A Genetic Programming Strategy to Induce Logical Rules for Clinical Data Analysis
Abstract
1. Introduction
2. Background
3. Materials and Methods
3.1. Evolutionary Strategy to Build Rule-Based Classifiers (ESRBC)
- Select the set of patterns from a new class i in the input dataset;
- Create an initial population of rules candidate to represent patterns in class i;
- Run the EA on to achieve a final population ;
- Add the most fit individual (rule) r of to the set of rules R (R is empty initially);
- Remove all patterns from class i holding rule r;
- If class i is not empty, then go to step 2;
- If there are more classes in the dataset, then and go to step 1;
- At the end of the process, R has a set of rules learned from each class of the input dataset.
3.2. Fitness Functions
3.3. Genetic Operators
- Mutation by clause, M1: Changes the attribute, comparison operator or value in a randomly chosen clause from the rule by others, also randomly selected;
- Mutation clause by clause, M2: This operator applies the M1 operator clause by clause to a rule. For each clause, the operator decides whether to mutate. If a mutation has been selected, then the operator decides what mutation type to perform. Namely, changing the attribute, the comparison operator or the value of the attribute in the current clause.
- Mutation by transformation, M3: This operator can remove a part of the rule, add a new rule, or apply the M1 operator to the rule. One of the three operations above is selected at random. In the first operation, a position in the rule is randomly selected to remove the left or right side. Then, it randomly selects the part of the rule to be removed. The second operation adds a new rule at the end of the current rule. The added rule is randomly created (by also choosing its size in a random way).
3.4. Running the Evolutionary Algorithm
- Running a genetic algorithm (GA) until it slows down, then letting a local optimizer take over the last generation (and/or best individual) of the GA. Hopefully, the GA is very close to the global optimal.
| Algorithm 1 ESLS |
| Input: POP, the population composed by the individuals in the last generation of the EA. MaxGeneration, the number of generations. MO applies one of the mutation operators given in Section 3.3, chosen at random. , fitness function given in Section 3.2. |
| Output: POP, as a result of improvement of the input. |
1.; 2.while MaxGeneration do 3. ; 4. for all rule r in POP do 5. % Computing fitness 6. f := (r); 7. % Applying mutation. 8. newr := MO(r); 9. % Evaluating new individual. 10. newf := (newr); 11. % Updating the improved individual. 12. if newf > f then r := newr; 13. end for 14.end while 15.end. |
4. Results
- Title: SPECTF Heart Dataset (DS1);
- Number of patterns: 267;
- Number of attributes: 22 plus the class attribute;
- Number of classes: 2 classes;
- Attribute type: binary;
- Missing attribute values: No missing values.
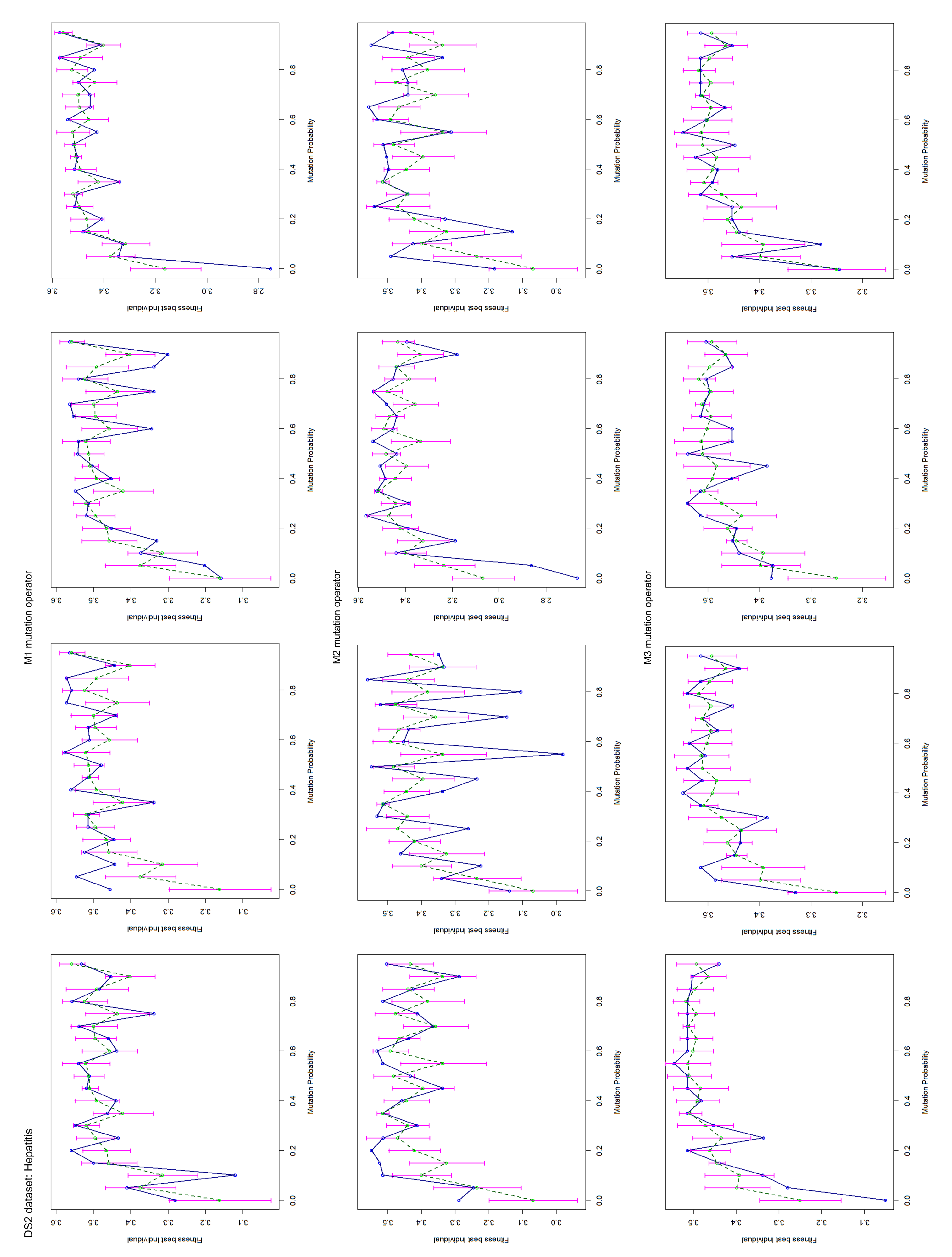
- Title: Hepatitis Domain (DS2);
- Number of patterns: 155;
- Number of attributes: 19 plus the class attribute;
- Number of classes: 2 classes;
- Attribute type: Categorical, Integer and Real;
- Missing attribute values: yes (10-nearest neighbor technique was used in this research for imputation of missing values).
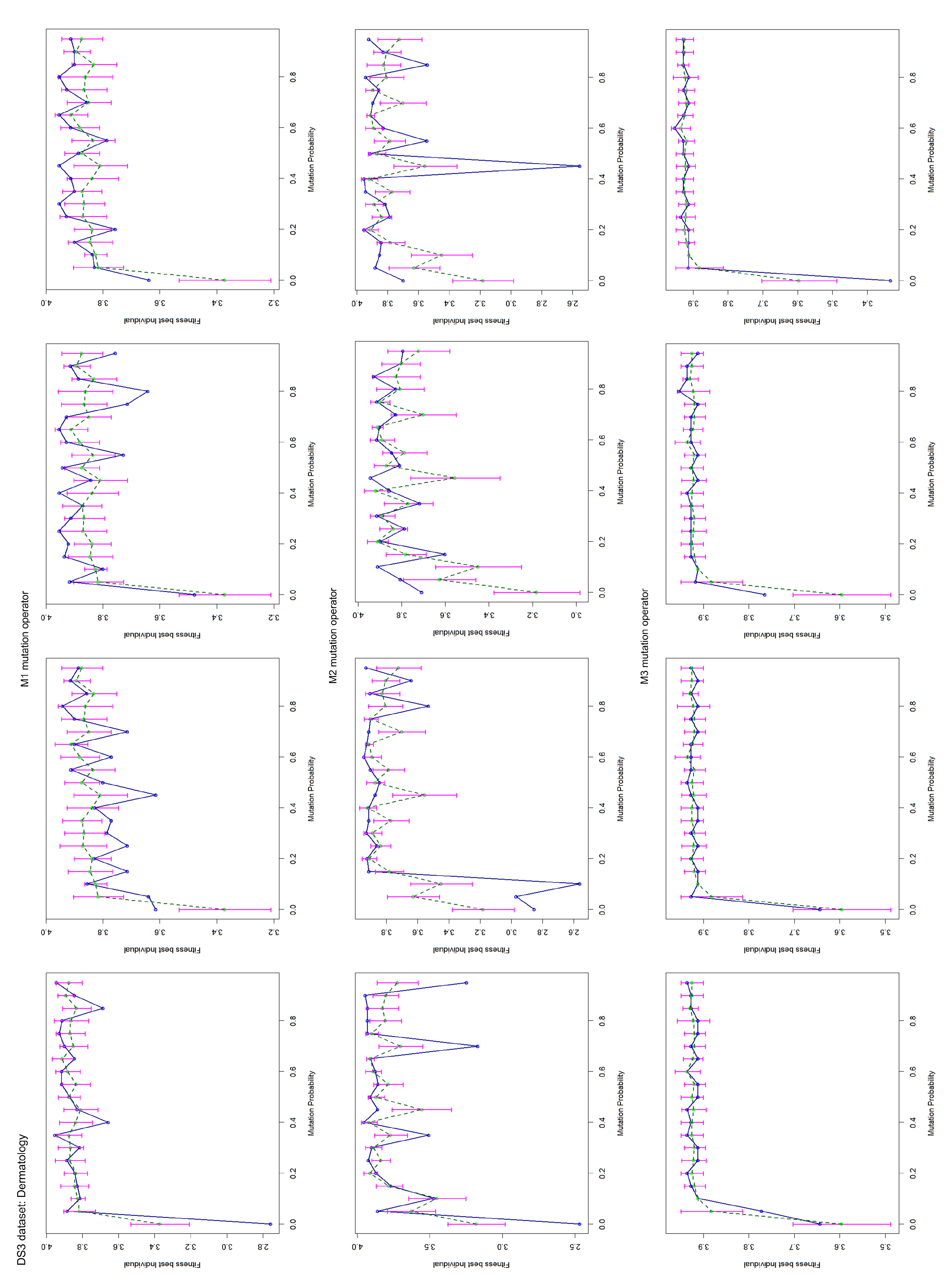
- Title: Dermatology Database (DS3);
- Number of patterns: 366;
- Number of attributes: 34 plus the class attribute;
- Number of classes: 6 classes;
- Attribute type: Categorical and Integer;
- Missing attribute values: yes (20-nearest neighbor technique was used in this research for imputation of missing values).
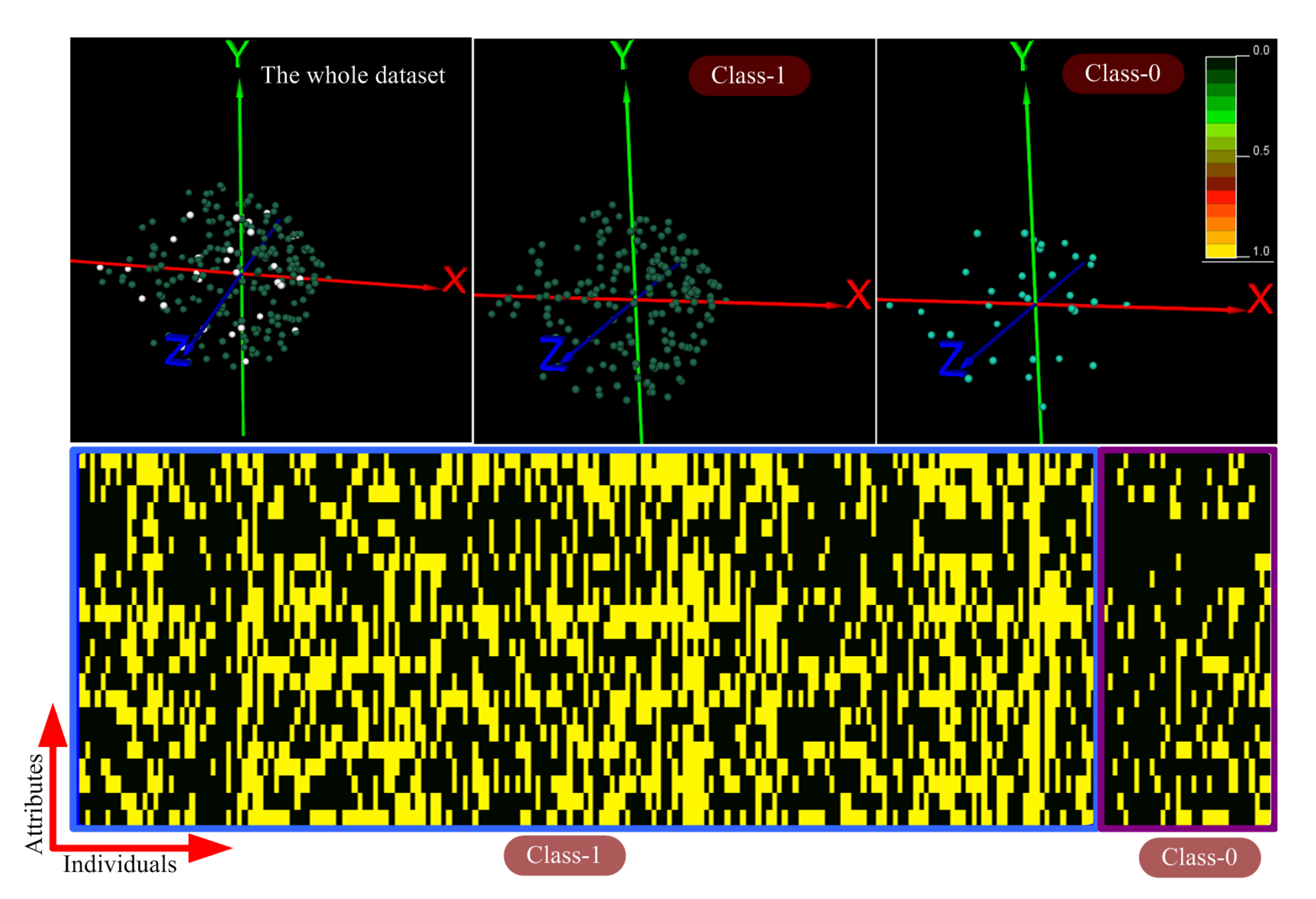
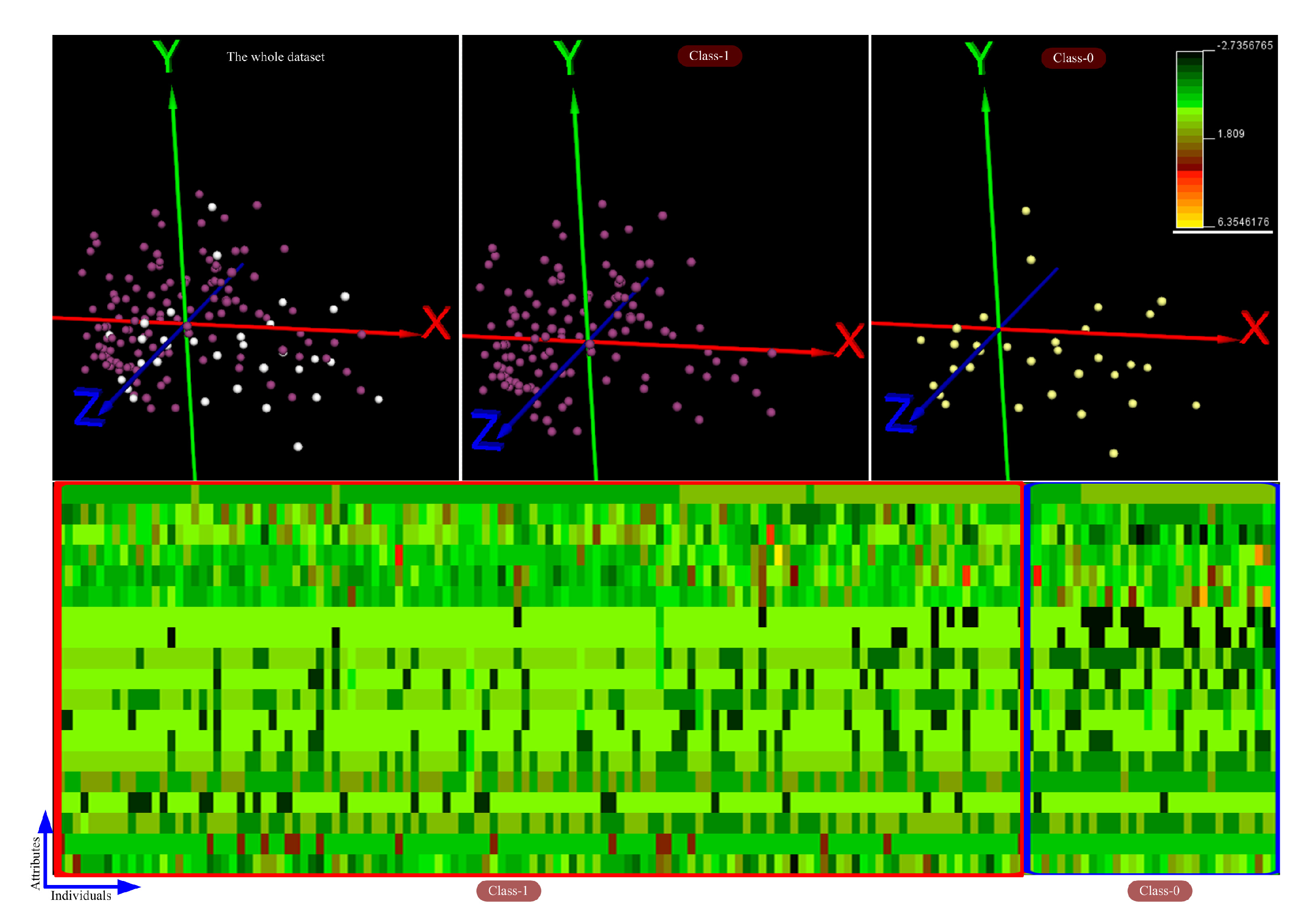
4.1. Exploring and Analyzing the Datasets
4.2. Mutation Operator Evaluation
4.3. Accuracy and Comparison of the Evolutionary Method
5. Discussion: Rule Analysis
- Class-0:
- patients holding ;
- Class-1:
- patients holding ;
- Class-2:
- patients holding ;
- Class-3:
- this class supports four age-related subgroups of patients, namely,, , and ;
- Class-4:
- patients holding ;
- Class-5:
- patients holding .
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A. Rule Based-Classifiers Rendered by the Evolutionary Method
Appendix A.1. Rules of the DS1 Dataset (Heart Dataset)
Appendix A.2. Rules of the DS2 Dataset (Hepatitis Dataset)
Appendix A.3. Rules of the DS3 Dataset (Dermatology Dataset)
Appendix B. Test Charts of the Mutation Operators

References
- Bandyopadhyay, S.; Pal, S.K. Classification and Learning Using Genetic Algorithms: Applications in Bioinformatics and Web Intelligence; Natural Computing Series; Springer: Berlin/Heidelberg, Germany, 2007. [Google Scholar]
- Bonelli, P.; Parodi, A. An efficient classifier system and its experimental comparison with two representative learning methods on three medical domains. In Proceedings of the 4th International Conference Genetic Algorithms (ICGA), San Diego, CA, USA, July 1991; pp. 288–295. [Google Scholar]
- Hong, J.H.; Cho, S.B. The classification of cancer based on DNA microarray data that uses diverse ensemble genetic programming. Artif. Intell. Med. 2006, 36, 43–58. [Google Scholar] [CrossRef] [PubMed]
- Kumar, T.P.; Iba, H. Prediction of Cancer Class with Majority Voting Genetic Programming Classifier Using Gene Expression Data. IEEE/ACM Trans. Comput. Biol. Bioinform. 2009, 6, 353–367. [Google Scholar]
- Kumar, R.; Verma, R. Classification Rule Discovery for Diabetes Patients by Using Genetic Programming. Int. J. Soft Comput. Eng. IJSCE 2012, 2, 183–185. [Google Scholar]
- Larranaga, P.; Calvo, B.; Santana, R.; Bielza, C.; Galdiano, J.; Inza, I.; Lozano, J.A.; Armañanzas, R.; Santafé, G.; Pérez, A.; et al. Machine learning in bioinformatics. Briefings Bioinform. 2006, 7, 86–112. [Google Scholar]
- Liu, K.H.; Xu, C.G. A genetic programming-based approach to the classification of multiclass microarray datasets. Bioinformatics 2009, 25, 331–337. [Google Scholar] [CrossRef] [PubMed]
- Maulik, U.; Bandyopadhyay, S.; Mukhopadhyay, A. Multiobjective Genetic Algorithms for Clustering: Applications in Data Mining and Bioinformatics; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar]
- na Reyes, C.A.P.; Sipper, M. Evolutionary computation in medicine: An overview. Artif. Intell. Med. 2000, 19, 1–23. [Google Scholar] [CrossRef]
- Podgorelec, V.; Kokol, P.; Stiglic, M.M.; Hericko, M.; Rozrnan, I. Knowledge discovery with classification rules in a cardiovascular dataset. Comput. Methods Programs Biomed. 2005, 1, 539–549. [Google Scholar] [CrossRef]
- Soni, J.; Ansari, U.; Sharma, D.; Soni, S. Intelligent and Effective Heart Disease Prediction System using Weighted Associative Classifiers. Int. J. Comput. Sci. Eng. IJCSE 2011, 3, 2385–2392. [Google Scholar]
- Tsakonas, A.; Dounias, G.; Jantzen, J.; Axer, H.; Bjerregaard, B.; von Keyserlingk, D.G. Evolving rule-based systems in two medical domains using genetic programming. Artif. Intell. Med. 2004, 32, 195–216. [Google Scholar] [CrossRef]
- Vargas, C.M.B.; Chidambaram, C.; Hembecker, F.; Silvério, H.L. Computational Biology and Applied Bioinformatics; Chapter A Comparative Study of Machine Learning and Evolutionary Computation Approaches for Protein Secondary Structure Classification; InTech: London, UK, 2011; pp. 239–258. [Google Scholar]
- Wolberg, W.H.; Mangasarian, O.L. Multisurface method of pattern separation for medical diagnosis applied to breast cytology. Proc. Natl. Acad. Sci. USA 1990, 87, 9193–9196. [Google Scholar] [CrossRef]
- Lucas, P. Analysis of notions of diagnosis. Artif. Intell. 1998, 12, 295–343. [Google Scholar] [CrossRef]
- Lucas, P. Prognostic methods in medicine. Artif. Intell. 1999, 15, 105–119. [Google Scholar] [CrossRef]
- Dong, X.; Yu, Z.; Cao, W.; Shi, Y.; Ma, Q. A survey on ensemble learning. In Frontiers of Computer Science; Springer: Berlin/Heidelberg, Germany, 2019. [Google Scholar] [CrossRef]
- Ramos, J.; Castellanos-Garzón, J.A.; González-Briones, A.; de Paz, J.F.; Corchado, J.M. An agent-based clustering approach for gene selection in gene expression microarray. In Interdisciplinary Sciences: Computational Life Sciences; Springer: Berlin/Heidelberg, Germany, 2017; Volume 9, pp. 1–13. [Google Scholar]
- Castellanos-Garzón, J.A.; Ramos, J.; González-Briones, A.; de Paz, J.F. A Clustering-Based Method for Gene Selection to Classify Tissue Samples in Lung Cancer. In 10th International Conference on Practical Applications of Computational Biology & Bioinformatics, Advances in Intelligent Systems and Computing; Saberi Mohamad, M., Rocha, M., Fdez-Riverola, F., Domínguez Mayo, F., De Paz, J., Eds.; Springer: Cham, Switzerland, 2016; Volume 477, pp. 99–107. [Google Scholar]
- Castellanos-Garzón, J.A.; Ramos, J. A Gene Selection Approach based on Clustering for Classification Tasks in Colon Cancer. ADCAIJ Adv. Distrib. Comput. Artif. Intell. J. 2015, 4, 1–10. [Google Scholar] [CrossRef]
- González-Briones, A.; Ramos, J.; De Paz, J.F. A drug identification system for intoxicated drivers based on a systematic review. ADCAIJ Adv. Distrib. Comput. Artif. Intell. J. 2015, 4, 83–101. [Google Scholar]
- Pappa, G.L.; Freitas, A.A. Evolving rule induction algorithms with multi-objective grammar-based genetic programming. In Knowledge and Information Systems; Springer: Berlin/Heidelberg, Germany, 2009; Volume 19, pp. 283–309. [Google Scholar]
- Alcalá-Fdez, J.; Sánchez, L.; García, S.; delJesus, M.J.; Ventura, S.; Garrell, J.M.; Otero, J.; Romero, C.; Bacardit, J.; Rivas, V.M.; et al. KEEL: A software tool to assess evolutionary algorithms for data mining problems. InSoft Computing; Springer: Berlin/Heidelberg, Germany, 2009; Volume 13, pp. 307–318. [Google Scholar]
- Fernández, A.; García, S.; Luengo, J.; Bernadó-Mansilla, E.; Herrera, F. Genetics-Based Machine Learning for Rule Induction: State of the Art, Taxonomy, and Comparative Study. IEEE Trans. Evol. Comput. 2010, 14, 913–941. [Google Scholar] [CrossRef]
- Oyebode, O.K.; Adeyemo, J.A. Genetic Programming: Principles, Applications and Opportunities for Hydrological Modelling. World Acad. Sci. Eng. Technol. Int. J. Environ. Ecol. Geol. Min. Eng. 2014, 8, 310–316. [Google Scholar]
- Ghaheri, A.; Shoar, S.; Naderan, M.; Hoseini, S.S. The applications of genetic algorithms in medicine. Oman Med. J. 2015, 30, 406–416. [Google Scholar] [CrossRef]
- Karnan, M.; Thangavel, K. Automatic detection of the breast border and nipple position on digital mammograms using genetic algorithm for asymmetry approach to detection of microcalcifications. In Computer Methods and Programs in Biomedicine; Elsevier: Amsterdam, The Netherlands, 2007; Volume 87, pp. 12–20. [Google Scholar]
- Gudmundsson, M.; El-Kwae, E.A.; Kabuka, M.R. Edge detection in medical images using a genetic algorithm. IEEE Trans. Med Imaging 1998, 17, 469–474. [Google Scholar] [CrossRef]
- Bhandarkar, S.M.; Zhang, Y.; Potter, W.D. An edge detection technique using genetic algorithm-based optimization. Pattern Recognit. 1994, 27, 1159–1180. [Google Scholar] [CrossRef]
- Jiang, J.; Yao, B.; Wason, A.M. A genetic algorithm design for microcalcification detection and classification in digital mammograms. In Computerized Medical Imaging and Graphics; Elsevier: Amsterdam, The Netherlands, 2007; Volume 31, pp. 49–61. [Google Scholar]
- Yao, B.; Jiang, J.; Peng, Y. A CBR driven genetic algorithm for microcalcification cluster detection. In International Conference on Knowledge Engineering and Knowledge Management; Springer: Berlin/Heidelberg, Germany, 2004; pp. 494–496. [Google Scholar]
- Bevilacqua, A.; Campanini, R.; Lanconelli, N. A distributed genetic algorithm for parameters optimization to detect microcalcifications in digital mammograms. In Workshops on Applications of Evolutionary Computation; Springer: Berlin/Heidelberg, Germany, 2001; pp. 278–287. [Google Scholar]
- Baum, K.G.; Schmidt, E.; Rafferty, K.; Król, A.; Helguera, A. Evaluation of novel genetic algorithm generated schemes for positron emission tomography (PET)/magnetic resonance imaging (MRI) image fusion. J. Digit. Imaging 2011, 24, 1031–1043. [Google Scholar] [CrossRef]
- de Carvalho Filho, A.O.; de Sampaio, W.B.; Silva, A.C.; de Paiva, A.C.; Nunes, R.A.; Gattass, M. Automatic detection of solitary lung nodules using quality threshold clustering, genetic algorithm and diversity index. Artif. Intell. Med. 2014, 60, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Asuntha, A.; Singh, N.; Srinivasan, A. PSO, Genetic Optimization and SVM Algorithm used for Lung Cancer Detection. J. Chem. Pharm. Res. 2016, 8, 351–359. [Google Scholar]
- Alshamlan, H.M.; Badr, G.H.; Alohali, Y.A. Genetic Bee Colony (GBC) algorithm: A new gene selection method for microarray cancer classification. Comput. Biol. Chem. 2015, 56, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Latkowski, T.; Osowski, S. Computerized system for recognition of autism on the basis of gene expression microarray data. Comput. Biol. Med. 2015, 56, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Arabasadi, Z.; Alizadehsani, R.; Roshanzamir, M.; Moosaei, H.; Yarifard, A.A. Computer aided decision-making for heart disease detection using hybrid neural network-Genetic algorithm. Comput. Methods Programs Biomed. 2017, 141, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yuan, D.; Ma, X.; Cui, D.; Cao, L. Genetic algorithm for the Optimization of Features and Neural Networks in ECG Signals Classification; Resreport 7, Scientific Reports; Springer Nature: Berlin/Heidelberg, Germany, 2017. [Google Scholar]
- Lin, T.; Huang, Y.; Lin, J.I.; Balas, V.E.; Srinivasan, S. Genetic algorithm-based interval type-2 fuzzy model identification for people with type-1 diabetes. In Proceedings of the IEEE International Conference on Fuzzy Systems (FUZZ-IEEE), Naples, Italy, 9–12 July 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Nguyen, L.B.; Nguyen, A.V.; Ling, S.H.; Nguyen, H.T. Combining genetic algorithm and Levenberg-Marquardt algorithm in training neural network for hypoglycemia detection using EEG signals. In Proceedings of the Engineering in Medicine and Biology Society (EMBC), 2013 35th Annual International Conference of the IEEE, Osaka, Japan, 3–7 July 2013; pp. 5386–5389. [Google Scholar]
- Ocak, H. A medical decision support system based on support vector machines and the genetic algorithm for the evaluation of fetal well-being. J. Med Syst. 2013, 37, 9913. [Google Scholar] [CrossRef] [PubMed]
- Nyathi, T.; Pillay, N. Automated Design of Genetic Programming Classification Algorithms Using a Genetic Algorithm. In Applications of Evolutionary Computation: 20th European Conference, EvoApplications 2017, Proceedings of the Part II, Amsterdam, The Netherlands, 19–21 April 2017; Squillero, G., Sim, K., Eds.; Chapter Applications of Evolutionary Computation. EvoApplications 2017. Lecture Notes in Computer Science; Springer International Publishing: Cham, Switzerland, 2017; Volume 10200, pp. 224–239. [Google Scholar] [CrossRef]
- Naredo, E.; Trujillo, L.; Legrand, P.; Silva, S.; Muńoz, L. Evolving genetic programming classifiers with novelty search. Inf. Sci. 2016, 369, 347–367. [Google Scholar] [CrossRef]
- Dick, G. Improving geometric semantic genetic programming with safe tree initialisation. In Genetic Programming: 18th European Conference, EuroGP 2015, Copenhagen, Denmark, 8–10 April 2015; Machado, P., Heywood, M.I., McDermott, J., Castelli, M., García-Sánchez, P., Burelli, P., Risi, S., Sim, K., Eds.; Chapter European Conference on Genetic Programming, EuroGP 2015: Genetic Programming; Springer International Publishing: Cham, Switzerland, 2015; pp. 28–40. [Google Scholar] [CrossRef]
- Alotaiby, T.N.; Alrshoud, S.R.; Alshebeili, S.A.; Alhumaid, M.H.; Alsabhan, W.M. Epileptic MEG Spike Detection Using Statistical Features and Genetic Programming with KNN. J. Healthc. Eng. Hindawi 2017, 2017, 7. [Google Scholar] [CrossRef]
- Wang, C.S.; Juan, C.J.; Lin, T.Y.; Yeh, C.C.; Chiang, S.Y. Prediction Model of Cervical Spine Disease Established by Genetic Programming. In Proceedings of the 4th Multidisciplinary International Social Networks Conference (MISNC ’17); ACM: New York, NY, USA, July 2017; pp. 381–386. [Google Scholar] [CrossRef]
- Burks, A.R.; Punch, W.F. Genetic Programming for Tuberculosis Screening from Raw X-ray Images. In Proceedings of the Genetic and Evolutionary Computation Conference (GECCO ’18), New York, NY, USA, 15–19 July 2018; pp. 1214–1221. [Google Scholar] [CrossRef]
- Tan, P.N.; Steinbach, M.; Kumar, V. Introduction to Data Mining; Addison-Wesley: Boston, MA, USA, 2006. [Google Scholar]
- Freitas, A.A. Soft Computing for Knowledge Discovery and Data Mining, Part II; Chapter A Review of Evolutionary Algorithms for Data Mining; Springer: Berlin/Heidelberg, Germany, 2008; pp. 79–111. [Google Scholar]
- Witten, I.H.; Frank, E. Data Mining: Practical Machine Learning Tools and Techniques, 2nd ed.; Morgan Kaufmann: Burlington, MA, USA, 2005. [Google Scholar]
- Pappa, G.L.; Freitas, A.A. Automating the Design of Data Mining Algorithms: An Evolutionary Computation Approach; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar]
- Espejo, P.G.; Ventura, S.; Herrera, F. A Survey on the Application of Genetic Programming to Classification. IEEE Trans. Syst. Man Cybern. Part Appl. Rev. 2010, 40, 121–144. [Google Scholar] [CrossRef]
- Freitas, A.A. A survey of evolutionary algorithms for data mining and knowledge discovery. Advances in Evolutionary Computation. In Advances in Evolutionary Computation; Ghosh, A., Tsutsui, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2002; pp. 819–845. [Google Scholar]
- Flach, P. MACHINE LEARNING: The Art and Science of Algorithms that Make Sense of Data; Cambridge University Press: Cambridge, UK, 2012. [Google Scholar]
- Witten, I.H.; Frank, E.; Hall, M.A. Data Mining: Practical Machine Learning, Tools and Techniques, 3rd ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Goldberg, D.E. Genetic Algorithms in Search, Optimization and Machine Learning; Addison-Wesley: Reading, MA, USA, 1989. [Google Scholar]
- Bacardit, J.; Goldberg, D.E.; Butz, M.V. Improving the performance of a Pittsburgh learning classifier system using a default rule. In Proceedings Revised Select Papers International Workshop Learning Classifier Systems; Springer: Berlin/Heidelberg, Germany, 2007; pp. 291–307. [Google Scholar]
- Haupt, R.L.; Haupt, S.E. Practical Genetic Algorithms, 2nd, ed.; John Wiley & Sons Inc.: Hoboken, NJ, USA, 2004. [Google Scholar]
- Castellanos-Garzón, J.A.; Díaz, F. An Evolutionary Computational Model Applied to Cluster Analysis of DNA Microarray Data. Expert Syst. Appl. 2013, 40, 2575–2591. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Blake, C.; Merz, C. Repository of Machine Learning Databases (UCI); Center for Machine Learning and Intelligent Systems: Irvine, CA, USA, 1998. [Google Scholar]
- Wu, X.; Kumar, V.; Ross Quinlan, J.; Ghosh, J.; Yang, Q.; Motoda, H.; McLachlan, G.; Ng, A.; Liu, B.; Yu, P.; et al. Top 10 algorithms in data mining. Knowl. Inf. Syst. 2008, 14, 1–37. [Google Scholar] [CrossRef]
- Moerland, P. Mixture for Latent Variable Models for Density Estimation and Classification; Technical Report; Dalle Molle Institution for Perceptual Artificial Intelligencie, IDIAP: Martigny, Switzerland, 2000. [Google Scholar]

| ESRBC Settings | DS1 | DS2 | DS3 |
|---|---|---|---|
| Population size | 70 | 50 | 100 |
| Number of generations per rule (fitness function-1) | 100,000 | 100,000 | 200,000 |
| Number of generations per rule (fitness function-2) | 200,000 | 200,000 | 400,000 |
| Mutation operator per rule for Algorithm 1 (local search) | 10,000 | 10,000 | 100,000 |
| Mutation operator | M3 | M1 and M3 | M3 |
| Crossover probability | 0.6 | 0.6 | 0.6 |
| Mutation probability | 0.2 | 0.3 | 0.2 |
| Maximum size of rules (maximum number of clauses) | 10 | 10 | 10 |
| Method | Description |
|---|---|
| SVM | Linear Support Vector Machine, which finds the best hyperplane separating both classes. |
| naiveBayes | This model computes the probability of each class given the values of all attributes and assuming the attribute conditional independence. |
| KNN | k-Nearest Neighbor classification. This is a lazy model which classifies the input pattern by using its k-Nearest Neighbors from the training set. |
| ANN | Artificial Neural Network implementing a Multilayer Perceptron, which uses a single intermediate layer for our case. Backpropagation and resilient backpropagation have been implemented. |
| ML-MPCA | Maximum Likelihood estimation with Mixture of Principal Component Analyzers. |
| Bayesian-MPCA | Bayesian approach with Mixture of Principal Component Analyzers. |
| Dataset | Method | Accuracy (%) | Youden Index (%) |
|---|---|---|---|
| DS1 | SVM | 78.95 | 12.45 |
| naiveBayes | 46.93 | 14.97 | |
| k-NN (k = 3) | 81.58 | 22.91 | |
| ANN | 79.39 | 12.95 | |
| ESRBC | 81.82 | 40.48 | |
| DS2 | SVM | 82.57 | 46.91 |
| naiveBayes | 75.47 | 37.56 | |
| k-NN (k = 5) | 75.47 | 16.47 | |
| ANN | 81.94 | 42.56 | |
| ESRBC | 81.32 | 41.67 | |
| DS3 | ML-MPCA | 94.9 | - |
| Bayesian-MPCA | 95.8 | - | |
| ESRBC | 95.92 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castellanos-Garzón, J.A.; Mezquita Martín, Y.; Jaimes Sánchez, J.L.; López García, S.M.; Costa, E. A Genetic Programming Strategy to Induce Logical Rules for Clinical Data Analysis. Processes 2020, 8, 1565. https://doi.org/10.3390/pr8121565
Castellanos-Garzón JA, Mezquita Martín Y, Jaimes Sánchez JL, López García SM, Costa E. A Genetic Programming Strategy to Induce Logical Rules for Clinical Data Analysis. Processes. 2020; 8(12):1565. https://doi.org/10.3390/pr8121565
Chicago/Turabian StyleCastellanos-Garzón, José A., Yeray Mezquita Martín, José Luis Jaimes Sánchez, Santiago Manuel López García, and Ernesto Costa. 2020. "A Genetic Programming Strategy to Induce Logical Rules for Clinical Data Analysis" Processes 8, no. 12: 1565. https://doi.org/10.3390/pr8121565
APA StyleCastellanos-Garzón, J. A., Mezquita Martín, Y., Jaimes Sánchez, J. L., López García, S. M., & Costa, E. (2020). A Genetic Programming Strategy to Induce Logical Rules for Clinical Data Analysis. Processes, 8(12), 1565. https://doi.org/10.3390/pr8121565






