidi, P.; Matteuzzi, D.; Bazzocchi, G.; Poggioli, G.; Miglioli, M.; Campieri, M. Oral bacteriotherapy as maintenance treatment in patients with chronic pouchitis: A doubleblind, placebo-controlled trial. Gastroenterology 2000, 119, 305–309. [Google Scholar] [CrossRef] [PubMed]
- Saran, S.; Gopalan, S.; Krishna, T.P. Use of fermented foods to combat stunting andfailure to thrive. Nutrition 2002, 18, 393–396. [Google Scholar] [CrossRef]
- Sazawal, S.; Hiremath, G.; Dhingra, U.; Malik, P.; Deb, S.; Black, R.E. Efficacy of probiotics in prevention of acute diarrhoea: A meta-analysis of masked, randomised, placebo-controlled trials. Lancet Infect. Dis. 2006, 6, 374–382. [Google Scholar] [CrossRef]
- Sheu, B.S.; Cheng, H.C.; Kao, A.W.; Wang, S.T.; Yang, Y.J.; Yang, H.B.; Wu, J.J. Pretreatment with Lactobacillus- and Bifidobacterium-containing yogurt can improve the efficacy of quadruple therapy in eradicating residual Helicobacter pylori infection after failed triple therapy. Am. J. Clin. Nutr. 2006, 83, 864–869. [Google Scholar] [CrossRef] [PubMed]
- Komatsuzaki, N.; Tsukahara, K.; Toyoshima, H.; Suzuki, T.; Shimizu, N.; Kimura, T. Effect of soaking and gaseous treatment on GABA content in germinated brown rice. J. Food Eng. 2007, 78, 556–560. [Google Scholar] [CrossRef]
- Chung, H.J.; Jang, S.H.; Cho, H.Y.; Lim, S.T. Effects of steeping and anaerobic treatment on GABA (γ-aminobutyric acid) content in germinated waxy hull-less barley. LWT Food Sci. Technol. 2009, 42, 1712–1716. [Google Scholar] [CrossRef]
- Hori, K.; Ishikawa, M. Microbial Adhesion to Surfaces Mediated by Bacterial Nanofibers. J. Environ. Biotechnol. 2010, 10, 3–7. [Google Scholar]
- Hori, K.; Matsumoto, S. Bacterial adhesion: From mechanism to control. Biochem. Eng. J. 2010, 48, 424–434. [Google Scholar] [CrossRef]
- Chaban, B.; Hughes, H.V.; Beeby, M. The flagellum in bacterial pathogens: For motility and a whole lot more. Semin. Cell Dev. Biol. 2015, 46, 91–103. [Google Scholar] [CrossRef]
- Beren, C.; Ellison, C.K.; Ducret, A.; Brun, Y.V. Bacterial adhesion at the single-cell level. Nat. Rev. Microbiol. 2018, 16, 616–627. [Google Scholar] [CrossRef]
- Morisaki, H.; Nagai, S.; Ohshima, H.; Ikemoto, E.; Kogure, K. The effect of motility and cell-surface polymers on bacterial attachment. Microbiology 1999, 145, 2797–2802. [Google Scholar] [CrossRef] [PubMed]
- Kouidhi, B.; Zmantar, T.; Hentati, H.; Bakhrouf, A. Cell surface hydrophobicity, biofilm formation, adhesives properties and molecular detection of adhesins genes in Staphylococcus aureus associated to dental caries. Microb. Pathog. 2010, 49, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Poncin-Epaillard, F.; Herry, J.M.; Marmey, P.; Legeay, G.; Debarnot, D.; Bellon-Fontaine, M.N. Elaboration of highly hydrophobic polymeric surface—A potential strategy to reduce the adhesion of pathogenic bacteria? Mater. Sci. Eng. C 2013, 33, 1152–1161. [Google Scholar] [CrossRef]
- Hao, X.; Chen, S.; Qin, D.; Zhang, M.; Fan, J.; Wang, C.; Dong, M.; Zhang, J.; Cheng, F.; Guo, Z. Antifouling and antibacterial behaviors of capsaicin-based pH responsive smart coatings in marine environments. Mater. Sci. Eng. C 2020, 108, 110361. [Google Scholar] [CrossRef] [PubMed]
- Boyd, R.D.; Verran, J.; Jones, M.V.; Bhakoo, M. Use of the atomic force microscope to determine the effect of substratum surface topography on bacterial adhesion. Adv. Colloid Interface Sci. 2012, 179, 142–149. [Google Scholar] [CrossRef]
- Matsumoto, S.; Ohtaki, A.; Hori, K. Carbon fiber as an excellent support material for wastewater treatment biofilms. Environ. Sci. Technol. 2012, 46, 10175–10181. [Google Scholar] [CrossRef] [PubMed]
- Song, F.; Koo, H.; Ren, D. Effects of Material Properties on Bacterial Adhesion and Biofilm Formation. J. Dent. Res. 2015, 94, 1027–1035. [Google Scholar] [CrossRef]
- Vural, H.C.; Ozgun, D. An improving DNA isolation method for identification of anaerobic bacteria in human colostrum and faeces samples. J. Med. Genet. Genomics 2011, 3, 95–100. [Google Scholar]
- Schmitz, A.; Riesner, D. Purification of nucleic acids by selective precipitation with polyethylene glycol 6000. Anal. Biochem. 2006, 354, 311–313. [Google Scholar] [CrossRef]
- Saeedi, M.; Shahidi, F.; Mortazavi, S.A.; Milani, E.; Yazdi, F.T. Isolation and identification of lactic acid bacteria in winter salad (Local Pickle) during fermentation using 16S rRNA gene sequence analysis. J. Food. Saf. 2015, 35, 287–294. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Higgins, D.G. The CLUSTAL X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The Neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Oda, K.; Imanishi, T.; Yamane, Y.; Ueno, Y.; Mori, Y. Bio-functional pickles that reduce blood pressure of rats. Biosci. Ciotechnol. Biochem. 2014, 78, 882–890. [Google Scholar] [CrossRef]
- Okamoto, A.; Matsumoto, E.; Yasuhara, T.; Koizumi, Y.; Yanagida, F. Angiotensin I-converting enzyme inhibitory action of fish sauce. Food Sci. Technol. Int. Tokyo 1995, 1, 101–106. [Google Scholar] [CrossRef][Green Version]
- Yokoyama, K.; Chiba, H.; Yoshikawa, M. Peptide inhibitors for angiotensin I-converting enzyme from thermolysin digest of dried bonito. Biosci. Biotechnol. Biochem. 1992, 56, 1541–1545. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Sakami, T.; Awai, K.; Shirakata, A. ZnO-based transparent conducting oxide films by reactive plasma deposition. Convertech 2004, 375, 68. [Google Scholar]
- Pereira, M.A.; Azeredo, J.; Mota, M.; Oliveira, R. Influence of physico-chemical properties of porous microcarriers on the adhesion of an anaerobic consortium. J. Ind. Microbiol. Biotechnol. 2000, 24, 181–186. [Google Scholar] [CrossRef][Green Version]
- Rajab, S.; Tabandeh, F.; Shahraky, M.K. The effect of lactobacillus cell size on its probiotic characteristics. Anaerobe 2020, 62, 102103. [Google Scholar] [CrossRef] [PubMed]
- Kitaoka, S.; Wada, T.; Osa, N.; Inagaki, H. Improvement of steam oxidation resistance and flame impingement resistance of glass having carbon coating film with controlled interfaces. In Prospect of Surface Treatment and Modification Technology in Inorganic Materials -Metal, Ceramics, and Glass-; Kamijo, E., Suzuki, Y., Fujisawa, A., Eds.; CMC Publisher: Tokyo, Japan, 2015; pp. 251–291. [Google Scholar]
- Uchida, H.; Fujitani, K.; Kawai, Y.; Kitazawa, H.; Horii, A.; Shiiba, K.; Saito, K.; Saito, T. A new assay using surface plasmon response (SPR) to determine binding of the Lactobacillus acidophilus group to human colonic mucin. Biosci. Biotechnol. Biochem. 2004, 68, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Horie, M.; Ishiyama, A.; Fujihira-Ukei, Y.; Sillanpaa, J.; Korhonen, T.K.; Toba, T. Inhibition of the adherence of Escherichia coli strains to basement membrane by Lactobacillus crispatus expressing an S-layer. J. Appl. Microbiol. 2002, 92, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.E.V.; Holmén Larsson, J.M.; Hansson, G.C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host–microbial interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 4659–4665. [Google Scholar] [CrossRef]
- Johansson, M.E.V.; Phillipson, M.; Petersson, J.; Velcich, A.; Holm, L.; Hansson, G.C. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc. Natl. Acad. Sci. USA 2008, 105, 15064–15069. [Google Scholar] [CrossRef]
- Holst, B.; Glenting, J.; Holmstrøm, K.; Israelsen, H.; Vrang, A.; Antonsson, M.; Ahrné, S. Molecular Switch Controlling Expression of the Mannose-Specific Adhesin, Msa, in Lactobacillus plantarum. Appl. Environ. Microbiol. 2019, 85, e02954-18. [Google Scholar] [CrossRef]
- Nishiyama, K.; Mukai, T. Colonization Properties of Lactic Acid Bacteria to Mucosal Surface of the Intestinal Tract. Kagakutoseibutsu 2016, 54, 471–477. [Google Scholar]
- Katsikogianni, M.; Missirlis, Y.F. Concise review of mechanisms of bacterial adhesion to biomaterials and of techniques used in estimating bacteria–material interactions. Eur. Cell Mater. 2004, 8, 37–57. [Google Scholar] [CrossRef]
- Ammar, Y.; Swailes, D.; Bridgens, B.; Chen, J. Influence of surface roughness on the initial formation of biofilm. Surf. Coat. Technol. 2015, 284, 410–416. [Google Scholar] [CrossRef]
- Hamadi, F.; Latrache, H.; Mabrrouki, M.; El Ghmari, A.; Outzourhit, A.; Ellouali, M.; Chtaini, A. Effect of pH on distribution and adhesion of Staphylococcus aureus to glass. J. Adhes. Sci. Technol. 2005, 19, 73–85. [Google Scholar] [CrossRef]
- Slullitel, P.A.; Buttaro, M.A.; Greco, G.; Oñativia, J.I.; Sánchez, M.L.; Mc Loughlin, S.; García-Ávila, C.; Comba, F.; Zanotti, G.; Piccaluga, F. No lower bacterial adhesion for ceramics compared to otherbiomaterials: An in vitro analysis. Orthop. Traumatol. Surg. Res. 2018, 104, 439–443. [Google Scholar] [CrossRef] [PubMed]
- Teughels, W.; Van Assche, N.; Sliepen, I.; Quirynen, M. Effect of material characteristics and/or surface topography on biofilm development. Clin. Oral Implants Res. 2006, 17, 68–81. [Google Scholar] [CrossRef] [PubMed]
- Gottenbos, B.; Busscher, H.J.; van der Mei, H.C.; Nieuwenhuis, P. Pathogenesis and prevention of biomaterial centered infections. J. Mater. Sci. Mater. Med. 2002, 13, 717–722. [Google Scholar] [CrossRef] [PubMed]
- Kirov, S.M. Bacteria that express lateral fragella enable dissection of the multifunctional roles of fragella in pathogenesis. FEMS Microbiol. Lett. 2003, 224, 151–159. [Google Scholar] [CrossRef]
- Taylor, R.L.; Verran, J.; Lees, G.C.; Ward, A.J.P. The influence of substratum topography on bacterial adhesion to polymethyl methacrylate. J. Mater. Sci. Mater. Med. 1998, 9, 17–22. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Taoka, Y.; Sakai, K.; Kinoshita, H.; Fukuyama, H.; Kobayashi, T.; Hirano, A.; Kuroki, N.; Kuroki, M.; Kimura, A. Evaluation of Rate of Adhesion of Lactobacillus namurensis Strain GYP-74 to Porous Fine Ceramics. Processes 2021, 9, 658. https://doi.org/10.3390/pr9040658
Taoka Y, Sakai K, Kinoshita H, Fukuyama H, Kobayashi T, Hirano A, Kuroki N, Kuroki M, Kimura A. Evaluation of Rate of Adhesion of Lactobacillus namurensis Strain GYP-74 to Porous Fine Ceramics. Processes. 2021; 9(4):658. https://doi.org/10.3390/pr9040658
Chicago/Turabian StyleTaoka, Yousuke, Kentaro Sakai, Hiroyuki Kinoshita, Hanako Fukuyama, Taichi Kobayashi, Atsushi Hirano, Noriaki Kuroki, Michiyo Kuroki, and Akihiko Kimura. 2021. "Evaluation of Rate of Adhesion of Lactobacillus namurensis Strain GYP-74 to Porous Fine Ceramics" Processes 9, no. 4: 658. https://doi.org/10.3390/pr9040658
APA StyleTaoka, Y., Sakai, K., Kinoshita, H., Fukuyama, H., Kobayashi, T., Hirano, A., Kuroki, N., Kuroki, M., & Kimura, A. (2021). Evaluation of Rate of Adhesion of Lactobacillus namurensis Strain GYP-74 to Porous Fine Ceramics. Processes, 9(4), 658. https://doi.org/10.3390/pr9040658




