Novel Purification Process for Amyloid Beta Peptide(1-40)
Abstract
1. Introduction
2. Results and Discussion
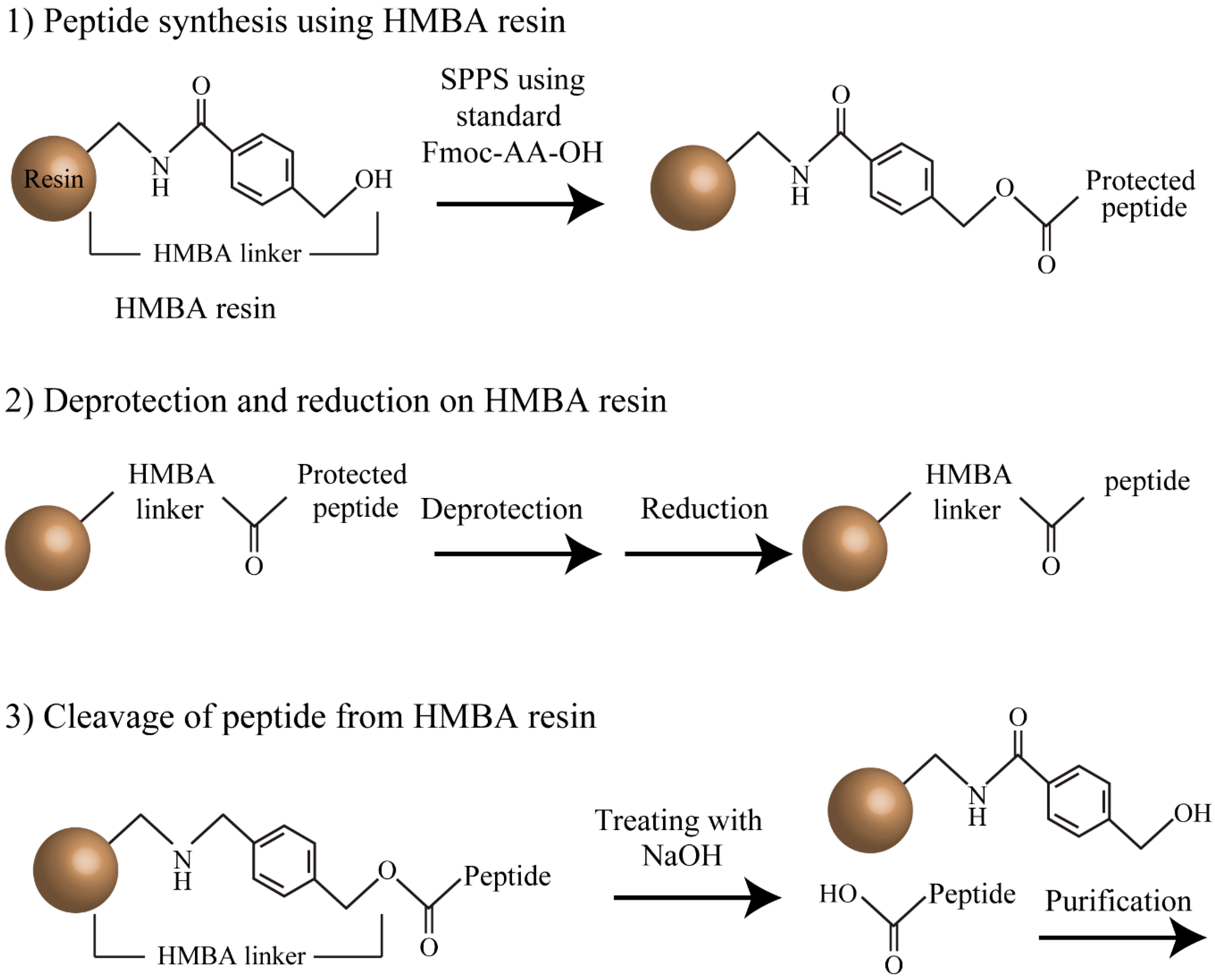
2.1. Fmoc Peptide Synthesis Using HMBA-Resin
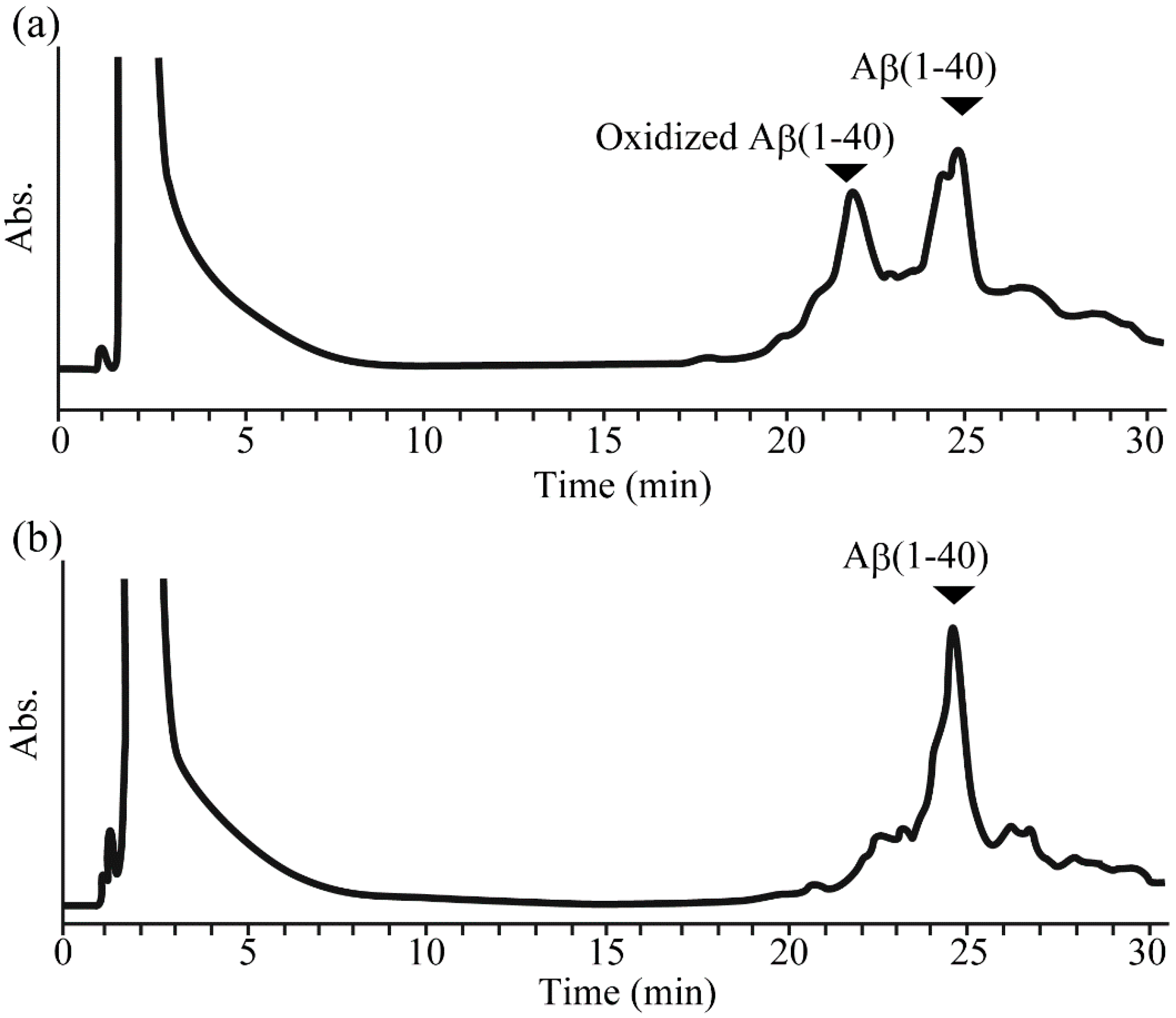
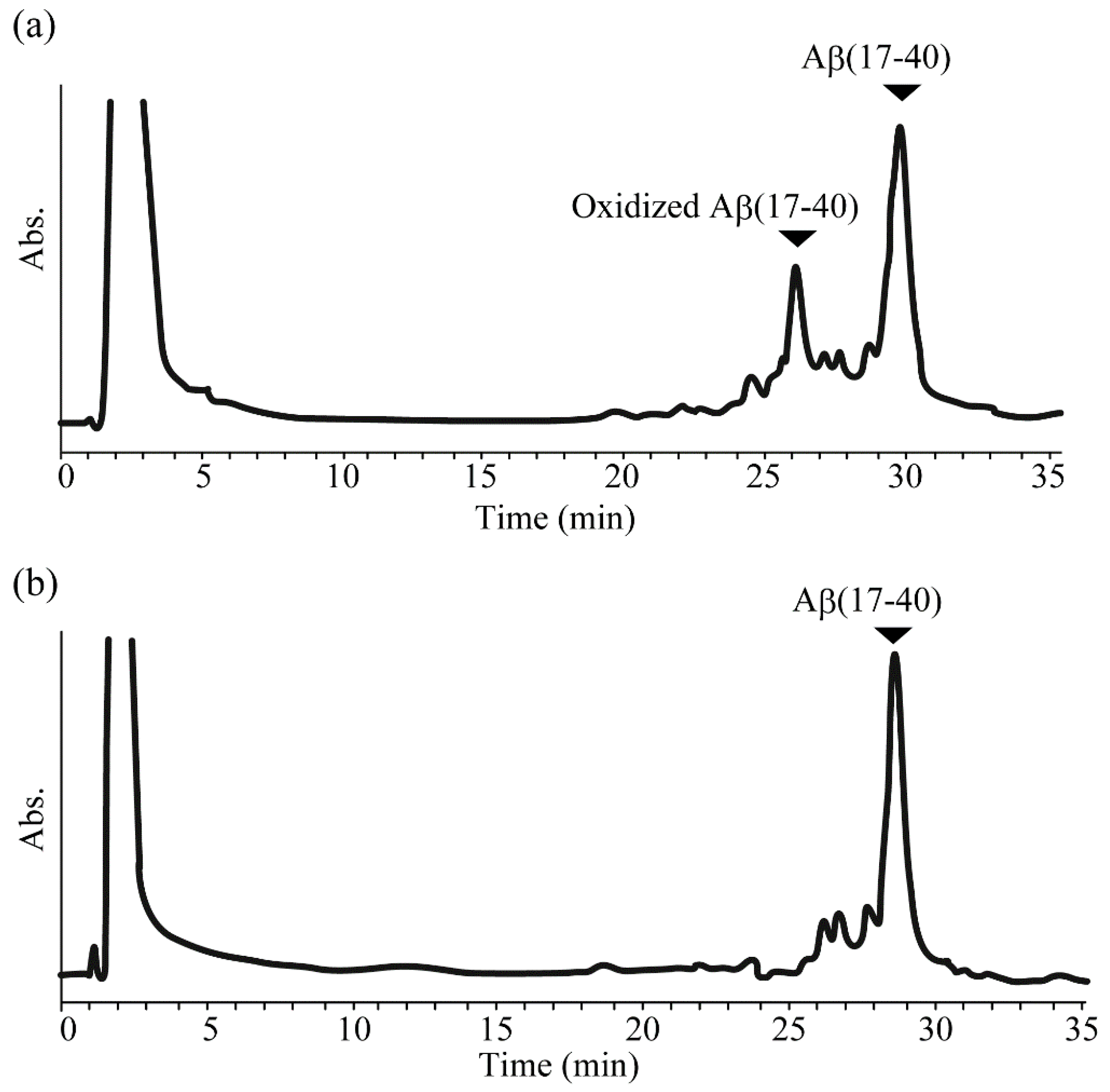
2.2. Reduction of Oxidized Aβ(1-40) and Aβ(17-40) on the Resin
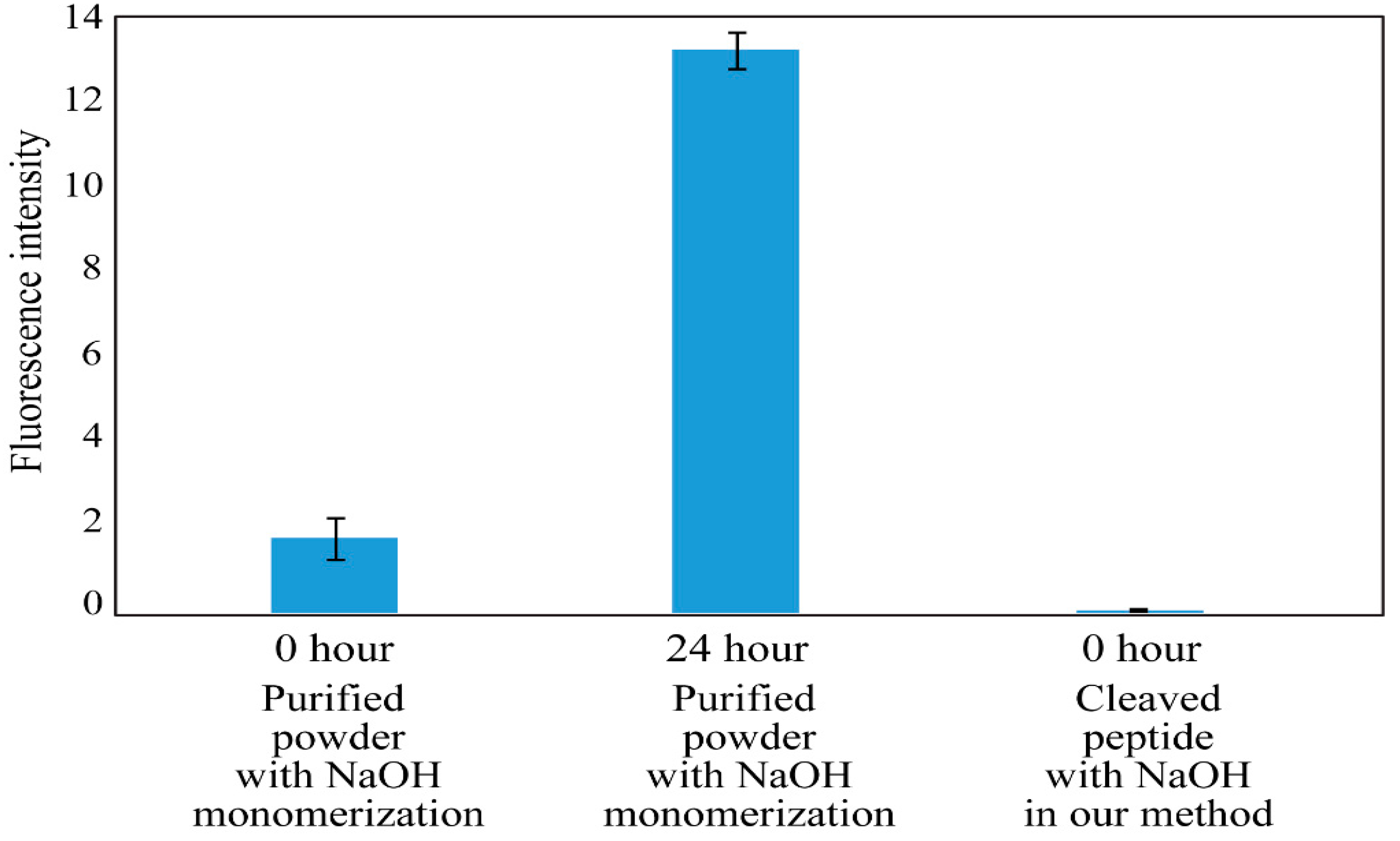
2.3. ThT Assay of Aβ (1-40) Cleaved from HMBA Resin
2.4. Optimization of the Purification Process of Aβ(1-40)
3. Materials and Methods
3.1. Materials
3.2. Fmoc Peptide Synthesis of Aβ(1-40) and Aβ(17-40) Using HMBA Resin
3.3. Deprotection of Aβ(1-40) and Aβ(17-40) Using HMBA Resin
3.4. Cleavage of Aβ(1-40) and Aβ(17-40) from HMBA Resin
3.5. Reduction of Oxidized Aβ(1-40) and Aβ(17-40) on the Resin
3.6. Analysis of Aβ(1-40) and Aβ(17-40)
3.7. Thioflavin T (ThT) Fluorescence Assay
3.8. Purification of Aβ (1-40)
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Knowles, T.P.; Vendruscolo, M.; Dobson, C.M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell. Biol. 2014, 15, 384–396. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.W. Alternative conformations of amyloidogenic proteins govern their behavior. Curr. Opin. Struct. Biol. 1996, 6, 11–17. [Google Scholar] [CrossRef]
- George, G.G. Alzheimer’s disease: Its proteins and genes. Cell 1988, 52, 307–308. [Google Scholar]
- Tanzi, R.E.; Bertram, L. Twenty years of the Alzheimer’s disease amyloid hypothesis: A genetic perspective. Cell 2005, 120, 545–555. [Google Scholar] [CrossRef]
- Hamley, I.W. The amyloid beta peptide: A chemist’s perspective. Role in Alzheimer’s and fibrillization. Chem. Rev. 2012, 12, 5147–5192. [Google Scholar] [CrossRef]
- Changiz, G.; Wu, C.-K.; Daniel, S.; Alfredo, L.; Menglan, Y.; Bruce, A.Y. Aging renders the brain vulnerable to amyloid β-protein neurotoxicity. Nat. Med. 1998, 4, 827–831. [Google Scholar]
- Tickler, A.K.; Clippingdale, A.B.; Wade, J.D. Amyloid-beta as a “difficult sequence” in solid phase peptide synthesis. Protein. Pept. Lett. 2004, 11, 377–384. [Google Scholar] [CrossRef]
- Usui, K.; Hulleman, J.D.; Paulsson, J.F.; Siegel, S.J.; Powers, E.T.; Kelly, J.W. Site-specific modification of Alzheimer’s peptides by cholesterol oxidation products enhances aggregation energetics and neurotoxicity. Proc. Natl. Acad. Sci. USA 2009, 106, 18563–18568. [Google Scholar] [CrossRef]
- Zarándi, M.; Soós, K.; Fülöp, L.; Bozsó, Z.; Datki, Z.; Tóth, G.K.; Penke, B. Synthesis of Abeta [1-42] and its derivatives with improved efficiency. J. Pept. Sci. 2007, 13, 94–99. [Google Scholar] [CrossRef]
- Usui, K.; Mie, M.; Andou, T.; Mihara, H.; Kobatake, E. Fluorescent and luminescent fusion proteins for analyses of amyloid beta peptide aggregation. J. Pept. Sci. 2017, 23, 659–665. [Google Scholar] [CrossRef]
- Guichou, J.-F.; Patiny, L.M. Mutter. Pseudo-prolines (Psi Pro): Direct insertion of Psi Pro systems into cysteine containing peptides. Tetrahedron Lett. 2002, 43, 4389–4390. [Google Scholar] [CrossRef]
- Johnson, T.; Quibell, M.; Sheppard, R.C. N,O-bisFmoc derivatives of N-(2-hydroxy-4-methoxybenzyl)-amino acids: Useful intermediates in peptide synthesis. J. Pept. Sci. 1995, 1, 11–25. [Google Scholar] [CrossRef] [PubMed]
- Martin, Q.; William, G.T.; Tony, J. Improved preparation of β-amyloid(1–43): Structural insight leading to optimised positioning of N-(2-hydroxy-4-methoxybenzyl)(Hmb) backbone amide protection. J. Chem. Soc. Perkin Trans. 1995, 1, 2019–2024. [Google Scholar]
- Kasim, J.K.; Kavianinia, I.; Ng, J.; Harris, P.W.R.; Birch, N.P.; Brimble, M.A. Efficient synthesis and characterisation of the amyloid beta peptide, Aβ1-42, using a double linker system. Org. Biomol. Chem. 2018, 17, 30–34. [Google Scholar] [CrossRef]
- Sohma, Y.; Sasaki, M.; Hayashi, Y.; Kimura, T.; Kiso, Y. Novel and efficient synthesis of difficult sequence-containing peptides through O-N intramolecular acyl migration reaction of O-acyl isopeptides. Chem. Commun. 2004, 124–125. [Google Scholar] [CrossRef]
- Sohma, Y.; Hayashi, Y.; Skwarczynski, M.; Hamada, Y.; Sasaki, M.; Kimura, T.; Kiso, Y. O-N intramolecular acyl migration reaction in the development of prodrugs and the synthesis of difficult sequence-containing bioactive peptides. Biopolymers 2004, 76, 344–356. [Google Scholar] [CrossRef]
- Sohma, Y.; Sasaki, M.; Hayashi, Y.; Kimura, T.; Kiso, Y. Design and synthesis of a novel water-soluble Aβ1-42 isopeptide: An efficient strategy for the preparation of Alzheimer’s disease-related peptide, Aβ1-42, via O–N intramolecular acyl migration reaction. Tetrahedron Lett. 2004, 45, 5965–5968. [Google Scholar] [CrossRef]
- Hansen, J.; Diness, F.; Meldal, M. C-Terminally modified peptides via cleavage of the HMBA linker by O-, N- or S-nucleophiles. Org. Biomol. Chem. 2016, 14, 3238–3245. [Google Scholar] [CrossRef]
- Fezoui, Y.; Hartley, D.M.; Harper, J.D.; Khurana, R.; Walsh, D.M.; Condron, M.M.; Selkoe, D.J.; Lansbury, P.T., Jr.; Fink, A.L.; Teplow, D.B. An improved method of preparing the amyloid beta-protein for fibrillogenesis and neurotoxicity experiments. Amyloid 2000, 7, 166–178. [Google Scholar] [CrossRef]
- Marta, V.; Ernesto, N.; Fina, C.; Ernest, G. Reduction of methionine sulfoxide with NH4I/TFA: Compatibility with peptides containing cysteine and aromatic amino acids. Tetrahedron 1998, 54, 15273–15286. [Google Scholar]
- Biancalana, M.; Koide, S. Molecular mechanism of thioflavin-T binding to amyloid fibrils. Biochim. Biophys. Acta. 2010, 1804, 1405–1412. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.C.; White, P.D. Fmoc Solid Phase Peptide Synthesis; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Usui, K.; Yokota, S.-i.; Iwata, K.; Hamada, Y. Novel Purification Process for Amyloid Beta Peptide(1-40). Processes 2020, 8, 464. https://doi.org/10.3390/pr8040464
Usui K, Yokota S-i, Iwata K, Hamada Y. Novel Purification Process for Amyloid Beta Peptide(1-40). Processes. 2020; 8(4):464. https://doi.org/10.3390/pr8040464
Chicago/Turabian StyleUsui, Kenji, Shin-ichiro Yokota, Kazuya Iwata, and Yoshio Hamada. 2020. "Novel Purification Process for Amyloid Beta Peptide(1-40)" Processes 8, no. 4: 464. https://doi.org/10.3390/pr8040464
APA StyleUsui, K., Yokota, S.-i., Iwata, K., & Hamada, Y. (2020). Novel Purification Process for Amyloid Beta Peptide(1-40). Processes, 8(4), 464. https://doi.org/10.3390/pr8040464





